ISSN: 0973-7510
E-ISSN: 2581-690X
Healthcare-associated infections (HCAIs) are considered adverse outcomes of confinement in a healthcare facility. Biofilm-producing, drug-resistant bacteria have further aggravated the problem with HCAIs. This study determined the prevalence, antibiotic susceptibility, and biofilm phenotype of coagulase-negative staphylococci (CoNS) isolated from a hospital environment in Northern Philippines. The identification of CoNS species and the determination of antibiotic susceptibility were done using an automated assay. Biofilm production was detected using tissue culture plate (TCP) and Congo red agar (CRA) methods. Out of 220 surfaces and 26 air samples collected, 103 (41.9%) CoNS strains were isolated, predominated by S. epidermidis with 30.1% prevalence. The medical ward was found to have the highest prevalence of CoNS at 64%. The CoNS isolates exhibited a variable resistance profile; the highest was found against penicillin (97.1%) and oxacillin (54.3%). Isolates manifesting resistance to linezolid and vancomycin were also detected. From the 103 CoNS isolates, 52 (50.5%) biofilm producers were detected using the TCP method, and 39 (37.9%) were detected by the CRA method. Statistically significant difference was found between the biofilm biomass and the slime-producing pattern. This study revealed the prevalence of biofilm-producing, drug-resistant strains of CoNS in a Level 3 hospital in Northern Philippines. This warrants further enhancement of infection prevention and control programs to avert the emergence of more biofilm-producing, drug-resistant bacterial strains that could pose formidable threats to public health.
Healthcare-associated infection, tissue culture plate method, Congo red agar method
The increase in the number of cases of healthcare-associated infections (HCAIs) has been considered as an imminent threat to public health worldwide.1 In industrialized countries, the prevalence is up to 12%. In comparison, developing countries might have up to 19%, but it is assumed to be higher since HCAIs are more prevalent in resource-limited countries,1,2 such as the Philippines. The problem with HCAIs has been aggravated by the increasing number of microorganisms that have acquired drug resistance making treatment more challenging.3,4 Problems are exacerbated when drug-resistant bacteria found in hospital environments contaminate medical devices and form biofilms.
Bacterial biofilms consist of bacteria that live together in clusters that produce an extracellular polymeric substance (EPS) serving as a protective coat.5,6 The EPS imparts the protective function against antimicrobials and the host’s immune system while enhancing bacterial pathogenicity through synergism.7,8 Bacterial biofilms have been found to form on different hospital surfaces and medical devices and are believed to have originated from patients, healthcare workers, or other environmental contaminants.9 Several studies have reported on drug-resistant strains of coagulase-negative staphylococci (CoNS) isolated from hospital environments and have exhibited biofilm-producing ability.10,11 Coagulase-negative staphylococci are frequent inhabitants of hospital environments. They have been classified as the most common cause of HCAIs, the pathogenesis of which is enhanced by the use of medical devices.12 From being simple contaminants in healthcare and non-healthcare settings, CoNS have slowly evolved into a drug-resistant group of microorganisms.13 Staphylococcus epidermidis is the most common CoNS implicated in device-associated HCAIs, and infections in neonates and immunocompromised individuals in ICUs.14,15 Considering the substantial role of biofilms in enhancing the pathogenicity of bacteria, these biofilm-forming, drug-resistant CoNS have become emerging threats to public health systems. In the Philippines, there is a dearth of studies describing the prevalence of biofilm-producing, drug-resistant CoNS in hospital environments.
During the Coronavirus disease 2019 (COVID-19) pandemic, healthcare institutions have enhanced their infection prevention and control (IPC) programs to curb the transmission of the Severe Acute Respiratory Syndrome Coronavirus 2 in their facilities. The enhanced IPC is expected to reduce the persistence of bacterial isolates from healthcare environments. In this study, the prevalence of biofilm-producing, drug-resistant CoNS from a Level 3 hospital environment in Northern Philippines was investigated. While much attention is being accorded to COVID-19, there is a need to remain vigilant in monitoring bacterial isolates with enhanced virulence that could emerge as formidable threats to public health in the near future.
Sample Collection
This cross-sectional, descriptive study determined the prevalence and species distribution of CoNS isolated from the air and surface samples obtained from the different wards (i.e., the four intensive care units [ICU], medical, palliative, Ob-Gyne, private room, and emergency room complex) of a Level 3 hospital in Northern Philippines. It also described the antibiotic susceptibility and biofilm phenotype of the isolated CoNS. The total number of samples collected in the study was 246, consisting of 220 surface samples and 26 air samples. Passive air sampling method, as previously described,16 was used to collect the air samples. In this method, settle plates containing Mannitol Salt Agar (MSA) were distributed in the different wards following the 1/1/1 scheme where plates were placed 1 meter above the floor, 1 meter away from any wall, for 1 hour.
The surface samples were collected from high-touch surfaces (i.e., supply carts, intravenous (IV) poles, beds and rails, door handles, and medical devices) using sterile cotton swabs immersed in sterile normal saline solution following the elution-dependent method.17 After the collection process, the swab samples were enriched using tryptone soy broth (TSB) for 24 hours at 37°C and then cultured on MSA for up to 48 hours to ensure recovery of small colony variants.
After incubation, yellow and white colonies were selected and characterized using Gram stain and biochemical tests (i.e.catalase test and coagulase test). All colonies that exhibited the characteristics of CoNS (i.e., Gram-positive cocci in clusters, positive catalase test, and negative coagulase test) were processed for species identification using the Thermo Fisher Scientific Sensititre Aris 2X ID automated system (TREK Diagnostic Systems, Inc., Cleveland, OH, USA). All pure cultures of CoNS were stored at -20°C in TSB supplemented with 20% glycerol for future testing.
Antibiotic Susceptibility Testing
The susceptibility of the CoNS isolates to several antibiotics (i.e., penicillin, oxacillin, erythromycin, clindamycin, tetracycline, ciprofloxacin, gentamicin, trimethoprim-sulfamethoxazole, rifampin, levofloxacin, moxifloxacin, chloramphenicol, daptomycin, linezolid, and vancomycin) was determined. This was done using the minimum inhibitory concentration (MIC) and breakpoint susceptibility system of the Thermo Fisher Scientific Sensititre Aris 2X AST automated system (TREK Diagnostic Systems, Inc., Cleveland, OH, USA). The nitrocefin disk test (Thermo Scientific Remel, KS, USA) was used to detect β-lactamase production in isolates with penicillin MIC of ≤0.12 µg/ml.
Determination of Biofilm Phenotype Using Tissue Culture Plate (TCP) Method
The quantitative biofilm production assay was performed using the tissue culture plate (TCP) method.18 In this method, the CoNS isolates were inoculated onto wells of a tissue culture plate containing TSB supplemented with 1% glucose then incubated at 37°C for 24 hours. Thereafter, the contents of the wells were discarded and washed 3x with phosphate-buffered saline. The biofilms that adhered to the walls of the microtiter plates were then fixed with 95% ethanol then stained with 0.1% (w/v) crystal violet. The dried, stained biofilms were eluted with glacial acetic acid, and the optical density (OD) of the eluted stained biofilm was measured using a microplate reader at 630 nm.19
The cutoff OD (ODcutoff), the mean OD of the negative control plus 3x its standard deviation, was computed for the interpretation of results. The following criteria were used to classify the biofilm-producing property of the CoNS isolates: ODisolate ≤ODcutoff =non-biofilm-producer; ODcutoff <ODisolate ≤2xODcutoff =weak biofilm-producer; 2xODcutoff <ODisolate≤4xODcutoff =moderate biofilm-producer; and ODisolate >4xODcutoff =strong biofilm-producer.20
Determination of Biofilm Phenotype Using Congo Red Agar (CRA) Method
The qualitative measurement of biofilm production was performed by determining slime production using the CRA method, as previously described.21 Briefly, CoNS isolates were inoculated on the CRA plates and incubated at 37°C for 24 hours. CoNS isolates that exhibited black non-crystalline or very black crystalline colonies were classified as slime producers, while red colonies were interpreted as non-slime producers.12
Statistical Analyses
All data were analyzed in Microsoft Excel 2016 and IBM SPSS Statistics software for Windows version 25 (SPSS Inc., Chicago, IL, USA) at a 95% confidence level. A two-sided Fisher’s Exact test was used to determine the association between the biofilm-producing ability of the CoNS isolates and their resistance to antibiotics. The one-way ANOVA test was used to compare biofilm-producing ability according to the source of isolates and the detection method. All differences with p-values <0.05 were considered significant. The sensitivity and specificity of the CRA method were determined using the TCP method as the standard.
A total of 246 environmental samples, comprised of 220 surfaces and 26 air samples, were collected in this study. The medical ward environment was the most contaminated with CoNS, wherein 64% of samples collected showed growth of at least 1 CoNS, followed by the Ob-Gyne ward, palliative ward, and the ER complex with 52.2%, 43.5%, and 41% prevalence, respectively (see Table 1).
Table (1):
Prevalence of coagulase-negative staphylococci in the different wards* of the hospital.
Source of samples |
NICU |
MICU |
NCCU |
SICU |
Ob-Gyne |
Palliative |
Medical Ward |
PR Annex |
ER Complex |
Total |
|---|---|---|---|---|---|---|---|---|---|---|
Supply cart |
1 |
2 |
2 |
2 |
2 |
2 |
2 |
2 |
4 |
19 |
Intravenous pole |
2 |
0 |
1 |
0 |
3 |
2 |
1 |
1 |
3 |
13 |
Beds and rails |
1 |
1 |
1 |
1 |
2 |
2 |
3 |
2 |
6 |
19 |
Door handles |
1 |
1 |
1 |
3 |
3 |
2 |
2 |
1 |
1 |
15 |
Medical devices |
2 |
1 |
3 |
1 |
2 |
2 |
5 |
0 |
6 |
22 |
Air |
1 |
2 |
1 |
1 |
0 |
0 |
3 |
2 |
5 |
15 |
Total CoNS isolated |
8 |
7 |
9 |
8 |
12 |
10 |
16 |
8 |
25 |
103 |
Total samples collected |
22 |
23 |
23 |
22 |
23 |
23 |
25 |
24 |
61 |
246 |
Prevalence of CoNS per ward (%) |
36.7 |
30.4 |
39.1 |
36.7 |
52.2 |
43.5 |
64.0 |
33.3 |
41.0 |
41.9 |
*NICU: Neonatal intensive care unit, MICU: Medical intensive care unit, NCCU: Neurological critical care unit, SICU: Surgical intensive care unit, Ob-Gyne: Obstetrics and gynecology ward, PR Annex: Private rooms complex, ER Complex: Emergency room complex.
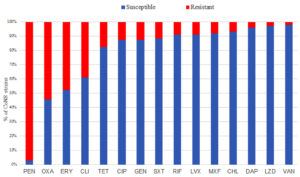
The overall prevalence of CoNS in the hospital was 41.9% (103/246), with S. epidermidis (30.1%) as the most predominant species, followed by S. warneri (24.3%), S. lugdunensis (13.6%), S. haemolyticus (12.6%), S. capitis (7.8%), S. hominis (6.8%), and S. saprophyticus (4.9%). All CoNS isolates were subjected to antibiotic susceptibility testing using 15 antibiotics recommended for testing and reporting stated in the Clinical and Laboratory Standards Institute (CLSI) guidelines.22 Results revealed variable susceptibility patterns among the isolated CoNS strains (see Fig. 1).
Fig. 1. Proportion of CoNS strains showing susceptibility and resistance to each of the tested antibiotics
Figure Legend: PEN- Penicillin, OXA- Oxacillin, ERY- Erythromycin, CLI- Clindamycin, TET- Tetracycline, CIP- Ciprofloxacin, GEN- Gentamicin, SXT- Trimethoprim-Sulfamethoxazole, RIF- Rifampin, LVX- Levofloxacin, MXF- Moxifloxacin, CHL- Chloramphenicol, DAP- Daptomycin, LZD- Linezolid, VAN- Vancomycin.
The isolates were highly susceptible to vancomycin (98.1%), linezolid (97.1%), daptomycin (96.1%), chloramphenicol (93.2%), moxifloxacin (92.2%), levofloxacin (91.3%), rifampin (91.3%), trimethoprim-sulfamethoxazole (88.3%), gentamicin and ciprofloxacin (87.4%), and tetracycline (82.6%). On the other hand, CoNS isolates exhibited high resistance rates to penicillin (97.1%), oxacillin (54.4%), erythromycin (47.5%), and clindamycin (38.8%). All CoNS isolates subjected to Nitrocefin disk test to confirm β-lactamase production showed positive results confirming penicillin resistance of the isolates.
The results of the CRA and TCP tests to determine the biofilm phenotype of the CoNS isolates are shown in Table 2. In the CRA method, out of the 103 CoNS isolates, 64 (62.1%) formed red or pink colonies; 28 (27.2%) formed black, non-crystalline colonies; and 11 (10.7%) formed very black crystalline colonies. The color change of colonies inoculated on CRA medium from red to black is a qualitative indicator of biofilm production.21 In the TCP method, 51 (49.5%) of the CoNS isolates did not form a biofilm, and 52 (50.5%) were able to form biofilms, which were further classified into 40 (38.8%) weak biofilm producers and 12 (11.7%) moderate biofilm producers. No isolate manifested strong biofilm production.
Table (2):
Biofilm-producing phenotype of the CoNS* isolated from the wards of the hospital.
| Biofilm production result† | SE N=31 |
SW N=25 |
SL N=14 |
SHa N=13 |
SC N=8 |
SHo N=7 |
SS N=5 |
Total N=103 |
|---|---|---|---|---|---|---|---|---|
| Congo red agar (CRA) method | ||||||||
| Red/pink | 17 (54.8) | 19 (76.0) | 9 (64.3) | 8 (61.5) | 6 (75.0) | 5 (71.4) | 0 (0.0) | 64 (62.1) |
| Black non-crystalline | 8 (25.8) | 6 (24.0) | 3 (21.4) | 4 (30.8) | 1 (12.5) | 2 (28.6) | 4 (80.0) | 28 (27.2) |
| Very black crystalline | 6 (19.4) | 0 (0.0) | 2 (14.3) | 1 (7.7) | 1 (12.5) | 0 (0.0) | 1 (20.0) | 11 (10.7) |
| Tissue culture plate (TCP) method | ||||||||
| Non biofilm producer | 14 (45.2) | 12 (48.0) | 7 (50.0) | 9 (69.2) | 3 (37.5) | 4 (57.1) | 2 (40.0) | 51 (49.5) |
| Weak biofilm producer | 14 (45.2) | 11 (44.0) | 6 (42.9) | 4 (30.8) | 3 (37.5) | 1 (14.3) | 1 (20.0) | 40 (38.8) |
| Moderate biofilm producer | 3 (9.7) | 2 (8.0) | 1 (7.1) | 0 (0.0) | 2 (25.0) | 2 (28.6) | 2 (40.0) | 12 (11.7) |
| Strong biofilm producer | 0 (0.0) | 0 (0.0) | 0 (0.0) | 0 (0.0) | 0 (0.0) | 0 (0.0) | 0 (0.0) | 0 (0.0) |
*SE: S. epidermidis, SW: S. warneri, SL: S. lugdunensis, SHa: S. haemolyticus, SC: S. capitis, SHo: S. hominis, SS: S. saprophyticus
†Value is no. (%), unless otherwise indicated
The difference between the biofilm producers and their non-biofilm producing counterparts and the antibiotic resistance profiles of the CoNS isolates was statistically determined. As shown in the last column of Table 3, all of the computed p-values were >0.05; hence, no statistically significant difference in the individual antibiotic resistance of the biofilm-producing and non-biofilm-producing CoNS isolates were found.
Table (3):
Comparison of the drug resistance profile of the CoNS isolates classified according to the biofilm phenotype.
| Antibiotics | No. of resistant isolates* | p-value (p ≤ 0.05, significant) |
|
|---|---|---|---|
| Biofilm-producer N = 52 |
Non-biofilm-producer N= 51 |
||
| Penicillin | 51 (98.1) | 49 (96.1) | 0.681 |
| Oxacillin | 29 (55.8) | 27 (52.9) | 0.844 |
| Erythromycin | 22 (42.3) | 27 (52.9) | 0.326 |
| Clindamycin | 19 (36.5) | 21 (41.2) | 0.688 |
| Tetracycline | 8 (15.4) | 10 (19.6) | 0.613 |
| Ciprofloxacin | 6 (11.5) | 7 (13.7) | 0.775 |
| Gentamicin | 6 (11.5) | 7 (13.7) | 0.775 |
| TMP-SXT | 6 (11.5) | 6 (11.8) | 1.000 |
| Rifampin | 2 (3.8) | 7 (13.7) | 0.092 |
| Levofloxacin | 5 (9.6) | 4 (7.8) | 1.000 |
| Moxifloxacin | 4 (7.7) | 4 (7.8) | 1.000 |
| Chloramphenicol | 1 (1.9) | 6 (11.8) | 0.060 |
| Daptomycin | 2 (3.8) | 2 (3.9) | 1.000 |
| Linezolid | 1 (1.9) | 2 (3.9) | 0.618 |
| Vancomycin | 1 (1.9) | 1 (2.0) | 1.000 |
*Value is no. (%), unless otherwise indicated.
Results also showed that there was no statistically significant difference in the mean biofilm biomass of the samples from the different wards as determined by one-way ANOVA (F(8,94) = 1.184, p = 0.317) (see Table 4).
Table (4):
The mean biofilm biomass (measured at OD630) of the CoNS isolates from the different wards* of the hospital.
| Biofilm formation result | NICU N=8 |
MICU N=7 |
NCCU N=9 |
SICU N=8 |
Ob-Gyne N=12 |
Palliative N=10 |
Medical ward N=16 |
PR Annex N=8 |
ER Complex N=25 |
Total N=103 |
|---|---|---|---|---|---|---|---|---|---|---|
| Mean Biofilm biomass (OD630) | 0.086 | 0.076 | 0.010 | 0.092 | 0.088 | 0.098 | 0.119 | 0.122 | 0.110 | – |
| p-value | 0.317 | |||||||||
*NICU: Neonatal intensive care unit, MICU: Medical intensive care unit, NCCU: Neurological critical care unit, SICU: Surgical intensive care unit, Ob-Gyne: Obstetrics and gynecology ward, PR Annex: Private rooms complex, ER Complex: Emergency room complex.
The slime-producing property of the CoNS isolates determined through the CRA method was the basis for the qualitative determination of biofilm production. There was a statistically significant difference in the mean biofilm biomass of the CoNS isolates when compared based on their slime-producing property as determined by one-way ANOVA (F(2,100) = 7.122, p = 0.001) (see Table 5 below). The increasing trend in the biofilm biomass was observed in the three slime-producing patterns. A post-hoc Tukey test showed the red or pink colonies, which manifested the lowest mean biofilm biomass at OD630, differed significantly with a p-value of <0.05 (see Table 5) compared with the mean biofilm biomass produced by the black and very black colonies. No difference was seen in the mean biofilm biomass of the black and very black colonies.
Table (5):
The slime producing property detected by the CRA method and biofilm biomass at OD630 of the CoNS isolates.
| Color of colonies on CRA | No. of isolates* | Mean biofilm biomass measured at OD630 | One-way ANOVA p-value | post-hoc Tukey test | |
|---|---|---|---|---|---|
| Comparison Groups | p-value | ||||
| Red or pink | 64 (62.1) | 0.090 | 0.001 | Red or pink vs. black, not crystalline | 0.004 |
| Black, not crystalline | 28 (27.2) | 0.122 | Red or pink vs. very black, crystalline | 0.039 | |
| Very black, crystalline | 11 (10.7) | 0.124 | Black not crystalline vs. very black, crystalline | 0.985 | |
*Value is no. (%)unless otherwise indicated.
Further, using comparative analysis (see Table 6) of the CRA and TCP methods, the sensitivity of CRA in detecting biofilm production in the CoNS isolates in this study was 48.1%, and the specificity was 72.5% using TCP as the reference method.
Table (6):
Sensitivity and specificity of CRA method using TCP method as standard.
| TCP method | |||
|---|---|---|---|
| Biofilm-producer | Non-biofilm-producer | ||
| CRA method | Black or very black colonies | 25 | 14 |
| Pink or red colonies | 27 | 37 | |
| Total | 52 | 51 | |
| Sensitivity: 48.1% | Specificity: 72.5% | ||
To the authors’ knowledge, this is the first study reporting on the prevalence and species distribution of CoNS isolated from environmental samples in a hospital in the Philippines during the COVID-19 pandemic, where healthcare facilities worldwide have enhanced their IPC measures. Based on the results, this study found that CoNS were prevalent in the different wards of the hospital.
Results of this study showed that of all the CoNS isolated, S. epidermidis was the most predominant. The isolation of S. epidermidis as the most common CoNS agrees with the findings of the hospital’s infection control committee in previous open cultures conducted before the COVID-19 pandemic. In previous studies, S. epidermidis was also reported as the most prevalent CoNS isolated from hospital environment samples13 and clinical isolates from patients in a hospital setting.23 Biofilm-producing S. epidermidis was also reported as the most commonly isolated CoNS from medical devices associated with HCAIs believed to have originated from hospital surface contaminants.14,24 Biofilm-producing CoNS could further aggravate HCAIs by making treatment more challenging because of their increased resistance to antibiotics and the defense mechanisms of the host.10
The medical, Ob-Gyne, and palliative wards and the ER complex of the hospital have shown the highest prevalence of CoNS contaminants in their environment. This could be because of the increased mobility of people in the said wards since there are more patients and healthcare workers in these areas than in other wards. The intensive care units have a relatively lower prevalence of CoNS which could be attributed to stricter implementation of the restricted mobility policy in these wards, especially during the COVID-19 pandemic. Nonetheless, despite the strict implementation of policies, including all personnel entering the ICUs being required to wash their hands, wear sterile lab gowns, and change their footwear, there were still substantial CoNS species that were isolated. In a study that described the contamination of healthcare environments by multidrug-resistant bacteria, the majority of the gram-positive isolates belonged to Staphylococcus species, and the medical ward was reported to have the highest prevalence of total isolated bacteria25 and is in concordance with this study.
To better characterize the CoNS isolated from the environmental samples of the hospital, antibiotic susceptibility profiling was done. The CoNS isolated from the air and surface samples of the selected hospital have shown drug resistance, with almost all isolates manifesting penicillin resistance. The resistance to penicillin revealed in this study was also reported in a previous study where nearly all hospital and community isolates exhibited resistance to penicillin.12 More than half of all the isolates are also resistant to oxacillin and are thus interpreted as methicillin-resistant CoNS.22 Erythromycin and clindamycin are still being used for infections caused by gram-positive bacteria, including the CoNS, especially in patients allergic to penicillin,26 and the high resistance shown by the CoNS isolates to these drugs is a cause for concern. The susceptibility profile exhibited by the CoNS isolates to the rest of the antibiotics is similar to results reported in other studies conducted in Spain27 and Egypt.28
Despite the high susceptibility to linezolid and vancomycin observed in most isolates, the presence of CoNS isolates that manifested resistance to these drugs is alarming. Vancomycin is the first choice to treat serious infections caused by CoNS because of the increasing resistance patterns of these microorganisms to many antibiotics such as the β-lactams.28 In the same way, linezolid is considered the last resort antibiotic for methicillin-resistant staphylococci and vancomycin-resistant enterococci. Although there have been reports of linezolid-resistant CoNS, these are still considered rare.27 Cases of HCAIs caused by strains of biofilm-producing S. epidermidis that exhibited drug resistance to vancomycin and linezolid have been reported in European Union member states.29
Interestingly, the CoNS strains that exhibited resistance to vancomycin and linezolid in the present study were isolated from an ICU. It was also revealed in a previous study that CoNS with decreased susceptibility to vancomycin were isolated from an ICU of a hospital.30 These findings mean that there are multidrug-resistant CoNS strains in the hospital environment that could pose grave public health threats in the near future if no immediate and appropriate response is made.
This study also characterized the biofilm-producing capability of the isolated CoNS. Using the TCP and CRA methods to determine the biofilm phenotype, the results gathered in this study revealed the presence of biofilm-producing CoNS strains in the hospital environment. As earlier stated, IPC measures in the hospitals were enhanced during the COVID-19 pandemic. However, the isolation of these strains showed that they could survive the disinfection procedures that were carried out in the study hospital. CoNS biofilms were previously reported to resist the effects of disinfectants such as sodium hypochlorite and could be attributed to the presence of the EPS matrix.31 Bacteria that can form biofilms produce an EPS matrix that protects them from antimicrobial agents such as antibiotics and disinfectants.32 This could explain the difficulty of eradicating these organisms from hospital surfaces despite frequent disinfection procedures. The presence of biofilm-producing CoNS species in this study may pose another public health hazard emanating from a hospital environment.
A comparison of the biofilm phenotypes was made according to the antibiotic resistance profile. No statistically significant differences were found when the biofilm-producers were compared with non-biofilm-producers based on antibiotic resistance. This suggests that, based on this study, the antibiotic resistance of the CoNS isolates in their planktonic state is not dependent on their ability to produce biofilms. Conversely, the biofilm-producing capacity is not associated with antibiotic resistance. This aligns with a study conducted in a hospital in Germany where no significant differences were found in the antibiotic susceptibility patterns between the biofilm-producers and the non-biofilm-producers.13 Thus, antibiotic resistance is not always directly associated with the amount of biofilm produced nor the capacity to form biofilms. The antibiotic resistance exhibited by biofilm-producing organisms could be more manifested in their sessile forms than in their planktonic state. This could be due to impaired drug penetration because of the biofilm protective effect rather than to any other biochemical processes that the individual organisms might possess.13
The biofilm phenotypes were further compared based on the isolates’ sources (i.e., the different wards). Results showing the absence of statistical difference in the mean biofilm biomass imply that the samples’ sources do not affect the degree of biofilm biomass being produced by the CoNS strains. This could be attributed to the uniform level of cleaning and disinfection being done in each of the hospital wards, particularly during the COVID-19 pandemic. A similar study found that the amount of biofilm biomass produced by S. haemolyticus did not correlate with the environmental sources of the samples.12
The biofilm-producing potential of the CoNS isolates detected through the TCP method was also compared with their slime-producing pattern using the CRA method. Results showed a statistically significant difference in the biofilm biomass produced when compared according to the slime-producing pattern. This implies that the CRA method correlated with the biofilm screening result of the TCP method, which is regarded as the gold standard for the detection of biofilm production. Other studies have also found a good correlation between the TCP method of biofilm determination and the CRA method.12,33
The biofilm detection capability of the CRA method was further characterized by obtaining the sensitivity and specificity of the assay using the TCP method as a reference since it is considered the gold standard for detection of biofilm production.34 The obtained sensitivity of the CRA method used in this study was considerably high compared with other studies.34,35 However, the specificity was relatively lower; the specificity of CRA in previous studies was higher34-36 compared with this study’s specificity of only 72.5%.
In conclusion, this study has determined the prevalence of biofilm-producing, drug-resistant CoNS in a Level 3 hospital in Northern Philippines. The isolated CoNS exhibited a high resistance profile to penicillin and oxacillin and were capable of biofilm production. These results imply the existence of biofilm-producing, drug-resistant CoNS in the hospital environment despite the implementation of enhanced disinfection measures. The presence of biofilm-producing, drug-resistant strains of CoNS in hospital environments warrants the formulation or enhancement of policies that consistently apply intensified infection prevention and control practices in healthcare facilities. Information on the presence of biofilm-producing, drug-resistant strains is also important for physicians for the appropriate management of infections.
ACKNOWLEDGMENTS
The authors would like to thank the Level 3 hospital in Northern Philippines for allowing the collection of environmental samples and Saint Louis University, Philippines for granting permission to use the Natural Sciences Research Unit and its laboratories for the experiments done in this study. The authors would also like to thank Drs. Nina Gloriani, Abubakar Asaad, Fresthel Monica Climacosa, and Glenn Sia Su for providing technical expertise to improve the final version of the dissertation where this article is a part of.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
AJCE performed the specimen collection and all laboratory experiments involved. AJCE and SYAMV, conceptualized the research, analyzed the gathered data, then wrote and thoroughly reviewed the final manuscript. Both authors read and approved the final manuscript for publication.
FUNDING
None.
ETHICS STATEMENT
This study did not involve human participants but was submitted for approval to the University of the Philippines Manila Research Ethics Board. The approval certificate (UPMREB Code: UPMREB 2020-188-EX) was subsequently granted on March 17, 2020. Since this study involved potentially infectious agents, approval was sought from the UP Manila Institutional Biosafety and Biosecurity Committee. The approval certificate (UPM-IBBC APPROVAL NO.: 2020-018) was subsequently granted on June 11, 2020.
AVAILABILITY OF DATA
The datasets generated during and or analyzed during the current study are available from the corresponding author on reasonable request.
- WHO. Healthcare-associated infections. Fact Sheet. 2020. http://www.who.int/gpsc/country_work/gpsc_ccisc_fact_sheet_en.pdf. Accessed on October 18, 2020.
- Danasekaran R, Mani G, Annadurai K. Prevention of healthcare-associated infections: protecting patients, saving lives. Int J Community Med Public Health. 2014;1(1):67.
Crossref - Lis DO, Pacha JZ, Idzik D. Methicillin resistance of airborne coagulase-negative staphylococci in homes of persons having contact with a hospital environment. Am J Infect Control. 2009;37(3):177-182.
Crossref - May L, Klein EY, Rothman RE, Laxminarayan R. Trends in Antibiotic Resistance in Coagulase-Negative Staphylococci in the United States, 1999 to 2012. Antimicrob Agents Chemother. 2013;58(3):1404-1409.
Crossref - European Centre for Disease Prevention and Control. Healthcare-associated infections acquired in intensive care units – Annual Epidemiological Report 2016 [2014 data]. 2017. https://ecdc.europa.eu/en/publications-data/infections-acquired-intensive-care-units-annual-report-2016. Accessed on November 26, 2020.
- Sievert DM, Ricks P, Edwards JR, et al. Antimicrobial-Resistant Pathogens Associated with Healthcare-Associated Infections Summary of Data Reported to the National Healthcare Safety Network at the Centers for Disease Control and Prevention, 2009-2010. Infect Control Hosp Epidemiol. 2013;34(1):1-14.
- Davies D. Understanding biofilm resistance to antibacterial agents. Nat Rev Drug Discov. 2003;2(2):114-122.
Crossref - Jamal M, Ahmad W, Andleeb S, et al. Bacterial biofilm and associated infections. J Chin Med Assoc. 2018;81(1):7-11.
Crossref - Murugan K, Selvanayaki K, Al-Sohaibani S. Urinary catheter indwelling clinical pathogen biofilm formation, exopolysaccharide characterization and their growth influencing parameters. Saudi J Biol Sci. 2016;23(1):150-159.
Crossref - Percival SL, Suleman L, Vuotto C, Donelli G. Healthcare-associated infections, medical devices and biofilms: risk, tolerance and control. J Med Microbiol. 2015;64(4):323-334.
Crossref - Donlan RM. Biofilms and device-associated infections. Emerg Infect Dis. 2001;7(2):277-281.
Crossref - Seng R, Kitti T, Thummeepak R, et al. Biofilm formation of methicillin-resistant coagulase negative staphylococci (MR-CoNS) isolated from community and hospital environments. PLoS One. 2017;12(8):e0184172.
Crossref - Wojtyczka RD, Orlewska K, Kepa M, et al. Biofilm formation and antimicrobial susceptibility of Staphylococcus epidermidis strains from a hospital environment. Int J Environ Res Public Health. 2014;11(5):4619-4633.
Crossref - Becker K, Heilmann C, Peters G. Coagulase-negative Staphylococci. Clin Microbiol Rev. 2014;27(4):870-926.
Crossref - Widerstrom M, Wistrom J, Edebro H, et al. Colonization of patients, healthcare workers, and the environment with healthcare-associated Staphylococcus epidermidis genotypes in an intensive care unit: a prospective observational cohort study. BMC Infect Dis. 2016;16(1):743.
Crossref - Getachew H, Derbie A, Mekonnen D. Surfaces and air bacteriology of selected wards at a referral hospital, Northwest Ethiopia: a cross-sectional study. Int J Microbiol. 2018;2018:6413179.
Crossref - Chai J, Donnelly T, Wong T, Bryce E. Environmental sampling of hospital surfaces: Assessing methodological quality. Can J Infect Control. 2018;33(3):138-145. https://ipac-canada.org/photos/custom/pdf/cjicArticleBryce.pdf. Accessed on November 16, 2020
- Christensen GD, Simpson WA, Younger JJ, et al. Adherence of coagulase-negative staphylococci to plastic tissue culture plates: a quantitative model for the adherence of staphylococci to medical devices. J Clin Microbiol. 1985;22(6):996-1006.
Crossref - Manandhar S, Singh A, Varma A, Pandey S, Shrivastava N. Evaluation of methods to detect in vitro biofilm formation by staphylococcal clinical isolates. BMC Res Notes. 2018;11(1):714.
Crossref - Stepanovic S, Vukovic D, Hola V, Di Bonaventura G, Djukic S, Cirkovic I, Ruzicka F. Quantification of biofilm in microtiter plates: overview of testing conditions and practical recommendations for assessment of biofilm production by staphylococci. APMIS. 2007;115(8):891-899.
Crossref - Freeman DJ, Falkiner FR, Keane CT. New method for detecting slime production by coagulase negative staphylococci. J Clin Pathol. 1989;42(8):872-874.
Crossref - Clinical and Laboratory Standards Institute (CLSI). Performance standards for antimicrobial susceptibility testing, 26th ed.; CLSI Supplement M100S; CLSI: Wayne, NJ, USA, 2016. M100S – American National Standards Institute [Internet]. https://webstore.ansi.org/preview-pages/CLSI/preview_CLSI+M100-S26.pdf. Accessed on October 19, 2020.
- Shrestha LB, Bhattarai NR, Khanal B. Comparative evaluation of methods for the detection of biofilm formation in coagulase-negative staphylococci and correlation with antibiogram. Infect Drug Resist. 2018;11:607-613.
Crossref - Ledwoch K, Dancer SJ, Otter JA, et al. Beware biofilm! Dry biofilms containing bacterial pathogens on multiple healthcare surfaces; a multi-centre study. J Hosp Infect. 2018;100(3):E47-E56.
Crossref - Chaoui L, Mhand RA, Mellouki F, Rhallabi N. Contamination of the surfaces of a health care environment by multidrug-resistant (MDR) bacteria. Int J Microbiol. 2019;2019:3236526.
Crossref - Bora P, Datta P, Gupta V, Singhal L, Chander J. Characterization and antimicrobial susceptibility of coagulase-negative staphylococci isolated from clinical samples. J Lab Physicians. 2018;10(04):414-419.
Crossref - Balandin B, Lobo B, Orden B, et al. Emergence of linezolid-resistant coagulase-negative staphylococci in an intensive care unit. Infect Dis (Lond). 2015;48(5):343-349.
Crossref - Mashaly GE-S, El-Mahdy RH. Vancomycin heteroresistance in coagulase negative Staphylococcus blood stream infections from patients of intensive care units in Mansoura University Hospitals, Egypt. Ann Clin Microbiol Antimicrob. 2017;16(1):63.
Crossref - European Centre for Disease Prevention and Control. Multidrug-resistant Staphylococcus epidermidis – 8 November 2018. Stockholm: ECDC. https://www.ecdc.europa.eu/sites/default/files/documents/15-10-2018-RRA-Staphylococcus%20epidermidis%2C%20Antimicrobial%20resistance-World_ZCS9CS.pdf. Accessed on November 20, 2020.
- Center KJ, Reboli AC, Hubler R, Rodgers GL, Long SS. Decreased vancomycin Susceptibility of Coagulase-Negative Staphylococci in a Neonatal Intensive Care Unit: Evidence of Spread of Staphylococcus warneri. J Clin Microbiol. 2003;41(10):4660-4665.
Crossref - Olwal CO, Angienda PO, Onyango DM, Ochiel DO. Susceptibility patterns and the role of extracellular DNA in Staphylococcus epidermidis biofilm resistance to physico-chemical stress exposure. BMC Microbiol. 2018;18(1):40.
Crossref - Stoica P, Chifiriuc MC, Rapa M, Lazar V. Overview of biofilm-related problems in medical devices. Biofilms and Implantable Medical Devices. Amsterdam: Elsevier; 2017:3-23.
Crossref - Panda PS, Chaudhary U, Dube SK. Comparison of four different methods for detection of biofilm formation by uropathogens. Indian J Pathol Microbiol. 2016;59(2):177:179.
Crossref - Halim RMA, Kassem NN, Mahmoud BS. Detection of biofilm producing Staphylococci among different clinical isolates and its relation to methicillin susceptibility. Open Access Maced J Med Sci. 2018;6(8):1335-1341.
Crossref - Hassan A, Usman J, Kaleem F, Omair M, Khalid A, Iqbal M. Evaluation of different detection methods of biofilm formation in the clinical isolates. Braz J Infect Dis. 2011;15(4):305-311.
Crossref - Mathur T, Singhal S, Khan S, Upadhyay D, Fatma T, Rattan A. Detection of biofilm formation among the clinical isolates of Staphylococci: an evaluation of three different screening methods. Indian J Med Microbiol. 2006;24(1):25-29.
Crossref
© The Author(s) 2022. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.