ISSN: 0973-7510
E-ISSN: 2581-690X
This study aimed to isolate and characterize bacteriophages, as a biocontrol agent, against certain antibiotic-resistant bacteria causing dental caries. Here, two dental caries-causing bacteria S. aureus and E. faecalis were isolated and characterized biochemically using the automated VITEK® 2 system. Antibiotic sensitivity pattern of the isolated dental caries bacteria was assessed against selection of antibiotics. The two isolates showed resistance against most of the tested antibiotics. To overcome this problem, two lytic phages vB_SauM-EG-AE3 and vB_EfaP-EF01 were isolated, identified, and applied to control the growth of S. aureus and E. faecalis, respectively. Phages were identified morphologically using TEM and showed that vB_SauM-EG-AE3 phage is related to Myoviridae and vB_EfaP-EF01 phage belongs to Podoviridae. The two phages exhibited high lytic activity, high stability, and a narrow host range. The one-step growth curve of phages showed burst sizes of 78.87 and 113.55 PFU/cell with latent periods of 25 and 30 minutes for S. aureus phage and E. faecalis phage respectively. In addition, the two phages showed different structural protein profiles and exhibited different patterns using different restriction enzymes. The genome sizes were estimated to be 13.30 Kb and 15.60 Kb for phages vB_SauM-EGAE3, vB_EfaP-EGAE1, respectively. Complete inhibition of bacterial growth was achieved using phages with MOIs of 103, 102 and 10 after 1, 3, 5, and 24 h of incubation at 37°C. Hence, this study indicates that the isolated bacteriophages are promising biocontrol agents that could challenge antibiotic-resistant dental caries bacteria to announce new successful alternatives to antibiotics.
Dental caries, Enterococcus faecalis, Staphylococcus aureus, multidrug-resistance, bacteriophage, biocontrol
Enterococcus is a genus of Gram-positive, non-spore-forming bacteria commonly found in the oral cavity, gastrointestinal tract, and vagina1. Though most species of Enterococcus in humans and animals are considered commensal2. Emergence of Enterococcus as human pathogens due to the presence of virulence factors such as adhesive production and aggregation compounds, or biogenic amine production in fermented foods3 and they resist different measures of disinfection and create a biofilm4. Enterococcus faecalis (E. faecalis) was not considered to belong to the normal oral flora. E. faecalis can act as an opportunistic pathogen that causes bacteremia, meningitis, endocarditis, or urinary tract and wound infection5,6. It has several survival mechanisms for living in unfavorable environments, such as grow in a low-oxygen environment at high pH, a broad range of temperatures between 10°C and 60°C and high salinity or a poor nutritional environment7. In patients with post-treatment apical periodontitis or refractory apical periodontitis, E. faecalis was located mainly in the root canals, suggesting an etiological role in the development of these diseases4,8. The excessive use or abuse of antibiotics has triggered a disturbing appearance of virulent antibiotics resistant pathogenic bacteria9. As important nosocomial pathogens, the therapeutic challenge of multi-drug resistant enterococci (MDR) strains with substantial resistance to two or more antibiotics, often including, though not limited to, vancomycin, has brought their role to a sharper focus10.
Staphylococcus aureus (S. aureus) is a gram-positive coccoid bacterium that causes human bacteremia, endocarditis, osteoarticular, skin and soft tissue, pleuropulmonary infection, and device-related infections11. S. aureus was isolated from the oral cavity12. Many oral infections are at least partly caused by S. aureus, as parotitis, and staphylococcal mucositis13. S. aureus has a range of virulence factors including adhesives, toxins, coagulase, and a variety of genes for resistance to antimicrobials14. Clinical S. aureus isolates exhibit recurrent multidrug resistance, with the most important being methicillin and vancomycin resistance15. Mature biofilm formation induces additional antibiotic tolerance16. Infections with multidrug-resistant S. aureus require new therapies with current interest focused on bacteriophages (phages).
Bacteriophages (phages) are viruses that invade bacteria; they interrupt the metabolism of bacteria and cause lysis of bacterial cells (lytic phages). Specifically, each type of phage attacks only certain bacteria as its host; it frequently does not affect total microbial biomass17. During-active infection, a strictly virulent phage generates typically more than 100 copies of itself18. If the numberof phages is lower than that of bacterial cells, after many generations the population of phages will surpass that of bacterial cells, ultimately the total bacterial cells will lyse17.
One alternative that has recently revived interest is phage therapy, first proposed in the early 20th century by Felix d’Herelle. Besides, several clinical studies have shown that the use of bacteriophages in both humans and animals is effective and without side effects19,20. Lytic phages are considered as potential alternative candidates to conventional antibiotics for a wide range of oral bacterial infections21.
The first phage therapy was reported concerning the treatment of S. aureus skin infections; also, phage therapy was used in some infections associated with burn wounds caused by S. aureus. Phages have been reported to be successful in treating various bacterial diseases such as infections of the skin caused by Staphylococcus, neonatal sepsis, infections of the urinary tract, and diabetic foot as a complication of diabetes22.
Polyvalent bacteriophage K, a well-characterized member of the Myoviridae family of viruses, is a candidate for combating S. aureus MDR infections23. Estrella et al(24) isolated and identified seven novel phages with broad lytic activities for S. aureus from untreated sewage. A phage able to parasitize E. faecalis has been described25. A total of 23 lytic phages, targeting E. fecalis, are based on previously published studies. They were isolated from several sources, including wastewater, effluent from farm animals and human feces26,21,27.
In vitro antimicrobial effect bacteriophages on human dentin infected with E. faecalis ATCC 29212 indicated that applying bacteriophage lysate to the roots after 6days incubation resulted in a major decrease in the survival of the bacteria28. Lee et al21 indicated that bacteriophage HEf13 has a lytic activity and is a therapeutic potential agent for treating or preventing infectious diseases associated with E. faecalis21. In this study, bacteriophage isolation and identification were achieved as novel candidates appropriate for bacteriophage therapy against E. faecalis and S. aureus as alternatives to conventional antibiotics.
Bacterial strains and growth conditions
This study was performed on two antibiotic-resistant dental caries isolates that were isolated previously from patients with dental plaques in Qalubiya governorate, Egypt. All strains were stored at -80°C in Brain-Heart-Infusion broth complemented with 20% (v/v) glycerol. Freshly overnight grown cultures were prepared by inoculating a single colony into 10 ml of Basal salt medium with yeast extract and incubating for 16 h at 37°C with shaking at 200 rpm.
Morphological and Biochemical identification of the dental caries bacteria
Morphological and Biochemical identification (Table S1, Supplementary data) of the dental caries bacteria were carried out according to Bergey’s Manual of Systematic Bacteriology. These isolates were confirmed by VITEK® 2 COMPACT automated instrument for ID/AST testing29.
Antibiotic sensitivity test
Antibiotics sensitivity testing was performed on Mueller-Hinton agar by the disc diffusion method30 for the following antibiotics (Oxoid, UK); Nitrofurantoin (F, 10 µg), Amoxicillin (AX, 25 µg), Chloramphenicol (C, 30 µg), Ciprofloxacin (CIP, 5 µg), Gentamicin (GN, 10 µg), Norfloxacin (NOR, 10 µg), Penicillin-G (P, 10 µg), Vancomycin (V, 30 µg), Ceftazidime (CAZ, 30 µg), Aztreonam (ATM, 30 µg), Cefotaxime (CTX, 30 µg) and Amikacin (AK, 30 µg). The results were interpreted conferring to the guidelines of the Clinical and Laboratory Standards Institute (CLSI)31.
Isolation of Bacteriophages
The clinical isolates S. aureus EG-AE3 and E. faecalis EF01 were used as hosts for the isolation, and propagation of bacteriophages. Saliva samples from 15 healthy individuals and 85 dental caries affected patients and drainage samples from dental clinics were collected from Benha city, Qalubiya governorate, Egypt for bacteriophage screening. The collected samples were centrifuged at 10,000 rpm for 10 min, supernatant fluids were collected, and membrane-filtered using 0.22 µm membrane filters (Millipore, Ireland).
Enrichment of phages and isolation were performed as described previously32. Briefly, 5 ml of a 0.22 µm-filtered sample was mixed with 20 ml double-strength Tryptic Soybean Broth (TSB) medium and 100 µl of a mid-log culture of each of S. aureus strain EG-AE3 and E. faecalis EF01 and incubated for 48 h at 37°C with shaking at 200 rpm. Later, bacteria were harvested by centrifugation at 10,000 rpm for 10 min, supernatant fluids were recovered, and membrane-filtered using 0.22-µm Millipore filters (Millipore, Ireland). Phages were screened by spotting five microliters of the enriched samples onto double-layered plates containing a lawn of the indicator bacteria strain and incubated for 48 h at 37°C33. Plates were inspected for the presence of clear lysis zones; the clear zone was cut and propagated in a fresh culture. This lysate was serially diluted, spotted onto double-layered plates, and incubated as described above.
Transmission electron microscopy of bacteriophages
Ten microliters of highly purified phage suspension were fixed onto 300- by 300-mesh copper grids (Electron Microscopy Sciences) supported by carbon-coated Formvar film34. After 5 minutes, fixed phages were negatively stained with 2% (w/v) aqueous phosphate tungsten acid, pH 7.2 for 1 min, and air-dried at room temperature for 1 h. Transmission electron microscope (A JEOL JEM-2100) was used for attaining the phage particle images at the Electron Microscope Facility, Al-Azhar University, Egypt.
One-step growth curve
Phages growth phases and burst size were determined as described previously33. A known number of an exponential-phase culture (ca. 1×107 CFU/ml) of each bacterial host was infected with each specific phage individually at an MOI of 1, phages were allowed to adsorb for 5 min at room temperature. The mixture was then centrifuged at 5,000 rpm for 5 min and the supernatant was decanted to remove free-unbound phages. The bacteria-phage pellet was then washed twice and resuspended in 10 ml of TSB and maintained at 37°C with continuous shaking. At appropriate times phage titers were enumerated using plaque assay35.
Determination of the bacteriophages host range
The host range for the two isolated phages vB_SauM-EG-AE3 and vB_EfaP-EF01 was determined against a collection of twenty-two bacterial isolates (Table S2, Supplementary data) as previously described with some modifications36,37. Ten microliters of each phage suspension (about 108 PFU/ml) were spotted, in duplicate, onto the TSA bacterial lawn plates and incubated at 37 °C for 16–18 h.
Thermal and pH stability
Thermal and pH-stability of the isolated phages were tested as described before33. For the assessment of thermal stability, 900 µl of pre-heated 0.22 µ m filter-sterilized SM buffer (5.8 g NaCl, 2.0 g MgSO4•7H2O, 50 ml 1 M Tris-HCl pH 7.4, in 1-liter dH2O) were added to 100 µl of each of phage lysates (8 log10 PFU/ml). Tubes were incubated at 10°C, 30°C, 40°C, 50°C, 60°C, 70°C, and 80°C for 1 h. Aliquots were collected After 60 min of incubation to determine phage titers. For pH-stability assessment, phage lysates (8 log10 PFU/ml) were added to tubes containing sterile SM buffer with pH values ranging from 2-13 adjusted with NaOH and HCl. The tubes were incubated at 37°C for 60 min. Subsequently, the phage solutions were serially diluted and the recovered phage titers were determined using bacterial hosts employing the double-layer agar method35. Each temperature and pH treatment was performed in triplicates and the average of triplicate counts was calculated.
Phage thermal/pH stability (%) = (Recovered phage titers following the treatment / Initial Phage titer before treatment) ×100%
Sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE)
Viral protein profiles were assed to Sodium Dodecyl Sulfate-Polyacrylamide Gel Electrophoresis (SDS-PAGE) using Agilent 2100 Bioanalyzer system (Agilent Technologies, Cat # G2939BA) which is a chip-based automated electrophoretic assay. Solutions and reagents were prepared according to the Agilent High Sensitivity Protein 230 Kit manual (Reference number 5067-1517). The protein assay was performed following the manufacturer’s instruction. The ladder ranged from 6 kDa (lower marker) to 240 kDa (upper marker). Results were analyzed and visualized using the 2100 Expert software provided by Agilent Technologies, USA.
Extraction of bacteriophages DNA and restriction analysis
DNA was extracted from purified high-titer stocks of phage using Wizard® Genomic DNA Purification Kit (Promega Corporation, Cat #A1120) according to the manufacturer’s instructions. DNA integrity was checked using 1% agarose gel electrophoresis and the image was captured using a gel documentation system (Gel Doc. BioRad). Concentration and purity of purified DNA were measured by BioTek Epoch2 Microplate reader (Thermo Scientific, USA). For all samples, DNA purity was >1.8 ± 0.20 under absorbance ratio A260/A280.
For profiling and comparing bacteriophage genomic DNA fragment patterns, genomic DNA was digested with different restriction enzymes (BamHI, EcoRI, DraI, PstI, and KpnI). The digestion process was performed following the manufacturer’s instructions of Promega Corporation for each enzyme. Restriction fragments were separated by electrophoresis using Agilent 2100 Bioanalyzer assay (Agilent Technologies, Cat # G2939BA) and following the manufacturer’s instructions of Agilent High Sensitivity DNA kit (Cat # G2938-90321). The ladder sizing range varied from 50 bp to 10380 bp for size determination of DNA fragments. Results were analyzed and visualized using the 2100 Expert software provided by Agilent Technologies, USA.
Biocontrol of dental caries-causing bacteria using bacteriophages
The efficacy of phages to inhibit the growth of their hosts was assessed in broth medium using different MOIs. Phages were separately mixed with bacterial suspension that was diluted to103 CFU/ml from an overnight culture of S. aureus and E. faecalis to obtain MOIs of 103, 102, and 10 incubated at 37°C for 24 h. Times of collecting samples were at 1, 3, 5, and 24 h. Surviving bacterial cells were counted using serial dilutions in sterile saline. Then, dilutions were plated onto nutrient plates and incubated at 37°C for 24 h. Bacterial growth was measured by monitoring optical densities at 600 nm.
Bacteria and antibiotic sensitivity testing
In the current study, S. aureus and E. faecalis were isolated formerly from infected patients with dental decays and cavities. The isolates were identified biochemically using conventional methods (Table S1, Supplementary data) and were confirmed by Vitek 2 system.
An antibiotic sensitivity test was performed for the two isolated bacteria against a selection of twelve antibiotics (Fig. S1, Supplementary data). Qualitative data from the antibiograms (Table 1) revealed that bothS. aureus EG-AE3 and E. faecalis EF01 were resistant to at least eight antibiotics with a resistance percentage of 75% and 66% against the tested antibiotics, respectively. S. aureus resisted Nitrofurantoin, Amox0icillin, Gentamicin, Norfloxacin, Penicillin-G, Vancomycin, Ceftazidime, Cefotaxime, and Amikacin but was sensitive to Chloramphenicol and Aztreonam. E. faecalis EF01 resisted Chloramphenicol, Gentamicin, Norfloxacin, Penicillin-G, Ceftazidime, Aztreonam, Cefotaxime, and Amikacin. While, this isolate was susceptible to Nitrofurantoin, Amoxicillin, Ciprofloxacin, and Vancomycin.
Table (1):
Antibiotic sensitivity pattern of the isolated dental caries bacteria against a selection of twelve antibiotics.
Bacteria |
F |
AX |
C |
CIP |
GN |
NOR |
P |
VA |
CAZ |
ATM |
CTX |
AK |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
S. aureus strain EG-AE3 |
R* |
R |
S* |
I* |
R |
R |
R |
R |
R |
S |
R |
R |
E. faecalis EF01 |
S |
S |
R |
S |
R |
R |
R |
S |
R |
R |
R |
R |
Nitrofurantoin (F, 10 µg), Amoxicillin (AX, 25 µg), Chloramphenicol (C 30 µg), Ciprofloxacin (CIP 5 µg), Gentamicin (GN, 10 µg), Norfloxacin (NOR, 10 µg), Penicillin-G (P, 10 µg), Vancomycin (V, 30 µg), Ceftazidime (CAZ, 30 µg), Aztreonam (ATM, 30 µg), Cefotaxime (CTX, 30 µg) and Amikacin (AK, 30 µg). * Denotes for Resistant (R), Intermediate (I), and Susceptible (S).
Morphology of bacteriophages
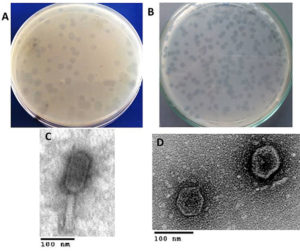
Two phages with different plaque morphologies (Fig. 1A & B) targeted dental caries causing bacteria, S. aureus, and E. faecalis were successfully obtained after a screening of 100 samples of saliva. Successive double soft-layer agar assays led to pure phages isolation,titrated and processed at 4°C. Fig. 1 shows clear plaques produced by vB_SauM-EG-AE3 and vB_EfaP-EF01 on S. aureus EG-AE3 and E. faecalis EF01 as host strains respectively. Phage vB_SauM-EG-AE3 produces large circular clear plaques with a diameter of 4 mm and large circular clear plaques were formed by vB_EfaP-EF01 but with 3 mm in diameter. The concentrations of phages were 2.5×1010 PFU/ml to 3.1×109 PFU/ml for vB_EfaP-EF01 and vB_SauM-EG-AE3 respectively.
Fig. 1. Plaques phenotypes and TEM morphology of the isolated phages. (A-B) Images of bacterial plaques formed by the isolated phage in top-agar lawns of S. aureus EG-AE3 and E. faecalis EF01, plaque appearance was detected and imaged after culturing 48 h on their hosts. (C-D) TEM micrographs of phages vB_SauM-EG-AE3 and vB_EfaP-Ef01 were negatively stained with 0.2% uranyl acetate as described in Materials and Methods. Scale bar= 100 nm.
Transmission Electron microscopy (Fig. 1C & D) allowed us to infer that vB_SauM-EG-AE3 and vB_EfaP-EF01 belong taxonomically to order Caudovirales. Dimensions of the isolated phages were measured and summarized in Table 2. The particle of vB_SauM-EG-AE3 had a contractile tail with 75 ±2 nm in length and head with a diameter of about 67 ±1 nm a typical member of Myoviridae family. While vB_EfaP-EF01 was a member of Podoviridae family whereas, the tail length was about 9 ± 0.5 nm and head diameter of about 58 ± 2 nm.
Table (2):
Dimensions of the isolated phages.
Phage |
Bacterial host |
Plaques diameter (mm) |
Head diameter (nm) |
Tail length (nm) |
Proposed family |
|---|---|---|---|---|---|
vB_SauM-EG-AE3 |
S. aureus EG-AE3 |
4 |
67 ± 1 |
75 ± 2 |
Myoviridae |
vB_EfaP-Ef01 |
E. faecalis EF01 |
3 |
58 ± 2 |
9 ± 0.5 |
Podoviridae |
Growth-kinetics and host range
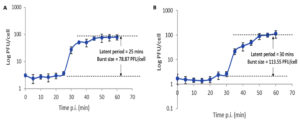
One-step growth kinetics of the isolated phages (Fig. 2) exhibited typical growth kinetics of most bacteriophages. Phages vB_SauM-EG-AE3 and vB_EfaP-EF01 gave burst sizes of 78.87 and 113.55 PFU/cell respectively with latent periods of 25 and 30 minutes. The host range of the isolated phages was estimated, and results were summarized in Table 3. Both phages established a narrow spectrum of lytic activity.
Table (3):
Host ranges of the isolated phages.
| Species | Strain ID number | Lysis by bacteriophage | |
|---|---|---|---|
| vB_SauM-EG-AE3 | vB_EfaP-Ef01 | ||
| S. aureus | EG-AE3 | + | – |
| EG-AE1 | + | – | |
| EG-AE2 | – | – | |
| SA101 | – | – | |
| SA1E | – | – | |
| BUH | – | – | |
| E. faecalis | Ef01 | – | – |
| Etfc1 | – | + | |
| Etfc2 | – | + | |
| Etfc3 | – | – | |
| Enterococcus faecium | 1 | – | – |
| 2 | – | – | |
| 3 | – | – | |
| Acinetobacter baumannii | Acint1 | – | – |
| Acint2 | – | – | |
| Acint3 | – | – | |
| E. coli | BE1 | – | – |
| BE2 | – | – | |
| BE3 | – | – | |
| Proteus spp. | Pr1 | – | – |
| Salmonella spp. | Sa1 | – | – |
| Shigella spp. | Sh1 | – | – |
“–,” no clearing; “+,” completely clear.
Fig. 2. One-step growth curves of phages vB_SauM-EG-AE3 (A) and vB_EfaP-Ef01 (B) on their corresponding hosts. Data shown are the mean of three replicates and error bars show the deviation in the values.
Thermal and pH stability
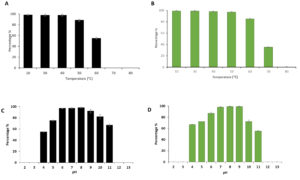
Thermal and pH stability patterns of phages vB_SauM-EG-AE3 and vB_EfaP-EF01 were tested based on residual phage titers after incubation under different pH values and temperatures (Fig. 3). The S. aureus phage (vB_SauM-EG-AE3) was thermostable at temperatures ranging from 10 to 60 °C, but no viable phages were detected after heating at ≥70°C for 60 min. Phage vB_SauM-EG-AE3 resisted a pH range between pH 4 and 11 for 1h. Similarly, no significant reduction in the E. faecalis phage (vB_EfaP-EF01) was observed at temperatures ranging from 10 to 60°C. But, after heating at 70°C for 60 min, phage titers decreased by 60% and no viable phages were detected after heating at 80°C for 60 min. Regarding the pH stability, both vB_SauM-EG-AE3 and vB_EfaP-EF01 phages were found to be resistant at a pH range of 4-11 after 60 min at 37 °C h (Fig. 3C-D).
Fig. 3. Thermal and pH tolerance test of phages vB_SauM-EG-AE3 and vB_EfaP-Ef01. (A-B) Thermal tolerance, and (C-D) pH stability of vB_SauM-EG-AE3 and vB_EfaP-Ef01 phages, respectively. Temperature experiments were performed for 1 h at pH 7. pH tolerance was performed for 60 min at 37°C. Data showed the percentages of the remaining phages after each treatment as normalized from the control. Data shown are the mean of three replicates and error bars show the deviation in the values.
Structural protein profiles
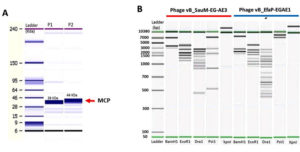
Molecular weights of bacteriophages’ structural polypeptides ranged from 10 to 81 kDa (Fig. 4A). Phages vB_SauM-EGAE3 and vB_EfaP-Ef01 harbored major structural proteins of 39 and 44 kDa, respectively. Additionally, four minor structural proteins were observed in each phage where three of them revealed similar profiles in vB_SauM-EGAE3 and vB_EfaP-Ef01, each phage-displayed unique specific band at varied position of 81 kDa and 32 kDa for vB_SauM-EGAE3 and vB_EfaP-Ef01, respectively.
Fig. 4. Agilent bioanalyzer gel-like images of (A) SDS-polyacrylamide gel electrophoretic profiles of vB_SauM-EG-AE3 and vB_EfaP-Ef01 structural proteins. Lane1: ladder ranged from 6 kDa to 240 kDa, lanes 2-3: Structural protein profiles of vB_SauM-EG-AE3 and vB_EfaP-Ef01 respectively, red rows represent the major capsid proteins of the isolated phages. (B) Restriction digestion profiles of the vB_SauM-EG-AE3 and vB_EfaP-Ef01 phages after digestion of DNA with BamHI, EcoRI, DraI, PstI, and KpnI. Lanes as shown on the figure, Ladder from 50 bp to 10380 bp.
Restriction digestion patterns
Phages vB_SauM-EGAE3 and vB_EfaP-EGAE1 exhibited different patterns of restriction digest profiles (Fig. 4B). The restriction endonucleases of BamHI, EcoRI, DraI, and PstI were able to digest the two genomes. While KpnI failed to cut both genomes (Fig. S2, Supplementary data), the digestion of bacteriophage genomes with DraI showed the most crowded pattern with total fragments of 15 bands for vB_EfaP-EGAE1 and 9 bands for vB_SauM-EGAE3. On other hand, BamHI showed the lowest digestion profile with 3 bands for vB_SauM-EGAE3 and 4 bands for vB_EfaP-EGAE1. The restriction enzyme patterns confirmed that these phages are double-stranded DNA viruses. For each phage, the genome size was predicted considering the total of the sizes of all fragments produced by restriction enzymes(Table S3, Supplementary data). The genome sizes were approx. 13.30 Kb and 15.60 Kb for phages vB_SauM-EGAE3, vB_EfaP-Ef01 respectively.
Biocontrol of dental caries-causing bacteria using the bacteriophages
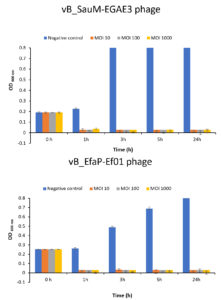
In the current study, different MOIs were used to control targeted pathogens in broth medium (Fig. 5). High values of MOI were used to be more enough to infect bacterial cells and to reduce the chance of bacterial cells regrowth. In broth medium, complete inhibition of bacterial growth was achieved using phages with higher and lower MOIs of 103, 102 and 10 after 1, 3, 5, and 24 hr at 37°C compared to control counts. Based on the stability and lytic activity of the isolated phages against S. aureus and E. faecalis, they were applied to control such hosts in broth medium using different MOIs. Findings showed that MOIs of 103, 102 and 10 appeared efficient to eradicate the bacterial growth and prevent their regrowth in broth medium. Thus, the involvement of these phages in therapy could be promising as alternatives to antibiotics.
Fig. 5. In vitro biocontrol assay of the isolated phages on their corresponding hosts at different MOIs. (A) Lytic activity of vB_SauM-EGAE3 phage on S. aureus EG-AE3, and (B) Lytic activity vB_EfaP-Ef01 phage on E. faecalis EF01. Each bacteria was challenged with the corresponding phage at different MOIs of 10, 102, and 103. Bacterial growth was determined by measuring the optical density at 600 nm.
Dental caries is one of a hygiene-related disease caused by decay-causing bacteria that produce acid resulting in damage in tooth enamel38. In the current study, two pathogenic bacteria, S. aureus and E. faecalis, which are related to dental caries, were isolated previously from infected patients with dental decays and cavities, which agrees with Ohara-Nemoto et al.39 and Wang et al.40. The isolated bacterial candidates were characterized microscopically and identified biochemically using conventional methods and were confirmed by Vitek 2 system according to previous studies41,42.
Recently, most bacteria have the potential to develop resistance against different classes of antibiotics. Antibiotic resistance is one of the top concerns that threaten global health43. Egypt is one of the countries where antibiotic remedies have fewer extreme limitations44,45 that enhances the chance for bacteria to resist antibiotics. In the current study, antibiotic sensitivity testing of S. aureus and E. faecalis against a selection of twelve antibiotics showed that the two isolates resisted at least eight of the tested antibiotics. Resistance mechanisms against antibiotics by S. aureus and E. faecalis were reported46,47.
Antibiotic resistance can be developed through mutations in chromosomal genes or by mobile genetic elements (horizontally acquired resistance)48. In that view, a resistance that is acquired through mutation, mechanism of horizontally acquired resistance, or overexpression of the drug efflux were discussed previously47. Hence, the current study used bacteriophages as an alternative strategy to control the spread of these organisms and mitigate dental caries.
Bacteriophages have been sought as one of the novel therapeutic approaches to control antibiotic-resistant pathogenic bacteria49,50. In the current study, two different phages (vB_SauM-EG-AE3 and vB_EfaP-EF01) targeted dental caries causing bacteria, S. aureus, and E. faecalis, respectively were isolated after a screening of 100 samples of saliva in concordance with previous study51. Phage vB_SauM-EG-AE3 belongs to the family Myoviridae, while vB_EfaP-EF01 belongs to Podoviridae.
Growth kinetics of the isolated phages exhibited typical growth kinetics of most bacteriophages. Phages vB_SauM-EG-AE3 and vB_EfaP-EF01 gave burst sizes of 78.87 and 113.55 PFU/cell respectively with latent periods of 25 and 30 minutes. These values, similar to those obtained in other studies with Staphylococcus and E. faecalis phages52,46,40 where the average burst size of such phages was approximately 36-122 PFU/ infected cell and latent period 25-50 min. additionally, a recent study showed a higher burst size of E. faecalis phage of 352 PFU/ infected cell21. Both vB_SauM-EG-AE3 and vB_EfaP-EF01 phages displayed a narrow spectrum of lytic activity. This could be attributed to the ability of the tested strains to develop resistance against these phages53,54. Narrow host range could be overcome using a cocktail of phages55. In addition to that cross infectivity of phages against different species and genera was investigated in the current study, and no lytic activity was shown.
The stability of phages under stressful environmental conditions promotes the application of phages as a bio-control agent in dental therapy. At temperatures between 10 and 60°C,and pH range between pH 4 and 11 for 1h.S. aureus phage was stable. Interestingly, no significant reduction in the E. faecalis phage was observed at temperatures ranging from 10 to 70° C or pH from 7 to 9. Previous studies showed stability of phages against S. aureus and E. faecalis under a wide range of temperature degrees (4–60°C) and resistancein acidic or alkaline pH (pH 3–12)21,56. These findings make the isolated phages highly potent to be used in clinical settings where they can be combined with alkaline disinfectants that are commonly used in the treatment of endodontic infections.
In the protein fractions obtained in the current study for two phages we detected the occurrence of main fractions with a molecular weight ranged from 10 to 81 kDa and the number of bands obtained ranged from 4 and 5 fractions for vB_SauM-EGAE3 and vB_EfaP-EGAE1 respectively. Seven protein bands were visualized on SDS-PAGE for purified S. aureus phage vB_SauS_SA2. The protein band of about 33 kDa was identified to be the major capsid protein by mass spectrometry, which is relatively smaller than the capsid protein (42 kDa) of phage SA257. While, SDS-PAGE analysis of the dissociated purified E. faecalis phage ϕEf11showed 11 well-resolved bands of proteins ranging from 27 to 85 kDa in size58. In taxonomic and phylogenic classifications, major capsid proteins, highly conserved among viruses, are often used.59,60. So, the close relation of vB_SauM-EGAE3 and vB_EfaP-Ef01 to the other phages of Gram-positive bacteria can, therefore, be speculated on.
Phages vB_SauM-EGAE3 and vB_EfaP-EGAE1 showed different patterns of restriction digest profiles and the genome sizes were approx. 13.30 Kb and 15.60 Kb for phages vB_SauM-EGAE3, vB_EfaP-Ef01 respectively. Kwiatek et al.61 characterized a virulent S. aureus phage (MSA6) which isolated from cow milk with mastitis and they indicated that the genome size of MSA6 of about 143 Kbp based on the KpnI and PstI restriction pattern and PFGE. Uchiyama et al.62 reported that the genome size of E. faecalis phage ϕEF24C was estimated to be about 143 kb. Phages vB_SauM-EGAE3 and vB_EfaP-EGAE1 were refractory for endonuclease enzyme digestion, such as KpnI. Interestingly, this mechanism, called an anti-restriction mechanism, is usual among phages of the Siphoviridae63,64 and Myoviridae65.
Phages developed various strategies of anti-restriction against restriction-modification (R-M) systems. Protection against DNA bacteriophages is given by staphylococcus (R-M) systems. These systems consist of methyltransferase DNA and endonuclease restrictions, which through methylation, protect self-DNA and inhibit foreign unmodified DNA in the same sequence. If the oligonucleotide sequence recognized by the enzyme is found, a DNA molecule will be hydrolyzed by the restriction endonucleases. As a consequence of point mutations or the acquisition in their genomes of the cognate methylase gene, a lack of endonuclease recognition sites in their genomes may be the strategy used by phages. Generally, methylation within the recognition sequence of one specific cytosine (C) or adenine (A) residue will prevent sequence cleavage by Type II restrictionendonuclease66,67.
Based on the stability and lytic activity of the isolated phages against S. aureus and E. faecalis, they were applied to control such hosts in broth medium using different MOI. Findings showed that MOI 10, 100 and 1000 appeared efficient to eradicate the bacterial growth and prevent their regrowth in broth medium. Thus, the involvement of these phages in therapy could be promising as alternatives to antibiotic. Using bacteriophages for the prophylaxis or treatment of oral infectious diseases was adopted in previous studies68,69,50. ListShieldTM and LISTEXTM P100, commercial phages, are approved to use and have gotten GRAS, Generally Recognized As Safe, status through FDA70, also a recent review studied possibilities of using bacteriophages against pathogenic bacteria with oral origin and found possible future uses of phages in dentistry71. A few promising studies attempted to use phages for treatment of dental infections caused by bacterial pathogens like S. aureus72 and E. faecalis28. Phages are overseen as a friendly biocontrol strategy and can be granted a commercial use. For example, ListShieldTM and LISTEXTM P100, commercial phages, are approved to use and have gotten GRAS status through FDA70. As a future work, we are trying to isolate more bacteriophages against dental caries causing bacteria and to produce a phage cocktail mix to be incorporated. Later this phage mix suspension can be used while dental flossing to reduce or eliminate dental caries bacteria into the interdental spaces. Moreover, the phage cocktail could be loaded on small sweety pleasant tablets that can be used daily, or merged with chewing gums and/ or incorporated on toothpaste.
Health concerns related to dental diseases have been exacerbated by antibiotic-resistant S. aureus and E. faecalis. Hence, bacteriophages could be proposed as an alternative strategy to mitigate the causative bacterial pathogens. In this study, two virulent phages were isolated for antibiotic-resistant S. aureus and E. faecalis, vB_SauM-EG-AE3belonged to family Myoviridae and vB_EfaP-EF01belonged to family Podoviridae morphologically. The two phages have narrow host ranges, low latent periods, strong pH and thermal resistance. More importantly, our findings demonstrate the efficacy of phages vB_SauM-EG-AE3 and vB_EfaP-EF01 for the inhibition of multidrug-resistant S. aureus and E. faecalis growth in vitro respectively. This research forms the basis for the therapeutic application of phages to manageS. aureus and E. faecalis infection.
Supplementary information
Additional file: Additional Table S1- S3. Additional Figure S1-S2.
ACKNOWLEDGMENTS
We gratefully acknowledge Dr. Omnia Badr, Faculty of Agricultural, Benha University, Egypt for the help in performing the restriction enzyme digestion experiment.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
AE designed and directed the project. MAN, NKE, YHE and AE performed the Experiments. MAN, NKE and AE analyzed the data. MAN, NKE and AE wrote the Initial draft. AE reviewed and edited the final manuscript. All authors discussed the Results and commented on the manuscript.
FUNDING
Funding for this paper was provided by Botany and Microbiology Department, Benha University, Egypt.
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the manuscript.
- Jett BD, Huycke MM, Gilmore MS. Virulence of Enterococci. Clin Microbiol Rev. 1994;7(4):462-478.
Crossref - Gu YH, Yamasita T, Kang KM. Subchronic oral dose toxicity study of Enterococcus Faecalis 2001 (EF 2001) in mice. Toxicol Res. 2018;34(1):55-63.
Crossref - Giraffa G. Enterococci from foods. FEMS Microbiol Rev. 2002;26(2):163-171.
Crossref - Alghamdi F, Shakir M. The Influence of Enterococcus faecalis as a Dental Root Canal Pathogen on Endodontic Treatment: A Systematic Review. Cureus. 2020;12(3):e7257.
Crossref - Arias CA, Contreras GA, Murray BE. Management of multidrug-resistant enterococcal infections. Clin Microbiol Infect. 2010;16(6):555-562.
Crossref - Weiner LM, Webb AK, Limbago B, et al. Antimicrobial-Resistant Pathogens Associated with Healthcare-Associated Infections: Summary of Data Reported to the National Healthcare Safety Network at the Centers for Disease Control and Prevention, 2011-2014. Infect Control Hosp Epidemiol. 2016;37(11):1288-1301.
Crossref - Narayanan L, Vaishnavi C. Endodontic microbiology. J Conserv Dent. 2010;13(4):233-239.
Crossref - Vidana R, Sullivan A, Billstrom H, Ahlquist M, Lund B. Enterococcus faecalis infection in root canals – host-derived or exogenous source? Lett Appl Microbiol. 2011;52(2):109-115.
Crossref - Walsh FM, Amyes SGB. Microbiology and drug resistance mechanisms of fully resistant pathogens. Curr Opin Microbiol. 2004:7(5):439-444.
Crossref - Huycke MM, Sahm DF, Gilmore MS. Multiple-drug resistant enterococci: The nature of the problem and an agenda for the future. Emerg Infect Dis. 1998;4(2):239-249.
Crossref - Tong SYC, Davis JS, Eichenberger E, Holland TL, Fowler VG. Staphylococcus aureus infections: Epidemiology, pathophysiology, clinical manifestations, and management. Clin Microbiol Rev. 2015;28(3):603-661.
Crossref - Smith AJ, Robertson D, Tang MK, Jackson MS, MacKenzie D, Bagg J. Staphylococcus aureus in the oral cavity: A three-year retrospective analysis of clinical laboratory data. Br Dent J. 2003;195(12):701-703.
Crossref - Bagg J, Sweeney MP, Wood KH, Wiggins A. Possible role of staphylococcus aureus in severe oral mucositis among elderly dehydrated patients. Microb Ecol Health Dis. 1995;8(2):51-56.
Crossref - Gordon RJ, Lowy FD. Pathogenesis of Methicillin-Resistant Staphylococcus aureus Infection. Clin Infect Dis. 2008;46(Suppl 5):S350-S359.
Crossref - Gardete S, Tomasz A. Mechanisms of vancomycin resistance in Staphylococcus aureus. J Clin Invest. 2014;124(7):2836-2840.
Crossref - Lister JL, Horswill AR. Staphylococcus aureus biofilms: Recent developments in biofilm dispersal. Front Cell Infect Microbiol. 2014;4:178.
Crossref - Wang I-N, Smith DL, Young R. Holins: The Protein Clocks of Bacteriophage Infections. Annu Rev Microbiol. 2000;54:799-825.
Crossref - Szafranski SP, Winkel A, Stiesch M. The use of bacteriophages to biocontrol oral biofilms. J Biotechnol. 2017;250:29-44.
Crossref - Burrowes B, Harper DR, Anderson J, McConville M, Enright MC. Bacteriophage therapy: Potential uses in the control of antibiotic-resistant pathogens. Expert Rev Anti Infect Ther. 2011;9(9):775-785.
Crossref - Patey O, McCallin S, Mazure H, Liddle M, Smithyman A, Dublanchet A. Clinical indications and compassionate use of phage therapy: Personal experience and literature review with a focus on osteoarticular infections. Viruses. 2019;11(1):18.
Crossref - Lee D, Im J, Na H, Ryu S, Yun CH, Han SH. The Novel Enterococcus Phage vB_EfaS_HEf13 Has Broad Lytic Activity Against Clinical Isolates of Enterococcus faecalis. Front Microbiol. 2019;10:2877.
Crossref - Kazmierczak Z, Piotrowicz A, Owczarek B, et al. Molecular imaging of T4 phage in mammalian tissues and cells. Bacteriophage. 2014;4(1):e28364.
Crossref - O’Flaherty S, Coffey A, Edwards R, Meaney W, Fitzgerald GF, Ross RP. Genome of Staphylococcal Phage K: A New Lineage of Myoviridae Infecting Gram-Positive Bacteria with a Low G+C Content. J Bacteriol. 2004;186(9):2862-2871.
Crossref - Estrella LA, Quinones J, Henry M, et al. Characterization of novel Staphylococcus aureus lytic phage and defining their combinatorial virulence using the OmniLog® system. Bacteriophage. 2016;6(3):e1219440.
Crossref - Bachrach G, Leizerovici-Zigmond M, Zlotkin A, Naor R, Steinberg D. Bacteriophage isolation from human saliva. Lett Appl Microbiol. 2003;36(1):50-53.
Crossref - Bolocan AS, Upadrasta A, De Almeida Bettio PH, et al. Evaluation of phage therapy in the context of enterococcus faecalis and its associated diseases. Viruses. 2019;11(4):366.
Crossref - Lossouarn J, Briet A, Moncaut E, et al. Enterococcus faecalis countermeasures defeat a virulent picovirinae bacteriophage. Viruses 2019;11(1):48.
Crossref - Paisano AF, Spira B, Cai S, Bombana AC. In vitro antimicrobial effect of bacteriophages on human dentin infected with Enterococcus faecalis ATCC 29212. Oral Microbiol. Immunol. 2004;19(5):327-330.
Crossref - Pincus DH. Microbial identification using the bioMerieux VITEK® 2 system. Encycl. Rapid Microbiol. Methods. 2010.
- Biemer JJ. Antimicrobial susceptibility testing by the Kirby-Bauer disc diffusion method. Ann Clin Lab Sci. 1973;3:135-140.
- Wayne P. Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing; 29th ed. CLSI supplement M100. ClinLab Stand Institute. 2019.
- Van Twest R, Kropinski AM. Bacteriophage enrichment from water and soil. Methods Mol Biol. 2009;501:15-21.
Crossref - Huang C, Shi J, Ma W, et al. Isolation, characterization, and application of a novel specific Salmonella bacteriophage in different food matrices. Food Res Int. 2018;111:631-641.
Crossref - Ackermann HW. Bacteriophage Electron Microscopy, in: Adv Virus Res. 2012;82:1-32.
Crossref - Kropinski AM, Mazzocco A, Waddell TE, Lingohr E, Johnson RP. Enumeration of bacteriophages by double agar overlay plaque assay. Methods Mol Biol. 2009;501:69-76.
Crossref - Clokie MRJ, Kropinski AM. Bacteriophages : methods and protocols. Methods in Molecular Biology. 2009;501.
Crossref - Danis-Wlodarczyk K, Olszak T, Arabski M, et al. Characterization of the newly isolated lytic bacteriophages KTN6 and KT28 and their efficacy against Pseudomonas aeruginosa biofilm. PLoS One. 2015;10(5):e0127603.
Crossref - CDC. Hygiene-related Diseases | Hygiene | Healthy Water | CDC. Cent. Dis. Control Prev. 2016.
- Ohara-Nemoto Y, Haraga H, Kimura S, Nemoto TK. Occurrence of staphylococci in the oral cavities of healthy adults and nasal-oral trafficking of the bacteria. J Med Microbiol. 2008;57(1):95-99.
Crossref - Wang QQ, Zhang CF, Chu CH, Zhu XF. Prevalence of Enterococcus faecalis in saliva and filled root canals of teeth associated with apical periodontitis. Int J Oral Sci. 2012;4(1):19-23.
Crossref - Ligozzi M, Bernini C, Bonora MG, De Fatima M, Zuliani J, Fontana R. Evaluation of the VITEK 2 system for identification and antimicrobial susceptibility testing of medically relevant gram-positive cocci. J Clin Microbiol. 2002;40(5):1681-1686.
Crossref - Nonhoff C, Rottiers S, Struelens MJ. Evaluation of the Vitek 2 system for identification and antimicrobial susceptibility testing of Staphylococcus spp. Clin Microbiol Infect. 2005;11(2):150-153.
Crossref - CDC. Biggest Threats and Data | Antibiotic/Antimicrobial Resistance | CDC. 2019.
- Esmael A, Hassan MG, Amer MM, et al. Antimicrobial activity of certain natural-based plant oils against the antibiotic-resistant acne bacteria. Saudi J Biol Sci. 2020;27(1):448-455.
Crossref - Esmat MM, Abdelhamid AG, Abo-ELmaaty SA, et al. Antibiotics and phage sensitivity as interventions for controlling Escherichia coli isolated from clinical specimens. J Pure Appl Microbiol. 2017;11(4):1749-1755.
Crossref - Guo Y, Song G, Sun M, Wang J, Wang Y. Prevalence and Therapies of Antibiotic-Resistance in Staphylococcus aureus. Front Cell Infect Microbiol. 2020;10:107.
Crossref - Miller WR, Munita JM, Arias CA. Mechanisms of antibiotic resistance in enterococci. Expert Rev Anti Infect Ther. 2014;12(10):1221-1236.
Crossref - Foster TJ. Antibiotic resistance in Staphylococcus aureus. Current status and future prospects. FEMS Microbiol Rev. 2017;41(3):430-449.
Crossref - El-Dougdoug NK, Cucic S, Abdelhamid AG, et al. Control of Salmonella Newport on cherry tomato using a cocktail of lytic bacteriophages. Int J Food Microbiol. 2019;293:60-71.
Crossref - Shlezinger M, Khalifa L, Houri-Haddad Y, et al. Phage Therapy: A New Horizon in the Antibacterial Treatment of Oral Pathogens. Curr Top Med Chem. 2017; 17(10):1199-1211.
Crossref - Edlund A, Santiago-Rodriguez TM, Boehm TK, Pride DT. Bacteriophage and their potential roles in the human oral cavity. J Oral Microbiol. 2015;7:27423.
Crossref - Alves DR, Gaudion A, Bean JE, et al. Combined use of bacteriophage K and a novel bacteriophage to reduce Staphylococcus aureus biofilm formation. Appl Environ Microbiol. 2014;80(21):6694-6703.
Crossref - Leon M, Bastias R. Virulence reduction in bacteriophage resistant bacteria. Front Microbiol. 2015;6:343.
Crossref - Petty NK, Evans TJ, Fineran PC, Salmond GPC. Biotechnological exploitation of bacteriophage research. Trends Biotechnol. 2007;25(1):7-15.
Crossref - Goodridge LD, Bisha B. Phage-based biocontrol strategies to reduce foodborne pathogens in foods. Bacteriophage. 2011;1(3):130-137.
Crossref - Ullah RS, Andleeb S, Raza T, Jamal M, Mehmood K. Effectiveness of a Lytic Phage SRG1 against Vancomycin-Resistant Enterococcus faecalis in Compost and Soil. Biomed Res Int. 2017;2017:9351017.
Crossref - Wang J, Zhao F, Sun H, et al. Isolation and characterization of the Staphylococcus aureus bacteriophage vB_SauS_SA2. AIMS Microbiol. 2019;5(3):285-307.
Crossref - Stevens RH, Porras OD, Delisle AL. Bacteriophages induced from lysogenic root canal isolates of Enterococcus faecalis. Oral Microbiol Immunol. 2009;24(4):278-284.
Crossref - Bamford DH, Grimes JM, Stuart DI. What does structure tell us about virus evolution? Curr Opin Struct Biol. 2005;15(6):655-663.
Crossref - Hambly E, Tetart F, Desplats C, Wilson WH, Krisch HM, Mann NH. A conserved genetic module that encodes the major virion components in both the coliphage T4 and the marine cyanophage S-PM2. Proc Natl Acad Sci USA. 2001;98(20):11411-11416.
Crossref - Kwiatek M, Parasion S, Mizak L, Gryko R, Bartoszcze M, Kocik J. Characterization of a bacteriophage, isolated from a cow with mastitis, that is lytic against Staphylococcus aureus strains. Arch Virol. 2012;157(2):225-234.
Crossref - Uchiyama J, Rashel M, Maeda Y, et al. Isolation and characterization of a novel Enterococcus faecalis bacteriophage φEF24C as a therapeutic candidate. FEMS Microbiol Lett. 2008;278(2):200-206.
Crossref - Kruger DH, Barcak GJ, Smith HO. Abolition of DNA recognition site resistance to the restriction endonuclease EcoRII. Biomed Biochim Acta. 1988;47(1):K1-5. PMID: 2839163.
- Marek A, Pyzik E, Stepien-Pysniak D, Urban-Chmiel R, Nowaczek A. Characterization of bacteriophages and their carriage in Staphylococcus aureus isolated from broilers in Poland. Br Poult Sci. 2019;60(4):373-380.
Crossref - Kesik-Szeloch A, Drulis-Kawa Z, Weber-Dabrowska B, et al. Characterising the biology of novel lytic bacteriophages infecting multidrug resistant Klebsiella pneumoniae. Virol J. 2013;10.
Crossref - Huang LH, Farnet CM, Ehrlich KC, Ehrlich M. Digestion of highly modified bacteriophage DNA by restriction endonucleases. Nucleic Acids Res. 1982;10(5):1579-1591.
Crossref - Lakhundi S, Zhang K. Methicillin-Resistant Staphylococcus aureus: Molecular Characterization, Evolution, and Epidemiology. Clin Microbiol Rev. 2018;13(4)e00020-18.
Crossref - Dalmasso M, De Haas E, Neve H, et al. Isolation of a novel phage with activity against Streptococcus mutans biofilms. PLoS One. 2015:10(9):e0138651.
Crossref - Parasion S, Kwiatek M, Gryko R, Mizak L, Malm A. Bacteriophages as an alternative strategy for fighting biofilm development. Polish J Microbiol. 2014;63(2)-137-145.
Crossref - Lu TK, Koeris MS. The next generation of bacteriophage therapy. Curr Opin Microbiol. 2011;14(5):524-531.
Crossref - Steier L, De Oliveira SD, De Figueiredo JAP. Bacteriophages in dentistry-state of the art and perspectives. Dent J. 2019;7(1):6.
Crossref - Gabisonia T, Giorgadze I, Topuria N, et al. Fersis-Phage against Purulent-Inflammatory (Staphylococcal, Streptococcal) Pathologies. Arch Clin Microbiol. 2019;10;1:89:1-6.
Crossref
© The Author(s) 2021. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.