ISSN: 0973-7510
E-ISSN: 2581-690X
The present study reports the isolation of Clostridium perfringens from samples comprising of freshwater fish and fish products from Kolkata city of India, and determining their genotypes and antibiogram profile. A total of 102 samples consisting of intestinal and gill samples from fresh water fish (n=69) and fish products including fish pickles, fish curry and fried fish (n=33) were collected randomly from retail shops and restaurants. On cultural isolation and biochemical characterization, 24 (23.52%) samples [17 (24.63%) from fresh water fish (n=69) and 07 (21.21%) from fish products (n=33)] were presumptively identified as C. perfringens positive. The genotyping of the recovered C. perfringens isolates was done by amplifying species specific 16S rRNA gene and four major lethal toxin genes viz., alpha toxin gene (cpa), beta toxin gene (cpb), epsilon toxin gene (etx) and iota toxin gene (itx). Apart from these, enterotoxin gene (cpe) and beta2 toxin gene (cpb2) were also targeted. In PCR assays, all the 24 (100%) isolates [17 (70.83%) from fresh water fish and 07 (29.17%) from fish products] were found to harbor species specific 16S rRNA and cpa toxin gene, however, 17 (70.83%) cpa positive isolates [12 (70.58%) from fresh water fish and 05 (71.42%) from fish products] were also found to harbor additional cpb2 toxin gene, while none of the isolates were found to be positive for cpb, etx, itx and cpe genes. Based on these results, all the isolates were confirmed as C. perfringens type A (containing only alpha toxin gene). On antibiotic sensitivity testing, 76.47% of the isolates were found to be multidrug resistant with ciprofloxacin and amoxicillin/clavulanic acid as being the most sensitive drugs. Report regarding isolation and molecular characterization of C. perfringens from fish and fish products especially from Kolkata does not exist as well as from other regions of India are very scanty. The present findings suggest that fish may be considered as a potential source of C. perfringens type A infection to human populations through food chain and the high antibiotic resistance observed may pose serious public health concerns. Further detailed molecular epidemiological and antibiogram studies are suggested for designing and adapting appropriate prevention and control strategies for countering this important pathogen and its food-borne zoonotic concerns.
C. perfringens, isolation, genotyping, 16S rRNA, lethal toxin genes, PCR, antibiotic sensitivity, fish, fish products
India is the second largest producer of fresh water fish in the world. Fisheries sector contributed about 0.9% to the National Gross Domestic Product (GDP) and 5.17% to the agricultural GDP (2014-15). West Bengal ranked second in the total production of fish after Andhra Pradesh; it has total fish production as increases from 1,447.26 x 103 tonnes (2007-08) to 1678.33 x 103 tonnes (2014-15).1 This aberrant growth in fish industry require careful monitoring since there is risk of bacterial contamination in fish, especially in the post harvesting storage period, a critical stage prior to human consumption. Whenever bacterially contaminated raw or partially cooked food is consumed by susceptible population, then there is a chance of detrimental impact on public health.2,3
Clostridium perfringens is one of the most important food-borne pathogen of humans and animals causing both histotoxic diseases and intestinal infections.4-7 It is Gram-positive, anaerobic, straight rod, spore former, found in the soil, dust, sewage, marine sediments and in the gastro-intestinal tract of humans and animals.8-10 Based on the production of four major lethal toxins i.e. alpha (a) beta (b), epsilon (e) and iota (i), C. perfringens is divided into five major toxino types/bio types (A-E)6,10,11. Apart from these, it also produces enterotoxin (CPE) and beta 2 toxin (CPB2).12 Symptoms of C. perfringens type A food poisoning include abdominal cramping, nausea and diarrhea, which usually begin 6–24 h after ingestion of contaminated food and then persist for 12–24 h.7,9 C. perfringens associated symptoms are caused by an enterotoxin which is produced during sporulation of the organism in the small intestine following ingestion of large number of vegetative cells of enterotoxin positive C. perfringens.9,13
Isolates originating from humans with gastrointestinal diseases carry most commonly CPE and sometimes newly discovered CPB2 toxin.12,14 Enterotoxigenic C. perfringens type A is also associated with antibiotic associated diarrhoea (AAD) and sporadic diarrhoea (SD) cases.15,16 The emerging problem of antimicrobial resistance between pathogenic and commensal bacteria is becoming more intense by intervention of environmental resistance phenomenon of bugs.17,18 Antibiotics represent one of the most successful forms of therapeutic regimens in medicine. But the overuse and misuse of antibiotics, mostly in developing countries, lead to growing number of antibiotic-resistant pathogens.19 Antibiotic resistance, which is associated with increased morbidity and mortality rates as well as increased treatment costs, is considered to be one of the major global public health problems and its magnitude recently prompted a number of national and international bodies to take actions to protect the public health20. Problem of antimicrobial resistance in pathogenic and commensal bacteria is increasing by the adoption of mobile genetic elements.21 Globally, there are reports of earlier studies on epidemiological characterisation of C. perfringens from fish samples,7, 22-31 but in Indian scenario there are limited studies carried out earlier on given aspects2,13,32 and there are no previous reports from the study area, therefore, it is necessary to carry out epidemiological studies in such prone areas to determine the types of C. perfringens prevalent in fish and fish products. Considering these facts, the present study was designed to isolate Clostridium perfringens from freshwater fish and fish products from Kolkata region of India, characterize these at molecular levels by determining their genotypes and study antibiotic sensitivity patterns.
Collection of samples
One hundred and two samples consisting of intestinal and gill samples from fresh water fish (n=69) and fish products including fish pickles, fish curry and fried fish (n=33) were collected from retail shops and restaurants from Kolkata city of India. All the collected samples were immediately transported to the laboratory in ice pack container and processed within 24 hours for isolation and molecular characterisation of C. perfringens using standard bacteriological and molecular procedures. The intestinal and gill samples from the fresh water fish were hygienically collected and processed for the bacteriological analysis. The fish products were aseptically collected in sterile container for the analysis.
Isolation of C. perfringens
All the samples (Intestine, gills and fish products) were inoculated aseptically in Robertson’s cooked meat (RCM) medium supplemented with glucose, hemin and vitamin K broth (HiMedia, Mumbai, India) and heated at 75°C for 20 minute for removal of vegetative cells of other competing microorganisms, followed by incubation at 37°C for 24-48 hours under anaerobic condition in McIntosh anaerobic jar using gas pack (HiMedia, Mumbai, India). Enriched 100 µl inoculums were taken in sterile Petridish, followed by pouring of sterile molten sulfite polymyxin sulfadiazine (SPS) agar over the inoculum with proper mixing by rotating the plate in clockwise and anticlockwise direction. Plates were incubated at 37°C for 24 h under anaerobic condition.33
Biochemical and phenotypical analysis of C. perfringens
Samples revealing characteristic black coloured colonies on SPS agar plates were subjected to biochemical tests for identification of C. perfringens.34 About 3-5 colonies from the culture plate for each sample were considered for Gram’s staining, lactose fermentation, nitrate reduction, gelatin liquefaction, lecithinase and indole tests.
Reference strains
Reference strains of C. perfringens were procured from Biological Standardization Division, ICAR-IVRI, Izatnagar, India.
Molecular characterization / genotyping of C. perfringens by PCR
All the presumptively identified C. perfringens isolates recovered from different sources were subjected to polymerase chain reaction (PCR) assays for the detection of species specific 16S rRNA35 and virulent toxin genes viz., cpa, cpb, etx, itx36, cpb2 and cpe genes37. The oligonucleotides used in present study were synthesized through M/S Xcelris Labs Limited, Ahmadabad, Gujarat (India). The details of primers used in present study are given in Table 1.38-40
Table (1):
Details of the primers for testing of samples for Clostridium perfringens by PCR.
| Target genes | Primer sequence 5’- 3’ | Product size (bp) | References |
|---|---|---|---|
| 16S rRNA | F-AAAGATGGCATCATCATTCAAC R-TACCGTCATTATCTTCCCCAAA | 279 | Yoo et al.38 |
| cpa | F-GTTGATAGCGCAGGACATGTTAAG R-CATGTAGTCATCTGTTCCAGCATC |
402 | |
| cpb | F-ACTATACAGACAGATCATTCAACC R-TTAGGAGCAGTTAGAACTACAGAC | 236 | |
| εtx | F-ACTGCAACTACTACTCATACTGTG R-CTGGTGCCTTAATAGAAAGACTCC |
541 | |
| ιtx | F-GCGATGAAAAGCCTACACCACTAC R-GGTATATCCTCCACGCATATAGTC |
317 | |
| cpb2 | F-AGATTTTAAATATGATCCTAACC R-CAATACCCTTCACCAAATACTC | 567 | Gibert et al.39 |
| cpe | F-GGAGATGGTTGGATATTAGG R-GGACCAGCAGTTGTAGATA |
233 | Czeczulin et al.40 |
The snap chill method41 was used to extract DNA from the isolates. About 200 µl of overnight culture of C. perfringens was taken in the microcentrifuge tube and the cell suspension was centrifuged for 10 min at 14,000 × g. The pellet was suspended in 100 µl of nuclease free water (NFW) by vortexing. The microcentrifuge tube was placed in hot water bath for 15 min at 100°C and immediately chilled in ice. An aliquot of 5 µl of the supernatant was used as the template DNA in the PCR assays.
The optimized PCR protocol for all the target genes with 25 µl reaction mixture volume included 1 µl (10 pmol/µl) each of forward and reverse primer sequences, 0.5 µl of 10 mM deoxyribonucleotide triphosphates (dNTPs), 2.5 µl of 10 × PCR buffer [(100 mM Tris-HCl (pH 8.8 at 25°C), 500 mM KCl, 0.8% (v/v) Nonidet P40)], 2.0 µl MgCl2 (25 mM), 0.20 µl of 5 U/µl of Taq DNA polymerase (Thermo scientific, USA), 5.0 µl DNA template and remaining NFW to make the final volume. The PCR reaction mixture was set for amplification in a thermocycler (Biometra Personal Cycler, Goettingen, Germany) with specific condition for each PCR assay. For 16S rRNA PCR assay, the initial denaturation condition was 94°C for 5 min. then amplified for 35 cycles (1.0 min at 94°C, 1.3 min. at 53°C, 1.3 min. at 72°C for denaturation, annealing and extension phases, respectively), followed by an additional period of extension for 10 min. at 72°C. Regarding multiplex PCR for cpa, cpb, etx and itx genes, there was initial denaturation at 95°C for 5 min. then amplified for 30 cycles (1.0 min. at 94°C, 1.0 min. at 55°C, 1.5 min. at 72°C for denaturation, annealing and extension phases, respectively), followed by an additional period of extension for 10 min. at 72°C. Similarly for cpb2 and cpe genes, there was initial denaturation at 94°C for 4 min. then amplified for 35 cycles (1.0 min. at 94°C, 1.2 min. at 55°C, 1.2 min. at 72°C for denaturation, annealing and extension phases, respectively), followed by an additional period of extension for 10 min. at 72°C.
Amplified PCR products (10 µl) were separated by electrophoresis for 45 to 60 min. at 80 V in a 1.2% (w/v) agarose gel with ethidium bromide (0.5 µg/ml) and a 100 bp DNA ladder (Thermo scientific, USA) was included in each agarose run. The resolution of amplified fragments in the gel was visualized by a UV trans-illuminator and digitally recorded by gel documentation system (UVP Gel Sequencing Software). Materials contaminated with ethidium bromide were disposed as per the local guidelines.
Antimicrobial sensitivity test
Antimicrobial sensitivity test was performed by using disc diffusion method as per the guidelines provided by Clinical Laboratory Standards Institute (CLSI).42 The antimicrobials included in the present study were selected based on the information gathered from local professionals (veterinarians, paediatrician) and available literature. The antimicrobials included were amikacin (AK, 30 µg), co-trimoxazole (COT, 25 µg), ceftazidime (CAZ, 30 µg), norfloxacin (NX, 10 µg), ceftriaxone (CTR, 30 µg), ciprofloxacin (CIP, 5 µg), tetracycline (TE, 30 µg) and amoxicillin/ clavulanic acid (AMC, 30µg) (HiMedia Laboratories Ltd, Mumbai, India).
Isolation and identification of C. perfringens by cultural and biochemical methods
On cultural and biochemical analyses, a total of 24 isolates [17 from fresh water fish (n=69) and 07 from fish products (n=33)] were found to be positive for C. perfringens.
Genotyping by PCR detection of virulence toxin genes
All the biochemically positive isolates were screened by PCR assays for detection of species specific 16S rRNA and virulence toxin genes viz., cpa, cpb2, cpe, cpb, etx and itx. All the 24 (100%) presumptively confirmed isolates of
C. perfringens [17 (70.83%) from fresh water fish and 07 (29.16%) from fish products] was also shown amplification of species specific 16S rRNA of product size 279 bp (Table 2, Fig 1) and a toxin gene of product size 402 bp (Table 2, Fig 2). However, 17 (70.83%) cpa positive isolates [12 (70.58%) from fresh water fish and 05 (71.42%) from fish products] were also positive for additional beta 2 toxin gene (cpb2) of 567 bp length (Table 2, Fig 3). The other virulence gene specific primers of cpb, etx, itx and cpe genes did not amplify any of the isolates. The virulence gene detection by PCR assay used in this study revealed that all the isolates were belonged to C. perfringens type A.
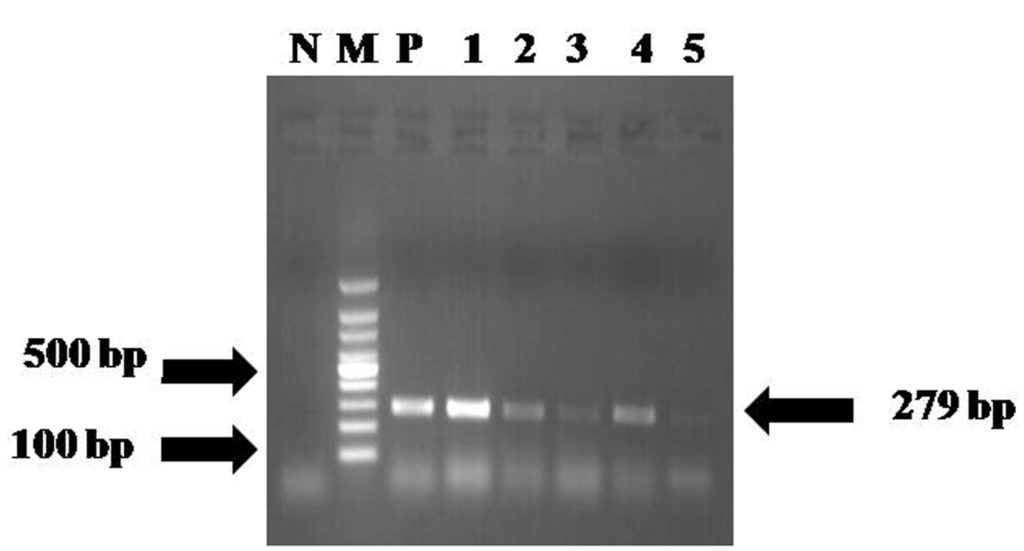
Fig. 1. PCR assays for detection of 16S rRNA of
C. perfringens
Lane N: Negative control; Lane M: 100 bp DNA ladder; Lane P: Reference strain of C. perfringens; Lane 1-5: C. perfringens isolates
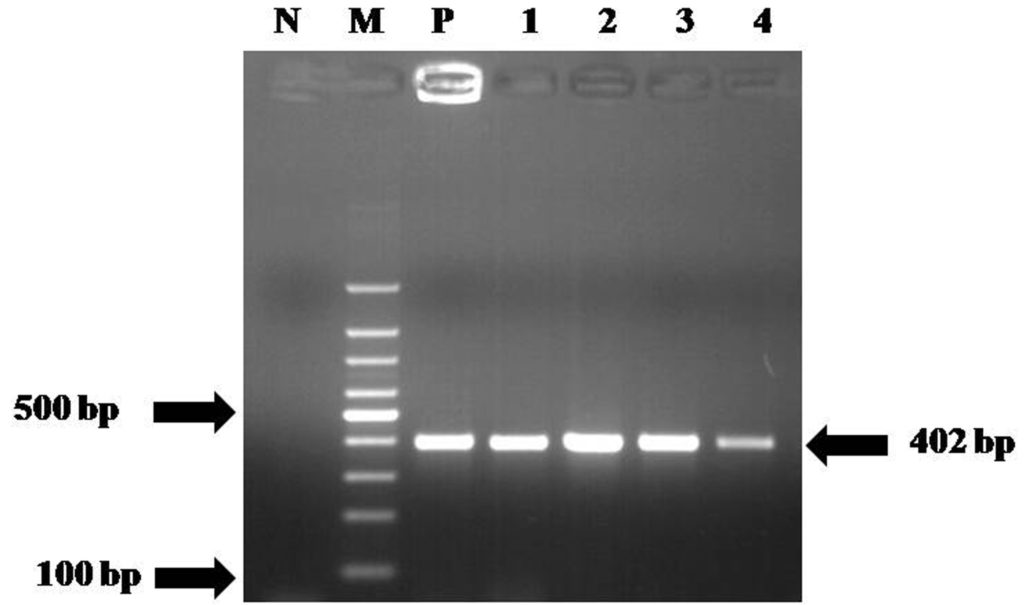 Fig. 2. PCR assays for detection of cpa gene of C. perfringens
Fig. 2. PCR assays for detection of cpa gene of C. perfringensLane N: Negative control; Lane M: 100 bp DNA Ladder; Lane P: Reference strain of C. perfringens; Lane 1-4: C. perfringens isolates
 Fig. 3. PCR assays for detection of cpb2 gene of C. perfringens
Fig. 3. PCR assays for detection of cpb2 gene of C. perfringensLane N: Negative control; Lane M: 100 bp DNA Ladder; Lane P: Reference strain of C. perfringens; Lane 1-3: C. perfringens isolates
Table (2):
Samples found positive for C. perfringens and their virulence genes by PCR.
| Sample source | Sample types | No. of C. perfringens positive samples (%) | No. of sample positive for virulence gene by PCR | |
|---|---|---|---|---|
| cpa | cpb2 | |||
| Fresh water fish (n=69) | Intestinal and gills | 17 (24.63) | 17 | 12 |
| Fish products (n=33) | Whole part | 07 (21.21) | 07 | 05 |
| 24 (100%) | 17 (70.83%) | |||
Antibiotic sensitivity testing
A total of 17 C. perfringens isolates (both cpa and cpb2 positive) were tested for antimicrobial sensitivity assays. Out of 17 isolates, 13 (76.47%) were found to be resistant to ³ 03 tested groups of antibiotics, indicating multidrug resistance phenomenon. Isolates showed highest resistance against co-trimoxazole (88%), followed by ceftriaxone (87%), cefazidime (53%), tetracycline (44%), norfloxacin (21%) and amikacin (11%), while all the isolates were found to be sensitive to ciprofloxacin and amoxicillin/clavulinic acid.
C. perfringens is considered as one of the most important food-borne pathogen and is ubiquitous in nature6-9, however, in Indian scenario there are limited studies occurred earlier on epidemiological and molecular aspects of C. perfringens in fresh water fish and fish products. In the present study, a total of 24 samples (23.52%) [17 (24.63%) from fresh water fish (n=69) and 07 (21.21%) from fish products (n=33)], were found to be positive for C. perfringens. It has been estimated that 17/24 isolates of C. perfringens (70.83%) were derived from the intestinal and gill samples of fresh water fish and 07/24 (29.17%) from fish products.
On PCR detection of species specific 16S rRNA and virulence toxin genes of the C. perfringens isolates it was found that all the isolates (24) were shown species specific amplicon of 16S rRNA and positive for a toxin gene (cpa). Apart from these, 17 cpa positive isolate (70.83%) [12 (70.58%) from fresh water fish and 05 (71.42%) from fish products], were also positive for beta2 toxin gene (cpb2). Previous studies have reported occurrence of C. perfringens from fish and sea food samples with a varying isolation rates of 1% to 84% from different countries2,7,13,22-32. Cai et al.27 reported that out of 75 isolates of C. perfringens from fresh water fish, 13 (17.3%) isolates were found to be positive for only a toxin gene, 58 (77.3%) isolates for a & b toxin genes positive (C. perfrigens type C) and 04 (5.3%) isolates for a, b and e toxin genes positive (C. perfringens type B), while 47 (62.70%) isolates have additional beta 2 toxin gene, however, none of the isolates were found to be positive for cpe and other virulence toxin genes, since, the presence of cpe gene in C. perfringens is very uncommon, and only <5% of global C. perfringens type A isolates were found to be cpe gene positive7,9. In concurrence to present results, the cpe– negative C. perfringens type A from the fish sample and clinical cases of gastroenteritis was reported previously2,13,31,32,43. In another study, out of 34 isolates of C. perfringens from fish samples, 31 were found to be positive for C. perfringens type A (only cpa gene positive) and 3 isolates were cpa as well as cpe genes positive, but none of the C. perfringens type A isolates were found to be positive for cpb2 gene2. In contradiction, present study reported the occurrence of cpb2 positive isolates in 70.83% of C. perfringens type A isolates. These findings highlights the high prevalence of cpb2 associated C. perfringens type A which is mainly associated with antibiotic associated diarrhoea (AAD) and sporadic diarrhoea (SD) cases12.
The antibiogram studies performed against isolates recovered from freshwater fish and fish products unveils an alarming public health concern due to higher resistance in recovered isolates. Highest resistance was observed against co-trimoxazole (88%), followed by ceftriaxone (87%), ceftazidime (53%), tetracycline (44%), norfloxacin (21%) and amikacin (11%), while all the isolates were found to be sensitive to ciprofloxacin and amoxicillin/clavulinic acid. These finding are found to be concurrent with earlier studies reporting a high emerging resistance toward various tested antibiotics44-52. These observations are major point of concern from public health perspectives, since most of the antibiotics drugs used in the present study are employed as a first line of treatment in diarrhoeal cases.
The present study suggested that PCR is a reliable molecular technique and useful tool for the detection of virulence genes typing of C. perfringens isolates recovered from fresh water fish and fish products. The presence of species specific 16S rRNA and cpa gene in all the isolates of C. perfringens suggest that C. perfringens type A (only have a toxin gene) is the most predominant type in fresh water fish and fish products in this study area. Primer specific to species specific 16S rRNA and cpa gene in PCR assays, used in present study, could be useful for the rapid identification of C. perfringens and provided a clues to the importance and improvement of the present method for the surveillance of C. perfringens from fresh water fish and fish products. On antibiogram profile, except for ciprofloxacin and amoxicillin/clavulinic acid, a high frequency of multiple drug resistance pattern was observed among the C. perfringens type A isolates. These emerging resistance trends are major point of concern from public health perspectives.
ACKNOWLEDGMENTS
The authors thank The Director, Indian Veterinary Research Institute, Izatnagar for providing necessary facilities for undertaking the research work.
- Department of Animal Husbandry, Dairying and Fisheries; Ministry of Agriculture and Farmers Welfare; Govt. of India Annual report: 2015-16. 65-141.
- Das, A. Jain, A. Clostridium perfringens type A from fresh water fishes. Int. J. Adv. Biotech. Res., 2012a: 3(3). 680-687.
- Dhama, K., Rajagunalan, S., Chakraborty, S., Verma, A.K., Kumar, A., Tiwari, R., Kapoor, S. Food-borne pathogens of animal origin-diagnosis, prevention, control and their zoonotic significance: a review. Pak. J. Biol. Sci., 2013: 16 (20): 1076-1085.
- McClane, B.A., Uzal, F.A., Miyakawa, M.F., Lyerly, D. Wilkins, T.D. The enterotoxic clostridia, 2006a: p 688 –752. In Dworkin, M., Falkow, S., Rosenburg, E., Schleifer, H. Stackebrandt, E. (ed), The prokaryotes, 3rd ed. Springer Press, New York, NY.
- McClane, B.A., Lyerly, D.M. Wilkins, T.D. Enterotoxic clostridia: Clostridium perfringens type A and Clostridium difficile, 2006b: p 703–714. In Fischetti, V.A., Novick, R.P., Ferretti, J.J., Portnoy, D.A. Rood, J. (ed), Gram-positive pathogens, 3rd ed. ASM Press, Washington, DC.
- Li, J., Adams, V., Bannam, T.L., Miyamoto, K., Garcia, J.P., Uzal, F.A., Rood, J.I. McClane, B.A. Toxin plasmids of Clostridium perfringens. Microbiol. Molec. Bio. Rev., 2013: 77(2): 208-233.
- Nashwa A.E., Ahmed M.A., Basma S., El. Haririr, M., Walaa S.O. Rapid detection of Clostridium perfringens in seafood. Adv. Environ. Biol., 2016: 10(4): 174-181.
- Wang, G., Zhang, P., Paredes-Sabja, D., Green, C., Setlow, P., Sarker, M.R. Li, Y.Q. Analysis of the germination of individual Clostridium perfringens spores and its heterogeneity. J. Appl. Microbiol., 2011: 111: 1212-1223.
- Uzal, F.A., Freedman, J.C., Shrestha, A., Theoret, J.R., Garcia, J., Awad, M.M., Adams, V., Moore, R.J.,, Rood, J.I. McClane, B.A. Towards an understanding of the role of Clostridium perfringens toxins in human and animal disease. Fut. Microbiolo., 2014: 9: 361-377.
- Silva, R.O.S., Lobato, F.C.F. Clostridium perfringens: a review of enteric diseases in dogs, cats and wild animals, London. Anaerobe. 2015: 33 : p. 14-17.
- Heikinheimo, A., Lindström, M., Liu, D. Korkeala, H. Clostridium. In: Liu, D. (Ed.), Molecular Detection of Foodborne Pathogens. CRC Press, Taylor and Francis Group, Boca Raton, FL, 2009: USA, pp. 145-155.
- Fisher, D.J., Miyamoto, K., Harrison, B., Akimoto, S., Sarker, M.R., McClane, B.A. Association of beta2 toxin production with Clostridium perfringens type A human gastrointestinal disease isolates carrying a plasmid enterotoxin gene. Mol. Microbiolo., 2005: 56(3): 747-762.
- Gurmu, E.B., Hazarika, R.A., Borah, P. Barua, A.G. Presence of enterotoxigenic Clostridium perfringens in foods of animal origin, Guwahati, India. J. Environ. Occup. Sci. 2013: 2(1): 45-50.
- Harrison, B., Raju, D., Garmory, H.S., Brett, M.M., Titball, R.W. Sarker, M.R. Molecular Characterization of Clostridium perfringens Isolates from Humans with Sporadic Diarrhea: Evidence for Transcriptional Regulation of the Beta2- Toxin-Encoding Gene. Appl. Environ. Microbiol., 2005: 71: 8362-8370.
- Banaszkiewicz, A., K¹dzielska, J., Gawroñska, A., Pituch, H., Obuch-Woszczatyñski, P., Albrecht, P., M³ynarczyk G., Radzikowski, A. Enterotoxigenic Clostridium perfringens infection and pediatric patients with inflammatory bowel disease. J. Crohn’s Colitis. 2014: 8: 276-281.
- Freedman, J.C., Shrestha, A., McClane, B.A. Clostridium perfringens Enterotoxin: Action, Genetics, and Translational Applications. Toxins. 2016: 8(3): 73.
- Wellington, E.M.H, Boxall, A.B.A., Cross, P., Feil, E.J., Gaze, W.H., Hawkey, P.M., Johnson-Rollings, A.S., Jones, D.L., Lee, N.M., Otten, W., Thomas, C.M. Williams, A.P. The role of the natural environment in the emergence of antibiotic resistance in Gram-negative bacteria. Lancet Infect. Dis., 2013: 13(2): 155-165.
- Garcia-Migura, L., Hendriksen, R. S., Fraile, L. Aarestrup, F. M. Antimicrobial resistance of zoonotic and commensal bacteria in Europe: The missing link between consumption and resistance in veterinary medicine. Vet. Microbiol., 2014: 170(1-2): 1- 9.
- Tiwari, R., Chakraborty, S., Dhama, K., Rajagunalan, S., Singh, S.V. Antibiotic resistance – an emerging health problem: causes, worries, challenges and solutions – a review. Int. J. Curr. Res., 2013: 5(07): 1880-1892.
- Lin, J., Nishino, K., Roberts, M.C., Tolmasky, M., Aminov, R.I. Zhang, L. Mechanism of antibiotic resistance. Front. Microbiolo., 2015: 6: 34.
- Dhaka, P., Vijay,, D., Vergis, J., Negi, M., Kumar, M., Mohan, V., Doijad, S., Poharkar, K.V., Malik, S.V.S., Barbuddhe, S.B., Rawool, D.B. Genetic diversity and antibiogram profile of diarrhoeagenic Escherichia coli pathotypes isolated from human, animal, foods and associated environmental sources. Infect. Ecol. Epidemiol., 2016: 6, http://dx.doi.org/10.3402/iee.v6.31055.
- Dalton, C.B., Gregory, J., Kirk, M.D., Stafford, R.J., Givney, R., Kraa, E. Gould, D. Foodborne disease outbreaks in Australia, 1995 to 2000.Commun. Dis. Intell. Q Rep., 2004: 28(2): 211-224.
- Qiyi, W. Bruce, A.M. Detection of Enterotoxigenic Clostridium perfringens Type A Isolates in American Retail Foods. Appl. Environ. Microbiol., 2004: 70(5): 2685-2691.
- Wen, Q. McClane, B.A. Detection of enterotoxigenic Clostridium perfringens Type A isolates in American retail foods. Appl. Env. Microbiol., 2004: 70: 2685–2691.
- Herrera, F.C., Santos, J.A., Otero, A. Garc, M.L. Occurrence of foodborne pathogenic bacteria in retail prepackaged portions of marine fish in Spain. J. Appl. Microbiol., 2006: 100: 527-536.
- Papadopoulou, C., Economou, E., Zakas, G., Salamoura, C., Dontorou, C., Apostolou, J. Microbiologicaland pathogenic contaminants of seafood in Greece. J. Food Quality. 2007: 30 (1): 28-42.
- Cai, Y., Gao, J., Wang, X., Chai, T., Zhang, X., Duan, H., Jiang, S., Zucker, B.A., Schlenker, G. Clostridium perfringens toxin types from freshwater fishes in one water reservoir of Shandong Province of China, determined by PCR. DtschTierarzt Wochenschr. 2008: 115(8): 292-297.
- Gamal, E.D. El-Shamery, M.R. Studies on contamination and quality of fresh fish meats during storage Egypt. Acad. J. biolog. Sci., 2010: 2(2): 65-74.
- Martha, I., Tracy, A. Barbara, E.M. David, L.S. American Society for Microbiology. All Rights Reserved. Epidemiology of Seafood-Associated Infections in the United States. Clin. Microbiol. Rev., 2010: 23(2): 399-411.
- El Shorbagy, M.M., Lamyaaand, M.R., Mona, H. Prevalence of Clostridium perfringens Alpha toxin in processed and unprocessed fish. Int. J. of Microbiol. Res., 2012: 3(3): 195-199.
- Sabry, M., Abd El-Moein, K., Hamza, E. Abdel Kader, F. Occurrence of Clostridium perfringens Types A, E, and C in Fresh Fish and Its Public Health Significance. J Food Prot., 2016: 79(6): 994-1000.
- Das, A., Jain, A. Genotyping of Clostridium perfringens from fresh water fish and fish pickles. J. Microbiol. Biotech. Food Sc., 2012b: 2(1): 162- 174.
- Sengupta, N., Alam, S.I., Kumar, R.B. Singh, L., Diversity and antibiotic susceptibility pattern of cultivable anaerobic bacteria from soil and sewage samples of India. Infect. Gen. Evol., 2011: 11: 64-77.
- Cowan and Steel’s Manual for the Identification of Medical Bacteria. 3rd edn. Ed GI Barrow, RKA Feltham. (pp 331; £40.) Cambridge University Press. 1993. ISBN 0-521-32611-7.
- Wu, J., Zhang, W., Xie, B., Wu, M., Tong, X., Kalpoe, J. Zhang, D. Detection and Toxin Typing of Clostridium perfringens in Formalin-Fixed, Paraffin Embedded Tissue Samples by PCR. J. Clin. Microbial., 2009: 47(3): 807-810.
- Osman, K.M., Soliman, Y.A., Amin, Z.M.S. Aly, M.A.K. Prevalence of Clostridium perfringens type A isolates in commercial broiler chickens and parent broiler breeder hens in Egypt. Rev. Sci. Tech. Off. Int. Epiz. 2012: 31(3): 931-941.
- Shome, B.R., Shom, R., Bujarbaruah, K.M., Das, A., Rahman, H., Sharma, G.D. Dutta, B.K. Investigation of haemorrhagic enteritis in pygmy hogs (Sus salvanius) from India. Rev. Sci. Tech. Off. Int. Epiz., 2010: 29(3): 687-693.
- Yoo, H.S., Lee S.U., Park K.Y. Park, Y.H. Molecular typing and epidemiological survey of prevalence of Clostridium perfringens types by multiplex PCR. J. Clin. Microbiol. 1997: 35: 228–232.
- Gibert, M., Jolivet-Reynaud, C. Popoff, M.R. Beta2 toxin, a novel toxin produced by Clostridium perfringens. Gene. 1997: 203: 65-73.
- Czeczulin J.R., Collie R.E., McClane B.A. Regulated expression of Clostridium perfringens enterotoxin in naturally cpe-negative Type A, B, and C isolates of C. perfringens. Infect. Immun. 1996: 64(8): 3301-3309.
- Croci L., Delibato E., Volpe G., Medici D.D., Palleschi G. Comparison of PCR, electrochemical enzyme-linked immunosorbent assays, and the standard culture method for detecting Salmonella in meat products. Appl. environ. Microbiol., 2004: 70: 1393-1396.
- CLSI, 2015. Performance Standards for Antimicrobial Susceptibility Testing; Twenty-Fifth Informational Supplement, M100-S25.Clinical and Laboratory Standards Institute. Pennsylvania: USA.
- Das, A., Mazumder, Y., Rajesh Lal LPM et al. Diagnosis of chronic enteritis in goats by polymerase chain reaction. Int. J. Biotech. Biochem., 2009: 5: 27-34.
- Singh, R.V., Bhilegaonkar, K.N. Agarwal, R.K. Studies on occurrence and characterization of Clostridium perfringens from select meats. J. Food Safety, 2005: 25: 146–156.
- Tansuphasiri, U., Matra, W., Sangsuk, L. Antimicrobial resistance among Clostridium perfringens isolated from various sources in Thailand. Southeast Asian J. Trop. Med. Public Health, 2005: 36(4): 954-961.
- Silva, R.O.S., Salvarani, F.M., Assis, R.A., Martins, N.R.S., Pires, P.S. Lobato, F.C.F. Antimicrobial susceptibility of Clostridium perfringens strains isolated from broiler chickens. Braz. J. Microbiol., 2009: 40: 262-264.
- Slavi´c, Ð., Boerlin, P., Fabri, M., Klotins, K.C., Zoethout, J.K., Weir, P.E., Bateman, D. Antimicrobial susceptibility of Clostridium perfringens isolates of bovine, chicken, porcine, and turkey origin from Ontario. Canad. J. Vet. Res., 2011: 75: 89-97.
- Osman, K.M. Elhariri. M. Antibiotic resistance of Clostridium perfringens isolates from broiler chickens in Egypt. Rev. Sci. Tech. Off. Int. Epiz., 2013: 32(3).
- Singh, R.V. Bist, B. Antimicrobial Profile of Clostridium Perfringens Isolates from Dairy Products. J. Ani. Res., 2013: 3(2): 147-151.
- Sarkar, M., Ray, J.P., Mukhopadhayay, S.K., Niyogi, D. Ganguly, S. Study on Clostridium perfringens type A infection in broilers of West Bengal, India. IIOABJ. 2013: 4(4): 1-3.
- Akhi, M.T., Asl, S.B., Pirzadeh, T., Naghili, B., Yeganeh, F., Memar, Y., Mohammadzadeh, Y. Antibiotic Sensitivity of Clostridium perfringens Isolated From Faeces in Tabriz, Iran. Jundishapur J. Microbiol., 2015: 8(7): e20863.
- Osama, R., Khalifa, M., Al-Toukhy, M. Al-Ashmawy, M. Prevalence and Antimicrobial Resistance of Clostridium perfringens in Milk and Dairy Products. World J. Dairy Food Sci., 2015: 10(2): 141-146.
© The Author(s) 2016. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.


