ISSN: 0973-7510
E-ISSN: 2581-690X
Parasitic biobank of Plasmodium falciparum is almost germinal in Côte d’Ivoire. However, several high-level research topics on this parasite involve the taking into account of nature isolates but also chemo-sensitive or resistant reference strains for a better validation of results. In addition, acquisition of these reference strains is still arduous for laboratories in developing countries due to complexity of administrative procedures. For those reasons, this study aimed in to combine several procedures into a consolidated one in order to enhance the multiplication of P. falciparum reference strains. Continuous culture of plasmodial strains was based on the Trager and Jensen procedures. The CELL culture protocols used are those of the Swiss TPH described by Sergio Wittlin; the “Growing Plasmodium falciparum cultures at high parasitemia” and the “Stockholm sorbitol method” of Methods in Malaria Research-6th edition 2013; and the INV-01 and INV-02 procedures of the Worldwide Antimalarial Resistance Network (WWARN). Reference Plasmodium falciparum strains NF54 sensitive to chloroquine (CQs) and K1 resistant to chloroquine (CQr) were received from the Swiss Tropical Institute and Public Health (Swiss TPH). The CQs NF54 strain reacted more quickly to the protocol unlike the CQr K1 strain. Parasitic densities (DP) obtained with NF54 strain were ranged from 0.4% at day zero (D0) to 11.4% at day eight (D8). Strain K1 finally adapted successfully after one month of follow-up. Related DPs ranged from less than 0.1% to more than 20% in just three growth cycles after adaptation. A joint protocol (from this work) called “CRLP-SwissTPH-Pasteur_001” is available and allows to efficiently multiply reference strains NF54 and K1. It is planned to spread out the tests to other plasmodial strains as well as to wild isolates in order to standardize this procedure.
Plasmodium falciparum, Reference strain, Parasitic biobank, In vitro CELL culture
In 2018, the World Health Organization reported 228 million malaria cases with 405,000 deaths. Africa remains the most affected continent with 93% of cases and 94% of deaths1. This situation is related to several causes. One of the most important is the resistance of Plasmodium falciparum (P.f.) to the currently used antimalarial drugs such as artemisinin derivatives, amodiaquine, halofantrine, mefloquine, chloroquine and sometimes quinine2-6. Indeed, there are 5 parasite species that cause malaria in humans (P. falciparum, P. vivax, P. malariae, P. ovale, and P. knowlesi)7,8. But only 2 of them (P.f. and P.v.) pose the greatest threat. In 2018, Plasmodium falciparum. accounted for 99.7% of estimated malaria cases in the WHO African Region 50% of cases in the WHO South-East Asia Region, 71% of cases in the Eastern Mediterranean and 65% in the Western Pacific, while P. vivax. is the predominant parasite in the WHO Region of the Americas, representing 75% of malaria cases9. Therefore, search for new therapies remains critical. In the process of developing new antimalarial drugs, studies or tests onto the incriminated parasite involve the taking into account of field isolates but also chemo-sensitive or chemo-resistant reference strains for better validation of results. Unfortunately, parasitic biobank of Plasmodium is almost non-existent in Cote d’Ivoire. Indeed, the Biological Resource Center (CeReB), biobank of the Pasteur Institute of Cote d’Ivoire and also for the Sub-Region of ECOWAS countries, was inaugurated only on 25th April 201910. In addition, acquisition of Plasmodium reference strains from specialized institutions such as BEI Resource, etc. is still very difficult for our local laboratories due to complexity of related administrative procedures. Moreover, even if some research teams have got strains, multiplication attempts using one or the other of the existing protocols came up against many failures such as the low multiplication and the loss of the strains after a maintenance that lasted at most a week. To face all these shortcomings, it was decided in this work to combine several procedures (used to multiply malaria parasites in vitro) into one harmonized protocol to efficiently multiply P. falciparum reference strains.
Laboratory material and reagents
Plasmodium falciparum reference strains used in this study were the NF54 strain sensitive to chloroquine (CQs NF54), and the K1 strain which is resistant to chloroquine (CQr K1) and several other antimalarial drugs such as amodiaquine, pyrimethamine, etc. The strains have been provided from the Swiss Tropical Institute and Public Health (Swiss TPH). Complete culture medium prepared with RPMI 1640 powder medium (GIBCO®, Invitrogen™, Ref: 51800-035, Batch N°1989282) stored at 4°C; Sodium bicarbonate (NaHCO3, FW 84.01), HEPES (C8H18N2O4S, FW 238.1), Hypoxanthine (C5H4N4O, FW 136.11), L-glutamine (C5H10N2O3, FW 146.1), Glucose (C6H12O6, FW 180.16), all stored at room temperature; Gentamicin (C21H43N5O7, FW 477.596) stored at -20°C; Sterile human serum (from BIOWEST®) stored at -20°C and Albumax (Albumax-II from GIBCO®, Batch N°326580A, Invitrogen) stored at + 4°C. As equipment, we used cryogenic conservator tank (Thermo Scientific™, Ref: BioCare20 (20 L) and Thermo30 (30 L), laminar flow hood (TELSTAR™ AV-100 & ESCO Class II BSC), binocular microscope (OLYMPUS® CX 22-RFS1), culture flask (type FALCON®; from Becton Dickinson), CO2 incubator (INCUBATOR CO2 JP-SELECTA S.A.), microscope slides (HEHYGLAS®, Chine), sterile filtration system with 0.22 µm membrane filters (STERICUP™, Ref. SCGPU05RE, Millipore®), water bath (Agibat-20, JP-SELECTA S.A.), 0.22 µm and 0.45 µm Millipore syringe filters (MINISART®, Sartorius Stedim Biotech), etc.
Harmonized procedure
The continuous culture of plasmodial strains was based on the procedure described by Trager and Jensen in 1976. Culture medium has been prepared following three protocols. That of the Swiss TPH described by Sergio Wittlin11, the “Growing Plasmodium falciparum cultures at high parasitemia” and the “Stockholm sorbitol method” of Methods in Malaria Research-6th edition 201312; the INV-01 and INV-02 procedures of the Worldwide Antimalarial Resistance Network (WWARN) 2010 and 201113.
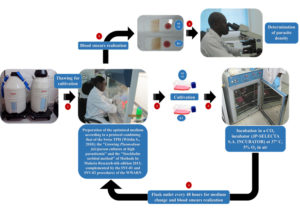
Thus, the parasites after being removed from the liquid nitrogen were rapidly warmed into a water bath at 37°C and transferred under a laminar flow hood for experiences. Thick film and thin smears have been made to determine parasitemia. Parasites were cultured according to a harmonized protocol (which we have called “CRLP-SwissTPH-Pasteur_001”) from the procedures mentioned above, based on several works14-16. The steps of the work described below, are sum up in the diagram on Fig. 1.
Fig 1. Process for the replication of Plasmodium falciparum strains NF54 and K1 following CRLP-SwissTPH-Pasteur_001 protocol
Procedure
Preparation of the different media and solutions
500 mL of RPMI (washing) medium will be prepared by measuring 400 mL of sterile distilled water in an Erlenmeyer flask to which 5.217 g of RPMI 1640 powder is added. After dissolving, 3g of HEPES is added, followed by 1g of sodium bicarbonate. The whole is homogenized and the final volume is adjusted to 500 mL with distilled water. The solution obtained is filtered at 0.22µm and 50 mL aliquots are made in conical tubes. To each aliquot 25 µL of gentamycin is added and the whole is stored at -20°C.
Then, 100 mL of albumax-hypoxanthine complex is prepared by measuring 90 mL of RPMI medium (without gentamycin) in an Erlenmeyer flask to which 5g of albumax powder is added. Dissolve and add 20 mg of hypoxanthine. The whole is homogenized and the volume is adjusted to 100 mL with RPMI medium. The solution obtained is filtered a first time at 0.45µm and then a second time at 0.22µm and stored at 2-4°C.
Finally, for this step, an optimized RPMI medium is prepared by measuring 250 mL of RPMI medium (without gentamycin) in an Erlenmeyer flask to which 1.57 mL of L-glutamine, 5 mL of decomplemented human serum, 25 mL of albumax-hypoxanthine complex, 500 mg of D-glucose and 500 mg of sodium bicarbonate are added. The whole is homogenized and filtered at 0.22µm and then 50 mL aliquots are made in conical tubes. Twenty-five (25) µL of gentamicin is added to each aliquot and stored at -20°C.
Thawing the strains followed by culture
The cryotube (containing the plasmodial strain) is taken out of the liquid nitrogen and rapidly heated to 37°C for one to two minutes. The content of the cryotube is then transferred into a 50 mL tube. While homogenizing, a volume of NaCl (12%) equivalent to 0.1 times that of the suspension inside the cryotube is added dropwise and slowly. Incubate at room temperature for five minutes, then slowly add a further volume of NaCl (1.6%) equivalent to 10 times that of the suspension in the cryotube dropwise while homogenizing. Incubate a second time at room temperature for five minutes and then the suspension is centrifuged at 2100 rpm for five minutes. The supernatant is removed and a quantity of washing RPMI equivalent to the volume removed is added. Homogenize again and centrifuge at 1500 rpm for five minutes. The supernatant is again removed and the red cell pellet is adjusted to 200 µL with red cells of the O+ group. An inoculum is then prepared at 2% by adding 10 mL of optimized RPMI medium. This inoculum is homogenized and transferred to a 24 cm2 culture flask and incubated at 37°C in a CO2 oven (set at 5% CO2 + air).
Renewal of the culture medium and preparation of the blood smear slides
Every 48 hours, the flask is carefully removed from the incubator and placed under the laminar flow hood. The supernatant (about 10 mL) is gently removed without touching the blood mat and discarded. The cell mat is slowly homogenized and 15 µL is collected for the blood smear. Then 10 mL of optimized RPMI medium is added to the flask. The solution is gently homogenized and the flask is returned to the CO2 incubator. The blood smears are fixed and stained and the slides are read with a binocular microscope to determine the different parasite densities (DP).
NF54 strain
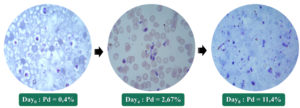
The CQs NF54 strain responded favorably and more quickly to culture conditions. From day zero (D0) to day four (D4), the parasite density (PD) was multiplied by approximately five; thus, rising from 0.4% to 2.67%. Likewise, from D4 to day eight (D8), this density increased from 2.67% to 11.4%; i.e. approximately 20 times the initial density in eight days of culture (Fig. 2).
K1 strain
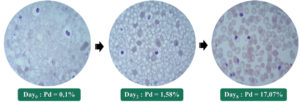
Strain CQr K1 reacted more slowly to culture conditions. In fact, a latency time of about one month was observed before perceiving the appearance of schizonts indicating the start of multiplication of the parasite. From this moment reconsidered as D0, up to two days later (D2), the parasite density (DP) increased from 0.1% to 1.58%; ie multiplied by approximately 15. Six days after D2 (ie D8), this density was multiplied by approximately 11, going from 1.58% to 17.07%. Thus, in eight days of culture after awakening of this strain K1, we obtained a parasite density of approximately 170 times the initial one (Fig. 3).
Comment
Overall, we note that this protocol recombination effectively increased the growth of the 2 strains of P. falciparum efficiently. The specific changes made in this protocol boil down to a subsequent modification of the growth medium. Optimization has targeted the association of specific elements related to each protocol; including the additional addition of L-Glutamine, Glucose and Sodium Bicarbonate and then the concomitant presence of decomplemented serum and Albumax-Hypoxanthine complex. Azar and collaborators in 2009, using the Albumax combined with a synchronization alternating Sorbitol and Percoll, were able to obtain 3 to 5 ml of 60% infected blood cells (at an initial parasite density of 10%) in just 1 week of culture17. With the Harmonized protocol, it has been possible to pass from 0.1 to about 17% infected blood cells in a week. In this study, the CQs NF54 strain has been more rapidly reactive to the protocol than the CQr K1 one. Indeed, Mogollon and collaborators in 2016 reported that CQs NF54 strain cultured in standard conditions (RPMI-1640 culture medium supplemented with L-Glutamine and 25mM HEPES (Gibco Life Technologies) + 50 mg/L hypoxanthine (Sigma), supplemented with 10% human serum and 0.225% NaHCO3, at a 5% hematocrit under 4% O2, 3% CO2 and 93% N2 gas-conditions at 75 rpm at 37°C) reacted as well as in the present study18. However, the delay in the reaction of CQr K1 is difficult to explain because it could depend on several factors. One of the elements of the culture medium that could be suspected of being a limiting factor in the growth of plasmodia is gentamicin because of its highly hydrophilicity. Nevertheless, Lawrenson et al (2018) showed that the hydrophilicity of some 4-aminoquinolines could therefore optimize activity against CQs NF54 strain, but unfortunately not to overcome chloroquine resistance in CQr K1 strain19. As well, the works of Basco et al (2004), Briolant et al (2007) and Duffy et al (2017) highlighted that some culture conditions (such as initial parasitemia, hematocrit, time of incubation, timepoint to add hypoxanthine, use of normal or substitute serum and gas mixture composition) have been singled out as influencing only the in vitro activity assay of drugs but not the ordinary growth of parasites20-22. But despite all this, CQr K1 has been more productive at the final.
A mixed protocol from this work that we entitled “CRLP-SwissTPH-Pasteur_001” is available and allows us to efficiently multiply reference P. falciparum strains CQs NF54 and CQr K1. It is planned to spread out the tests to other P. f. strains as well as to wild isolates in order to standardize this process.
ACKNOWLEDGMENTS
We would like to express our heartfelt thanks to Doctor KOUASSI Dinard, head of the National Institute of Public Health of Cote d’Ivoire (INSP-CI) for helping on the whole work. The authors are grateful to Professor MENAN E. I. Herve, head of the Center for Diagnosis and Research on AIDS and other infectious diseases (CeDReS) in Abidjan for allowing us to use their Cell culture room for the experimentation. We would like to thank the Swiss Center for Scientific Research in Cote d’Ivoire (CSRS-CI) and the Pasteur Institute of Cote d’Ivoire (IPCI) for the excellent collaboration to achieve the study.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
TKD contributed to the multiplication of parasites strains, conception and design of the study, analysis and interpretation of data, drafting the article and final version for approval. KAJ and DMT contributed to the multiplication of parasites strains and data analysis. SKD and TK contributed in providing Plasmodium falciparum reference strains and some specific reagents and consumables. NBC, BEMA and YI contributed to the multiplication of parasites strains. MEIH contributed to designing of methods used and a deep revision of the manuscript. YW contributed to study design, draft correction and final version approval.
FUNDING
None.
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the manuscript.
- World Health Organization. World Malaria Report 2019. World Health Organization. 2019.
- Plowe CV. The evolution of drug-resistant malaria. Trans R Soc Trop Med Hyg. 2009;103(1):S11-S14.
Crossref - Yavo W, Bla KB, Djaman AJ, et al. In vitro susceptibility of Plasmodium falciparum to monodesethylamodiaquine, quinine, mefloquine and halofantrine in Abidjan (Cote d’Ivoire). Afr Health Sci. 2010;10(2):111-116.
- Lim P, Dek D, Try V, et al. Ex Vivo Susceptibility of Plasmodium falciparum to Antimalarial Drugs in Western, Northern, and Eastern Cambodia, 2011-2012: Association with Molecular Markers. Antimicrob Agents Chemother. 2013;57(11):5277-5283.
Crossref - Agarwal P, Anvikar AR, Pillai CR, Srivastava K. In vitro susceptibility of Indian Plasmodium falciparum isolates to different antimalarial drugs & antibiotics. Indian J Med Res. 2017;146(5):622-628.
Crossref - Chaorattanakawee S, Saunders DL, Sea D, et al. Ex Vivo Drug Susceptibility Testing and Molecular Profiling of Clinical Plasmodium falciparum Isolates from Cambodia from 2008 to 2013 Suggest Emerging Piperaquine Resistance. Antimicrob Agents Chemother. 2015;59(8):4631-4643.
Crossref - Cox-Singh J, Davis TME, Lee K-S, et al. Plasmodium knowlesi Malaria in Humans Is Widely Distributed and Potentially Life Threatening. Clin Infect Dis. 2008;46(2):165-171.
Crossref - Calderaro A, Piccolo G, Gorrini C, et al. Accurate identification of the six human Plasmodium spp. causing imported malaria, including Plasmodium ovale wallikeri and Plasmodium knowlesi. Malar J. 2013;12(1):321.
Crossref - Fact sheet about Malaria. Accessed January 4, 2021. https://www.who.int/news-room/fact-sheets/detail/malaria
- Le Centre de ressources Biologiques (CeReB) / BIOBANQUE de l’Institut Pasteur de Cote d’Ivoire (IPCI). Accessed September 9, 2019. http://www.pasteur.ci/index.php/cereb-biobanque
- Wittlin S, Ekland E, Craft JC, et al. In Vitro and In Vivo Activity of Solithromycin (CEM-101) against Plasmodium Species. Antimicrob Agents Chemother. 2012;56(2):703-707.
Crossref - Methods In Malaria Research. 499. https://www.beiresources.org/portals/2/MR4/Methods_In_Malaria_Research-6th_edition.pdf
- Delves M, Plouffe D, Scheurer C, et al. The Activities of Current Antimalarial Drugs on the Life Cycle Stages of Plasmodium: A Comparative Study with Human and Rodent Parasites. Beeson JG, ed. PLoS Med. 2012;9(2):e1001169.
Crossref - Trager W, Jensen J. Human malaria parasites in continuous culture. Science. 1976;193:673-675.
Crossref - Tritten L, Matile H, Brun R, Wittlin S. A new double-antibody sandwich ELISA targeting Plasmodium falciparum aldolase to evaluate anti-malarial drug sensitivity. Malar J. 2009;8(1):226.
Crossref - Vargas S. Developpement d’une methodologie et optimisation d’un test colorimetrique pour la recherche de substances antimalariques d’origine vegetale. Published online 2009.
https://doi.org/10.13097/archive-ouverte/unige:12744 - Radfar A, Mendez D, Moneriz C, et al. Synchronous culture of Plasmodium falciparum at high parasitemia levels. Nat Protoc. 2009;4(12):1899-1915.
Crossref - Mogollon CM, van Pul FJA, Imai T, et al. Rapid Generation of Marker-Free P. falciparum Fluorescent Reporter Lines Using Modified CRISPR/Cas9 Constructs and Selection Protocol. Pradel G, ed. PLoS ONE. 2016;11(12):e0168362.
Crossref - Lawrenson AS, Cooper DL, O’Neill PM, Berry NG. Study of the antimalarial activity of 4-aminoquinoline compounds against chloroquine-sensitive and chloroquine-resistant parasite strains. J Mol Model. 2018;24(9):237.
Crossref - Basco Lk. Molecular Epidemiology Of Malaria In Cameroon. Xx. Experimental Studies On Various Factors Of In Vitro Drug Sensitivity Assays Using Fresh Isolates Of Plasmodium Falciparum. Am J Trop Med Hyg. 2004;70(5):474-480.
Crossref - Briolant S, Parola P, Fusai T, et al. Influence of oxygen on asexual blood cycle and susceptibility of Plasmodium falciparum to chloroquine: requirement of a standardized in vitro assay. Malar J. 2007;6(1):44.
Crossref - Duffy S, Avery VM. Plasmodium falciparum in vitro continuous culture conditions: A comparison of parasite susceptibility and tolerance to anti-malarial drugs throughout the asexual intra-erythrocytic life cycle. Int J Parasitol Drugs Drug Resist. 2017;7(3):295-302.
Crossref
© The Author(s) 2021. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.