ISSN: 0973-7510
E-ISSN: 2581-690X
The goal of the current study is to evaluate the potential of marine phages for therapy of motile Aeromonas septicemia caused by Aeromonas hydrophila in Nile tilapia (Oreochromis niloticus). To achieve this goal, four different Aeromonas phages namely AP1, AP2, AP3 and AP4 were isolated from seawater samples and tested for their potentiality as antibacterial agents against culture community in aquaculture water using RAPD-PCR. Results showed significant variation in their antibacterial effect. Characterization of the isolated phages based on restriction digestion using EcoRI, Bam HI, HindIII and Eco RV restriction enzymes and RAPD-PCR proved that the phages were not identical. Plackett Burman experimental design was applied for optimization of culture conditions to improve the antibacterial activity of the most promising AP2 phage against A. hydrophila. Results achieved 94% elimination of A. hydrophila comparing to phage infectivity under basal conditions. Invivo efficiency of AP2 against A. hydrophila invading the aquaria of Nile tilapia was investigated. Elimination of A. hydrophila in the rearing water was detected after 24h. Moreover, morphological and histological examination of the tested liver and gills tissues of Nile tilapia proved the promising effect of AP2 for therapy of motile Aeromonas Septicemia induces by A. hydrophila.
Phages, A. hydrophila, Phage therapy, Nile tilapia
Aeromonas sp. is attributed to many diseases by producing cytotoxins, hemolysins and resistance to many antibiotics1. The most serious bacterial pathogen Aeromonas hydrophila in fish is responsible for many pathological conditions, such as, septicemia, tail rot and epizootic ulcerative syndrome 2.
Control of infection in Aquaculture was dependent on the use of chemical compounds, which resulted in the arising of antibiotic resistance. Therefore there is an urgent need to have alternative tools for pathogen control in aquaculture3.
Phages are distributed in nature with increased numbers than their hosts. They can exist in aquatic system, wastewater, animal gut and springs 4. They have been estimated to kill 20-40% of marine bacteria every day5. They are encouraged to be used in therapeutic applications due to their specific bactericidal capability.
In recent years, natural antimicrobial agents using lytic bacteriophages is a promising new alternative therapy to face the growing bacterial resistance to antibiotics 6,7, which has caused problems in different fields and aquaculture. Different studies concerned the use of Aeromonas phages in treatment of bacterial infection 8,9.
Different factors affect the success of phage frightening against the target pathogenic bacteria such as susceptibility to target bacteria, environmental conditions (temperature, pH, target bacteria /bacteriophages ratio, the time and mode of treatment10.
Therefore the present study was planned to isolate and characterize different marine Aeromonas phages in addition to studying the potential antibacterial of the most potent phage against A. hydrophila under the optimized culture conditions as an alternative to the traditional chemotherapeutic agents.
Samples
Sea water samples were collected from different sites along Mattrouh seashores using sterile screw capped bottles, transferred to the lab and stored at 4 oC.
Media
The following media (g/l) were used throughout the present work (g/l): Nutrient agar: Peptone, 5; yeast extract, 3; beef extract, 1 and agar, 20. Luria Bertani (LB): Peptone, 10; yeast extract, 5. They were supplied from LAB M, England. The pH of each medium was adjusted to 7.0 ± 0.2 before autoclaving at 121°C for 15 min.
Isolation and purification of marine Aeromonas phages
Isolation, purification and determination of bacteriophage titer was carried out as previously described11. First, filtration of sea water samples was done using membrane filter (0.45 µm), followed by serial dilution of the filtrate. Diluted filtrates (0.5 mL) and 1 mL of the host A. hydrophila (in early log phase) were both added on 3 mL soft agar, the mixture was left to solidify. Plates were checked after 24 h of incubation at 30 oC for plaque formation. In order to gain homogenous plaques, each plaque was collected, and used for repeated steps of A. hydrophila infection using double-layer method 12.
Antibacterial activity of the isolated phages against bacterial community structure using Random Amplified Polymorphic DNA (RAPD-PCR)
The isolated marine Aeromonas phages (AP1, AP2, AP3 and AP4) at 4.1 × 106 PFUmL-1 .were added separately to Erlenmeyer flasks containing 100 mL of aquaculture water collected from El-Mex fish farm. Another set without phage addition was kept as control (C). All sets were incubated at 30 °C for 24 h. After incubation, samples were filtered using 0.22 µm pore-size filters. The bacterial cells retained on the membranes were suspended in 2 mL of TE buffer (10 mM Tris HCl, 1 mM ethylenediamine tetraacetic acid (EDTA), pH 8.0) and centrifuged13. In all cases, the extraction of DNA was carried out for further differentiation among different treatments compared to the control (un treated sample) using RAPD-PCR. PCR amplification was carried out by using a PCR Thermocycler (T100TM Thermal Cycler BIO RAD) in the subsequent setting: 1 round of 95°C for 5 min; 35 cycles of 95°C for 30s, 30°C for 30s, and 72°C for 1 min. The following primers were used:
P1: GAATTTTTTTAGGAGGACACTATGAGTG;
P2: AGAGTTTGATCMTGGCTCAG;
P3: TACGGYACCTTGTTACGACTT;
P4: GAGTCGTATAAGATGAATAAGGGGGAATG;
P5: ATCATATGTCTACATCCCTGAAGTAC;
P6: TAGAGCTCTTACAGTCCCTTCGCTTGC
Molecular characterization of the isolated phages using RAPD-PCR
DNA of the isolated Aeromonas phages was extracted using (thermo) kit. The differentiation among phages was studied using RAPD-PCR, based on the occurrence or lack of an exacting RAPD band14. PCR amplification was carried out by using a PCR Thermocycler (T100TM Thermal Cycler BIO RAD) in the subsequent setting: 1 round of 95°C for 5 min; 35 cycles of 95°C for 30s, 30°C for 30s, and 72°C for 1 min using the following primers:
P1: GAATTTTTTTAGGAGGACACTATGAGTG;
P2: AGAGTTTGATCMTGGCTCAG
P4: GAGTCGTATAAGATGAATAAGGGGGAATG;
P7: TAGGCCTGTAAGCTTTTGTCTTGCACAATAAGT;
RFLP analysis
RFLP were take place by restriction digestion analysis for the purified DNA isolated from phage isolates. The digestion reaction was performed using fifty units of EcoRI, Bam HI, HindIII and Eco RV (promega, Southampton, UK), the products of the reactions were analyzed by agarose gel (1%) electrophoresis15.
Host specificity of the isolated phages
Different bacterial strains (Table 1) kindly, provided by Marine Microbiology Department, NIOF, Alexandria, Egypy were tested for their specificity to the isolated phages. Briefly, host cells were grown to log phase and plated on nutrient agar plates. Different purified phage stocks were separately spotted onto the plates, and the plates were incubated overnight at 30 oC. Production of clear or turbid plaques was considered as positive record16.
Table (1):
Host specificity of the isolated Aeromonas phages.
Host |
AP1 |
AP2 |
AP3 |
AP4 |
|---|---|---|---|---|
S.aureus |
+ |
+ |
– |
+ |
S. epidermidis |
– |
– |
+ |
– |
V. parahaemolyticus |
+ |
+ |
– |
– |
V. alginolyticus |
– |
+ |
– |
– |
E. coli |
+ |
+ |
+ |
– |
E. faecalis |
– |
– |
– |
+ |
P. aeruginosa |
– |
+ |
– |
– |
V. damsela |
– |
+ |
– |
+ |
Optimization of culture conditions for improving the antibacterial activity against A. hydrophila using Plackett-Burman experimental design
Effect of different factors on infectivity of AP2 to A.hydrophila was investigated by applying Plackett- Burman experimental design17. In this experiment, seven factors (Table 2) were screened in 8 combinations according to the Plackett Burman design matrix (Table 3). Increase of the original component level is shown by the (+) sign, while decrease of the original component level is shown by the (-) sign. The main effect of each factor was determined using the following equation:
Exi = (Mi+ – Mi-) / N
Where Exi is the variable main effect, and Mi+ and Mi- are optical density (OD at 600 nm) in the trials, where the independent variables were present in high and low concentrations, respectively, and N is the number of trials divided by two. Microsoft Excel was used to calculate statistical t-values for equal unpaired samples and significance of the tested variables. From main effect results, an optimized conditions were predicted which will give maximum inhibition of bacterial growth, followed by verification step to confirm the accuracy of the optimized medium.
Table (2):
Growth factors affecting antibacterial activity of AP2 against A. hydrophila and their levels in the Plackett-Burman experiment.
| Factor | Symbol | Levels | ||
|---|---|---|---|---|
| -1 | 0 | 1 | ||
| Peptone (g/l) | P | 5 | 10 | 15 |
| Yeast (g/l) | Y | 2.5 | 5 | 7.5 |
| Inoculum’s size of host bacteria (ml) | ISb | 0.5 | 1 | 1.5 |
| Inoculum’s size (ml) of phage | ISp | 0.5 | 1 | 1.5 |
| Sea water concentration (%) | C | 50 | 100 | 150 |
| pH | pH | 6 | 7 | 8 |
| Temperature (oC) | T | 25 | 30 | 37 |
Electron microscopy examination
Specimen was negatively stained with sodium tungstate (2%) in bidistilled water at pH 6-7.5. and dropped onto a carbon coated grid. The excess liquid was removed with filter paper after 1 min. Five µl of dye solution were added and after 1 min, the grid was dried. The examination of grids were carried out and electron micrographs were taken with Transmission Electron Microscope (TEM) (JEOL 100 CX) operating at 80 kv.
Table (3):
The applied Plakett-Burman experimental design for the seven culture variables with its antibacterial activity (expressed as bacterial growth (O.D.) at 600 nm).
| Factors symbol | Response | (O.D. at 600 nm) at different time intervals (h) | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Trials | P | Y | SW | pH | Isb | ISp | T | 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 |
| 1 | -1 | -1 | -1 | 1 | 1 | 1 | -1 | 0.020 | 0.030 | 0.045 | 0.060 | 0.160 | 0.600 | 0.210 | 0.230 | 0.280 |
| 2 | 1 | -1 | -1 | -1 | -1 | 1 | 1 | 0.020 | 0.105 | 0.190 | 0.350 | 0.500 | 0.490 | 0.910 | 0.790 | 0.800 |
| 3 | -1 | 1 | -1 | -1 | 1 | -1 | 1 | 0.020 | 1.180 | 0.010 | 0.100 | 0.120 | 0.670 | 0.030 | 0.010 | 0.010 |
| 4 | 1 | 1 | -1 | 1 | -1 | -1 | -1 | 0.020 | 0.160 | 0.320 | 0.390 | 0.420 | 0.620 | 0.440 | 0.260 | 0.460 |
| 5 | -1 | -1 | 1 | 1 | -1 | -1 | 1 | 0.020 | 0.170 | 0.320 | 0.490 | 0.510 | 0.590 | 0.540 | 0.140 | 0.160 |
| 6 | 1 | -1 | 1 | -1 | 1 | -1 | -1 | 0.020 | 0.032 | 0.040 | 0.280 | 0.230 | 0.560 | 0.390 | 0.010 | 0.010 |
| 7 | -1 | 1 | 1 | -1 | -1 | 1 | -1 | 0.020 | 0.490 | 1.870 | 1.850 | 1.840 | 2.230 | 1.490 | 1.530 | 1.520 |
| 8 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0.020 | 0.710 | 1.830 | 1.360 | 1.270 | 3.050 | 2.110 | 1.610 | 1.700 |
| 9 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0.020 | 0.042 | 0.030 | 0.200 | 0.560 | 0.800 | 0.920 | 1.70 | 0.450 |
| Main effect |
0.19 | 0.56 | -0.215 | 0.94 | 0.5 | -0.025 | 0.13 | |||||||||
| t-value | 0.25 | 1.24 | -0.43 | 2.8 | 1.07 | -0.05 | 0.26 | |||||||||
In vivo efficiency of AP2 phage against A. hydrophila
In vivo efficiency was carried out using a modified method8. Nile tilapia (O. niloticus) (weight range: 5-9 g) were used in the study. All fishes were kept in tanks (47 cm x 33 cm x 30 cm) with approximately 5 L de-chlorinated tap water (10 fish per tank), acclimatized for 1 weak prior to the experiment and fed with organic feeds. The aquaria were maintained at 28 ± 1 °C with a pH of 7. Bacterial inoculum of A. hydrophila was prepared using a 24-h-old culture of A. hydrophila inoculated in LB. Four aquaria (T1-T4) were prepared (Table 4), T1,T2 and T3 were infected with 28×104 CFU ml1 of A. hydrophila. On the initial day, AP2 was added to T2, T3 with the ratio of 1:1, 1:2 (AP2: A. hydrophila), respectively, while the first tank (T1) was kept without phage inoculation and T4 was kept as control. In all cases, the total bacterial viable count (TVC) and count of Aeromonas spp. were monitored daily during 72 h with platting of 100 µl of fish culture water on nutrient agar and m-Aeromonas agar medium with ampicillin selective supplement (SRO136), respectively and incubating at 37oC for 24 h. The experiment was carried out in duplicate and the results were taken as average ± SD. Two apparently healthy fish were collected from each tank for histological examination. Any dead fish was daily recorded and removed.
Table (4):
Mean count of total bacteria and Aeromonas spp. in rearing water of O.niloticus.
| TVC x104 | Aeromonas spp. x104 | |||||||
|---|---|---|---|---|---|---|---|---|
| Zero time | 24h | 48h | 72h | zero time | 24h | 48h | 72h | |
| T1 | 28±1.27d | 4000±2.22a | 400±2.6.67b | 280±1.333c | 7±0.33d | 25±1.56c | 48±2.40b | 90±6.43a |
| T2 | 28±2.33a | 2±0.11d | 7±0.47c | 13.6±0.65b | 7±0.58a | 2.5±0.16b | 1.8±0.09bc | 1.2±0.09c |
| T3 | 28±1.56a | 0.2±0.01b | 24 ±1.60a | 24±1.14a | 7±0.39a | 2.5±0.16b | 1.25±0.06c | 1.19±0.09c |
| T4 | 17 ± 0.78 a | 30± 0.15 b | 34±0.16 b | 44 ± 0.19c | 5 ± 0.20a | 11±0.25b | 19±0.37b | 26±0.46c |
a, b, c, d indicate significant difference (p< 0.05).
Histological examination
After fish dissection, the liver and gills were removed, thoroughly washed with a physiological saline (0.9% NaCl) solution and blotted on filter paper then buffered formalin 10%. The fixed specimens were processed using a conventional paraffin embedding technique. This was followed by staining 5 mm thick sections of the prepared paraffin blocks using Hematoxylin and Eosin (HE) for light microscopic examination18.
Natural antibacterial agents are recommended due to their biocompatibility and safety for environment and human health. In the present study, phages specific to A. hydrophila were isolated from sea water samples using soft-agar overlay method. Among the isolated phages AP1, AP2, AP3, and AP4 representing different size and plaque morphology were selected for further study.
Screening for the antibacterial activity of the isolated phages against bacterial community in aquaculture water
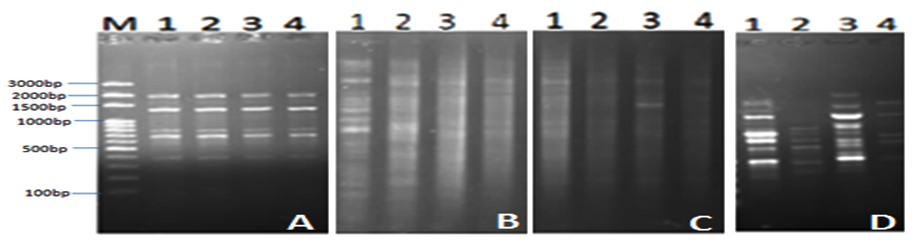
In order to study the antibacterial impact of the isolated phages (AP1, AP2, AP3 and AP4) on the bacterial community in the collected aquaculture water from El-Mex fish farm, DNA extraction & RAPD profiles of the bacterial community was compared before and after addition of phages. Data indicated that all phage isolates showed a principal DNA with size larger than 10 kb. Concentration of DNA in the treated samples was 369 µg/µl, 68 µg/µl, 656 µg/µl and 454 µg/µl for treatment with Ap1, AP2, AP3 and AP4, respectively compared to 1937 µg/µl for the control (without phage addition), this variation was according to the effect of phage on the bacterial cells. Results of RAPD (Fig. 1) indicated that the addition of phages altered the bacterial ribotype diversity represented as difference in the pattern of bands, which indicated that the isolated phages are promising candidate as antibacterial agent in Aquaculture. In accordance with the current investigation, Pereira et al. (2011)13 reported that Aeromonas salmonicida phage showed moderated effect on the structure of bacterial community after addition to aquaculture water.
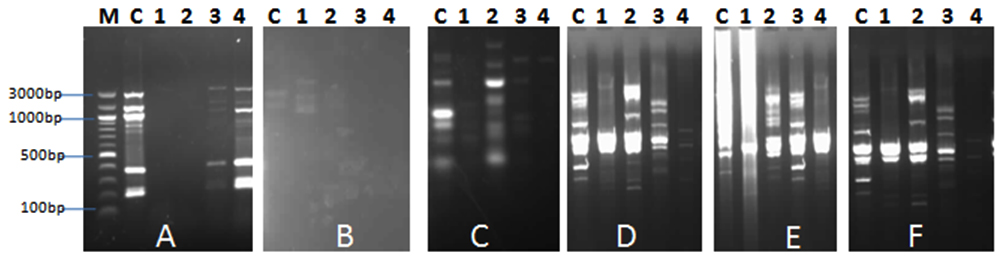 Fig. 1. RAPD PCR fragments using the primers: A, B, C, D, E. and F. after addition of AP1, AP2, AP3 and AP4 phages (Lanes 1-4) to bacterial community of the aquaculture system.. M: molecular weight marker, C: water samples (without addi-tion of phages)
Fig. 1. RAPD PCR fragments using the primers: A, B, C, D, E. and F. after addition of AP1, AP2, AP3 and AP4 phages (Lanes 1-4) to bacterial community of the aquaculture system.. M: molecular weight marker, C: water samples (without addi-tion of phages)
Characterization of the isolated Aeromonas phages
Differentiation of phage DNA by RAPD-PCR
RAPD-PCR was used to detect differences among the isolated phages. As shown in Fig. 2, the RAPD-PCR profile of the isolated phages using different primers was varied which indicate that the AP1, AP2, AP3 and AP4 are not identical. RAPD was documented as quick typing for differentiation of phage isolates19, 20.
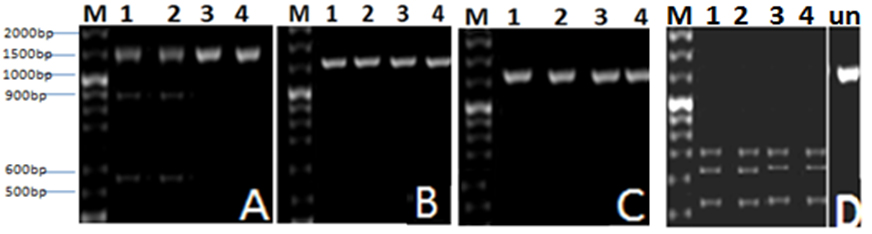
Fig. 2. Agarose gel (1 %) electrophoresis of RAPD products from AP1, AP2, AP3 and AP4 template DNA with the primers: A, B, C and D. Lanes 1-4 represent (AP1-AP4) phages. M is molecular weight marker
Restriction Fragment Length Polymorphism (RFLP)
In this experiment a PCR reaction was followed by RFLP analysis with EcoRI, Bam HI, HindIII and Eco RV. According to (RFLP) pattern, phage isolates were classified. The results (Fig. 3) revealed that none of the tested PCR products showed a change in size after applying HindIII and Bam HI restriction enzyme; this means that there is no site for this enzyme within the tested fragments. Further digestion of PCR products with the other restriction enzymes lead to presence of variation between phage isolates. Data indicated that there are different positions for restriction digestion with these enzyme and the developed RFLP patterns of phage isolates are varied.
Fig. 3. Agarose gel (1 %) electrophoresis for DNA of Aeromonas phages AP1, AP2, AP3 and AP4 indicated by Lanes (1-4), cut with: A: EcoRI, B: Bam HI,C: HindIII and D: Eco RV restriction enzymes. M is molecular weight marker, (Un) is uncut PCR product
Cluster analysis
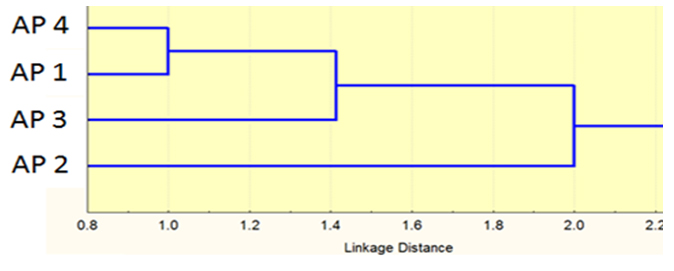
Tree of the isolated phages was carried out based on RAPD and RFLP showing different similarity levels and relationships between the four isolates; it was obvious that AP2 phage was not identical to the other phages (Fig. 4).
Host specificity
Host specificity of the isolated phages was studied against eight bacterial strains (Table 1) using the spot test. Results indicated variation in host specificity, AP2 exhibited broad spectra of host specificity as it was able to infect all the tested pathogens except for S. epidermidis and E. faecalis. Consequently AP2 was choosen to complete the study. Host specificity is often associated with the differences in tail fiber proteins 21. Another factor is conservative structure nature of phage receptors on the outer membrane of bacterial cell 22. Another study reported that some marine phages are specific and lyse only the original host bacterium23. El-Araby et al. (2016)8 studied the phage host specificity of two isolated marine Aeromonas phages namely qZH1 and qZH2 and stated that among the tested bacterial pathogens, the phages were specific to infect Aeromonas and do not have the ability to infect the other bacteria.
Optimization of culture conditions for improving the antibacterial activity of AP2 against A.hydrophila
Physicochemical parameters are important factors affecting survival and infectivity of bacteriophages24.Thus the aim of this experiment is to recognize the most important factors affecting the infectivity of Aeromonas phage using Plackett-Burman experimental design. The screened factors and their levels were presented in Table 2. Results in Table 3 showed varied results of the antibacterial activity of AP2 against A. hydrophila (expressed as elimination of bacterial growth) during the period of study and 7 hours were the most suitable time for higher antibacterial activity. The main effect of the tested variables on the infectivity of AP2 and their corresponding t-values were illustrated in Table 3. The current study indicated that increased concentrations of peptone and yeast extract exhibited positive effect on bacterial inactivation. It was reported that bacteria are more susceptible to phage infection in nutrient-rich conditions 25-28.
pH and temperature finding of the current study showed positive effect of AP2 in reduction of A. hydrophial as pH and temperature can interfere with phage attachment, thus preventing phages to infect the host. Infectivity of some bacteriophages was sensitive to pH values lower than five and more than ten29. Langlet et al. (2007)30 reported that virus exhibited stability at wide range of pH. 37 oC was the most suitable temperature for phage infectivity which is in agreement with El-Araby et al. (2016)8 showing that ÖZH1 and ÖZH2 Aeromonas phages survived better at 37°C and Taj et al. (2014)28 who confirmed that 37°C was ideal temperature for infectivity for T4 bacteriophage. The negative effect of sea water concentration in the current study is coincide with a study by Fennema (1996)31who reported the denaturation of phage proteins at high sea water concentrations. Silva (2005)32 showed that salt concentration affected infectivity of V. vulnificus and V. parahaemolyticus phages. Negative effect of phage inoculum was also detected, which is coincide with a study by Ly-Chatain (2014)10who stated that active biocontrol relies on the addition of a small amount of phages, where elimination of bacteria, in this case, supposes the replication of phages over several generations.
The predicted optimum conditions were: Peptone, 15; yeast extract, 7.5; sea water concentration, 50%; bacterial inoculum size, 1.5; inoculum size of phage, 0.5; pH, 8 and incubated at 37 oC. According to values predicted using t-test (Table 3), pH and yeast extract concentration were highly significant variables.
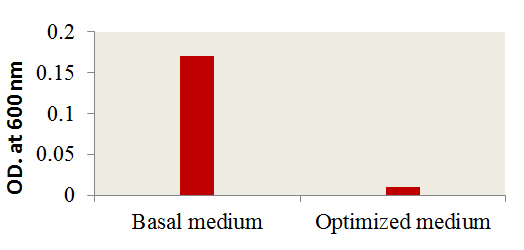
Fig. 5. Effect of optimized culture conditions versus basal me-dium on the antibacterial activity of AP2
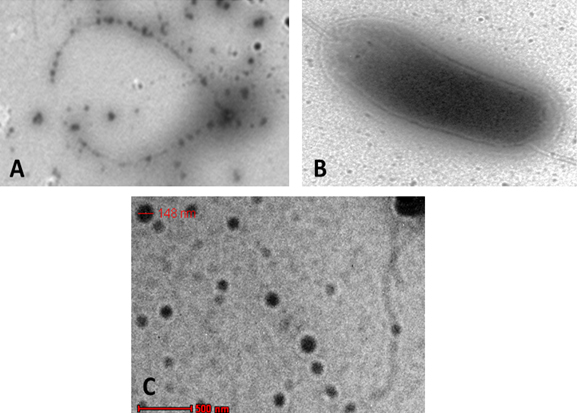
A verification tests were performed to assure the validation of Plackett-Burman experimental design using the predicted optimized media against the basal condition. Results (Fig. 5) showed the increase in the infectivity of AP2 in terms of bacterial growth elimination by about 94% comparing to the phage infectivity under basal conditions. The effect of AP2 grown under the optimized conditions on A. hydrophila was monitored by TEM. As shown a rupture of A. hydrophila cell was observed after phage addition (Fig. 6A) compared to the control A. hydrophila (Absence of AP2) (Fig. 6B). Fig. 6 C showing the morphology of AP2 infecting A. hydrophila. Elimination Aeromonas hydrophila by action of Aeromonas phages was previously reported 8, 33.
Fig. 6. TEM showing AP2 mediated lysis of A. hydrophi-la (A); A. hydrophila cell before lysis (B) and AP2 phage (C)
In vivo efficiency of AP2 in elimination of A. hydrophila
In vivo efficiency of AP4 as antibacterial agent against A. hydrophila in aquaculture was investigated using different ratios of AP2: A. hydrophila (1:2, 1:1). As shown in Table 4, significant elimination (P<0.05) of TVC from 28×104±2.33 to 2×104±0.11 in rearing water of O.niloticus post 24h of AP2 addition was observed using 1:2 concentration of AP2: A. hydrophila (T2 treatment). Similarly, the antibacterial effect of AP2 against Aeromonas started 24h post treatment and prolonged until 72 h with gradual decrease in Aeromonas counts to reach 1.2±0.09 after 72 h compared to 90±6.43 CFU/ml in the infected aquaria. Also the mortality reached 40% during the treatment period. Protective effect of Aeromonas phages against A. hydrophila causing motile aeromonad septicemia was reported in different studies 8, 19, 34.
Morphological symptoms
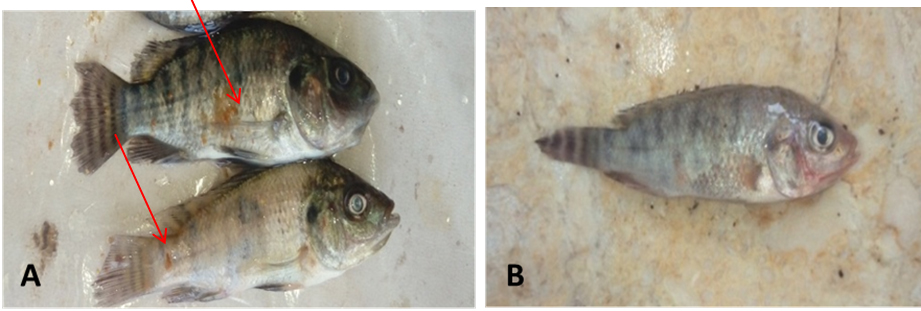
Morphology of O.niloticus and infection signs were observed during 15 days. As shown in Fig. 7 A, B darkness in skin with fin rot in the caudal fin, black spot and red spots on the operculum were observed in the infected fish (shown by red arrows), while these signs were disappeared in the healthy fish (Fig. 7C). Similar signs were reported and attributed to infection with A. hydrophila 35, 36.
Fig. 7. Morphological symptoms: (A): infected O.niloticus with A. hydrophila showing darkness in skin with fin rot in the cau-dal fin, black spot on the operculum (shown by red arrows); (B) healthy O. nilot-icus
Liver histopathological alterations in O. niloticus
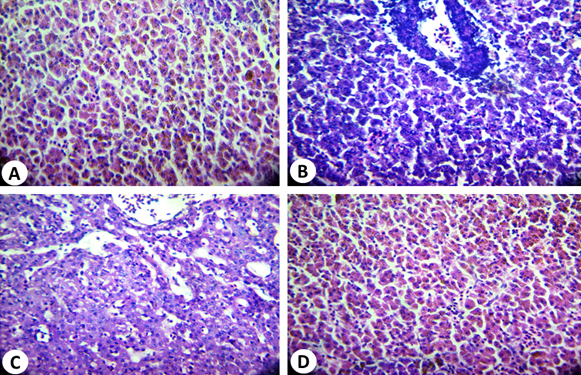
Histological sections of the liver (control) of O. niloticus (Fig. 8A) showed a prominent central vein, with cords of parenchyma cells (hepatocytes) leading to it. Blood vessels appear between the hepatic cells and surrounded by a thin layer of smooth muscle fibers. Nuclei usually centrally located with one prominent central nucleolus. On the other hand, it was noted that in liver of the untreated O. niloticus, small vacuoles of different sizes are scattered in the cytoplasm of some hepatocytes. The infected fish showed disruption of the normal hepatic cord-like pattern, a great reduction in size and number of hepatocytes. The hepatocytes show swelling with foamy cytoplasm and smaller nuclei and a prominent nucleolus which pushed aside. Most blood vessels were empty with red blood cells suffering from necrosis and haemolysis (Fig. 8B). The degree of degeneration is more prominent in case of O. niloticus treated with 1:1 (AP2: A. hydrophia) (Fig. 8 C). At O. niloticus, the hepatocytes had not great difference as compared to the control. However, it was noticed that nuclear degeneration was apparent and cytoplasm appeared more vacuolated than the control as shown in Figure 8D. Similar results have been shown for septicemia caused by A. hydrophila 36-38.
Fig. 8. Histopathological alterations of O.niloticus liver: (A): Normal liver tissue (control); (B): infected liver tissue showing disruption of the normal he-patic cord-like pattern, a great reduction in number and size of hepatocytes, (C): O.niloticus liver tissue treated with 1:1 (AP2 : A. hydrophila) and (D) O.niloticus liver tissue treated with 1:2 (AP2 : A. hydrophila) (D) (H&E: X, 400)
Gill histopathological alterations in O. niloticus
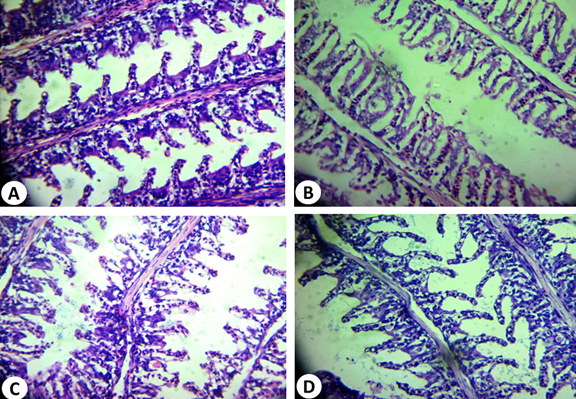
The structure of gills in the control fish was similar to that reported in previous study39. Two specialized epithelia with distinct blood compartments could be distinguished in a frontal section (Fig. 9A), the primary lamellae devoted mainly to ionic regulation and the secondary lamellae responsible for gas transfer39.The respiratory lamellae are lined by squamous epithelial layer one cell thick. In addition, there are a number of pillars, chloride and mucous cells. It is important to mention that; hyperplasia was observed in the epithelial cells of the primary lamellae in most control gills examined.
Fig. 9. Histopathological alterations of O. niloticus gills tissue: (A) normal gills tissue; (B): infected gills tissue showed separation of the epithelial lining of the secondary lamellae from its basement membrane; (C): O.niloticus gills tissue treated with 1:1 (AP2 : A. hydrophila) and (D) O.niloticus gills tis-sue treated with 1:2 (AP2 : A. hydrophila) (H&E: X, 400)
The infected fish gills showed separation of the epithelial lining of the secondary lamellae from its basement membrane as shown in Fig. 9B. Hyperplasia and hypertrophy of the epithelium of the primary lamellae and fusion of the adjacent secondary lamellae are shown in treated fish with 1:1 (AP4: A. hydrophila) (Fig. 9C). The adjacent respiratory lamellae were shortened due to curling or twisting and fused together making one side of the gill filament “solid” structure with no respiratory surface. The gills in case of 1:2 (AP2: A. hydrophila) treatment may recover its shape of primary and secondary lamellae as shown in Fig. 9 D. These results are in accordance with previous reports40,41.
The current study suggested the use of marine Aeromonas phages as an efficient and economical tool to control Aeromonas spp. The marine Aeromonas phage AP2 showed broad range of host specificity, which improves its potential as antibacterial agent. AP2 provided protective effects for Nile tilapia directed to motile Aeromonas Septicemia.
ACKNOWLEDGMENTS
None.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
- Praveen, P. K., Debnath, C., Shekhar, S., Dalai, N., Ganguly, S. Incidence of Aeromonas spp. infection in fish and chicken meat and its related public health hazards: A review, Veterinary World, 2016; 9(1): 6-11.
- Janda, M.J., Abbot, S.L. The genus Aeromonas: Taxonmoy, pathogenicity and infection. Clin. Microbiol. 2010; 23(1): 35-73.
- Pasharawipas, T., Manopvisetcharean, J., Flegel, T.W. Phage treatment of Vibrio harveyi: A general concept of protection against bacterial infection. Res. J. Microbiol. 2011; 6: 560-567.
- Williamson, K. E., Fuhrmann, J. J., Wommack, K. E., Radosevich, M., 2017. Viruses in soil ecosystems: an unknown quantity within an unexplored territory. Annu. Rev. Virol, 2017; 4: 201–219.
- Suttle, C.A. Viruses in the sea. Nature. 2005; 437(7057): 356-361.
- Schooley, R. T., Biswas, B., Gill, J. J., Hernandez-Morales, A., Lancaster, J., Lessor, L., et al. Development and use of personalized bacteriophage-based therapeutic cocktails to treat a patient with a disseminated resistant Acinetobacter baumanniiinfection. Antimicrob. Agents Chemother. 2017; 61: e00954–17.
- Xu, Y., Yu, X., Gu, Y., Huang, X., Liu, G., Liu, X. Characterization and genomic study of phage vBEcoS-B2 infecting multidrug-resistant Escherichia coli. Front. Microbiol., 2018; 9: 1-13.
- El-Araby, D.A., El-Didamony, G., Marihan Megahed, T. H. New approach to use phage therapy against Aeromonas hydrophila induced motile Aeromonas Septicemia in Nile Tilapia. J. Marine Sci. Res. 2016; 6(3): 1-6.
- Donna, M. A., Dela Cruz-Papa, Justine, M., R., Baquiran, Christelle, J. Pineda, Lindley, C. S., Rey Donne, S., P. The treatment of motile Aeromonad Septicemia in Nile Tilapia (Oreochromis niloticus) using phage cocktail therapy with notes on the isolation and description of a novel phage B614. Phillip. Agric. Sci. Microbiol. 2017; 148: 2283-2291.
- Ly-Chatain, M. The factors affecting effectiveness of treatment in phages therapy. Front. Microbiol. 2014; 51: 1-7.
- Clokie, M.R., Kropinski, A.M. Bacteriophages Methods and Protocols. Volume 1: Isolation, Characterization, and Interactions. New York: Humana Press. 2009, 307 pp.
- Adams, N.A. Bacteriophages; John Wiley and Sons Inc.: New York, NY, USA, 1959.
- Pereira, C., Silva, Y., Santos, A., Cunha, A., Gomes, N., Almeida, A. Bacteriophages with potential for inactivation of fish pathogenic bacteria: Survival, host specificity and effect on bacterial community structure. Mar. Drugs., 2011; 9: 2236-2255
- Cateno-Anolles, G., Bassam, B.J., Gresshoff, P.M. High resolution DNA amplification fingerprinting using very short arbitrary oligonucleotide primers. Biotechnol. 1971; 9: 553-557.
- Sambrook, J., Fritsch, E. F., Maniatis, T. Molecular cloning: a laboratory manual, 2nded. Cold Spring Habor laboratory Press, Cold Spring Harbor, N.Y. 1989.
- Xie, Y., Wahab, l., Gill, J. Development and validation of a microtiter plate-based assay for determination of bacteriophage host range and virulence. Viruses, 10(14): 189.
- Plackett, R.L., Burman, J.P. The design of optimum multifactorial experiments. Biometrika. 1946; 33: 305–25.
- Culling, C. F.. Handbook of histopathological and histochemical techniques. Third Ed. Butterworth, London, 1983.
- Le, T.S., Nguyen, T. H., Vo, H.P., Doan, V.C., Nguyen, H.L., Tran , M.T., Tran, T.T., Southgate, P.C., and Kurtböke, D. Protective effects of bacteriophages against Aeromonas hydrophila causing Motile Aeromonas Septicemia (MAS) in Striped Catfish. Antibiotics. 2018; 7: 16.
- Gutiérrez, D, Martín-Platero, A.M., Rodríguez, A., Martínez-Bueno, M., García, P., Martínez, B.Typing of bacteriophages by randomly amplified polymorphic DNA (RAPD)-PCR to assess genetic diversity. FEMS Microbiol Lett. 2011; 322(1):90-7.
- Yosef, I., Goren, M. G., Globus, R., Molshanski-Mor, S., Qimron, U. Extending the host range of bacteriophage particles for DNA transduction. Mol.Cell. 2017; 66: 721–728.
- Rakhuba, D.V., Kolomiets, E., Dey, E.S., Novik, G.I. Bacteriophage receptors, mechanisms of phage adsorption and penetration into host cell. Pol. J. Microbiol. 2010; 59(3): 145-55.
- Hassan, S.W. Isolation and characterization of some bacteriophages and their associated bacteria in sea food: Phage-host interaction. J. High. Inst. Public Health. 2011; 41(4): 403-424.
- Srinivasan, P., Ramasamy, G., Brennan, P., Hanna, R. E. Inhibitory effects of bacteriophages on the growth of Vibrio sp., pathogens of shrimp in the Indian Aquaculture Environment. Asian. Animal. Vet Adv 2007; 2: 166-183.
- Chibani-Chennoufi, S., Bruttin, A., Dillmann, M.L., Brüssow, H. Phage-host interaction: an ecological perspective. J. Bacteriol. 2004; 186: 3677–3686.
- Almeida, A., Cunha, Â., Gomes, N.C.M., Alves, E., Costa, L., Faustino, M.A.F. Phage therapy and photodynamic therapy: low environmental impact approaches to inactivate microorganisms in fish farming plants. Mar. Drugs. 2009; 7: 268–313.
- Silva,Y., Costa, L., Pereira, Ângela, C., Cunha, Calado, R., Newton, C. M. et al., Influence of environmental variables in the efficiency of phage therapy in aquaculture. Microbial Biotechnol. 2013; 7: 401–413.
- Taj, M. K., Ling, J. X., Bing, L. L., Qi, Z., Taj, I., Hassani, T. M., Samreen, Z., Yunlin, W. Effect of dilution, temperature and pH on the lysis activity of T4 phage against E. coli BL21. J. Animal. Plant Sci. 2014; 24(4): 1252-1255.
- Jepson, C.D., March, J.B. Bacteriophage lambda is a highly stable DNA vaccine delivery vehicle. Vaccine. 2004; 22: 2413-2419.
- Langlet, J., Gaboriaud, F., Gantzer, C. Effects of pH on plaque forming unit counts and aggregation of MS2 bacteriophage. J. App. Microbiol. 2007; 103(5): 1632–1638.
- Fennema, O.R. Food Chemistry, 3rd edn. New York, NY, USA: Marcel Dekker, Inc.1996.
- Silva, L.V.A. Control of Vibrio vulnificus and Vibrio parahaemolyticus in oysters. Dissertation. Department of Food Science, Louisiana State University, 2005.
- Chen, L., Yuan, S., Liu, Q., Mai, G., Yang, I., Deng, D., Zhang, B., Liu, C., Ma, Y. In vitro design and evaluation of phage cocktails against Aeromonas salmonicida. Front Microbiol. 2018; 9: 1476.
- Pasaribu, D., Sukenda, S., Nuryati, S .The efficacy of Nile Tilapia (Oreochromis niloticus) Broodstock and larval immunization against Streptococcus agalactiae and Aeromonas hydrophila. Fishes. 2018; 3: 16.
- Güvener, R.P., Timur, G. A study on determination of the Aeromonad infections in some aquarium fish. Istanbul Univ. J. Fish. Aqua. Sci. 2005; 19: 27-39.
- Parvez, N., Mudarris, M.S.A. Investigation on the bacterial haemorrhagic septicemia disease of Cyprinus carpio and Channa striatus. Poult. Fish Wildl .Sci. 2014; 2: 2.
- Stratev, D., Stoev, S., Vashin, I., Daskalov, H. Some varieties of pathological changes in eximentalper infection of carps (Cyprinus carpio) with Aeromonas hydrophila. J. Aquacult. Eng. Fish. Res. 2015; 1: 191–202.
- Abdelhamed, H., Ibrahim, I., Baumgartner, W., Lawrence, M.L., Karsi, A. Characterization of histopathological and ultrastructural changes in channel catfish experimentally infected with virulent Aeromonas hydrophila. Front. Microbiol. 2017; 8: 1519.
- Newstead, J. D. Fine structure of the respiratory lamellae of teleostean gills. Z. Zellforsch. Mikrosk Anat. 1967; 79: 396-428.
- AlYahya, S. A., Ameen, F., Al-Niaeem, K., Al-Sa’adi, B., Hadi, S., Mostafa, A. Histopathological studies of experimental Aeromonas hydrophila infection in blue tilapia, Oreochromis aureus. Saudi J. Biolo. Sci. 2018; 25(1): 182-185.
- Kumar R., Pande V., Singh L., Sharma L., Saxena N. Pathological findings of experimental Aeromonas hydrophilainfection in golden mahseer (Tor putitora) Fish Aquacult. J. 2016; 7:160.
© The Author(s) 2018. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.