ISSN: 0973-7510
E-ISSN: 2581-690X
Successive use and misuse of disinfectant against certain microbes may trigger resistance, causing the problem of ineffectiveness of disinfectant due to development of resistance strains. Ultimately, it is speculated to raise public health issues worldwide. Keeping in view the said problem an experiment was designed to test the efficacy of different disinfectants against some pathogenic bacteria isolated from selected restaurants and cafeterias kitchen. A total of 45 isolates were purified and identified. Amongst them, three isolates; Klebsiella pneumoniae, Macrococcus caseolyticus, Staphylococcus sciuri were selected for their pathogenicity while Terribacillus halophilus were selected for significant tolerance to high temperature and salt regimes. Different parameters including Minimum Inhibitory Concentrations (MIC) and Minimum bactericidal concentration (MBC) were determined for disinfectant; sodium hypochlorite, benzalkonium chloride and chloroxylenol. MIC values was: 2500µg/ml to 5000µg/ml for sodium hypochlorite; 0.54µg/ml to 17.3µg/ml for Benzalkonium chloride; 18.7µg/ml to 150µg/ml for chloroxylenol. MBC values was: 5000µg/ml to 33320µg/ml for sodium hypochlorite; 8.6µg/ml to 138µg/ml for Benzalkonium chloride; 75µg/ml to 300µg/ml for chloroxylenol. Conclusively, K.pneumoniae showed high resistance to sodium hypochlorite and benzalkonium chloride compared to other bacteria used in this study. Among the studied, Staphylococcus sciuri were the most tolerant and resistance to chloroxylenol.
Disinfectant, Resistance, Pathogenic bacteria.
Killing or eliminating bacteria through Disinfectants and Antiseptics particularly in microbiological laboratory, hospitals, food industry, humans and animal care centers are generally exercised for the purpose1,2. Selection of disinfectant and antiseptic counter to pathogenic microorganisms is become a problem due to extensive use to check and control the growth of microbes in living and non-living entities. The extensive application of disinfectant and antiseptic products has encouraged the assumption: promotion of resistance in microbes especially in cross resistance to antibiotics3-6. Currently, chemicals are available in the market in the form of disinfectants and antiseptics having ingredients like halogen compounds, alcohol, chlorohexidine, sodium hypochlorite, peroxides, phenols and quaternary ammonium compounds (QAC) such as benzalkonium chloride7,8. Ethanol, Chloroxylenol, Chlorohexidine are commonly used in microbiology laboratories9. Disinfectant in food industry is important to keep hygiene and to protect the product in food processing and production areas2. Quaternary ammonium compounds, chlorine-based biocides and peroxides are the most disinfecting agent that are used in food industry4. The use of disinfectant is increasing to keep high level of hygienic in food industry environment. The wide use of disinfectant impels selective pressure on bacteria and can lead to emergence of disinfectant-resistance microorganisms10. Disinfectant antimicrobial characteristics towards some of the pathogenic bacteria have been documented. However, microorganisms are developing resistance to these disinfectant and antiseptic, continuously2. Hence, for suitable selection of chemical it is essential to assess the usefulness of disinfectant or antiseptic against particular pathogen11,12. All over the world resistance of pathogenic bacteria against antimicrobial products has been detected and will become a major health threat which lead to treatment failure in human and animal infections13. Bacteria have the ability to adapt new environment rapidly even in the presence of antimicrobial molecules14. Ultimately, increases resistance in pathogenic bacteria along antimicrobial uses15. It is difficult to eradicate these pathogens with antibiotics due to new resistant strains development and new virulent pathogens appearance. New formulations with some non-antibiotic agents have been introduced which are mainly focusing breaking the chain of infections in home, industries and hospitals16.
In this study, we investigated the presence of pathogenic and disinfectant resistant bacteria in some food facility providers (Restaurants, cafeterias) located in south Makkah city. This study will Determine the efficacy and deficiencies in hygiene practices of antimicrobial product in food facilities, the presence of pathogenic and disinfectant resistance bacteria at food preparation rooms.
Samples collection
Samples was collected using cotton swabs from five different location including three restaurants and two cafeterias located in south Makkah from different kitchen accessories including food preparation place, Cutting boards, spoons and cutting tools. A total of 25 swabs were collected from these locations. Further, Swabs was cultured in nutrients agar media at 30°C for 24 hours.
Identifying the bacteria isolates using conventional methods
Isolates were tested by using conventional methods. Gram stain was done, which classifies bacteria into gram-positive bacteria and gram-negative bacteria. The ability to grow on different range of temperature (40 °C, 50°C and 60°C) and the ability to grow in high salt concentration by culturing bacteria isolates on halophilic agar (HiMedia) at 35°C.
Molecular studies
Bacteria was identified using DNA extraction assay; bacterial selected isolates were cultured overnight in enriching broth media. DNA was extracted following the protocols for further PCR. PCR amplification of 16S rDNA gene and sequencing. Bacteria was identified using molecular studies. DNA isolated using Thermo Scientific GeneJET Genomic DNA Purification Kit, following the protocol bacteria cell DNA were extracted for further PCR. The universal primers (Forward primer 5′ – AGA GTT TGA TCC TGG CTC AG -3′ and reverse primer 5′ – GGT TAC CTT GTT ACG ACTT-3′) were used for the amplification of the 16S rRNA gene fragment. The PCR cycling conditions were as follows: 94°C for 5 minutes, followed by 30 cycles of 91°C for 1 min, variable annealing temperature 55°C for 1 min and 72°C for 2 min and then final extension for 5 min at 72°C. The amplified DNA fragments were gel-purified using QIAquick™ GelExtraction Kit. DNA fragment were sent to Macrogen Inc (Seoul, Korea). and sequence analyses were determined by the BLAST.
Determination of the Minimum Inhibitory Concentrations and Minimum Bactericidal Concentration
Using commercially available biocides, biocide tolerance was measured to determine the Minimum Inhibitory Concentrations for each of the selected biocides using a macro-dilution broth method following the Clinical and Laboratory Standards Institute guidelines17. The Minimum Inhibitory Concentrations is the lowest concentration of an antimicrobial product that inhibits the visible growth of a microorganism after overnight incubation. Minimum bactericidal concentration (MBC) was measured by transferring the bacterial culture of each tube by using 1µl loops to a nutrient agar plate after 24 hours. By using growth method 3 to 5 colonies were picked up and resuspend into tube containing 4-5 ml of muller hinton broth by using loops and incubated at 35oc until it achieves the turbidity of 0.5 McFarland standard by measuring absorbance using a spectrophotometer. The absorbance at 625 nm was from 0.08 to 0.13. Inoculum were diluted in MHB, each tube contained approximately 5 x 105 CFU/ml. 1ml of prepaid inoculum were added to each test tube by using test tube, two-fold disinfectant was prepared which was the desired final concentration. 1ml of the disinfectant solution were added to the first tube. The final volume was 1ml in each tube for the test.
Identifying the bacteria isolates using conventional methods
Samples obtained from these locations were cultured in nutrient agar and the growth was observed after 24 hours. After culturing the samples, 45 Individual different colonies were phenotypically selected, isolated and purified. The plates were incubated at 30°C and subsequent subculture was done until pure distinguishable single bacteria colonies were obtained. Bacteria isolates were identified using conventional method. All isolate was microscopically observed for gram staining and cell morphology. 40 isolates (88.8%) were found gram positive and 5 isolates (11.1%) were found negative gram. Growth tolerance of the 45 isolates were also tested for temperature stress (40°C, 50°C and 60°C) and tolerance to NaCl stress using halophilic agar. All 45 isolates were inoculated on nutrient agar at 40°C for 24 hours, after 24 hours we found that all the bacteria isolates were able to grow at 40°C. At 50°C Bacteria isolates was inoculated on nutrient agar, we found that 10 isolates (22.2%) was unable to grow at 50°C while 35 isolates (77.7%) grew after 24 hours except for 1 isolate grew after 48 hours. At 60°C we found that 20 isolates (44.4%) was unable to grow, 18 isolates (40%) was able to grow after 24 hours, 3 isolates (6.6%) grew after 48 hours while 4 isolates (8.8%) grew after 72 hours and 16 isolates were selected for further studies.
Molecular studies
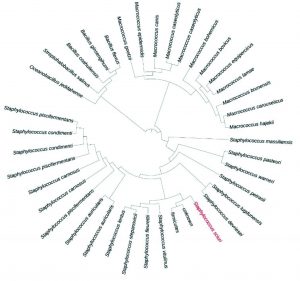
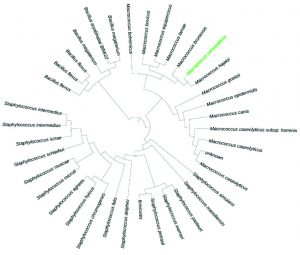
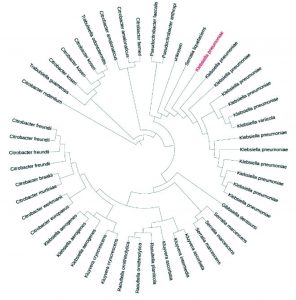
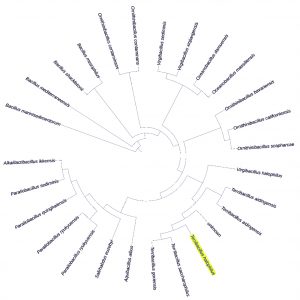
About 45 bacteria were isolated from different areas of south Makkah city. These strains were maintained at 35°C for 24 hours. Out of these three of them was identified as pathogenic based on 16s rRNA primer as Staphylococcus sciuri, Macrococcus caseolyticus and Klebsiella pneumoniae as shown in Fig. 1, 2 and 3. One of them which grow in high salt media was identified as Terribacillus halophilus as shown in Fig. 4. The phylogenetic tree of this strain was constructed against the other pseudomonas species retrieve from NCBI website. Phylogenetic tree was constructed using online iTol software as shown in (Fig. 1, 2, 3 and 4).
Fig. 1. Phylogenetic analysis of the Staphylococcus sciuri, isolates with different Staphylococcus strains from GenBank.
Fig. 2. Phylogenetic analysis of the Macrococcus caseolyticus, isolates with different Macrococcus strains from GenBank.
Fig. 3. Phylogenetic analysis of the Klebsiella pneumoniae, isolates with different Klebsiella strains from GenBank.
Fig. 4. Phylogenetic analysis of the Terribacillus halophilus, isolates with different Terribacillus strains from GenBank.
Determination of the Minimum Inhibitory Concentrations and Minimum Bactericidal Concentration
The minimum inhibitory concentration (MIC) values were investigated. 3 bacterial isolates were selected according to their pathogenicity, Klebsiella pneumoniae, Macrococcus caseolyticus, staphylococcus sciuri and one isolate Terribacillus halophilus were selected for exhibiting high tolerance to growth factors at high Temperature and high salt concentrations.
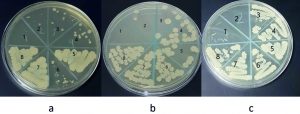
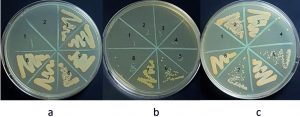
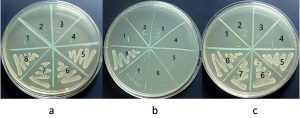
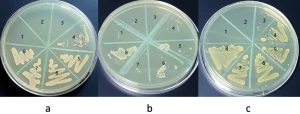
Minimum inhibitory concentration of sodium hypochlorite values was: 5000µg/ml for S. sciuri, 2500µg/ml for M. caseolyticus and 4165µg/ml for both K. pneumoniae and T. halophilus. MBC values were: 33320µg/ml for K. pneumoniae, 5000µg/ml for M. caseolyticus, 8330µg/ml for T. halophilus and 10000µg/ml for S. sciuri. Benzalkonium chloride MIC values were: 0.54µg/ml for M. caseolyticus, and for K. pneumoniae, 1.08µg/ml for S. sciuri and 17.3µg/ml for T. halophilus. MBC values were: 34.7µg/ml for both M. caseolyticus and T. halophilus, 8.6µg/ml for S. sciuri, 138 µg/ml for K. pneumoniae. The data is shown in Fig. 5, 6, 7, 8 and table 1. Moreover, Chloroxylenol MIC values were: 18.7µg/ml for both M. caseolyticus and T. halophilus, 37.5µg/ml for K. pneumoniae and 150µg/ml for S. sciuri. MBC values were: 75µg/ml for K. pneumoniae, 150µg/ml for both M. caseolyticus and T. halophilus, 300µg/ml for S. sciuri.
Fig. 5. K. pneumoniae MBC (a) Sodium hypochlorite #1 = 33320µg/ml, (b) Benzalkonium chloride #1 = 138 µg/ml and (c) Chloroxylenol #2 = 75µg/ml.
Fig. 6. S. sciuri MBC, (a) Sodium hypochlorite #2 = 10000 µg/ml, (b) Benzalkonium chloride #5 = 8.6 µg/ml and (c) Chloroxylenol #1 = 300 µg/ml.
Fig. 7. T. halophilus MBC, (a) Sodium hypochlorite #4 = 8330 µg/ml, (b) Benzalkonium chloride #4 = 34.7µg/ml and (c) Chloroxylenol #2 = 150 µg/ml.
Fig. 8. M. caseolyticus MBC, (a) Sodium hypochlorite #3 = 5000 µg/ml, (b) Benzalkonium chloride #3 = 34.7µg/ml and(c) Chloroxylenol #2 = 150 µg/ml.
Bacteria exposed to sodium hypochlorite showed high tolerance comparing with other disinfectant used in this study. K.pneumoniae were the highest resistance bacteria as MBC result were 33.3 mg/ml. Benzalkonium chloride were highly effective to inhibit bacteria growth according to the result showed on table 1 except for T. halophilus which was 17.3µg/ml, however regarding MBC result K. pneumoniae were high up to 138µg/ml. Chloroxylenol were less effective than Benzalkonium chloride and S. sciuri were the highest tolerance and resistance comparing with other bacteria. The results indicate that the misuse of disinfectant products for disinfection processes may increase bacteria resistance to disinfectant which will lead to disinfection failure and cause threat to public health as previous study by18 confirmed that three strains of K.pneumoniae showed reduced susceptibility against Chlorhexidine, Benzalkonium chloride and Trigene, and another study by19 about three clinical isolates K.pneumoniae that exposed to sub-inhibitory concentration of chlorhexidine showed increase in MIC values from 4-8µg/ml to 128 µg/ml for all three strains. The presence of pathogenic bacteria that can tolerate disinfection process’s and contaminate food facilities and the prevalence of antimicrobial resistance (AMR) bacteria considered as a major public threat and could lead to treatment failure.
Table (1):
Minimum Inhibitory Concentrations and Minimum Bactericidal Concentration Of Disinfectant Agents For Selected Isolates.
| Bacteria | K. pneumoniae | T. halophilus | S. sciuri | M. caseolyticus | |||||
|---|---|---|---|---|---|---|---|---|---|
| MIC Chemical Ag. | MIC µg/ml | MBC µg/ml | MIC µg/ml | MBC µg/ml | MIC µg/ml | MBC µg/ml | MIC µg/ml | MBC µg/ml | |
| Sodium hypochlorite | 4165 | 33320 | 4165 | 8330 | 5000 | 10000 | 2500 | 5000 | |
| Benzalkonium chloride | 0.54 | 138 | 17.3 | 34.7 | 1.08 | 8.6 | 0.54 | 34.7 | |
| Chloroxylenol | 37.5 | 75 | 18.7 | 150 | 150 | 300 | 18.7 | 150 | |
ACKNOWLEDGMENTS
Thanks to Deanship of Scientific Research (DSR), King Abdulaziz University, Jeddah. The authors, therefore, gratefully acknowledge DSR technical support.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
Authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
None.
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the manuscript.
- McDonnell, G. and P. Burke, Disinfection: is it time to reconsider Spaulding? Journal of Hospital Infection, 2011; 78(3): 163-170.
Crossref - SCENIHR, Assessment of the Antibiotic Resistance Effects of Biocides: Scientific Committee on Emerging and Newly Identified Health Risks, 2009.
- Gadea, R., et al., Effects of exposure to quaternary-ammonium-based biocides on antimicrobial susceptibility and tolerance to physical stresses in bacteria from organic foods. Food Microbiol., 2017; 63: 58-71.
Crossref - Vazquez-Sanchez, D., et al., Biofilm-forming ability and resistance to industrial disinfectants of Staphylococcus aureus isolated from fishery products. Food Control, 2014; 39: 8-16.
Crossref - Alekshun, M.N. and S.B. Levy, Molecular mechanisms of antibacterial multidrug resistance. Cell, 2007; 128(6): 1037-50.
Crossref - Couto, N., et al., Biocide and antimicrobial susceptibility of methicillin-resistant staphylococcal isolates from horses. Vet Microbiol., 2013; 166(1-2): 299-303.
Crossref - Fraise, A.P., Historical Introduction, in Russell, Hugo & Ayliffe’s: Principles and Practice of Disinfection, J.-Y.M. Adam P. Fraise, and Syed A. Sattar, Editor, Blackwell Publishing Ltd., 2013; p. 2-4.
Crossref - Sheridan, ְ., et al., The potential for biocide tolerance in Escherichia coli and its impact on the response to food processing stresses. Food Control, 2012; 26(1): 98-106.
Crossref - Jang, H.-H., et al., Use of hydrogen peroxide as an effective disinfectant to Actinobacillus ureae., 2008; 43: 225-228.
Crossref - Langsrud, S., et al., Bacterial disinfectant resistance—a challenge for the food industry. International Biodeterioration & Biodegradation, 2003; 51(4): 283-290.
Crossref - Manivannan, G. and A. Widmer, Disinfection and Decontamination: Principles, Applications and Related Issues:Disinfection and Decontamination: Principles, Applications and Related Issues, 2008; 47: 299-300.
Crossref - Wisplinghoff, H., et al., Resistance to disinfectants in epidemiologically defined clinical isolates of Acinetobacter baumannii. J. Hosp. Infect., 2007; 66(2): 174-81.
Crossref - Ryu, S., The new Korean action plan for containment of antimicrobial resistance. J. Glob. Antimicrob. Resist., 2017; 8: 70-73.
Crossref - Brown, A.E., Microbiological Applications: Laboratory Manual in General Microbiology: McGRAW-Hill Companies, 2005; 35.
- WHO, A safer future: Global public health security in the 21 century, 2007.
- Quinn, P.J.C., M.E.; Markey, B. and Carter GR, Clinical veterinary microbiology: An imprint of Elsevier limited, 2004.
- CLSI, Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria That Grow Aerobically; Approved Standard-Ninth Edition. CLSI document M07-A9. Wayne, PA: Clinical and Laboratory Standards Institute, 2012.
- Abuzaid, A., A. Hamouda, and S.G. Amyes, Klebsiella pneumoniae susceptibility to biocides and its association with cepA, qacDeltaE and qacE efflux pump genes and antibiotic resistance. J. Hosp. Infect., 2012; 81(2): 87-91.
Crossref - Zhang, Y., et al., Chlorhexidine exposure of clinical Klebsiella pneumoniae strains leads to acquired resistance to this disinfectant and to colistin. International Journal of Antimicrobial Agents, 2019; 53(6): 864-867.
Crossref
© The Author(s) 2019. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.