ISSN: 0973-7510
E-ISSN: 2581-690X
The study was designed to compare the extent of bacterial colonisation on the surface of Polylactic acid/Polyglycolic acid copolymer and expanded Polytetrafluoroethylene based Guided tissue regeneration (GTR) membrane in an in vitro model by spot analysis and scanning electron microscopy. Earlier in vitro models have aimed to study the barrier function against the bacterial penetration across just one surface of the GTR membranes. No such study is present in the literature which aimed to quantify bacterial adhesion over both the surfaces of the membrane. Sterile Resorbable PLA/PGA copolymer and non-resorbable microporous ePTFE based GTR membrane were used in the study. Both groups were subdivided into two subgroup (n=10) based on incubation period of 24 and 48 hours. Incubated in Todd Hewitt broth with Streptococcus mutans, the samples were vortexed and analysed for bacterial count using spot test and scanning electron microscopy. Between PLA/PGA and ePTFE GTR membrane at 24 hours incubation time period, there was a significant difference in terms of Colony forming units (p = <0.001), with the median Colony forming units being highest in the PLA/PGA GTR membrane. At 48 hours incubation time period, there was a significant difference in terms of Colony forming units (p = <0.001), with the median Colony forming units being highest in the PLA/PGA GTR membrane. Our findings depict that ePTFE based GTR membrane showed significantly lesser bacterial colonisation on its surfaces as compared to PLA/PGA at both the incubation periods i.e., 24 hours and 48 hours as shown by both spot test and SEM.
Guided Tissue Regeneration, Bacterial Adhesion, PLA/PGA Membrane, EPTFE Membrane, Spot Test, Scanning Electron Microscopy
The Guided tissue regeneration (GTR) helps prevent migration of epithelial cells along the cemental wall of the pocket and maintains space for clot stabilization thereby facilitating the periodontal ligament and perivascular cells to proliferate in the space created because of which there is possible gain in the periodontal attachment apparatus.1 The GTR procedure involves a barrier placed from the crest of the remaining bone till the cemento-enamel junction of the tooth. As a result, the cemental surface is secluded from the gingival and connective tissue wall. These barriers (membranes) may be either resorbable or non-resorbable. Consequently, epithelial migration is prevented during postsurgical healing, at the same time proliferation of the area by cells from periodontal ligament takes place, provided that infection does not occur due to the surrounding biofilm.2-3
The ideal physical properties of a GTR membrane includes the ability to seclude the cemental wall from the epithelial cells and maintain a space for appropriate cells of periodontal ligament, osteoblasts, or cementoblasts to repopulate the periodontal apparatus, to increase the attachment level and to prevent biofilm production, thereby isolation of the region against any bacterial contamination.4 Thus, a membrane must maintain its structural integrity during early wound healing along with surface characteristics that potentiate regeneration of periodontium and at same time prevent bacterial proliferation and infection.
Commercially GTR membranes were earlier available as non-absorbable materials like expanded polytetrafluorethylene (ePTFE). GTR procedures using these as a barrier requires a second surgical intervention for removal of these membranes from the surgical site. Natural or synthetic bioabsorbable materials for production of GTR membranes was thus explored to overcome the need of second surgery.5-6
Bacterial contamination of the regenerating wound represents one of the leading factors causing failure of the procedure.7-9 Species of bacteria,10-11 bacterial count,12 and the area of bacterial contamination found on the GTR membrane13-14 are some of the factors that may affect the gain in clinical attachment after the procedure. The bacterial species found on GTR membranes include various periodontal pathogens.15 A clinical study by Nowzari et al. found that GTR membranes were contaminated within 3 minutes after the GTR placement in situ.16-17 In vivo studies showed that bacteria adhere to and proliferate on various kinds of GTR membranes easily.12 Surface characteristics and material of GTR membranes will possibly give different niches to the biofilm forming pathogens and a difference will exist in the extent of adhesion to various membranes being used as a barrier in GTR procedures.18
The earlier in vitro studies have analysed the difference in bacterial penetration across the GTR membranes. These studies have considered the regenerative site to be sterile and GTR membranes to behave as a perfect barrier against the biofilm producing bacteria and hence just analysed the penetration across the surface of the membrane.11,18,19 The region of interest in these in vitro studies was only one surface of a GTR membrane which exhibited bacterial colonisation threat while the other surface was considered sterile. These in vitro studies in different experimental setups successfully demonstrated a difference in the penetration of the biofilm producing bacteria.11,18,19
The purpose of our in vitro study is to evaluate quantitatively by Spot test (Colony forming unit count) and scanning electron microscopy the efficacy of resorbable (PLA/PGA) and non-resorbable (ePTFE) based membranes to prevent bacterial colonisation on the GTR membranes. We hypothesized that both sides of membranes surface are at equal risk of bacterial colonisation considering the fact that there is no sterile environment where barrier against bacterial contamination can be achieved in any regenerative procedures.
The entire in vitro study was performed at Dr. Ziauddin Ahmed Dental College in collaboration with Department of Microbiology, Jawaharlal Nehru Medical College, Aligarh Muslim University, Uttar Pradesh, Aligarh, India. Clearance for the study was issued by the Institutional Ethics Committee, JNMCH (Regd. under the Central Drugs Standard Control Organization, Ministry of Health and Family Welfare, Government of India).
GTR Membranes
Sterile Resorbable Polylactic acid/Poly glycolic copolymer based GTR membrane, CYTOFLEXR RESORB (Lot: 040920-1 Unicare Biomedical Inc.) (Figure 1A) and sterile Non resorbable Microporous expanded Polytetrafluoroethylene based GTR membrane, CYTOFLEXR TEF – GUARD (Lot: 072920-1 Unicare Biomedical Inc.) were used in the study
(Figure 1A and B, respectively).
The membranes were divided into two groups according to the following protocol:
- Resorbable PLA/PGA polymer based GTR membranes (RP group)
- Non resorbable ePTFE based GTR membrane (NRE group)
Each group was further subdivided into two subgroups based on duration of incubation of 24h and 48h as following:
(a) RP24 (n=10)
(b) RP48 (n=10)
(c) NRE24 (n=10)
(d) NRE48 (n=10)
The membrane corresponding to each subgroup was carefully cut into ten 1mm/1mm square sections using a sterile surgical scissors.
Duplicate samples were also made of each group for analysis of bacterial adhesion under scanning electron microscopy.
Bacterial strain and inoculum preparation
A bacterial strain of the common oral bacterium Streptococcus mutans ATCC 25175-0266P was used to evaluate the membranes for bacterial adhesion and biofilm formation. An inoculum of 105 Colony forming units per millilitres (CFU/mL) of the strain Streptococcus mutans ATCC 25175-0266P was made at Department of Microbiology, AMU as follows:
The frozen stock was taken and was plated on a 5% (v/v) blood agar plate and incubated for 24 hours at 37°C. Isolated colonies were made to suspend in pre-warmed Todd Hewitt Broth (THB) to produce an optical density (OD546nm) of 0.28, which corresponds to a concentration of 108 CFU/mL, and thereafter further dilutions were made in THB until a final concentration of 105 CFU/mL was obtained.
Biofilm formation of Streptococcus mutans on GTR membranes
Four sterile 96 well microplates were used for the experiment, one for each subgroup. This was done to ensure a sterile isolated and contained environment for the adhesion between the bacteria and membrane to take place. The wells in each of the microplates were first numbered and one well was designated to each of 10 samples in the subgroup. In all microplates ten 1mm/1mm membrane, earlier measured and cut were placed within the designated well for each sample. One millilitre inoculum (105 CFU/mL) was added on top of each material inside the microplates using a micropipette and incubated after covering with sterile cellophane at 37°C (Figure 1C).
Estimation of Colony Forming Units (CFU counting) using spot test
As the incubation time period was reached, each GTR membrane sample was collected and rinsed with 0.9% sterile saline using micropipette to remove non adherent cells. Then the materials were transferred to sterilized glass test tubes containing 1 mL saline. The adhered bacterial colonies onto the surface of the membranes were dislodged from the surface by subjecting the material to a high-speed vortex at 2000 rpm for 1 min (Figure 1D). The number of bacterial counts of the vortex suspension was assessed by quantitative culture using the spot test method. Before proceeding to the serial dilution (spot test), the stock solution was tested for any unwanted bacterial growth by plating 10 microliters of it on a blood agar plate and observing the growth under light microscope after Gram staining (Figure 1E).
The vortex suspension was diluted in a series of five 1:10 dilution (1ml of stock solution in 9 ml of diluent solution) in 0.9% saline (Figure 1F). Before counting the colonies, ten microliters from each dilution were then plated on blood agar plate using micropipette and incubated at 37°C for 24 hours. (Figure 2) In the series of five dilutions of saline plated on blood agar plates (1:10), fourth dilution was chosen to deduce the CFU/mL as the counting was much easier because the number of colonies were between the range of 30-300 CFU. Hence total dilution factor for CFU counting was of 104.
For calculating the Colony Forming Unit per millilitre (CFU/ml) the following derivation was used –
Number of colonies × Total dilution factor / Volume of culture (mL).
Where total dilution factor was 104 and the volume of saline solution plated was 10 microliters or 0.01 millilitres.
Scanning electron microscopy
The tested GTR membranes of each group (duplicate samples other than being subjected to vortex) containing Streptococcus mutans were taken and were rinsed with 0.9% saline to dislodge non-adhered bacteria from surface. Primary fixation was then done in 2.5% phosphate-buffered glutaraldehyde (maintaining pH at 7.3) for 10 hours.
The specimens were then dehydrated in a graded series of ethanol (50%, 70%, 90%, and 100%). After being critical point-dried in CO2 and sputter coated with a thin layer of gold (Figure 1G), the specimens were observed by Scanning electron microscope JEM-2100 JEOL, JAPAN (Figure 1H) installed at University Sophisticated Instrument Facility, Aligarh Muslim University, at the 1500X and 5000X original magnification at accelerating voltage ¬15 KV to evaluate the bacterial adhesion to the GTR membranes.
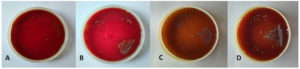
Figure 1. Cytoflex tef-guard ePTFE non – resorbable GTR membrane (A), Cytoflex PLA/PGA resorbable GTR membrane (B), 96 well microplate containing streptococcus mutans strain within to Hewitt broth (C), vortex used to break the adhering bacterial colonies to the membrane within sterile saline (D), light microscopy of stock solution after plating on blood agar to check sterility of experiment (E), serial dilution of the stock solution obtained after vortex in sterile saline (F), Sample preparation for scanning electron microscopy gold sputtering (G), scanning electron microscope JBM 6510 LV, Jeol Japan (H)
Figure 2. Plating the saline solution after series dilution onto blood agar plate and incubating for 24 hours and 48 hours A, B respectively (ePTFE membrane) and C, D respectively (PLA/PGA membrane)
Statistical analysis
The normality of the data was assessed using Shapiro Wilk test. Mean values ± standard error of the mean (SEM) of the observations were calculated and outlined in tables. The difference in the CFU present on the surfaces of GTR membrane pertaining to resorbable and non-resorbable group and also for the two different time period of incubation i.e. 24 and 48 hours were analysed separately using Wilcoxon Mann Whitney U test. The strength of association was deduced using Point Biserial correlation. The p values < 0.05 were considered statistically significant. The analysis was done using IBM SPSS Version 2.0 software, Chicago, USA.
CFU by serial dilution and spot test
The mean CFU of two groups i.e. after 24 hours and 48 hours incubation is shown in Table 1 and Table 2, respectively. On comparing resorbable GTR membrane group with the non resorbable group corresponding to the two incubation period i.e. 24 hours and 48 hours separately a significant difference was found, CFU were considerably more in the resorbable GTR group at both the incubation periods. Between PLA/PGA GTR membrane (RP24) and ePTFE GTR membrane (NRE24) at 24 hours incubation time period (Table 1), there was a significant difference in terms of CFU (W = 100.000, p = <0.001), with the median CFU being highest in the PLA/PGA GTR membrane (RP24). Between PLA/PGA GTR membrane (RP48) and ePTFE GTR membrane (NRE48) at 48 hours incubation time period (Table 2), there was a significant difference in terms of CFU (W = 100.000, p = <0.001), with the median CFU being highest in the PLA/PGA GTR membrane (RP48).
Table (1):
Comparison of the 2 Subgroups of the Variable Type of Material in Terms of Colony forming units (Time Period: 24 Hours)
| Colony forming units | Type of Material | Wilcoxon-Mann-Whitney U Test | ||
|---|---|---|---|---|
| Resorbable (RP24) | Non-Resorbable (NRE24) | W | p value | |
| Mean (SD) | 2.34 (0.37) | 1.06 (0.26) | 100.000 | <0.001 |
| Median (IQR) | 2.33 (2.01-2.55) | 0.98 (0.88-1.25) | ||
SD: Standard deviation
IQR: Interquartile range
Table (2):
Comparison of the 2 Subgroups of the Variable Type of Material in Terms of Colony forming units (Time Period: 48 Hours)
| Colony forming units | Type of Material | Wilcoxon-Mann-Whitney U Test | ||
|---|---|---|---|---|
| Resorbable (RP48) | Non-Resorbable (NRE48) | W | p value | |
| Mean (SD) | 4.16 (0.63) | 1.50 (0.23) | 100.000 | <0.001 |
| Median (IQR) | 4.1 (3.86-4.68) | 1.52 (1.36-1.65) | ||
SD: Standard deviation
IQR: Interquartile range
The CFU increased significantly in both non resorbable and resorbable groups of membranes between the two incubation time periods i.e. from 24 hours to 48 hours in all the samples. (W = 0.000, p = <0.001) With the median CFU being highest in case of PLA/PGA membrane at 48 hours.
Scanning electron microscopy analyses
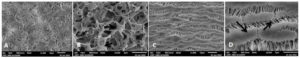
The GTR membranes pertaining to both the groups were subjected to scanning electron microscopy to analyse the initial structure prior to the experiment (Figure 3). All the samples prepared for SEM analysis were analysed at 1500X and 5000X magnification and multiple images of 5-micron area of the specimen were taken at each magnification.
Figure 3. Surface of PLA/PGA GTR membrane under SEM at 1500x magnification (A), surface of PLA/PGA gtr membrane under 5000x magnification showing fibrous structure and interlaying spaces (arrows) (B), surface of ePTFE membrane under 1500x (C), surface of ePTFE membrane under 5000x showing nodes connected by fibrillar areas (arrows) (D)
PLA/PGA Copolymer
At 1500X magnification (Figure 3A) the copolymer GTR membrane reveals cross-linked fibre structure with ample spaces between the crosslinking fibres hence rendering a highly porous material. The fibres have no definite course and the crosslinking is also showing no definite pattern which can be appreciated at both 1500X and 5000X; all the crosslinking fibres have a variable thickness and the dimension is highest at the junction where multiple crosslinks were present.
At 5000X magnification (Figure 3B) the spaces in between this fibrous structure can easily be appreciated throughout the structure, volume of which is also varying remarkably. Each region rendered a much varying picture owing to haphazard orientation of fibres and multiple crosslinks. This was unique for this material as compared to the much definite and monotonous view of the ePTFE material which showed nodes and fibrils throughout. This structure of crosslinking fibres and spaces within is believed to impart the PLA/PGA copolymer more surface area.
ePTFE
At 1500X magnification (Figure 3C) under SEM ePTFE membrane showed a typical expanded structure with fibrillar and lamellar orientation. The structure is characterised by parallel arrangement of nodes interconnected by transversely placed fibrils connecting two adjacent nodes. This structure of ePTFE gives the material high degree of porosity.
At 5000X magnification (Figure 3D), the fibrillar structure lying between the nodes can be appreciated more distinctly. Fibres run parallel to each other from one node to the next, the orientation of each fibre is somewhat linear with some fibres lying haphazardly as well but there is no break in continuity of a fibre traversing between two adjacent nodes. Although the fibres are tightly packed hence filling the space between two nodes, there is space within the fibre bundles which can be appreciated as well. The porosity can be increased or decreased by altering the distance between the nodes to yield desired properties.
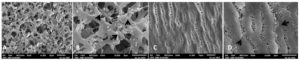
SEM analyses for bacterial adherence on the surface of GTR membrane
The GTR membranes pertaining to the groups which were subjected to incubation with the bacterial broth were also subjected to scanning electron microscopy to analyse the bacterial adherence on the surface.
PLA/PGA COPOLYMER
At 1500X magnification (Figure 4A), the crosslinking fibrous structure of the PLA/PGA membrane can be appreciated even after incubation of 48 hours. This emphasized no change in membrane structure. However, multiple aggregated clusters of elecron-lucent cocci can be seen along with the fibriform extracellular matrix like structure. Most of the cocci were seen almost separated from the biofilm at 5000X magnification (Figure 4B). The most significant finding in the SEM image of PLA/PGA copolymer membrane was the amount of adhering cocci on the surface which were almost covering the 5 micron area of the surface.
ePTFE
At 1500X magnification (Figure 4C) the expanded structure was appreciated along with electron-lucent aggregated cocci and fibriform extracellular matrix. Most of the surface showed the cells almost separated from the biofilm. The cocci are mostly seen in pairs and at few locations are forming an aggregated cluster.
There was a fascinating peculiarity regarding the location of the adhering cocci on the surface of the ePTFE membrane when seen under 5000X magnification (Figure 4D). The cocci were prescent on the juntion between the fibrillar area and the node. This finding might delineate the affinity of cocci to bind on a certain location which render them favaurable for proliferation. It was quite clear that the number of cocci seen are far lesser in the SEM images of ePTFE GTR membrane as compared to the PLA/PGA copolymer at each 5 micron area examined.
Figure 4. Bacterial adhesion on surface of PLA/PGA gtr membrane under 1500x (A), bacterial adhesion on surface on surface of PLA/PGA gtr membrane under 5000x magnification showing adhering bacteria to the fibrous area forming aggregated clusters (arrows) (B), surface of ePTFE membrane showing adhering bacteria colonies at 1500x magnification (C), surface of ePTFE membrane under 5000x magnification showing adhering bacteria at the junction of node and fibrillar region (D)
Non resorbable ePTFE GTR membranes have lower bacterial colonisation over its surface as compared to the resorbable PLA/PGA membrane. Better clinical outcomes can be achieved after periodontal regenerative procedures using ePTFE membranes as a barrier because its inherent structure renders lesser chances of post-operative infection of the regenerative site.
A novel in vitro experiment was used in the study to assess the colonisation of S. mutans over the ePTFE and PLA/PGA GTR membranes after 24 and 48 hours of incubation periods. The extent of colonisation was then analysed by SEM and was quantified using spot test method. Unlike the previous studies that have analysed the bacterial penetration across the GTR membranes only, this study demonstrates the extent of bacterial colonisation over both the surfaces of GTR membranes. Our study has showed the structural difference of GTR membranes may be responsible for preventing bacterial colonisation.
The comparison when made in the colony-forming unit count found over GTR membranes and the incubation time period; it was evident in almost all samples that the number of colonies of bacteria significantly increased as the incubation period was increased from 24 hours to 48 hours (p<0.001). This unique finding is in accordance with that of Cheng et al.11 in their penetration experiment, where the colony forming unit count across the membrane was seen to increase when the incubation time period was increased. In contrast, the findings made by Margarita Trobos et al18 concluded that the colony forming unit bacterial count decreased when the incubation time period was raised from 24 hours to 48 hours. The basis for this increase in the colonisation of S. mutans over GTR membranes with longer incubation time period can be cell multiplication only as strict aseptic experimental model setup was used.
It is known that S. mutans adhere to tooth surfaces by insoluble glucan formation and other glucan-independent mechanisms where adhesins bind with the salivary components present in the acquired enamel pellicle.20 Many studies have shown the mechanism of S mutans adherence in which the basic model used was hydroxyapatite or saliva-coated hydroxyapatite for enamel or saliva-coated enamel, respectively.20-21
There is no clear picture available in the literature regarding how the biofilm formation takes place on a model which does not use saliva-coated hydroxyapatite. Hence binding and proliferation of S. mutans over a non-organic material such as the GTR membrane’s surface is not clear. The colonisation of S. mutans on the surface of these membranes is thought to be due to glucan-mediated interaction20 as revealed by scanning electron microscopy images. The S. mutans adherence was clearly seen more on the fibrillar structure on the two different membranes used in our study. Therefore, fibrillar content may be associated to a greater degree of bacterial adherence.
The number of CFU of S. mutans found on the surface of the PLA/PGA membrane was significantly more than that on the ePTFE membrane at both 24 hours. (p<0.001) and 48 hours. (p<0.001) after incubation. This can be explained by the surface morphology of these membranes. The polymer membrane has a completely fibrillar structure as seen in the SEM images, whereas the ePTFE has an expanded structure with fibrillar areas that alternate with the lamellar regions, thereby contributing to lesser bacterial adherence. These findings were in accordance to the study by Cheng et al.,21 where they have shown significant difference in the bacterial penetration across the PLA/PGA and ePTFE membranes. However, in the study by Cheng et al, the membranes were loaded with antibiotic but only one surface of the membrane was analysed for bacterial colonisation. In the SEM analysis, our study demonstrated greater affinity of the S. mutans to adhere to the fibres which are present just at the junction of the fibrillar region and the nodes in the ePTFE membrane (Figure 4D). Previous studies have not reported such findings, Future studies evaluating the surface characteristics of ePTFE may shed insight with respect to this association.
It can be concluded that the surface characteristics of the GTR membrane affects the bacterial colonisation over it. The haphazard arrangement of fibres with considerable amount of dead space of the membrane favours the bacterial colonisation. The more organised expanded structure of the ePTFE membrane do not favour bacterial colonisation. Hence, ePTFE membranes can further be modified to induce more of the nodes and less of fibres to decrease the bacterial colonisation and chances of infection in regenerative procedures.
Although scanning electron microscopy was used to analyse the surfaces of membrane both before and after bacterial adhesion and the bacterial morphology as well as the biofilm. However, the full potential of this resource can’t be utilised because quantification of the adhering bacteria was not done under SEM due to lack of resources. The quantification done by CFU count provides a crude picture only.
Future research experiments can be made to quantify the bacterial adhesion onto the surface of various commercially available GTR barrier membranes directly under SEM by making use of computer software (U-Net convolutional networks for biomedical images), which detects the morphology of various species of bacteria and quantifies them with ease.
Finally, this study clearly signifies that the surface of ePTFE membrane resists the adhesion of S. mutans which is the most common oral bacteria. Hence, we suggest that there are lesser chances of infection in the regenerating wounds if ePTFE membranes are used in GTR procedures. It is for the clinician’s discretion to choose between the lesser contaminations while using ePTFE, though there will be the need of second surgery to retrieve the non-resorbable ePTFE membrane or to prescribe systemic antibiotics to prevent high levels of contamination while using highly porous PLA/PGA GTR membranes.
ACKNOWLEDGMENTS
None.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
None.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
This study was approved by the Institutional Ethics Committee, Jawaharlal Nehru Medical College & Hospital, Faculty of Medicine, Aligarh Muslim University, Aligarh, India, with reference number IECJNMC/445.
- Retzepi M, Donos N. Guided Bone Regeneration: biological principle and therapeutic applications. Clin Oral Implants Res. 2010;21(6):567-576.
Crossref - Nyman S, Lindhe J, Karring T, Rylander H. New attachment following surgical treatment of human periodontal disease. J Clin Periodontol. 1982;9(4):290-296.
Crossref - Gottlow J, Nyman S, Lindhe J, Karring T, Wennstrom J. New attachment formation in the human periodontium by guided tissue regeneration. Case reports. J Clin Periodontol. 1986;13(6):604-616.
Crossref - Elgali I, Omar O, Dahlin C, Thomsen P. Guided bone regeneration: materials and biological mechanisms revisited. Eur J Oral Sci. 2017;125(5):315-337.
Crossref - Chiapasco M, Zaniboni M. Clinical outcomes of GBR procedures to correct peri-implant dehiscences and fenestrations: a systematic review. Clin Oral Implants Res. 2009;20(Suppl 4):113-23.
Crossref - McAllister BS, Haghighat K, Gonshor A. Histologic evaluation of a stem cell-based sinus-augmentation procedure. J Periodontol. 2009;80(4):679-686.
Crossref - Cortellini P. Reconstructive periodontal surgery: a challenge for modern periodontology. Int Dent J. 2006;56(Suppl 4):250-5.
Crossref - Selvig KA, Nilveus RE, Fitzmorris L, Kersten B, Khorsandi SS. Scanning electron microscopic observations of cell populations and bacterial contamination of membranes used for guided periodontal tissue regeneration in humans. J Periodontol. 1990;61(8):515-520.
Crossref - Rossa ML, Lima LA, Pustiglioni FE, et al. SEM analyses of bacterial contamination of e-PTFE membranes and GTR clinical results. J Int Acad Periodontol. 2006;8(4):115-124. PMID : 17042167
- MacDonald ES, Nowzari H, Contreras A, Flynn J, Morrison J, Slots J. Clinical and microbiological evaluation of a bioabsorbable and a nonresorbable barrier membrane in the treatment of periodontal intraosseous lesions. J Periodontol. 1998;69(4):445-453.
Crossref - Cheng CF, Lee YY, Chi LY, Chen YT, Hung SL, Ling LJ. Bacterial penetration through antibiotic-loaded guided tissue regeneration membranes. J Periodontol. 2009;80(9):1471-1478.
Crossref - Nowzari H, Slots J. Microorganisms in polytetrafluoroethylene barrier membranes for guided tissue regeneration. J Clin Periodontol. 1994;21(3):203-210.
Crossref - De Sanctis M, Zucchelli G, Clauser C. Bacterial colonization of barrier material and periodontal regeneration. J Clin Periodontol. 1996;23(11):1039-1046.
Crossref - Wang HL, Yan K, Burgett F, Shyr Y, Syed S. Adherence of oral microorganisms to guided tissue membranes. An in vitro study. Journal of Periodontology, 1994; 65.
Crossref - Tempro PJ, Nalbandian J. Colonization of retrieved polytetrafluoroethylene membranes: morphological and microbiological observations. J Periodontol. 1993;64(3):162-168.
Crossref - Nowzari H, Matian F, Slots J. Periodontal pathogens on polytetrafluoroethylene membrane for guided tissue regeneration inhibit healing. J Clin Periodontol. 1995;22(6):469-474.
Crossref - Zucchelli G, De Sanctis M, Clauser C. Integrated Connective Tissue in Bioabsorbable Barrier Material and Periodontal Regeneration. J Periodontol. 1997;68:996-1004.
Crossref - Trobos M, Juhlin A, Shah FA, Hoffman M, Sahlin H, Dahlin C. In vitro evaluation of barrier function against oral bacteria of dense and expanded polytetrafluoroethylene (PTFE) membranes for guided bone regeneration. Clin Implant Dent Relat Res. 2018;20(5):738-748.
Crossref - Cheng C-F, Wu KM, Chen YT, Hung SL. Bacterial adhesion to antibiotic-loaded guided tissue regeneration membranes e A scanning electron microscopy study. J Formos Med Assoc. 2015;114(1):35-45.
Crossref - Bleiweis AS, Oyston PC, Brady LJ. Molecular, immunological and functional characterization of the major surface adhesin of Streptococcus mutans. Adv Exp Med Biol. 1992;327:229-41.
Crossref - Brady LJ, Piacentini DA, Crowley PJ, Oyston PC, Bleiweis AS. Differentiation of salivary agglutinin-mediated adherence and aggregation of mutans streptococci by use of monoclonal antibodies against the major surface adhesin P1. Infect Immun. 1992;60(3):1008-17.
Crossref
© The Author(s) 2023. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.