ISSN: 0973-7510
E-ISSN: 2581-690X
Non-fermenting gram-negative bacteria (NFGNB) endeavouring as major pathogen in infectious disease, predominantly in urinary tract infection (UTI) and increased resistance in NFGNB are matter of concern. This study aimed to evaluate the frequency of NFGNB, antibiotics resistance pattern, plasmid profiling, and antibacterial efficacy of Moringa oleifera against NFGNB. NFGNB were isolated from clinically suspected UTI patients. Identification of isolates and their antibiotics sensitivity pattern were analyzed according to conventional method, and Vitek 2 automated system. Moreover, NFGNB were evaluated for biofilm production and presence of plasmid. Furthermore, antibacterial activity of Moringa oleifera was evaluated against NFGNB. P. aeruginosa (86.0%), and A. baumannii (10.0%) were the most frequent NFGNB followed by Providencia rettgeri 2.0%, Stenotrophomonas maltophilia 1.0%, myroides species 1.0%. 68.6% P. aeruginosa and 60.0% A. baumannii were biofilm producers whereas imipenem and meropenem were the most effective antibiotics. Isolated NFGNB showed multiple bands of plasmid. Furthermore, Moringa oleifera leaves extract showed antibacterial activity against tested NFGNB. MDR-NFGNB presents challenges in treatment and Moringa oleifera leaf extract may be used as an alternative medicine. However, the therapeutic role of specific ingredients present in extract needs further investigation and purification.
Multi-drug Resistant, Uropathogen, Non-fermenting Gram-negative Bacteria, Plasmid Profile Analysis, Moringa oleifera
Non-fermenting gram-negative bacilli (NFGNB) are taxonomically diverse group of aerobic, non-sporing bacteria utilise glucose as an energy source via the oxidative pathway rather than the fermentative process.1
NFGNB comprises organisms from diverse genera like Pseudomonas, Acinetobacter, and Burkholderia. Several studies have been conducted to identify the prevalence of these microorganisms. Isolation rate of non-fermenting gram-negative bacteria (NFGNB) in urine samples was 10.8%- 11.0% with the maximum distribution of isolates as Pseudomonas aeruginosa (72.0%-87.0%), followed by Acinetobacter baumannii (10.4%- 18.0%) and Acinetobacter lwoffii (0.7%).2 NFGNB present as saprophytes in the environment. Many of them come forward as significant Hospital acquired pathogens even though habitually considered as contaminants causing opportunistic infections in immunocompromised hosts. NFGNB causes various infections, including urinary tract infections (UTI), meningitis, pneumonia, wound infections, osteomyelitis, septicaemia etc.3 NFGNB contributes majorly to nosocomial infections with risk factors for infections, including the long duration of hospitalization, prolonged antibiotic therapy, and chronic infection.4 Urinary tract infections are one of the most widespread urological diseases globally. Even though the accessibility of antimicrobial agents alongside UTIs is copious, they are still left as one of the most predominant diseases occurring in humans.5 So, this study was designed to identify the incidence of NFGNB in UTI patients, their antibiotic sensitivity pattern, biofilm production, Plasmid profiling, and analysis of In vitro antibacterial efficacy of Moringa oleifera against NFGNB.
The present research was conducted from August 2019 to September 2021 on urine samples received from clinically suspected patients of urinary tract infections (UTI) at MMIMSR, MMDU, Mullana. Only NFGNB obtained from clinical specimens were recruited in this study.
Sample collection
The midstream urine specimen was collected from clinically suspected uncomplicated UTI patients in a sterile, wide-mouthed universal container. Samples were then sent to the microbiology lab for further processing. Microscopic examination of urine was done by wet mount preparation. Furthermore, samples were inoculated on Cysteine Lactose Electrolyte Deficient Agar (CLED) and incubated at 37°for 24 hours.
Identification of bacterial strains
As per established standard microbiological Procedures, identification of microbes was done using Gram staining followed by series of biochemical assays. Additionally, automated VITEK® 2 system was employed to identify bacterial species. Phenotypically confirmed NFGNB.
Antibiotic susceptibility testing (AST)
AST of the NFGNB strains was done using Kirby Bauer disc diffusion method as per clinical laboratory standard institute (CLSI) and Vitek® 2 automated system as previously established and standardized disc diffusion method. Antibiotics tested against isolates using the standard protocol. The phenotypic analysis of extended-spectrum β Lactamase (ESBL) and metallo β Lactamase (MBL) production was identified by Double-Disc Diffusion method and Combined Disc Method, respectively.6
Double-disc diffusion test (DDDT)
All Ceftazidime resistant isolates of Pseudomonas species and Acinetobacter species were evaluated for ESBL production by employing phenotypic confirmatory test (DDDT). As per CLSI guidelines, bacterial suspension (0.5 MacFarland’s) was lawn cultured on MHA plate. After that ceftazidime (CAZ) disc (30 µg) and ceftazidime + clavulanic acid (CAC) (30 µg/ 10 µg) discs were placed at a distance of about 15mm and incubated at 37°C for 24 hours. The isolates showing zone of inhibition ≥ 5mm in CAC compared to CAZ was identified as ESBL producer.
Combined disc diffusion test (CDDT)
All imipenem resistant isolates were evaluated for MBL production. In brief, bacterial suspension (0.5 MacFarland’s) was lawn cultured on MHA plate. After that Imipenem disks (10µg) alone and another imipenem disc (10µg) in combination with EDTA were placed at a distance of 20mm on MHA plate and incubated at 37°C for 24 hours. The strains exhibited increased zone of inhibition ≥7mm around imipenem + EDTA discs compared to Imipenem alone were considered as MBL producing.
Biofilm production test
Identification of biofilm production by recruited organism was analysed by tube method, briefly 5ml of 1% glucose-containing brain heart infusion (BHI) broth were used to inoculate a loop full of test organisms. The tubes were then incubated for 24 hours at 37°C. Following incubation, the bacterial suspension was removed, cleaned with phosphate buffer saline (pH 7.2), and dried. After that, crystal violet (0.1%) was used to stain the tubes. With distilled water, the extra discoloration was removed. Tubes were further dried by inverted drying. When a discernible film was seen on the tube’s bottom and wall, biofilm development was deemed successful.7
Isolation of plasmid & its profiling
Plasmid DNA extraction was performed using previously established alkaline lysis method, and further profiling was done by agarose gel electrophoresis by following protocol described earlier.8, 9
Antimicrobial efficacy of moringa oleifera
Agar well diffusion assay was employed to evaluate the antibacterial efficacy of Moringa oleifera extract against P. aeruginosa & A. baumannii. In brief, the bacterial suspension (0.5 McFarland) was prepared, and lawn cultured on MHA plate after that 8mm well was punched using sterile filter tips. 50µl of Moringa oleifera leave extract at various concentrations (20mg, 40mg, 80, 160mg, and 320mg of Moringa oleifera leave extract per ml) were dispensed in punched well and incubated at 37oc for 24 hours. Zone of inhibition was evaluated to know antibacterial potency of extract.10
Identification of microbial isolates
In this study, total of 1538 samples were collected from clinically suspected patients of UTI, out of which 729 (48.4%) culture was sterile, and 809 samples were culture positive. Among total isolates, 193 were gram-positive cocci, 511 were gram-negative bacteria, and 105 were candida. Out of 511 GNB, 100 bacterial isolates were NFGNB. Out of 100 NFGNB, 86 P. aeruginosa, 10 A. baumannii, 2 P. rettgeri, 1 S. maltophilia and 1 Myroides species were isolated.
Antibiotic sensitivity pattern
In current research, we observed that Imipenem showed maximum efficacy against all NFGNB bacterial isolates except S. maltophilia which have intrinsic resistance to imipenem, meropenem, amikacin, and gentamycin. Meropenem too exhibited noteworthy susceptibility, in opposite, ceftazidime & cefepime (cephalosporins) and amikacin showed the least susceptibility to all NFGNB. Specifically, in the case of P. aeruginosa meropenem, imipenem, piperacillin tazobactam and gentamycin had a susceptibility rate of 79.0% 74.4%, 73.2%, and 55.8%, respectively. In the case of A. baumannii, Imipenem and meropenem showed an 80.0% susceptibility rate, while amikacin and gentamycin showed a susceptibility of 70.0%. All isolated NFGNB showed resistance to one antibiotic of at least three different classes, hence considered multidrug-resistant strains (Table 1).
Table (1):
Antibacterial susceptibility pattern of NFGNB.
| Antibiotics used | Antibacterial sensitivity pattern of different non-fermenter isolates | ||||
|---|---|---|---|---|---|
| Pseudomonas aeruginosa (86) | Acinetobacter baumannii (10) | Providencia rettgeri (2) | Stenotrophomonas maltophilia (1) | Myroides species (1) | |
| Piperacillin+ Tazobactam | 73.25% | 30% | 0% | I/R | 0% |
| Ceftriaxone | I/R | 20.0% | 0% | I/R | 0% |
| Ceftazidime | 8.1% | 20.0% | 0% | 0% | 0% |
| Cefipime | 8.1% | 20.0% | 0% | 0% | 0% |
| Imipenem | 74.4% | 80.0% | 100% | I/R | 100% |
| Meropenem | 79.0% | 80.0% | 100% | I/R | 0% |
| Amikacin | 48.8% | 70.0% | 0% | I/R | 100% |
| Gentamicin | 55.8% | 70.0% | 0% | I/R | 100% |
| Tobramycin | 50.0% | 30.0% | 0% | I/R | 0% |
| Netilmycin | 50.0% | 30.0% | 0% | 0% | 0% |
| Polymyxin B | 100% | 100% | 100% | 100% | 100% |
Note: I/R: Intrinsic Resistance
ESBL, MBL and biofilm production
Out of total P. aeruginosa 66.2% of were ESBL producers, whereas 60.0% A. baumannii were ESBL producers. MBL was identified in 25.5% of P. aeruginosa, followed by A. baumannii (20.0%). P. rettgeri, S. maltophilia and Myroides species were not tested for ESBL and MBL production (Figure 1). Among NFGNB, 68.6% P. aeruginosa, and 60.0% A. baumannii were biofilm producer, while P. rettgeri, S. maltophilia, and Myroides species were not biofilm producers (Figure 2).
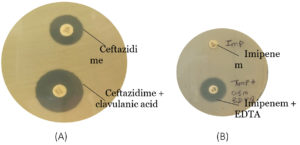
Figure 1. Phenotypic Identification of ESBL and MBL Producing NFGNB. (A) Double-Disc Diffusion method showed increase in zone of inhibition with addition of clavulanic acid in ceftazidime; (B) Similarly, Inhibition enhances with synergy of EDTA and Imipenem
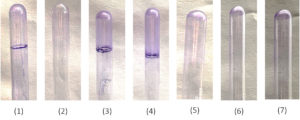
Figure 2. Identification of Biofilm Producing NFGNB by tube method; 1: Positive control; 2: Negative control; 3: A. baumannii; 4: P. aeruginosa; 5: P. rettgeri; 6: S. maltophilia; 7: Myroides species
Plasmid profiling
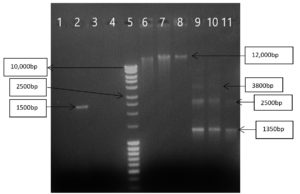
Among total isolated Pseudomonas species, 25 strains showed the presence of 2 plasmids, and 12 strains had 3 plasmids of size 3800 bp, 2500 bp, and 1350 bp, respectively. In the case of Acinetobacter baumannii, 6 strains showed 1 plasmid of size 12,000 bp. One isolate of Providencia rettgeri showed the presence of 1 plasmid of size 2500 bp (Figure 3).
Figure 3. Agrose gel electrophoresis of plasmid isolated from NFGNB; 1, 2- P. rettgeri; 3- S. maltophilia; 4- Myroides species; 5 ladder; 6, 7, 8- A. baumannii; 9, 10, 11- P. aeruginosa
Antibacterial activity of Moringa oleifera
Isolated strains of P. aeruginosa and A. baumannii exhibited resistance to conventional antibiotics, therefore, we examined if an extract of Moringa oleifera leaves could control these pathogens. The antimicrobial activity of Moringa oleifera was investigated using the agar well diffusion assay. Zones of inhibition (mm) were noted around each well containing different concentrations (20 mg, 40 mg, 80 mg, 160 mg, 320 mg per ml) of Moringa oleifera leave extract. Colistin (20 µl/ml) was taken as a positive control. At concentration of 160mg/ml, average zone of inhibition was 8mm against all tested MDR strains of P. aeruginosa and A. baumannii whereas at 320mg/ml concentration average zone of inhibition was 13mm and 15mm against all tested MDR strains of P. aeruginosa and A. baumannii respectively. While at lower concentration (20mg/ ml, 40mg/ml and 80mg/ml) leaf extract of M. oleifera did not exhibit antibacterial activity (Table 2, Figure 4).
Table (2):
Antimicrobial Activity of extract of Moringa oleifera by agar well diffusion method.
| MDR isolates | Zone of Inhibition at various concentrations of plant extract | Colistin 20µl/ml |
||||
|---|---|---|---|---|---|---|
| 20mg/ml | 40mg/ml | 80mg/ml | 160mg/ml | 320mg/ml | ||
| Pseudomonas aeruginosa | No activity | No activity | No activity | 8mm | 13mm | 17 mm |
| Acinetobacter baumannii | No activity | No activity | No activity | 8mm | 15mm | 16 mm |
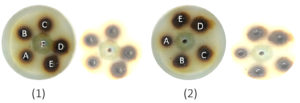
Figure 4. Agar well diffusion assay (1) P. aeruginosa (A) 20mg/ml, (B) 40mg/ml, (C) 80mg/ml, (D) 160mg/ml, (E) 320mg/ml concentration of Moringa oleifera (F) Colistin 20µl/ml; (2) A. baumannii (A) 20mg/ml, (B) 40mg/ml, (C) 80mg/ml, (D) 160mg/ml, (E) 320mg/ml concentration of Moringa oleifera (F) Colistin 20µl/ml
Throughout the world, UTIs is one of the most common infections affecting human of all ages and gender. The frequency of UTIs is elevated more in women as compared to men because of the shorter urethra. The current study examined the frequency of NFGNB in suspected UTI patients. It was observed that out of 1538 urine samples, 809 samples were culture positive. In contrast to our study, Milud Ahmed Salem et al. conducted a study in which 1423 urine specimens were tested, and 291 (20.4%) specimens were culture positive.11 Similarly, Guesh Gebre mariam et al. collected 341 urine samples during their study period and found that 72 were culture positive.12 Among total isolates, 193 were gram-positive cocci, 511 were gram-negative bacteria, and 105 were candida. Out of 511 GNB, 100 bacterial isolates were NFGNB. Further, it was observed that Pseudomonas species (86.0%) was the most common NFGNB, followed by A. baumannii (10.0%), P. rettgeri (2.0%), Stenotrophomonas maltophilia (1.0%) and Myroides species (1.0%). Similarly, Malini et.al, Rit K et al. and Wang H et al. reported Pseudomonas as prominent NFGNB (64.0%, 69.4%, and 46.9%) followed by A. baumannii (22.0%, 45.0 %, 31.0%), and S. maltophilia (5.0%, 2.0% and 9.2%) respectively in UTI patients.4,13,14 Besides this, antibiotic sensitivity profiling revealed that imipenem was the most effective drug with a sensitivity rate of 88.6%, followed by meropenem (64.7%) against isolated NFGNB. Particularly in the case of Pseudomonas species, meropenem (79.0%) was the most effective drug, followed by imipenem (74.4%) and piperacillin-tazobactam (73.25%). In contrast, ceftazidime and cefepime were least effective antibiotics and showed 8.1% sensitivity against NFGNB. In the case of A. baumannii, amikacin and gentamycin also showed a 70.0% sensitivity rate. P. rettgeri and Myroides species showed resistance to maximum antibiotics except imipenem, meropenem, amikacin, and gentamycin, whereas Stenotrophomonas maltophilia showed intrinsic resistance to maximum tested antibiotics but 100% sensitivity observed in case of Polymyxin B only. Dash M et al. illustrated that most P. aeruginosa isolates were sensitive to Imipenem, meropenem, and piperacillin-tazobactam.15 Different studies reported higher susceptibility rates for Imipenem, meropenem, and piperacillin-tazobactam than other drugs. In the case of A. baumannii, imipenem and meropenem were 80.0% sensitive; besides this amikacin and gentamicin showed a 70.0% sensitivity rate. P. rettgeri showed 100% sensitivity against imipenem, and meropenem. Myroides species showed 100% sensitivity against imipenem, amikacin, and gentamicin. Parallel to our study, Hadadi A et al. found imipenem was 77.0% sensitive, followed by piperacillin-tazobactam (54.5%),16 whereas in another study, Dimple et al. observed that 96.0% of Acinetobacter species was resistant to penicillin, 91.0% resistant against carbapenems and cephalosporin.17 Mohammad et al. reported that Acinetobacter species was 100% resistant to amikacin, gentamycin, imipenem, and meropenem.18 Another aspect on concerning a therapeutic challenge to clinicians in treating infections with NFGNB due to their increasing level of resistance against several classes of antibiotics. Cephalosporin and carbapenems group of antibiotics considered as the most favoured drug for treatment resulting in development of several enzymes by microbial pathogens which degrade its efficacy.19 In our study, Among NFGNB, 66.2% of Pseudomonas species were ESBL producers, followed by A. baumannii (60.0%). Metallo β-Lactamase was identified in 25.5% of Pseudomonas species, followed by A. baumannii (20.0%). ESBL &MBL production was not tested in P. rettgeri, S. maltophilia and Myroides species. Similarly, Peshattiwar PD et al. observed production of ESBL in 22.2% and MBL in 7.8% of Pseudomonas spp. out of 126 species.20 Kotwal et al. found that 6.0% of Pseudomonas spp. was ESBL producers, and 4% were MBL producers.21 On the other hand, Kaur et al. showed that 27.5% of Acinetobacter baumannii were ESBL, and 44.8% were MBL producers. This resistance delivers significant difficulties in the treatment; hence these drugs should be used judiciously to avoid the emergence of resistance.22 The capability of Gram-negative non-fermenting bacteria of the genera Pseudomonas and Acinetobacter to quickly attach to solid surfaces and form biofilm is a critical trait that allows them to survive in less-than-ideal environments and to colonize in a wide range of ecological niches.23 We observed that 68.6% of the P. aeruginosa and 60.0% of A. baumannii were biofilm producer, while P. rettgeri, S. maltophilia, and Myroides species were not biofilm producer. Similar to our finding, Falk KN et al. reported that 55.8% of uropathogen were biofilm producers.24 Subramanian P et al reported 75.0% P. aeruginosa and 50.0% Acinetobacter were biofilm producer.25 In contrast to our study, Mohammed et al. reported that 17% of Pseudomonas species was biofilm producer.26 It was previously reported that Plasmids could be the possible carrier of antibiotic-resistant genes among bacteria in the environment. Plasmids are extra-chromosomal DNA generally present in a free state in the cytoplasm of bacteria and associated with carrying resistant genes and confers the drug resistance.27 In our study, among total isolated P. aeruginosa, 25 strains showed presence of 2 plasmids, and 12 strains had 3 plasmids of size 3800 bp, 2500 bp, and 1350 bp. Talukder A. et al. showed that 60.0% of the Pseudomonas isolates had double plasmids of 1000–2000 bp sizes.28 Another finding of Akingbade et al. showed two P. aeruginosa isolates had double plasmids, and six of them contained a single plasmid.29 In case of A. baumannii 6 strains showed 1 plasmid band of 12,000 bp. Compared with the study of Monjurul Haque et al., where among the 4 Acinetobacter isolates, 1 isolate showed no band and 2 showed only a single plasmid band and 1 isolate showed multiple band.30 Bacterial strains harbouring plasmid showed resistance to Ceftazidime, Cefepime, Amikacin, Gentamicin, Tobramycin. Various studies have shown that bacterial pathogens lose their resistance markers after losing their plasmids. Thus, it can be concluded as plasmid may facilitate drug resistance among bacterial pathogens. Plasmids, a type of mobile genetic material, is one of the main culprits for spreading resistance genes among bacterial community and serve as a transporter for additional resistance mechanisms.31 Moringa oleifera is considered as the medicinal plant due to its pharmacological activities. M. oleifera has several Phyto-chemical compounds, glycosylates, flavonoid, alkaloids, tannins, saponins, and triterpenoids. Many of these compounds have potent anti-helminthic, anti-fungal and antibacterial activity. As per WHO survey, 80% of the world population favours herbal extract and their active components as traditional medicine.32 Furthermore, in vitro antibacterial activity of leaf extract of M. oleifera against MDR strains of NFGNB (P. aeruginosa and A. baumannii) was evaluated by agar well diffusion assay and interestingly M. oleifera exhibits impressive antibacterial activity against all isolated MDR strains of P. aeruginosa and A. baumannii. While Muhuha et al.33 study revealed that extract of M. oleifera at 400 mg/ml concentration exhibited the highest zone of inhibition with a diameter of 8.5 mm against MDR P. aeruginosa.
The current study comes to the conclusion that the NFGNB, once thought of as contaminants or commensals, are now emerging as significant bacteria capable of producing UTIs. Because of the diversity in the NFGNB’s antibiotic sensitivity profile, it is essential to identify them and keep track of their susceptibility profile in order to manage UTIs effectively. Moreover, bacterial resistance is largely influenced by the production of beta-lactamases (ESBL and MBL) and biofilms. Initially, it should be routinely advised that markers of ESBLs or MBL be detected in order to combat such growing resistance. Better infection control practices and antibiotic stewardship programmes will be required to stop or slow the establishment and spread of antibiotic-resistant NFGNB because, these bacteria have high potential for survival in the healthcare settings. Recent research also revealed that the evolution and spread of antibiotic resistance among isolates can also be determined using plasmid and chromosomal profiling. In addition to this, leaf extract of Moringa oleifera may be used as an alternative medication. However, more research and purification are required to determine the therapeutic value of specific compound present in the extract of Moringa oleifera.
ACKNOWLEDGMENTS
The authors would like to thank Maharishi Markandeshwar (Deemed to be University), Mullana, Ambala, Haryana India, for their support.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
SS conceptualized the study. SS and SA designed the study. JD and MH collected resources. JD, MH, H and CK data collection. RB carried out the initial analyses. SA, JD, MH, H and CK data interpretation. NK and DM coordinated and supervised data collection. SA and SS wrote the manuscript. SS, JC, RB and DM reviewed the manuscript. SS, JC and RB revised the manuscript. All authors read and approved the final manuscript for publication.
FUNDING
None.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
This study was approved by the Institutional Ethics Committee of MMIMSR, MMDU, with IEC number: 1410.
- Aprameya I. Non-Fermenting Gram-Negative Bacilli (NFGNB) Other Than Pseudomonas. J Acad Clin Microbiol. 2013;15:59-61.
Crossref - Yadav SK, Jha UK, Sherchan JB. Urinary Tract Infection Due to Non-fermentative Gram-Negative Bacilli in Tribhuvan University Teaching Hospital, Kathmandu, Nepal. NUTA Journal. 2020;7(1-2):71-78.
Crossref - Juyal D, Prakash R, Shanakarnarayan SA, Sharma M, Negi V, Sharma N. Prevalence of non-fermenting gram negative bacilli and their in vitro susceptibility pattern in a tertiary care hospital of Uttarakhand: A study from foothills of Himalayas. Saudi J Health Sci. 2013;2:108-12.
Crossref - Malini A, Deepa EK, Gokul BN, Prasad SR. Nonfermenting Gram-Negative Bacilli Infections in a Tertiary Care Hospital in Kolar, Karnataka. J Lab Physicians. 2009;1(2):62-6.
Crossref - Berwal A, Shobha KL, Gupta R, et al. Non-Fermenting Gram Negative Bacteria as Uropathogens In Causing Urinary Tract Infection And Its Antimicrobial Susceptibility Pattern At A Tertiary Care Centre Of South India. J Pure Appl Microbiol. 2020;14:2033-8.
Crossref - Aman S, Mittal D, Shriwastav S, et al. Prevalence of Multidrug-Resistant Strains in Device Associated Nosocomial Infection and Their In Vitro Killing by Nanocomposites. Ann Med and Surg. 2022;78:103687.
Crossref - Hassan A, Usman J, Kaleem F, Omair M, Khalid A, Iqbal M. Evaluation Of Different Detection Methods Of Biofilm Formation In The Clinical Isolates. Braz J Infect Dis. 2011;15:305-11.
Crossref - J.Doly, Imboim HC. A Rapid Alkaline Extraction Procedure for Screening Recombinant Plasmid DNA. Nucl Acids Res. 1979;7:1513-23.
Crossref - Yang CH, Su PW, Moi SH, Chuang LY. Biofilm formation in Acinetobacter Baumannii: genotype-phenotype correlation. Molecules. 2019;24(10):1849.
Crossref - Valgas C, de Souza SM, Smânia EFA, Smânia A. Screening Methods to Determine Antibacterial Activity of Natural Products. Braz J Microbiol. 2007;38:369-80.
Crossref - Derese B, Kedir H, Teklemariam Z, Weldegebreal F, Balakrishnan S. Bacterial Profile Of Urinary Tract Infection And Antimicrobial Susceptibility Pattern Among Pregnant Women Attending At Antenatal Clinic In Dil Chora Referral Hospital, Dire Dawa, Eastern Ethiopia. Ther Clin Risk Manag. 2016;12:251.
Crossref - Gebremariam G, Legese H, Woldu Y, Araya T, Hagos K, GebreyesusWasihun A. Bacteriological Profile, Risk Factors and Antimicrobial Susceptibility Patterns Of Symptomatic Urinary Tract Infection Among Students Of Mekelle University, Northern Ethiopia. BMC Infect Dis. 2019;19(1):950.
Crossref - Rit K, Nag F, Raj HJ, Maity PK. Prevalence and Susceptibility Profiles Of Nonfermentative Gram-Negative Bacilli Infection In A Tertiary Care Hospital Of Eastern India. Indian J Clin Pract. 2013;24:5.
- Wang H, Chen MJ. Changes Of Antimicrobial Resistance Among Nonfermenting Gram-Negative Bacilli Isolated from Intensive Care Units From 1994 To 2001 In China. Zhonghua Yi XueZaZhi. 2003;83:385–90. PMID: 12820914
- Dash M, Narasimham M, Padhi S, Pattnaik S. Antimicrobial Resistance Pattern of Pseudomonas Aeruginosa Isolated From Various Clinical Samples In A Tertiary Care Hospital, South Odisha, India. Saudi J Health Sci. 2014;3:15.
Crossref - Hadadi A, Rasoulinejad M, Maleki Z, Yonesian M, Shirani A, Kourorian Z. Antimicrobial Resistance Pattern of Gram-Negative Bacilli of Nosocomial Origin At 2 University Hospitals In Iran. Diagn Microbiol Infect Dis. 2008;60:301-5
Crossref - Dimple R, Nupur S, Mahawal BS, Ankit K, Ajay P. Speciation And Antibiotic Resistance Pattern Of Acinetobacter Species In A Tertiary Care Hospital In Uttarakhand. Int J Med Res& Health Sci. 2016;5:89–96.
- Mohammed MA, Alnour TMS, Shakurfo OM, Aburass MM. Prevalence and Antimicrobial Resistance Pattern Of Bacterial Strains Isolated From Patients With Urinary Tract Infection In Messalata Central Hospital, Libya. Asian Pac J Trop Med. 2016;9:771-6.
Crossref - McGowan JE. Resistance In Nonfermenting Gram-Negative Bacteria: Multidrug Resistance To The Maximum. Am J InfectControl. 2006;34:S29-37.
Crossref - Peshattiwar PD, Peerapur BV. ESBL And MBL Mediated Resistance in Pseudomonas Aeruginosa: An Emerging Threat To Clinical Therapeutics. J Clin Diagnostic Res. 2011;5(8): 1552-4. http://hdl.handle.net/123456789/1850
- Kotwal A, Biswas D, Kakati B, Singh M. ESBL And MBL In Cefepime Resistant Pseudomonas Aeruginosa: An Update from A Rural Area In Northern India. J Clin Diagnostic Res. 2016;10(4):DC09.
- Kaur A, Singh S. Prevalence of Extended Spectrum Betalactamase (ESBL) And Metallobetalactamase (MBL) Producing Pseudomonas Aeruginosa And Acinetobacter Baumannii Isolated From Various Clinical Samples. J Pathog. 2018;2018.
Crossref - Donlan RM, Costerton JW. Biofilms: Survival Mechanisms of Clinically Relevant Microorganisms. Clin Microbiol Rev. 2002;15(2):167-93.
Crossref - Falk KN, Satola SW, Chassagne F, Northington GM, Quave CL. Biofilm Production by Uropathogens In Postmenopausal Women With Recurrent And Isolated Urinary Tract Infection. Female Pelvic Med Reconstr Surg. 2022;28(4):e127-32.
Crossref - Subramanian P, Shanmugam N S, Sivaraman U, Kumar S, Selvaraj S. Antiobiotic Resistance Pattern of Biofilm-Forming Uropathogens Isolated from Catheterised Patients in Pondicherry, India. Australas Med J. 2012;5(7):377.
- Almalki MA, Varghese R. Prevalence of Catheter Associated Biofilm Producing Bacteria and Their Antibiotic Sensitivity Pattern. J King Saud Univ Sci. 2020;32(2):1427-33.
Crossref - Bennett PM. Plasmid Encoded Antibiotic Resistance: Acquisition and Transfer of Antibiotic Resistance Genes in Bacteria. In: Br J Pharmacol. 2008;153(S1):S347-57.
Crossref - Talukder A, Rahman M, Chowdhury MM, Mobashshera TA, Islam NN. Plasmid profiling of multiple antibiotic-resistant Pseudomonas aeruginosa isolated from soil of the industrial area in Chittagong, Bangladesh. Beni-Suef Univ J Basic Appl Sci. 2021;10(1):1-7.
- Akingbade OA, Balogun S, Ojo DA, et al. Plasmid profile analysis of multidrug resistant Pseudomonas aeruginosa isolated from wound infections in South West, Nigeria. World Appl Sci J. 2012;20(6):766-75.
Crossref - M Haque FK, Akhter MZ. Plasmid Profile Analysis of Multi Drug Resistant Escherichia coli and Acinetobacter from Clinical Samples. Bang J Med Sci. 2015:20-21:67-73
- Beige F, Salehi MB, Bahador N, Mobasherzadeh S. Plasmid mediated antibiotic resistance in isolated bacteria from burned patients. Jundishapur J Microbiol. 2015;8.
- Amabye TG, Tadesse FM. Phytochemical and Antibacterial Activity of Moringa Oleifera Available in the Market of Mekelle. J Anal Pharm Res. 2016;2(1): 00011.
Crossref - Muhuha AW, Kang’ethe SK, Kirira PG. Antimicrobial Activity of Moringa oleifera, Aloe vera and Warbugia ugandensis on MultiDrug Resistant Escherichia coli, Pseudomonas aeruginosa and Staphylococcus aureus. J Antimicrob Agents. 2018;4(2):162-8.
© The Author(s) 2023. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.