ISSN: 0973-7510
E-ISSN: 2581-690X
Petroleum crude oil is transported on a global scale through marine vessels and barges by the sea route. Oil spills into the marine environment are known to cause long term effects on the health of marine life and human beings in addition to harming the eco-system. In this study, petroleum refinery effluent samples were collected and analyzed for their physico-chemical properties. Thirty two bacterial strains were isolated by direct isolation and post enrichment in crude oil. Among these, fourteen isolates could utilize petroleum hydrocarbons as sole carbon source on Bushnell Hass Agar plates supplemented with crude oil within 2 days. Out of these fourteen strains, four could decolorize 2,6-dichlorophenol indophenol within 36 h completely. They were identified as Bacillus cereus WD22, Pseudomonas aeruginosa WD23 and WDE11 (2 strains) and Acinetobacter baumannii OCB1 by 16sRNA sequencing. All four strains could tolerate salinity up to 4.0%w/v. The reduction in total petroleum hydrocarbons in sea water spiked with crude oil (1.0%v/v) supplemented with trace amounts of glucose and yeast extract was studied by gravimetric analysis. P. aeruginosa WD23 degraded 27.25% of supplied petroleum crude oil under limited nutrient conditions in seawater in 15 days.
Petroleum crude oil, Sea water, Petroleum refinery effluent, Acinetobacter baumannii, Bacillus cereus, Pseudomonas aeruginosa.
The occurrence of oil spills is now a common issue of concern around the world. Some of the most well-known oil spills that have taken place in last four decades include the Exxon Valdez in Alaska (1989), the ‘Gulf War’ in Kuwait (1991), the Erica in France (1999); the Aegean Sea in Galicia, Spain (1992) and the Prestige in Spain and France (2002)1. Recently, oil spills have occurred in India such as in Goa, West Coast of India in 20052, Mumbai, Maharashtra in 20103 and Ennore, Tamil Nadu in 20174. Also, the transfer of petroleum crude oil (PCO) from Arabian countries such as Saudi Arabia and Iran to India is by the sea route through tanker vessels.
The release of petroleum hydrocarbons into the marine environment has impact on marine ecosystems, adverse affects on human health and results in death of marine life as some of the polyaromatic hydrocarbons are known to be potent carcinogens, teratogens and mutagens5.The potential hazardous nature of components of PCO in oil spills continues to harm marine life and human health and damages the ecosystem until they are removed from the seawater. Study conducted by Han et al5 showed high concentration of polyaromatic hydrocarbons in the sea water at Ennore sea coast after 2-3 months of the oil spill, inspite of using physical methods to clear up the spill.
There are various physico-chemical methods employed for treatment of oil spills such as advection, diffusion, dissolution and emulsification but these methods offer more disadvantages over advantages6. Physical methods of treatment are expensive whereas the chemical methods are less efficient and alter the natural ecosystem7.
Biodegradation of petroleum hydrocarbons in oil spills by indigenous microorganisms is presently gaining significance as a potential treatment method. Microorganisms utilize the components of PCO as a source of nutrition and convert them into simpler compounds. The growth of microorganisms in stressful conditions can be stimulated by addition of easily assimilated carbon and nitrogen sources8. Bioremediation offers benefits such as economy, reliability and eco-friendly nature over the physico-chemical methods9.
The present study deals with the isolation and screening of petroleum hydrocarbon degrading bacteria from petroleum refinery effluent (PRE). The physico-chemical characteristics of PRE were studied. The best degraders were identified by 16S RNA sequencing and characterized. The ability of the identified bacteria to degrade petroleum hydrocarbons in seawater supplemented with PCO and minimal amount of glucose and yeast extract was determined by gravimetric analysis.
Collection of PCO and PRE
PCO was collected from a petroleum refinery industry dumping yard situated in Mangalore, Karnataka, India (12.9951°N, 74.8094°E). PRE from refinery industry is subjected to treatment involving oil separation, activated sludge process and tertiary treatment post which it is sent to the refinery industry dumping yard. PRE was collected from the dumping yard in sterile plastic containers at two different seasons (winter and rainy).
Substrates and Chemicals
The redox indicator 6-dichlorophenol indophenol (DCPIP) was obtained from SRL (SRL chemicals, Bangalore, India). All other chemicals, solvents and reagents used in the study were of analytical grade, unless mentioned.
Physico-chemical characterisation of PRE
The physico-chemical characteristics of PRE at both seasons such as pH, total organic carbon (TOC), total dissolved solids (TDS), phosphates, nitrates, phenolics, conductivity and metal composition were analysed. The pH was determined by digital pH meter. The TOC, TDS, phosphates and nitrates were determined7. Conductivity was determined using Conductivity/TDS meter (Wagtech WEDIST6). The metal composition was analysed by using standard atomic absorption spectrophotometer (Elico, India).
Quantitative Analysis of Bacteria
The quantitative analysis of bacteria in the PRE samples was performed to estimate the percentage of oil utilizing bacteria (OUB) in total viable count (TVC)10.
Determination of TVC
PRE was diluted (1:1) and spread plated (50µl) onto nutrient agar medium. The plates were incubated at 37±1°C for 24 – 48 h. The bacteria were enumerated and TVC was determined as CFU/ml.
Determination of OUB
Diluted PRE (50µl) was spread plated onto Bushnell Haas Agar (BHA) medium spread with PCO (100µl). The plates were incubated at 37±1°C for 24 – 48 h. The bacteria were enumerated and OUB was determined as CFU/ml.
Percentage of OUB in TVC
Percentage of OUB in TVC was calculated as per the following formula:
Percentage of OUB in TVC = (OUB / TVC) x 100 …(1)
Isolation of bacteria from PRE
Isolation Post Enrichment in PCO
The bacteria in the PRE was enriched by addition of PRE (1 ml) to nutrient broth supplemented with 1.0%v/v PCO for 2 days followed by enrichment in Bushnell Haas (BH) broth with 1.0%v/v PCO as the sole carbon source for 6 days10. These steps were carried out three times. Enrichment was done at 27°C and 37°C. The enriched medium was spread plated onto nutrient agar medium and incubated at 37±1°C for 72h. Pure cultures of the isolates were sub cultured and stored as glycerol stocks at -20°C.
Growth of enriched isolates on Malachite Green Agar
Bacteria isolated post enrichment were streaked onto Malachite Green Agar, selective for Pseudomonas aeruginosa and incubated at 37±1°C for 48h.
Direct Isolation
PRE was serially diluted till 10-12 and spread plated onto nutrient agar plates. The plates were incubated at 37±1°C for 24 – 48h. Distinct bacterial colonies were isolated based on morphological characteristics and stored as glycerol stocks at -20°C.
Preliminary screening of PCO degraders
The isolates obtained from direct isolation as well as post enrichment were streaked on BH plates overlaid with 100µl of PCO and incubated at 37°C for 6 days. The isolates which grew on the agar plates were confirmed as hydrocarbon degraders. The zone of clearance around the degraders, grown on BHA plate was observed.
Screening of bacteria with good hydrocarbon remediating potential
Isolates showing zone of clearance within 2 days of incubation were selected for the estimation of oil degrading efficiency. The isolates were grown in BH broth supplemented with 1.0%v/v PCO along with DCPIP indicator (0.4%w/v) at 37±1°C for 5 days. During microbial oxidation of hydrocarbons, the electrons are transferred to DCPIP which becomes colourless (reduced) from blue (oxidised). This is estimated by measuring the absorbance at 600 nm11.
Identification and Biochemical Characterization of best degraders
Best degraders were identified by 16S RNA sequencing. The 16S sequence obtained was analyzed by BLAST. Phylogenetic tree was constructed using neighbour joining method considering 16S sequences of bacteria showing maximum homology using Bioedit software. The 16S sequences were then deposited in NCBI-GenBank and accession numbers were obtained.
Growth of identified bacteria at different culture conditions such as temperature (4°C, 25°C, 37°C and 45°C), pH (2.0 -12) and NaCl (4.0 – 12.0%w/v) was checked. Also, the different biochemical tests such as oxidase, catalase, urease, methyl red, Voges Proskauer, Indole, H2S production, ortho-Nitrophenyl-β-galactoside (ONPG), nitrate reduction, esculin hydrolysis, utilization tests (citrate, lysine, ornithine, malonate) and sugar hydrolysis tests (arabinose, xylose, adonitol, rhamnose, cellobiose, melibiose, sachharose, raffinose, trehalose, glucose, maltose, fructose, sucrose and lactose) were performed12.
Degradation of petroleum hydrocarbons of PCO in sea water by identified bacteria
Identified bacteria (16 h old, 5.0%v/v) containing 1010 CFU/ml cells were inoculated into sterile sea water (100 ml) supplemented with 1%v/v PCO, 1 g/L glucose and 0.05 g/L yeast extract at concentration. These were incubated at 27°C for 15 days at an agitation speed of 100 rpm. Sea water with the supplemented nutrients and without the inoculum was used as control.
After 15 days, the growth of bacteria was checked by measuring optical density at 600 nm. PCO from the culture broth as well as the control was extracted with hexane in the ratio 20:1 in an extraction funnel. The solvent fraction was passed through anhydrous sodium sulfate to remove moisture content and was evaporated completely at 50°C13. The weight of the residual PCO was noted down. The percentage degradation of PCO was calculated as follows:
% Degradation = Wi – Wf/Wi x 100 …(2)
Where Wi and Wf are the weights of added and residual PCO after 15 days of incubation
Physico-chemical characterisation of PRE
Physico-chemical characterisation of treated PRE collected at summer (PRE1) and winter (PRE2) showed that all the parameters tested were high for PRE1 and low for PRE2 (Table 1). This is because the sump where the treated PRE is stored in the dumping yard is uncovered and rain water could have diluted the treated PRE.
The treated PRE were rich in TDS, TOC and phosphates, whereas the amount of nitrates was found to be relatively less. The phenol content was also measured to be low (Table 1). Out of the heavy metals tested, high concentration of magnesium and cadmium, low concentration of copper, nickel, manganese and zinc and trace amount of lead were found to be present. Mercury was observed at below detection limits
Table (1):
Profile of PRE from refinery industry at Mangalore.
| Parameter | Amount | |
|---|---|---|
| PRE1 | PRE2 | |
| Physico chemical | ||
| pH | 7.3 | 7.9 |
| Conductivity (mS) | 3.05 | 1.4 |
| TDS (mg/L) | 1943 | 770 |
| Inorganics | ||
| Phosphates (mg/L) | 48.22 | 31.8 |
| Nitrates (mg/L) | 0.061 | 0.016 |
| Phenolics (mg/L) | 1.29 | BDL |
| Organics | ||
| TOC (mg/L) | 10.76 | 10.67 |
| Heavy metals | ||
| Mercury | BDL* | BDL |
| Lead | 0.001 | BDL |
| Cadmium | 2.173 | 1.091 |
| Manganese | 0.232 | 0.075 |
| Nickel | 0.084 | 0.045 |
| Magnesium | 16.89 | 15.67 |
| Zinc | 0.082 | 0.045 |
| Copper | 0.103 | 0.045 |
Quantitative Analysis of Bacteria
TVC and OUB in PRE was more in summer than in the winter (Table 2). This is because the temperature of Mangalore in summer (29-30°C) is higher than in the rainy season (25-26°C). The TVC content in PRE in summer and winter was 1×105 and 5×103 CFU/ml respectively. The TVC content in untreated effluent from ONGC sites was found to be in the range 3×106 – 4×108 CFU/ml10. The percentage average of OUB in TVC in PRE in summer and winter was 22.1% and 20% respectively.
Table (2):
Bacterial load of PRE from refinery industry at Mangalore.
Sample |
TVC (CFU/ml) |
OUB (CFU/ml) |
% |
|---|---|---|---|
PRE1 |
1×105 |
2.21×104 |
22.1 |
PRE2 |
5×103 |
1×103 |
20 |
Isolation post enrichment in PCO
Five isolates designated as 1, 2, 3, 4 and 5 were isolated post enrichment in PCO based on colony characteristic differences on the nutrient agar (Table 3). Isolates 1-3 and 4-5 were obtained from enrichment studies at 27°C and 37°C respectively. All the isolates were Gram negative bacilli. When they were grown in nutrient broth they produced a blue green pigment within 24 h of incubation. This is a characteristic of P. aeruginosa, a Gram negative bacillus which produces pyocyanin, a blue green phenazine pigment14.
Table (3):
Morphological and Colony characteristics of isolates obtained post enrichment in PCO.
| Sl No | Grams stain | Shape | Colony Morphology | ||||
|---|---|---|---|---|---|---|---|
| Form | Elevation | Margin | Color$ | Size# | |||
| 1 | – | B* | Circular | Convex | Entire | Cr | ++ |
| 2 | – | B | Circular | Flat | Entire | Cr | + |
| 3 | – | B | Circular | Convex | Entire | Cr | ++ |
| 4 | – | B | Circular | Convex | Entire | W | +++ |
| 5 | – | B | Circular | Convex | Und* | P | + |
*B – Bacilli
*Irregul – Irregular, Crater – Crateriform, Und – Undulate
$Colour: Cr – Cream, W – White, P – Pink
#Size: + – Very Small, ++ – Small, +++ – Big
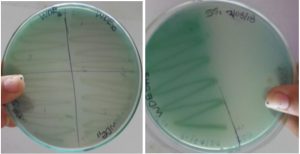
Growth of enriched isolates on Malachite Green Agar
The enriched isolates grew on Malachite Green agar, selective for P. aeruginosa (Fig. 1). P. aeruginosa is known to exhibit different morphologies such as small-colony, rough small-colony, wrinkled, autoaggregating cells and rugose on solid medium15. Pyocyanin is a known potent antimicrobial inhibitor that can act against a wide range of microorganisms16. Bacteria belonging to Pseudomonas genera are reported to be one of the most predominant polyaromatic hydrocarbons (PAH) degrading bacteria isolated from PAH contaminated environmental sites17-19.This could be because pyocyanin, produced by P. aeruginosa inhibits most of the bacteria growing with it during selective enrichment. Hence, direct isolation of bacteria from PRE samples was carried out to observe the diversity of its bacterial population
Direct isolation
On direct plating of PRE samples on nutrient agar a total of 27 bacterial isolated were selected on the basis of difference in colony characteristics (Table 4). They were designated as WD1 to WD26 and OCB1. Isolates WD1-WD15 were obtained from PRE1 whereas the rest were isolated from PRE2.
Table (4):
Morphological and Colony charecteristics of isolates obtained from direct isolation.
| Sl NO | Grams stain | Shape* | Colony Morphology | ||||
|---|---|---|---|---|---|---|---|
| Form | Elevation | Margin | Colour$ | Size# | |||
| WD1 | – | B | Irregul | Crater | Und | O | +++ |
| WD2 | – | B | Circular | Convex | Entire | Cr | ++ |
| WD3 | + | B | Circular | Flat | Curled | W | ++ |
| WD4 | – | B | Circular | Flat | Entire | W | + |
| WD5 | – | B | Circular | Raised | Entire | O | ++ |
| WD6 | – | B | Irregul | Crater* | Und* | Cr | +++ |
| WD7 | – | B | Circular | Flat | Entire | Y with O centre | ++ |
| WD8 | – | B | Circular | Convex | Entire | W with O center | +++ |
| WD9 | + | C | Circular | Raised | Curled | O | +++ |
| WD10 | – | B | Circular | Convex | Entire | W | +++ |
| WD11 | – | B | Circular | Flat | Entire | Cr | + |
| WD12 | + | SC | Circular | Raised | Lobate | Bright Y | + |
| WD13 | – | B | Irregul | Crater | Und | W | +++ |
| WD14 | + | B | Circular | Raised | Entire | Bright P | ++ |
| WD15 | + | B | Irregul | Crater | Und | O with P center | ++ |
| WD16 | – | B | Irregul | Crater | Lobate | W | +++ |
| WD17 | + | C | Circular | Convex | Entire | W mucoid | + |
| WD18 | – | B | Circular | Convex | Entire | Cr | ++ |
| WD19 | – | B | Circular | Convex | Entire | W with O center | + |
| WD20 | – | B | Irregul | Crater | Und | W | +++ |
| WD21 | + | SC | Round | Raised | Lobate | Pale Y | + |
| WD22 | + | B | Irregul | Crater | Lobate | Cottony W | +++ |
| WD23 | – | B | Circular | Convex | Entire | W with Bl center | + |
| WD24 | – | B | Circular | Convex | Und | P | + |
| WD25 | + | B | Circular | Raised | Lobate | Bright O | ++ |
| WD26 | + | SC | Circular | Convex | Entire | Light Y with Bl center | ++ |
| OCB1 | – | CB | Circular | Convex | Entire | Pale Y mucoid | + |
*Shape B – Bacilli, C – Cocci SC – Staphylococci, CB – coccobacilli
*Irregul – Irregular, Crater – Crateriform, Und – Undulate
$Colour Cr – Cream, W – White, O – Orange, P – Pink, Bl – Black, Y -Yellow
#Size + – Very Small, ++ – Small, +++ – Big
On direct isolation of bacterial isolates (27), it was observed that bacterial population was dominated by Gram negative bacilli (16). Eighty four percent of isolated bacteria from Brazilian soil, water and marine water contaminated with PAH were found to be Gram negative20. The rest of the isolates were found to be Gram positive bacilli (5), Gram positive cocci (5) and Gram negative coccobacilli (1). Gram positive rods with pink, orange and yellow pigmentation could be carotenoid producing Bacillus species21. Majority of the bacteria isolated from petroleum refinery waste of IOCL oil refinery, Assam, India collected during June also belonged to Pseudomonas and Bacillus genera20.
On comparison of colony characteristics of enriched isolates with direct isolates, it was observed that isolates 1, 2, 3, 4 and 5 had colony characteristics similar to WD2, WD11, WD18, WD13 and WD24 respectively. Hence they were re-designated as WDE2, WDE11, WDE18, WDE13 and WDE24 respectively.
Preliminary screening of PCO degraders
The bacterial isolates obtained post enrichment and from direct isolation were grown on BHA plates spread with PCO and observed periodically for 2 days till 6 days. Fourteen isolates (Fig. 2) grew on the BHA plate with in 2 days. The isolates WD6, WD7, WD24 and OCB1 gave the maximum zone of clearance. It was observed that WD22 and WD23 grew on the BH plates but failed to clear PCO around their colonies. Since, PCO is the sole carbon source that is supplied to the bacteria, all the bacteria could utilise PCO as the sole source of carbon. Surprisingly, only one enriched bacterial isolate (WDE11) grew with 2 days on the plate. Also, the rest of the isolates (19) could utilise PCO as sole carbon source by 6 days.
Fig. 2. Bacterial isolates exhibiting growth and zone of clearance on BH plates supplemented with PCO, at 2 days of incubation.
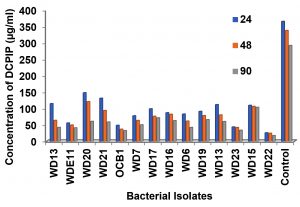
Screening of bacteria with good hydrocarbon remediating potential
The isolates which grew on BHA plates by 2 days were screened further by growing them in nutrient broth containing 1.0%v/v PCO and DCPIP (0.4%w/v). The reduction in color of DCPIP was measured spectrophotometrically at 600 nm. From Fig. 3 it was observed that culture WD22, WD23, WDE11 and OCB1 decolorised DCPIP rapidly within 24h. Complete decolorisation of DCPIP can be considered at a concentration of 30µg/ml. Similar studies showed bacterium HUB1 isolated from untreated petroleum refinery effluent could decolorize DCPIP at 120 h10.
Identification of best degraders of PCO
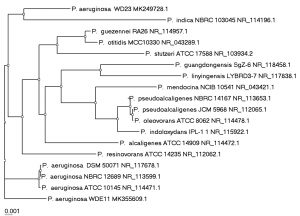
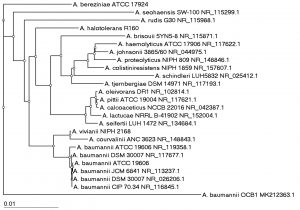
The identification of WD22, WD23, OCB1 and WDE11 by molecular sequencing of 16S RNA gene was performed and the gene was analyzed by BLAST and a phylogenetic tree was plotted using isolates showing maximum homology. The sequence of WD22 showed maximum similarity (80.78%) to all the strains of B. cereus isolates (Fig. 4a) hence was identified as B. cereus WD22. The isolates WD23 and WDE11 showed similarity to strains belonging to Pseudomonas genera with maximum similarity (97.67% and 98.05% respectively) to P. aeruginosa from BLAST analysis (Fig. 4b). They were identified as P. aeruginosa WD23 and P. aeruginosa WDE11. The isolate OCB1 showed similarity to strains belonging to Acinetobacter genera with maximum similarity of 86.67% to three strains of A. baumannii (Fig. 4c). Hence it was identified as A. baumannii OCB1. Acinetobacter strains are observed to be Gram negative coccobacilli and OCB1 also demonstrated the same morphology.
Fig. 4a. Phylogenetic tree showing the relationship of isolated WD22 with isolates showing homology as obtained from BLASTn.
Fig. 4b. Phylogenetic tree showing the relationship of isolated WD23 and WDE11 with isolates showing homology as obtained from BLASTn.
Fig. 4c. Phylogenetic tree showing the relationship of isolated OCB1 with isolates showing homology as obtained from BLASTn.
The identified isolates formed a separate clade in the phylogenetic tree that indicated significant differences from the isolates showing similarity through BLAST analysis. This could be because of evolution as they are found in PRE with low nutrient conditions (Table 1). The partial 16s sequences of the isolated cultures i.e. B. cereus WD22, A. baumannii OCB1, P. WD23 and P. WDE11 have been deposited in NCBI-GenBank with the accession numbers MK355614, MK212363, MK249728 and MK355609 respectively.
Table (5):
Growth of isolated and identified best degraders of PCO at different cultural conditions Culture.
| Culture Condition | OCB1 | WD22 | WD23 | WDE11 |
|---|---|---|---|---|
| Temperature (°C) | ||||
| 4 | – | – | – | – |
| 10 | – | – | – | – |
| 25 | + | + | + | + |
| 37 | + | + | + | + |
| 45 | + | + | + | + |
| pH | ||||
| 2.0 | – | – | – | – |
| 4.0 | ++ | – | + | + |
| 6.0 | +++ | +++ | +++ | +++ |
| 8.0 | +++ | +++ | +++ | +++ |
| 10.0 | +++ | ++ | ++ | ++ |
| 12.0 | – | – | – | – |
| NaCl (%w/v) | ||||
| 4 | ++ | + | ++ | ++ |
| 6 | + | – | + | + |
| 8 | – | – | – | – |
| 10 | – | – | – | – |
| 12 | – | – | – | – |
Biochemical charecteristisation of best degraders of PCO
All the identified isolates grew at temperatures 27-45°C but failed to grow at lower temperatures (Table 6). All isolates except B. cereus WD22 grew at pH from 4.0 to 10.0. All isolates could tolerate NaCl concentration up to 4%v/v, hence they can be used for treatment of oil spills in sea water (NaCl% = 3.5). Most of the biochemical characteristics of all the identified isolates matched with the standard results (Table 6). All the isolates were positive for catalase, urease, hydrogen sulphide production and hydrolysis of esculin. None of the tested isolates could hydrolyse lysine and ornithine whereas only B. cereus WD22 could utilize malonate. All the isolates could hydrolyse glucose, fructose and xylose (Table 7). P. aeruginosa cannot hydrolyse maltose, sucrose and lactose and similar results were observed in this study.
Table (6):
Biochemical characterisation of isolated and identified best degraders of PCO.
| Tests | OCB1 | WD22 | WD23 | WDE11 |
|---|---|---|---|---|
| Biochemical Tests | ||||
| Oxidase | – | + | + | + |
| Catalase | + | + | + | + |
| Urease | + | + | + | + |
| Nitrate Reduction | – | – | – | – |
| H2S Production | + | + | + | + |
| ONPG | – | – | – | – |
| Phenylalanine | – | – | – | – |
| Esculin Hydrolysis | + | + | + | + |
| Methyl Red | + | – | – | – |
| Voges Proskauer | – | + | – | – |
| Indole | – | – | – | – |
| Utilisation Tests | ||||
| Lysine | – | – | – | – |
| Ornithine | – | – | – | – |
| Citrate | + | + | – | – |
| Malonate | – | + | – | – |
Table (7):
Sugar hydrolysis results of isolated and identified best degraders of PCO.
Sugar |
OCB1 |
WD22 |
WD23 |
WDE11 |
|---|---|---|---|---|
Glucose |
+ |
+ |
+ |
+ |
Maltose |
+ |
+ |
– |
– |
Fructose |
+ |
+ |
+ |
+ |
Sucrose |
+ |
+ |
– |
– |
Lactose |
– |
– |
– |
– |
Xylose |
+ |
+ |
+ |
+ |
Arabinose |
+ |
+ |
– |
– |
Raffinose |
– |
– |
– |
– |
Rhamnose |
– |
– |
– |
– |
Cellobiose |
– |
– |
+ |
– |
Melibiose |
– |
+ |
+ |
+ |
Saccharose |
– |
– |
+ |
+ |
Adonitol |
– |
– |
– |
– |
Trehalose |
– |
– |
– |
– |
Degradation of petroleum hydrocarbons of PCO in sea water by identified bacteria
All the isolates were then grown in sea water supplemented with additional carbon and nitrogen sources and spiked with PCO for 12 days. The temperature was maintained at 27°C as in the marine environment as the average temperature of Indian seawater is observed to be between 25-28°C5. Spillage of oil into marine and fresh water environment causes an increase in carbon content and a drastic decrease in nitrogen content23. It has been reported that the addition of a simple carbon source such as glucose increased the biodegradation rate of total petroleum hydrocarbons (TPH) in contaminated samples24. Hence in the present study the growth of identified isolates and the biodegradation of PCO were studied under the presence of trace amounts of glucose and yeast extract.
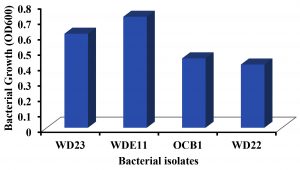
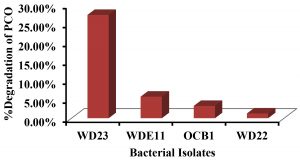
Among the four isolates P. aeruginosa WD23 showed the maximum degradation of 1.0%v/v PCO (27.25%) whereas the least degradation was performed by B. cereus WD22 (1.25%) (Fig. 5). Even though P. aeruginosa WDE11 showed the maximum growth (OD600 = 0.72) it could degrade only 5.68% of PCO. This indicates that WDE11 utilised the supplied glucose and yeast extract as nutrients for its growth whereas on the other hand WD23 also degrades PCO when other nutrients are supplied. A. baumannii OCB1 and B. cereus WD22 showed moderate growth (OD600 = 0.45 and 0.41) with very low degradation of crude oil (Fig. 6).
In the present study, all the isolates obtained from PRE post enrichment in PCO from petrochemical refinery industry in Mangalore, India were P. aeruginosa strains. On direct isolation, other PCO degrading strains were obtained. Among the 32 isolates tested, four best degraders i.e B. cereus WD22, A. baumannii OCB1, P. aeruginosa WD23 and P. aeruginosa WDE11 were identified and characterised. P. aeruginosa WD23 could degrade 27.25% of PCO in sea water supplemented with glucose and yeast extract. From this study we can conclude that the bacteria isolated from petroleum hydrocarbon contaminated sites have the potential to be exploited for treatment of oil spills in sea water.
ACKNOWLEDGMENTS
Authors would sincerely like to thank Department of Biotechnology, NMAM Institute of Technology for providing necessary requirements for the research work.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
This research work has been funded by Vision Group of Science and Technology, Govt of Karnataka (VGST-GoK) under RGS-F grant (GRD No 846/315) for the FY 2018-19 awarded to LCG.
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
AVAILABILITY OF DATA
All datasets obtained or studied during this study are incorporated in the manuscript.
- De la Huz R, Lastra M, Lopez J Oil Spills, Editor(s): JO Nriagu. Encyclopedia of Environmental Health, Elsevier, 2011; 251–255.
Crossref - Rekadwad BN, Khobragade CN. A case study on effects of oil spills and tar-ball pollution on beaches of Goa (India). Mar. Pol. Bull., 2015; 100: 567–570.
Crossref - Sukumaran S, Mulik J, Rokade MA, Kamble K. Impact of ‘Chitra’ oil spill on tidal pool macrobenthic communities of a tropical rocky shore (Mumbai, India) Estuaries and Coasts, 2014; 37(6): 1415-1431.
Crossref - Han Y, Nambi IM, Clement TP. Environmental impacts of the Chennai oil spill accident – A case study. Sci. Total Environ., 2018; 626: 795–806.
Crossref - Wang Q, Zhang S, Li Y, Klassen W. Potential approaches to improving biodegradation of hydrocarbons for bioremediation of crude oil pollution. J. Environ. Prot., 2011; 2: 47–55.
Crossref - Obi EO, Kamgba FA, Obi DA. Techniques of oil spill response in the sea. IOSR Journal of Applied Physics., 2014; 1(10): 36–41.
Crossref - Ghoreishi G, Alemzadeh A, Mojarrad M, Djavaheri M. Bioremediation capability and characterization of bacteria isolated from petroleum contaminated soils in Iran. Sustain. Environ. Res., 2017; 27: 195–202.
Crossref - Al-Hawash AB, Alkooranee JT, Abbood HA, Zhang J, Sun J, Zhang X, Ma F. Isolation and characterization of two crude oil-degrading fungi strains from Rumaila oil field, Iraq. Biotechnology Reports, 2018a; 17: 104–109.
Crossref - Kachienga L, Momba MNB. Kinetics of petroleum oil biodegradation by a consortium of three protozoan isolates (Aspidisca sp., Trachelophyllum sp. and Peranema sp.). Biotechnology Reports, 2017; 15: 125–131.
Crossref - Varjani SJ, Rana DP, Bateja S, Upasani VN. Isolation and screening for hydrocarbon utilizing bacteria (hub) from petroleum sample. Int. J. Curr. Microbiol. App. Sci., 2013; 2(4): 48–60.
- Hanson KG, Desai JD, Desai AJ. A rapid and simple screening technique for potential crude oil degrading microorganisms. Biotechnol. Tech., 1993; 7(10): 745–748.
Crossref - Liang J, Cheng T, Huang Y, Liu J. Petroleum degradation by Pseudomonas sp. ZS1 is impeded in the presence of antagonist Alcaligenes sp. CT10. AMB Express, 2018; 8(1): 88-98
Crossref - Sanchez I, Palop L, Ballesteros C. Biochemical characterization of lactic acid bacteria isolated from spontaneous fermentation of ‘Almagro’ eggplants. Int. J. Food Microbiol., 2000; 59(1-2): 9-17.
Crossref - Varjani SJ, Upasani VN. Core flood study for enhanced oil recovery through ex-situ bioaugmentation with thermo- and halo tolerant rhamnolipid produced by Pseudomonas aeruginosa NCIM 5514. Biores. Technol., 2016; 220: 175–182.
Crossref - Kirisits MJ, Prost L, Starkey M, Parsek MR. Characterization of colony morphology variants isolated from Pseudomonas aeruginosa biofilms. Appl. Environ. Microbiol., 2005; 71(8): 4809-4821.
Crossref - Abdul-Hussen SR, Atia SS. Antimicrobial effect of pyocyanin extracted from Pseudomonas aeruginosa. European Journal of Experimental Biology, 2016; 6(6): 6–13.
- Ma Y, Wang L, Shao Z. Pseudomonas, the dominant polycyclic aromatic hydrocarbon-degrading bacteria isolated from Antarctic soils and the role of large plasmids in horizontal gene transfer. Environ. Microbiol., 2006; 8(3): 455–465.
Crossref - Saadoun I, Mohammad MJ, Hameed KM, Shawaqfah M. Microbial populations of crude oil spill polluted soils at the Jordan-Iraq dessert (the Badia region). Brazil. J. Microbiol., 2008; 39: 453–456.
Crossref - Belhaj A, Desnoues N, Elmerich C. Alkane biodegradation in Pseudomonas aeruginosa strains isolated from a polluted zone: identification of alkBand alkB-related genes. Research in Microbiology, 2002; 153: 339–344.
Crossref - Batista SB, Mounteer AH, Amorim FR, Totola MR. Isolation and charecterisation of biosurfactant/bioemulsifier producing bacteria from petroleum contaminated sites. Biores. Technol., 2006; 97(6): 868–875.
Crossref - Nguyen HT, Nguyen TT, Pham HTT, Nguyen QTM, Tram MT, Nguyen AH. Fate of carotenoid producing Bacillus aquimaris SH6 colour spores in shrimp gut and their dose-dependent probiotic activities. PLoS ONE, 2018; 13(12): 209–215.
Crossref - Roy A, Dutta A, Pal S, Gupta A, Sarkar J, Chatterjee A, Saha A, Sarkar P, Sar P, Kazy SK. Biostimulation and bioaugmentation of native microbial community accelerated bioremediation of oil refinery sludge. Biores. Technol., 2018; 253: 22–32.
Crossref - Al-Hawash AB, Dragh MA, Li S, Alhujaily A, Abbood HA, Zhang X, Ma F. Principles of microbial degradation of petroleum hydrocarbons in the environment. Egyptian Journal of Aquatic Research, 2018b; 44: 71–76.
Crossref - Farag S, Soliman NA, Abdel-Fattah YR. Statistical optimization of crude oil bio-degradation by a local marine bacterium isolates Pseudomonas sp. sp48. Journal of Genetic Engineering and Biotechnology, 2018; 16: 409–420.
Crossref
© The Author(s) 2020. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.