ISSN: 0973-7510
E-ISSN: 2581-690X
Keratinase is a proteolytic enzyme capable on degrading the hardy polymeric biomolecule or keratin. In recent days, the utilization of keratinolytic microorganisms is seen as a promising way in recycling the keratin wastes from the avian and mammalians into valuable derived products. Previous study has reported the presence of a keratinolytic bacterium, later identified as Azotobacter chroococcum B4 obtained from dump soils. The present study investigates the enzyme characteristics of keratinase produced by this strain based on the physical appearance of final degraded product using SEM, the molecular weight of keratinase using SDS-PAGE, the effects of nutrition (C/N-source) on strain production of keratinase and the enzyme stability in metal ions solution. The molecular weight of keratinase produced by A. chroococcum B4 was about 30 kDa. Both sucrose and tryptone supplementation increase the keratinase activity by 71.7 and 97.8 U/mL after 96 h of cultivation. Metal ions, Ca2+, Mg2+, Mn2+, Na+, and K+ are regarded as activators by increasing the relative activity of keratinase by 117, 166, 111, 113, and 112% respectively, while phenylmethylsulfonyl flouride (PMSF) is regarded as inhibitor by decreasing the relative activity down to 31%. Based on the metal ion characteristics, this strain produced a serine-protease type of keratinase which may further studied for its application in the field.
Azotobacter chrocoocum, keratinase, sucrose, tryptone
Poultry-processing industries generate more than 4 billion pounds of feathers annually as waste. In chicken feather weight accounts up to 5-7% of its body chicken. Chicken feather contains about 90% of keratin1–2.
Keratin is a stable and insoluble structural protein tightly packed by the numerous polypeptide chained with disulfide cross-linkage (S=S bonds) which increased the hydrophobic nature of this polymer. It is highly resistant to degradation by conventional proteases. The stability of keratin depends on number of helical chains (a-keratin) and β-sheet (b-keratin) of proteins and their cystine bridge. Biodegradation of keratin involving microbial ketainase is an easy and economical method for conversion of keratin waste into useful products as source of amino acids and proteins, which are typically utilized as animal feed additives as well as nitrogen source for plants3–5. Keratinase can be utilized as biocatalysts in leather and textile industry, and waste recycling4.
Keratinase is produced by various types of fungi and bacteria, including Gram-positive bacteria of Bacillus6–7 and Gram-negative, such as Vibrio sp. strain kr28, Aeromonas hydrophila strain FB38, Stenotrophomonas spp.9–11, Chryseobacterium spp12–13. A fungal isolate from crocodilian feces was reported to degrade chicken feather waste within 10 days14.
During a screening of keratinolytic bacteria from various sources, we reported a new strain of Azotobacter chroococcum B411 which produced a considerably potential keratinolytic activity. The purpose of this study was then to obtain the biological characteristics of keratinase produced by A. chroococcum strain B4 for a better understanding of its nature in the future application.
Isolate and growth medium
Azotobacter chroococcum B4 was grown on feather meal broth (FMB) composed of chicken feather powder (15 g), K2HPO4 (0.7 g), KH2PO4 (0.4 g), NaCl (0.5 g), MgSO4 (0.1 g) in 1000 mL distilled water14. Growth medium was sterilized at 121°C, 1 atm for 15 min.
Chicken feather waste was collected from slaughterhouses. Feathers were washed using detergent-water and cut into small pieces (± 2 cm) prior soaking in acetone for 24 h. Washed feathers were dried in oven at 40°C for 72 h and subjected to be grinded to make chicken feather powder.
Observation using scanning electron microscope (SEM)
Whole chicken feathers were inserted into A. chroococcus B4 liquid culture and incubated for 48–96 h, 37°C under agitation of 180 rpm. After incubation, a feather or strand sample was soaked into a 2% (w/v) sodium coccodylate buffer for 1 h. The sample was further soaked into a 1% (w/v) tetraoxide solution for 1 h. The sample was removed and dipped into following EtOH concentrations of: 70, 80, 90 and 100%, each for 30 sec. The sample was removed and immersed into a n-butanol solution prior coating into gold-coated metal plate. The plate was placed inside SEM apparatus operating at 20 kV while tubes were conditioned in vacuum (0 Pa).
Effect of C- and N-sources
Variation in carbon and nitrogen sources was used to promote bacterial keratinolytic activity. Glucose, sucrose, starch, fructose, lactose, and sorbitol (1% w/v) was tested as C-source, while casein, gelatin, KNO3, peptone, tryptone, NaNO3, and yeast extract (0.5% w/v) was used as N-source. pH and temperature were adjusted as described in previous study10. Keratinase activity was measured at 24, 48, 72, 96, 120, 144, and 168 hours.
Purification of keratinase
Crude keratinase was harvested from an overnight culture by centrifugation at 8000×g, 4°C for 20 min. Supernatant was precipitated using 70% (w/v) (NH4)2SO4 at 5°C for 24 h and subjected to further centrifugation at 8000×g, 4°C for 20 min. Ten milligrams of pellets were dissolved by adding 5 mL buffer A (2:1, 25 mM Tris-HCl, pH 8) and dialyzed inside a cellulose membrane (10 kDa cut-off). Dialysis membrane was then submerged into 600 mL buffer B (50 mM Tris-HCl, pH 8) following slow stir for 24 h at 5°C. The buffer was periodically changed in the interval of 8 h. Enzyme solution was further purified using a Sephadex G-50 (Sigma-Aldrich) gel filtration chromatography column (fractionation range 1.5 to 30 kDa). Partially-purified protein solutions were gently poured into the column and eluted with 25 mM Tris-HCl, pH 8.0. Fractions were stored in vials and subjected to further experiments.
Keratinase assay
Keratinase assay was performed as previously described10 with some modifications. Chicken feather powder was used a keratin source and substrate in the keratinase assay. The keratin substrate (4 mg) was dissolved in 1 mL 50 mM Tris–HCl, pH 8.0. Keratin solution (250 µL) was reacted with 500 µL enzyme solution in 250 μL 50 mM Tris–HCl buffer. Solution was placed on a hot plate for 30 min at 40°C. The reaction was stopped by mixing 1 mL 10% (w/v) trichloroacetic acid solution followed by incubation for 10 min at 4°C. The reaction mixture was centrifuged for 10 min at 10,000×g. Free polypeptides in the supernatant were measured at λ 280 nm. A standard was prepared using tyrosine. One unit of enzyme activity was defined as the amount of keratinase that caused an increase of 0.01 in absorbance at λ 280 nm12. Measurement of protein content was based on colorimetry method using Bradford reagent. After mixing 500 µL samples and 750 μL Bradford reagent, protein concentration was determined using a spectrophotometer at A595nm.
Effect of metal and inhibitor to keratinolytic activity
Effect of metal ions on keratinolytic activity was examined using 5 and 10 mM chloride salts of Ca2+ (CaCl2), Mg2+ (MgCl2), Ba2+ (BaCl2), Na+ (NaCl), K+ (KCl), Co2+ (CoCl2), and Mn2+ (MnCl2). Solutions of keratinase and ions were incubated at 37°C for 1 h in 50 mM phosphate buffer (pH 7.5)15. Inhibitors i.e. pepstatin A, benzamidine, phenylmethylsulfonyl fluoride (PMSF), ethylenediaminetetraacetic acid (EDTA), dithiothreitol (DTT), 2-mercaptoethanol, soybean trypsin inhibitor, N-tosyl-L-lysine chloromethyl ketone (TLCK), bromoacetic acid, chymostatin, and iodoacetic acid were tested. Inhibitor was mixed with purified keratinase at 1 and 5 mM concentration. All reactions were stopped using 500 µL of 10% (w/v) TCA16. Pellet was separated by spinning at 8000×g for 30 min at 4°C. Reaction with no metal and inhibitor addition was used as a control.
Determination of keratinase molecular weight
Molecular weight was measured using SDS-PAGE. SDS-PAGE was conducted in 10% (w/v) polyacrylamide resolving and 5% (w/v) stacking gel. Sample of 15 µL (protein conc. 2 µg/mL) dissolved in a loading dye were incubated at ±100°C for 5 min for protein denaturation. The samples were inserted into wells and run at 110 V for 90 min. After separation, gel was removed and stained using Coomassie® Brilliant Blue (CBB) R-250 in orbital shaker. Consecutively, the gel was soaked in methanol acetate solution (250 mL distilled water, 200 mL methanol, and 50 mL acetic acid). The molecular weight (kDa) of purified keratinase was determined by measuring the relative migration distance (Rf) between protein standards and the unknown protein analysed using linear regression method. The presence of multiple bands was confirmed through Zymogram analysis using keratin as specific substrate for enzyme detection. Zymogram was performed using a 10% separating gel added with 0.2% (w/v) keratin powder. The gel was immersed in 2.5% Triton® X-100 for 1 h at 37°C and subsequently in a 50 mM Tris-HCl buffer (pH 8) overnight. The gel was stained using 0.05% (w/v) CBB for 2 h followed by destaining until white band was formed visually.
Chicken feather degradation by Azotobacter chroococcum B4
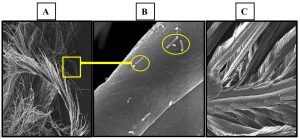
Newly keratinolytic bacterial isolate A. chroococcum B4 ability in degradation of chicken feather keratin and production of keratinase activity was assessed in this study. A. chroococcum B4 was inoculated in liquid media containing chicken feathers as nutrient sole source. It was observed that A. chroococcum B4 successfully colonized and degraded chicken feathers starting from 2-days of incubation until 5 days of observation. Gupta and Singh5, and Fakhfakh et al.16 reported similar result in Bacillus pumilus A1. However, Jeong et al.10 showed that their isolate Stenotrophomonas maltophilia R13 took 6 days to completely degrade chicken. Feather degradation was observed using scanning electron microscopy. Visualization of the chicken feather degradation during cultivation was observed by scanning electron microscopy (SEM) (Fig. 1).
Fig. 1. SEM images (Magnification at 1000×) of feather degradation by A. chrocoocum B4. (A) Barbules and barbs degradation after 5 days; (B) Colonization of Azotobacter chrocoocum B4 on feather surface in 5 d (yellow circles); (C) Whole chicken feather and feather barbs being uninoculated.
Bacterial ability in degrading feather was associated with two enzymes, serine protease and disulphide reductase as showed by Stenotrophomonas sp. D-15,17,18. As far as we know, no report of A. chroococcum was as keratinolytic bacteria. Microbial degradation of keratin releases peptides and amino acids including lysine, alanine, glycine, cysteine, valine, serine, and small quantities of tryptophan and methionine19, which can be used as N and C sources, and ultimately producing ammonium20.
SDS-PAGE and zymogram analysis of B4 keratinase
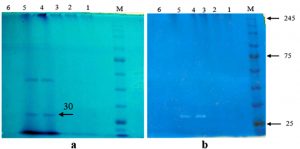
Molecular weight of keratinase was measured using SDS-PAGE method. The keratinase samples were from 23, 24, 25, and 35 fractions of Sephadex G-50 chromatography. SDS-PAGE analysis showed two major bands corresponding to fractions 24 and 25 (Fig. 2a). No protein bands were observed from other samples. SDS-PAGE result was reconfirmed by zymogram using keratin as a substrate dissolved in polyacrylamide gel. A single hydrolysis band of 24 and 25 fractions was formed in the gel (Fig. 2b). It was confirmed that molecular weight of B4 keratinase was 30 kDa.
Fig. 2. Visualization of SDS-PAGE (a) and zymography (b) of keratinase from A. chrocoocum strain B4. Five samples of keratinase purification protocol were analysed: lane 1= ammonium sulfate (70%), lane 2= dialysis, lane 3= fraction (23th), lane 4= fraction (24th), lane 5= fraction (25th), lane 6= fraction (35th). M = Protein Marker, Color Prestained Protein Standard (BioLabs Inc, New England).
In our study, we did not observe any presence of protein bands both in 70% ammonium sulphate precipitated and dialyzed crude keratinase. The phenomenon was due to the presence of ammonium salts which disrupted the keratinase activity of A. chrococcum B4 during partial purification. In the beginning of dialysis, majority of salts were removed but not all of them, as shown from fraction 23th revealing a clear and thin band. As a result, fraction 24 and 25th produced a thick band with the highest keratinase activity. The thick bands indicated a relatively high concentration of protein. The presence of non-keratinase protein bands was assumed to be other protease in our samples. Therefore, a zymogram analysis was performed to selectively estimate the molecular weight of our purified keratinase by using chicken feather as its substrate of hydrolisis.
Previous study showed that molecular weight of keratinases might vary between18-240 kDa21,22. In general, keratinases have molecular weight lower than 40 kDa, such as 22 kDa of B. subtilis23, 28 kDa of B. licheniformis24, 32.8 and 35.5 kDa of B. licheniformis YJ425, 36 kDa of S. maltophilia N426, 35.2 kDa of S. maltophilia DHHJ27. Larger keratinase was found in F. pennavorans with molecular weight of 130 kDa28. The result of this study suggested that B4 keratinase showed to have similar molecular weight to those of Bacillus and Meiothermus29,30.
Effect of C and N sources on keratinase production
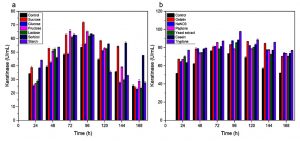
Keratinase of A. chroococcum strain B4 was produced by optimization of submerged fermentation (SmF) as described previously31 using different C and N sources. Sucrose showed to increase keratinase production by 71.7 U/ml in 96 h of cultivation (Fig. 3a). On the other hand, monosaccharide such as glucose resulted to decrease keratinase production. Inhibition of protease biosynthesis by certain monosaccharides is likely correlated with catabolite repression mechanism32. However, not all keratinase production by bacteria was inhibited by glucose. Keratinase production by B. subtilis33-34, B. licheniformis35 and Stenotrophomonas sp. D-117 were inhibited by glucose. Seemingly, glucose might stimulate synthesis keratinases in B. pseudofirmus AL-8936 and Bacillus sp.37.
Various result of keratinase production was demonstrated due to different N sources. The addition of several nitrogen sources by 0.5% (w/v) resulted in an increase in keratinase synthesis by A. chroococcum B4 (Fig. 3b). High keratinase activity of 97.8 U/mL was observed when tryptone was added. Although the only source of C and N in controls were chicken feathers, the additional sources of C and N can help bacterial cell growth before the keratin is hydrolyzed. Having complex structure of keratins, exogenous nutrient source may contribute to keratinolytic bacteria earlier in feather degradation process. Consider that sucrose and tryptone are expensive, other C and N sources from other waste can be examined for increasing keratinase production.
Fig. 3. Keratinase activity during growth of A. chroococcum B4 on chicken feathers supplemented with different C- and N- sources. (a) C- source. (b) N-source. Values are the means ± SD of three replicates.
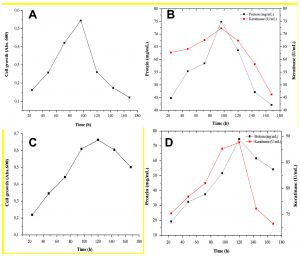
Measurement of bacterial growth as optical density at 600 nm was carried out using spectrophotometer from 24 to 168 h (Fig. 4). The initial growth phase occurs after 24 h of cultivation, for addition both sucrose and tryptone. The logarithmic phase observed between 24 and 96 h of cultivation, while the stationary phase was seemingly not well defined. The bacterial cell growth was correlated with the keratinase activity and soluble protein produced. Data showed that maximum keratinase activity and soluble protein were at 96 h in sucrose (Fig. 4a-b) and 120 h in tryptone medium (Fig. 4c-d).
Fig. 4. Bacterial growth, keratinase activity and soluble protein release growth of A. chroococcum B4 on chicken feathers supplemented with C- and N- sources. Bacterial growth in medium supplemented with (a) 1% (w/v) sucrose (c) with 0.5% (w/v) peptone (b) Soluble protein and enzyme activity in medium supplemented with 1% (w/v) sucrose and (d) 0.5% (w/v) peptone.
The increase of bacteria cell growth depends on the provision of growth substrate. More efficient production of primary metabolites was expected to occur in the logarithmic phase. Bacteria growth rate significantly affect the product during the fermentation process; the faster the growth rate, the less the fermentation process time, which then also leads to less production cost38.
Effect of metal ions and inhibitors on keratinase B4
Metal ions have a significant role in the microbial metabolism, affecting the activity of many enzymes. The increase of keratinase B4 activity was observed when Ca2+, Mg2+, Mn2+, Na+, and K+ were added, with relative activities of 117%, 166%, 111%, 113%, and 112%, respectively (Table 1).
Table (1):
Effect of metal ions and inhibitors on keratinolytic activity of Azotobacter chroococcum B4.
| Metal ions and Inhibitors | Relative activity (%) | |
|---|---|---|
| 5 Mm | 10 Mm | |
| Control | 100 | 100 |
| Ca2+ | 115 ± 0.9 | 117 ± 0.7 |
| Mg2+ | 112 ± 0.6 | 166 ± 1.0 |
| Ba2+ | 85 ± 1.2 | 110 ± 1.0 |
| Mn2+ | 109 ± 0.7 | 111 ± 0.8 |
| Co2+ | 51 ± 0.7 | 40 ± 0.8 |
| Na+ | 103 ± 0.9 | 113 ± 0.6 |
| K+ | 102 ± 1.0 | 112 ± 1.2 |
| 1Mm | 5 Mm | |
| Pepstatin A | 51 ± 1.0 | 33 ± 1.5 |
| PMSF | 65 ± 0.9 | 31 ± 0.9 |
| DTT | 42 ± 0.7 | 22 ± 0.7 |
| EDTA | 52 ± 0.7 | 110 ± 0.7 |
| N-tosyl-L-lysine | 72 ± 1.5 | 64 ± 0.9 |
| Bromoacetic acid | 87 ± 1.5 | 35 ± 0.9 |
| Chymostatin | 15 ± 0.9 | 14 ± 0.7 |
| Iodoacetic acid | 51 ± 0.7 | 33 ± 1.0 |
| Benzamidine | 57 ± 0.7 | 12 ± 0.9 |
| Soybeane trypsin inhibitor | 75 ± 0.9 | 66 ± 1.3 |
| 2-mercaptoethanol | 75 ± 1.3 | 73 ± 0.9 |
However, Co2+ decreased the relative activity of keratinase B4 to 40%. In present report, Mg2+ can also be considered as significant factor in order to increase B4 keratinase activity, as also shown by keratinase of Kocuria rosea39, B. subtilis23 and Paracoccus sp. WJ-9840. However, metal ions may have different effect in different strains. For example, Ca2+ increased S. brevicaulis keratinase production41, but decreased keratinase of B. licheniformis RG142.
It was known that various inhibitors dicreased relative activity of keratinases. Inhibitor assays aims to determine keratinase type, considering that protease groups are very sensitive to presence of specific inhibitors43. It has been reported previously that microbial keratinase was mostly related to serine proteases and metalloproteases20,44, except that of yeast group that produced aspartic proteases45. The keratinase B4 was inhibited by PMSF, benzamidine, chymostatin, which are well-known serine protease inhibitors. This indicated that B4 keratinase B4 belonged to serine protease family. DTT, iodoacetic acid, and 2-mercapthoethanol also inhibited the enzyme, suggesting that cysteine residues were relevant for the activity of B4 keratinase as well.
Reported strain A. chroococcum B4 was able to degrade whole chicken feathers after 5 days of cultivation as evidenced by scanning electron microscopy. The analysis conducted by SDS-PAGE and zymogram showed that A. chroococcum B4 keratinase had a molecular weight of about 30 kDa. The keratinase activity increased after the addition of sucrose (1% w/v) as carbon source and tryptone (0.5% w/v) as nitrogen source during cultivation on feathers, reaching 97.8 U/mL. The enzyme activity enhanced by addition of Ca2+, Mg2+, Mn2+, Na+, and K+, but it was inhibited by PMSF and other inhibitors indicating its belongingness to serine-protease group.
ACKNOWLEDGMENTS
We would like thank to Program of Master’s Degree toward Doctoral Degree for Excellent Graduates (PMDSU) Scholarship from Ministry of Research, Technology and Higher Education, Indonesia for providing graduate scholarship.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
DS: concept of the main idea, developing the supporting theory, performing the experiment, writing of manuscript. EM: verifying the analytical methods. JM.: developing the theoretical formalism, performing the experiment. AZM: providing the laboratory necessity, supervising the findings of this project.
FUNDING
This study was supported by DRPM, Direktorat Jenderal Penguatan Risetdan Pengembangan, Kemenristekdikti RI, Grant# 003/SP2H/LT/DRPM/IV/2017 tanggal 20 April 2017.
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
AVAILABILITY OF DATA
All datasets generated and analyses are provided in this manuscript. The raw data may be requested upon acceptance by the authors.
- Martinez-Hernandez AL, Velasco-Santos C. Nova Science Publishers pp. 2011.
- Manivasagan P, Venkatesan J, Sivakumar K, Kim SK. Production, characterization and antioxidant potential of protease from Streptomyces sp. MAB18 using poultry wastes. Biomed Res. Int., 2013.
Crossref - Sangali S, Brandelli A. Feather keratin hydrolysis by a Vibrio sp. strain kr2. J. Appl. Microbiol., 2000.
Crossref - Syed DG, Lee JC, Li WJ, Kim CJ, Agasar D. Production, characterization and application of keratinase from Streptomyces gulbargensis. Bioresource Technol. 2009.
Crossref - Gupta S, Singh R. Hydrolyzing Proficiency of Keratinases in Feather Degradation. Indian J Microbiol. 2014.
Crossref - Fakhfakh-Zouari N, Haddar A, Hmidet N, Frikha F, Nasri M. Application of statistical experimental design for optimization of keratinases production by Bacillus pumilus A1 grown on chicken feather and some biochemical properties. Process Biochem, 2010.
Crossref - Tork SE, Shahein YE, El-Hakim AE, Abdel-Aty AM, Aly MM. Production and characterization of thermostable metallo-keratinase from newly isolated Bacillus subtilis NRC 3. Int J Biol Macromol. 2013.
- Mamangkey J, Suryanto D, Munir E, Mustopa AZ. Isolation, Molecular Identification and Verification of Gene Encoding Bacterial Keratinase from Crocodile (Crocodylus porosus) Feces. in IOP Conference Series: Earth and Environmental Science, 2019.
Crossref - Bhange K, Chaturvedi V, Bhatt R. Feather degradation potential of Stenotrophomonas maltophilia KB13 and feather protein hydrolysate (FPH) mediated reduction of hexavalent chromium. 3 Biotech. 2016.
Crossref - Jeong JH et al. Production of keratinolytic enzyme by a newly isolated feather-degrading Stenotrophomonas maltophilia that produces plant growth-promoting activity. Process Biochem. 2010.
Crossref - Suryanto D, Walida H, Nasution SK, Munir E. Partial characterization of keratinolytic activity of local novel bacteria isolated from feather waste. J Pure Appl Microbiol. 2017.
Crossref - Lv L, Sim M, Li Y, Min J, Feng W, Guan W. Production, characterization and application of a keratinase from Chryseobacterium L99 sp. Nov. Process Biochem. 2010.
Crossref - Gurav RG, Tang J, Jadhav JP. Sulfitolytic and keratinolytic potential of Chryseobacterium sp. RBT revealed hydrolysis of melanin containing feathers. 3 Biotech. 2016;6.
Crossref - Mamangkey J, Suryanto D, Munir E, Mustopa AZ. Keratinolytic fungi isolated from Asam Kumbang Crocodile Breeding Farm, Medan, North Sumatra. in IOP Conference Series: Earth and Environmental Science, 2019.
Crossref - Tiwary E, Gupta R. Medium optimization for a novel 58 kDa dimeric keratinase from Bacillus licheniformis ER-15: biochemical characterization and application in feather degradation and dehairing of hides. Bioresour Technol. 2010.
Crossref - Fakhfakh N et al. Total solubilisation of the chicken feathers by fermentation with a keratinolytic bacterium, Bacillus pumilus A1, and the production of protein hydrolysate with high antioxidative activity. Process Biochem. 2011.
Crossref - Yamamura S, Morita Y, Hasan Q, Yokoyama K, Tamiya E. Keratin degradation: A cooperative action of two enzymes from Stenotrophomonas sp. Biochem. Biophys. Res Commun. 2002.
Crossref - Gupta S, Singh SP, Singh R. Synergistic effect of reductase and keratinase for facile synthesis of protein-coated gold nanoparticles. J Microbiol Biotechnol. 2015.
Crossref - Grazziotin A, Pimentel FA, De Jong EV, Brandelli A. Nutritional improvement of feather protein by treatment with microbial keratinase. Anim Feed Sci Technol. 2006.
Crossref - Kumar D, Savitri Thakur N, Verma R, Bhalla TC. Microbial proteases and application as laundry detergent additive. Research Journal of Microbiology. 2008.
Crossref - Brandelli A, Daroit DJ, Riffel A. Biochemical features of microbial keratinases and their production and applications. Appl Microbiol Biotechnol. 2010.
Crossref - Verma A et al. Microbial keratinases: industrial enzymes with waste management potential. Crit Rev Biotechnol. 2017.
- Gupta S, Nigam A, Singh R. Purification and characterization of a Bacillus subtilis keratinase and its prospective application in feed industry. Acta Biol. Szeged. 2015.
- Okoroma EA, Garelick H, Abiola OO, Purchase D. Identification and characterisation of a Bacillus licheniformis strain with profound keratinase activity for degradation of melanised feather. Int Biodeterior Biodegrad. 2012.
Crossref - Yin LJ, Lin HH, Xiao ZR. Purification and characterization of a cellulase from Bacillus subtilis YJ1. J Mar Sci Technol. 2010.
- Jankiewicz U, Frk M. Identification and properties of a keratinase from Stenotrophomonas maltophilia N4 with potential application in biotechnology. Ecol Quest. 2017.
Crossref - Cao ZJ et al. Characterization of a novel Stenotrophomonas isolate with high keratinase activity and purification of the enzyme. J Ind Microbiol Biotechnol. 2009.
Crossref - Friedricht AB, Antranikian G. Keratin degradation by feividobacterium pennavomns, a novel thermophilic anaerobic species of the order thermotogales. Appl Environ Microbiol. 1996.
Crossref - Kazi YF, Kumar P, Soomro IH. Characterization of the keratinolytic activity of indigenous Bacillus subtilis keratinase. J Chem Pharm Res. 2015. Available online www.jocpr.com.
- Wu WL et al. The discovery of novel heat-stable keratinases from Meiothermus taiwanensis WR-220 and other extremophiles. Sci Rep. 2017.
Crossref - Amit V, Hukum S, Mohammad SA, Mohammad W A, Sanjeev A. Production of alkaline protease from a haloalkaliphilic soil thermoactinomycete and its application in feather fibril disintegration. African J Microbiol Res. 2014.
Crossref - Gupta R, Beg Q, Lorenz P. Bacterial alkaline proteases: Molecular approaches and industrial applications. Appl Microbiol Biotechnol. 2002.
- Imtiaz A, Rehman A. Bacillus subtilis BML5 Isolated from Soil Contaminated with Poultry Waste has Keratinolytic Activity. Pak J Zool. 2018.
Crossref - Kole MM, Draper I, Gerson DF. Production of protease by bacillus subtilis using simultaneous control of glucose and ammonium concentrations. J Chem Technol Biotechnol. 1988.
- Abdel-Fattah AM, El-Gamal MS, Ismail, SA, Emran MA and Hashem AM. Biodegradation of feather waste by keratinase produced from newly isolated Bacillus licheniformis ALW1. J Genet Eng Biotechnol. 2018.
Crossref - Gessesse A, Hatti-Kaul R, Gashe BA, Mattiasson B. Novel alkaline proteases from alkaliphilic bacteria grown on chicken feather. Enzyme Microb Technol. 2003.
Crossref - Prakasham RS, Rao SCh, Sarma PN. Green gram husk—an inexpensive substrate for alkaline protease production by Bacillus sp. in solid-state fermentation. Bioresour Technol. 2006.
Crossref - Pandey A. Solid-state fermentation. Biochem Eng J. 2003.
- Bernal C, Diaz I, Coello N. Response surface methodology for the optimization of keratinase production in culture medium containing feathers produced by Kocuria rosea. Can J Microbiol. 2006.
Crossref - Lee YJ, Kim JH, Kim HK, Lee JS. Production and characterization of keratinase from Paracoccus sp. WJ-98. Biotechnol. Bioprocess Eng. 2004.
Crossref - Anbu P, Gopinath SCB, Hilda A, Lakshmipriya T, Annadurai G. Optimization of extracellular keratinase production by poultry farm isolate Scopulariopsis brevicaulis. Bioresource Technol. 2007.
Crossref - Ramnani P, Gupta R. Optimization of medium composition for keratinase production on feather by Bacillus licheniformis RG1 using statistical methods involving response surface methodology. Biotechnol Appl Biochem. 2004.
- Rao MB, Tanksale AM, Ghatge MS, Deshpande VV. Molecular and biotechnological aspects of microbial proteases. Microbiol Mol Biol Rev. 1998.
Crossref - Gupta R, Ramnani P. Microbial keratinases and their prospective applications: An overview. Appl Microbiol Biotechnol. 2006.
Crossref - Monod M et al. Secreted proteases from pathogenic fungi. Int J Med Microbiol. 2002.
© The Author(s) 2020. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.