ISSN: 0973-7510
E-ISSN: 2581-690X
In this study, an Actinomycetota Streptomyces mediolani strain CF51DZ was isolated from a beetle Protaetia morio collected in a cork oak forest in northeast Algeria, to extract, purify and characterize the antibacterial compound. The strain was isolated in specific ISP culture media, identified via morphological, biochemical, physiological and molecular characteristics and then studied its antibacterial activities against pathogenic bacteria using the well diffusion technique. The extraction, purification and characterization of the bioactive compound was carried out using chromatographic and spectroscopic methods. Morphological, physiological, and biochemical properties, as well as the 16S ribosomal DNA sequence of the strain, were found close to those of Streptomyces mediolani. The isolated strain produced an active compound having potent antibacterial properties against Escherichia coli, Pseudomonas aeruginosa, and Staphylococcus aureus. This compound was isolated and purified with thin layer chromatography and high-performance liquid chromatography, and chemically identified by mass spectroscopy and NMR spectroscopy. In addition, the chemical structure elucidation of the antibacterial metabolite by APCI-MS analysis revealed a peak at m/z 375 [M+H]+, a molecular formula of C19H18O8. This compound was identified as methyl rosarinate: 3-(3,4-dihydroxyphenyl)-1-methoxy-1-oxopropan-2-yl (E)-3-(3,4-dihydroxyphenyl)acrylate based on the IUPAC nomenclature. Moreover, the antibacterial agent was purified from Streptomyces mediolani strain CF51DZ that was for the first time, in Algeria, isolated from the beetle Protaetia morio. This supports further characterization of this promising chemical to ensure the best use for antibacterial applications.
Actinomycete, Beetle Protaetia morio, Streptomyces mediolani strain CF51DZ, 16S Ribosomal DNA Sequence, Antibacterial Activity, Chemical Structure
In Algeria, cork oak forests are mainly important as an essential element of the physical and climatic balance.1 Also, they are omnipresent in Algeria and cover an extended area of more than 278,000 ha along with great ecological value and entomofauna biodiversity, and accordingly, major assets become highly important to ensure their conservation.1 The conditions prevailing in this type of habitat mean that they are populated by a specialized fauna, rich in the order Coleoptera, known as the richest species on earth and the major biodiversity element.2 Furthermore, Protaetia morio is a species occupying a wide variety of functional niches and microhabitats across a wide range of spatial and temporal scales, and fed mainly on plant organic matters.3
The drug resistance possibly solved by the research of new bioactive antibiotics which can be primarily produced by many types of microorganisms. Highly promising prospects for potentially producing new substances are novel or non-cultivable species; unexplored environments represent a possible source of novel microorganisms.4,5 Actinomycetes are one of the most diverse bacterial groups in nature, able to generate bioactive molecules.6,7 Discovery of new secondary metabolites as bioactive molecules in unexplored habitats is pointed out as strategic actions to fight against the growing number of microorganisms that are resistant to a wide range of antibiotics obtained from actinomycetes.8 Many actinobacteria, including Streptomycetes have been isolated as appropriate defensive partners associated withvarious types of insects,9 and in this regard, Streptomyces and Pseudonocardia were reported to protect the leaf-cuttingants against pathogenic fungus.10 Additionally, these insect-associated actinomycetes are possibly found in various female beewolf wasps, providing a specialized antennal reservoir for cultivating Streptomyces philanthi, and cover the cocoon wall to protect larva against pathogens.11,12
The current investigation was therefore, undertaken to isolate and identify the actinomycete symbionts of beetle Protaetia morio collected in northeastern Algeria, and among the isolated and identified strains, only Streptomyces mediolani strain CF51DZ has been studied. Also, the study issued the hypothesis that this species produces antibacterial compounds having probably have a health benefit, and noteworthy, the research aimed to discover novel biologically active secondary metabolites.
Description of the studied site and sampling
Twenty beetle samples were collected in Spring 2022, from cork oak trees in the national forest of Ouled Bechih region belonging to the municipality of Mechrouha, and Souk Ahras city (North-East of Algeria) at coordinates of 36°27’06.7″ N and 7°56’35.4″ E. Samples were transported in sterile vials for the experimental use. The beetles were separately smashed within sterilized 10% phosphate-buffered saline (PBS) and then decimal dilution series were prepared.13 ISP2 agar composed of 10 g malt extract, 4 g glucose, 4 g yeast extract, 20 g agar, and 1000 ml H2O, with a pH 7.2 (25 μg/ml nystatin was added to the medium to prevent the development of fungi and 10 μg/ml nalidixic acid to ovoid Gram-negative and some Gram-positive bacteria), as previously described by Shirling and Gottlieb,14 was inoculated with 100 μl of each dilution. After that, at 30°C, the plates were incubated for 21 days, and mean while, they were inspected daily for growth. The isolate was stored at -20°C by cultivating it with 15% glycerol in ISP2 broth for 7 days.6
Identification of strains CF51DZ
The strain CF51DZ was identified using ISP1, ISP2, ISP3, ISP4 and ISP5 culture media, and melanoid pigments were screened on ISP6 and ISP7 for 7-14 days at 30°C.14 The morphology of hyphae with the spore chain was observed under optic microscopy and scanning electron microscope (Carl Zeiss, SMT GmbH, Oberchoken, Germany). Analytical profiling index (API) 50CHB and ZYM strips tests were applied according to the instructions provided by the manufacturer (bioMerieux, France).
Using forward and reverse primers, the 16S rRNA gene was amplified by polymerase chain reaction (PCR), created using the protected zones inside the rRNA operon of Escherichia coli.15 The used forward primer (27F) (5′-AGAGTTTGATCCTGGCTCAG-3′) and reverse primer (1525R) (5′-AAGGAGGTGATCCAAGCC-3′) are extended, respectively from base position 8 to 27, and 1,541 to 1,525. The Wizard® Genomic DNA Purification Kit (Promega, Madison, WI, USA) was used to purify the CF51DZ strain’s genomic DNA, and applied to amplification of PCR (40 cycles, 94°C for 45 s denaturation, 60°C for 90 s primer annealing, and 72°C for 60 s extension). 16S rRNA gene nucleotide sequences were performed at least three times on both strands with the automated DNA sequencer ABI PRISM® 3100-Avant Genetic Analyser (Applied Biosystems, Foster City, CA, USA) and BigDye Terminator cycle sequencing ready reaction kits. Sequencing data were grouped by the STADEN (version 4.5) and DNASTAR (DNASTAR Inc., Madison, WI, US) software packages. Nucleotide sequence alignment was accomplished by BioEdit version 7.0.2 software. The nucleotide sequence data were analyzed by the Softberry Gene Finding tool, while those of the studied 16S rRNA (1432 bp of strain CF51DZ) gene were registered in the GenBank/ENL/DDPJ under accession no. OQ976966.
Antibacterial activity of strain CF51DZ
The antibacterial activity of CF51DZ was tested by diffusion on agar wells.8 In brief, wells of 6 mm in diameter were formed by a sterile well driller on Muller Hinton agar medium previously inoculated with pathogenic bacteria (Escherichia coli ATCC 25922, Pseudomonas aeruginosa ATCC 27853 and Staphylococcus aureus ATCC 29213) obtained from the Laboratory of Organic Chemistry and Interdisciplinary (Research’s Team: valorization of active biomolecules), University of Souk-Ahras, Algeria. After centrifugation at 8000 rpm/20 min, 80 μl of supernatant) of liquid culture of strain CF51DZ was deposited in well. After allowing for diffusion over night at 4°C, the plates were incubated for 24 hours at 37°C. A caliper was used to measure the inhibitory zones. The experiment was performed in triplicate.
Extraction of crude antibacterial compound
The strain was inoculated into 100 ml of ISP2 broth in an Erlenmeyer flask, maintained in an incubator shaker at 30°C for 7 days, then centrifuged to eliminate biomass. The cell-free supernatant was extracted with ethyl acetate. To obtain the crude extract, the solvent phases were evaporated at 70°C by a vacuum evaporator (Heidolph Laorota, 4001). 1 ml of methanol was used to dissolve the crude residue for further use.6,16 The obtained compound was used to determine the antibacterial activity via the agar well diffusion method as previously described. 24 hours after the 37°C incubation period, the Petri dishes were inspected for the inhibition zones.
Chromatographic analysis of active extracts
To purify the biomolecules, the organic phase produced after extraction from the supernatant of the culture of CF51DZ strain was exposed to various chromatographic stages: TLC, UV spectrophotometry, HPLC.17 The crude extract was then fractionated on TLC silica gel (Merck, Germany) by eluting with n-butanol, acetic acid, and water in the ratio of 3:3:1 as mobile phase. The TLC plates were air-dried and visualized under UV (254 nm and 365 nm) with an anisaldehyde-sulphuric acid reagent.18 The active substances appeared on silica gel plates, and the value of the retention factor (RF) was calculated. Further, the absorption spectra of active fraction obtained from TLC were spectrophotometrically (Shimadzu 2600 UV/VIS spectrophotometer), evaluated in the UV range (200-500 nm). The column chromatography, Medium Pressure Chromatography System was plied using reverse phase C18 column. Additionally, analytical HPLC-UV/VIS profiles by HPLC System (Agilent Technologies, USA) were applied to count the secondary metabolites in the form of chromatographic peaks.
The chemical structures were determined by mass spectrometer with an ion trap analyzer and ESI source under Atmospheric Pressure Chemical Ionization (SM-APCI) conditions, and nuclear magnetic resonance (NMR) whose spectra were acquired on a Bruker Avance 500 MHz. Deuterated solvent of NMR technique (ppm; CD3OD δH = 3.29 ppm, δC = 48.0 ppm) was supplied by Sigma-Aldrich and Euriso-top, and its chemical shifts (δ) were expressed in ppm to the deuterated solvent used as internal standard. The coupling constants (J) were expressed in Hertz and their multiplicity was represented by the following abbreviations: m (multiplet), q (quadruplet), t (triplet), d (doublet), and s (singlet).
Identification of strain CF51DZ
The isolated strain CF51DZ was identified on morphological, biochemical, physiological, and molecular characteristics. Based on the Bergey’s Manual of Systematic Bacteriology, these bacterial properties shown in Table 1 proved that the isolated and the identified CF51DZ strain is a Gram-positive, long filamentous aerial mycelium, rectifiable. Electronic microscopy revealed straight, smooth, spiral spores. Strain CF51DZ revealed good growth on ISP2, ISP4, ISP5, ISP6, and ISP7 medium and small growth on ISP1. On ISP2, ISP3, and ISP4, aerial mycelium was prevalent, moderately on ISP5 and ISP6, and absent on ISP1 and ISP7. It was white on ISP2, ISP3, and ISP4 media, and grey on ISP5, ISP6, and ISP7 media. The actinomycete properties described in Bergey’smanual of systematic bacteriology19 proved that Streptomyces is the genus CF51DZ was found to be an aerobic, catalase-positive, and oxidase-positive strain. It is able to utilize glucose, fructose, mannitol, mannose, maltose, sucrose, cellobiose, raffinose, D-xylose, D-ribose, L-arabinose, galactose and fucose, but not starch, lactose, inositol, adonitol, erythritol and L-rhamnose based on the obtained data using API 50 CHB gallery (Table 1).
Table (1):
Morphological, biochemical and physiological characteristics of strain CF51DZ
| Properties | Characteristics | Properties | Characteristics |
|---|---|---|---|
| Morphological Features | Sucrose | + | |
| Spore morphology | Spirale, straight | Xylose | + |
| Spore mass | Smooth white | Galactose | + |
| Soluble pigment | – | Mannose | + |
| Biochemical Features | Optimum pH | 5-11 | |
| Gram Staining | Positive | Optimum Temperature | 25-37°C |
| Oxidase | – | Resistant to NaCl | 5-10% |
| Catalase | + | Fucose | + |
| Hydrolysis of Starch | – | Arabinose | + |
| Hydrolysis of Gelatin | + | Ribose | + |
| Production of Indole | + | Lactose | – |
| VogesProskauer | + | Erythrol | – |
| Activity of Urease | + | Adonitol | – |
| Utilization of Citrate | + | Inositol | – |
| Production of H2S | – | alkaline phosphatase | + |
| Production of Nitrate | + | esterase lipase (C8) | + |
| Production of Melanin | – | leucine arylamidase | + |
| Hydrolysis of Casein | + | valine arylamidase | + |
| Dextrose | + | lipase (C14) | – |
| Fructose | + | trypsin | – |
| Glucose | + | α-chymotrypsin | – |
| Maltose | + | N-acetyl-β-lucosamidase | – |
| Mannitol | + | β-glucuronidase | – |
| Rhamnose | – | α-mannosidase | – |
| Raffinose | + | α-fucosidase | – |
| Cellubiose | + | ||
The production of several enzymes by the isolated strain was examined. The strain enzymes identified by API ZYM tests exhibited leucine arylamidase, esterase lipase (C8), and valine arylamidase alkaline phosphatase, but not N-acetyl-β-lucosamidase, α-mannosidase, α-fucosidase activity, lipase (C14), trypsin, α-chymotrypsin and β-glucuronidase. CF51DZ strain was halophilic and grew up to 10% w/v NaCl. The optimal tolerance range for NaCl was 2.5% to 5%, while the optimal pH was 7 and the optimal temperature was 25 to 37°C.
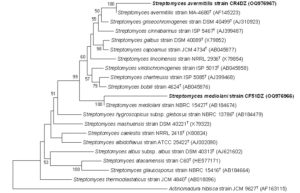
The phylogenetic tree (Figure 1) mentioned that the new isolates were clustered with members of the Streptomyces, the closest neighbor being Streptomyces mediolani NBRC 15427 T (GenBank accession no.: AB184674) (97.52% sequence similarity) for strain CF51DZ. The obtained results suggested that these isolates should be assigned as Streptomyces mediolani strain CF51DZ.
Figure 1. Phylogenetic tree illustrating the relations between strains CF51DZ (GenBank accession no.: OQ976966) and closely associated species that have been previously reported, as deduced from 16S rRNA gene sequences of the genus Streptomyces. Based on a comparison of around 1200 bp, the tree was built using the neighbour-joining (NJ) method in the MEGA program, and it was visualized using NJplot. Numbers at nodes or branch points (> 50%) indicate support for the internal branches within the tree obtained by bootstrap analysis (percentages of 1000 bootstraps). Actinomadura hibisca strain JCM 9627T (GenBank accession no.: AF163115) is the outgroup. Bar, 0.01% sequence divergence. The NCBI accession numbers are provided in parenthesis
Antibacterial activity of strain CF51DZ
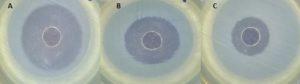
Significant inhibition was observed in the culture filtrate of strain CF51DZ, particularly against pathogenic Staphylococcus aureus ATCC 29213, Pseudomonas aeruginosa ATCC 27853 and Escherichia coli ATCC 25922, with an inhibition zone of 40, 20 and 18 mm respectively (Figure 2).
Figure 2. Antibacterial activity of bioactive compound produced from Streptomyces mediolani strain CF51DZ against (A) Staphylococcus aureus ATCC 29213, (B) Pseudomonas aeruginosa ATCC 27853, (C) Escherichia coli ATCC 25922, by agar well diffusion method
Production, extraction and purification of active compound
Production was conducted for 7 d at 30°C using ISP2 broth as a production medium, and the supernatant containing the active metabolite was then extracted with ethyl acetate. It has previously been documented that actinomycete’s crude extracts may be extracted using ethyl acetate.20 The crude extracts were dissolved in methanol and used for subsequent analysis by TLC and HPLC. Two different bioactive spots were obtained, spot 1 with RF = 0.81 and spot 2 with RF = 0.95.
TLC bioautography test detected an active spot with RF = 0.81 against Escherichia coli ATCC 25922, Pseudomonas aeruginosa ATCC 27853 and Staphylococcus aureus ATCC 29213. Of note, purified antibiotic was designated 51A, whose purity exhibited antibacterial activity as confirmed by HPLC, showing a single peak with a retention time of 39.670 min. The analysis was then performed on the active fraction, showing the highest activity against Staphylococcus aureus ATCC 29213 and Pseudomonas aeruginosa ATCC 27853 with inhibitory zones of 30 mm and 19 mm, respectively. Our results suggested that the actinomycete strains associated with the intestinal beetle Protaetia morio protect them from infection by Gram-positive and negative bacteria, inhabitants, and presumably more abundant bacteria in the same environment.
Structure elucidation of compound
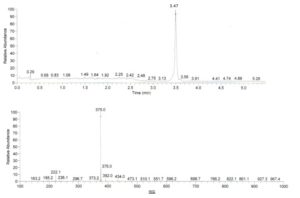
Analysis of this compound by mass spectrometry (SM-APCI) in a positive mode revealed a molecular ion m/z 375.0 [M+H]+ (Figure 3A). The compound’s 1H NMR spectra revealed two ABX systems corresponding to 1,3,4- trisubstituted phenyl rings, at δ 6.70 (d, J = 2.0 Hz, H-2), δ 6.69 (d, J = 8.0 Hz, H-5) and δ 6.57 (dd, J = 2.0; 8.0 Hz, H-6) on the one hand, and δ 7.04 (d, J = 2.0 Hz, H-2′), δ 6.91 (dd, J = 2.0; 8.5 Hz, H-6′) and δ 6.75 (d, J = 8.5 Hz, H-5′) on the other hand. Four oxygenated aromatic carbons at δ 146.1, δ 145.6, δ 147.3 and δ 150.1, corresponding to the phenol groups present in positions 3, 4, 3′ and 4′ are confirmed to be present by the 13C spectra.
This compound is a yellow powder soluble in methanol, and reacts with Neu’s reagent, displaying light green fluorescence under UV at 365 min. The signals at δ 148.6 and δ 114.1 belong to the conjugated double bond, since the signal at δ 74.8 corresponds to an alcohol function, and signals at δ 168.2 and δ 172.1 are attributed to the presence of two ester functions (Table 2). The information obtained from the compound’s 1H and 13C NMR spectra also showed an integrating signal for a methyl proton at δ 52.8 corresponding to a methyl group in this compound (Figure 3B).
Table (2):
Chemical shifts of the 1H and 13C NMR spectrum of compound 1 in deuterated methanol
Signals |
δC |
δH (m ; J en Hz) |
|---|---|---|
1 |
128.4 |
– |
2 |
117.8 |
6.70 (d; 2.0) |
3 |
146.1 |
– |
4 |
145.6 |
– |
5 |
116.9 |
6.69 (d; 8.0) |
6 |
122.8 |
6.57 (dd ; 8.0 ; 2.0) |
7a |
38.0 |
3.06 (dd; 14.5 ; 5.5) |
7b |
38.0 |
3.00 (dd; 14.5 ; 5.5) |
8 |
74.8 |
5.11 (dd ; 7.5 ; 5.0) |
9 |
172.1 |
– |
1’ |
127.5 |
– |
2’ |
115.3 |
7.04 (d ; 2.0) |
3’ |
147.3 |
– |
4’ |
150.1 |
– |
5’ |
116.1 |
6.75 (d ; 8,5) |
6’ |
123.3 |
6.91 (dd; 8.5 ; 2.0) |
7’ |
148.6 |
7.52 (d; 15.5) |
8’ |
114.1 |
6.26 (d; 15.5) |
9’ |
168.2 |
– |
-OCH3 |
52.8 |
3.72 s |
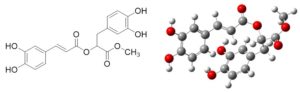
The active antimicrobial compound was identified as methyl rosmarinate with a chemical name of 3-(3,4-dihydroxyphenyl)-1-methoxy-1-oxopropan-2-yl(E)-3-(3,4-dihydroxyphenyl) acrylate, and medicinal properties owed to its presence in the medicinal plant Hyptis atrorubens.
Despite the great success in the discovery and development of the production process of new antibacterial molecules, the risks of infectious diseases resulting in worldwide death are still increasing.21 This is due to one of the main reasons involving the development of antibiotic resistance among pathogenic microbes, leading to the need to progressive search for new and more effective biomolecules.22 To our knowledge, the study is the first to investigate insects (Protaetia morio) –associated actinomycetes (symbionts) in Algeria. Studies have reported that actinomycetes belong to a symbionts group associated with bark beetles and a pine-infesting woodwasp in North America.13,23
It was found that the extent of actinobacteria resembled the observational pattern, while the growth pattern demonstrated that most isolates were cultured in ISP2, where the substrate hyphae varied from yellowish to brown color on all tested media.24 As reported,25 the majority of the antibacterial compound producers that appeared in color series varied from yellow and grey, and similarly, a recent study has reported the highest percentage of active isolates showing white and grey colors in series, since no diffusible pigment was noticed in ISP6 and ISP7 media.
As previously reported,26 the genus Streptomyces possessed stable morphological, physiological, and biochemical characteristics. The strain consumed the most tested sugars, indicating a diversified mode of carbon use. These results are highly in accordance with those of Holt.27 Interestingly, the spore morphology differing substantially between species is the key criteria of Streptomyces identification which differs. It was mentioned that among the typical features of Streptomyces, the smooth spores were arranged in chains flexibly.28 Actinomycetes were reported to have various biochemically active enzymes, meanwhile, Streptomyces species were found to produce important enzymes, such as amylase, protease, and lipase having commercial and industrial applications.
Actually, termites-associated actinobacteria improve the acquisition of nutrients from different polysaccharide arrays including cellulose.29,30 It was reported also that 78% of Streptomyces isolates from the feces of rhinoceros beetle larvae (Allomyrina dichotoma L.) are able to hydrolyze cellulose and casein owed to their typical biochemical properties.31
Recently, the actinomycetes were molecularly identified by using 16S rRNA gene analysis of actinomycetes.32
Similar previous works have reported the utility of molecular analysis of 16S rRNA in the identification of actinomycetes.26,33,34 In this context, Hye-Won et al.31 have isolated and identified 53 strains of Streptomyces from the feces of rhinoceros beetle larvae (Allomyrina dichotoma L.). These isolates exhibited antimicrobial activities against phytopathogenic fungi and Gram-positive bacteria. These results proved that diverse and valuable actinomycetes could be isolated from insect fecal samples, as well as the insect intestines may be rich sources of novel bioactive compounds.
The study of antibacterial activity revealed that the strain CF51DZ is inhibitory to the pathogenic bacteria tested: Staphylococcus aureus, Pseudomonas aeruginosa and Escherichia coli.
While this regard S. aureus, are among the etiological agents widely involved in human and veterinary medicine.35,36 Also, E. coli is responsible for diarrhea which, sometimes develops into bloody diarrhea (hemorrhagic colitis). Other health conditions, including urinary tract infection, bronchopulmonary infection, and bacteremia are associated with P. aeruginosa. Conclusively, the Gram-negatives are intrinsically more resistant to antibiotic compound than Gram-positives, and this is likely due to the existence of an outer membrane. In addition, the Gram-negative have been assigned to the exclusion of antimicrobial metabolites by transmembrane efflux pumps.37 However, our results showed that our strain inhibited both types of bacteria. The strain Streptomyces antibiotics ess_am A8 isolated from the ant Tapinoma simrothi was tested for the antimicrobial activity, inhibiting the Escherichia coli ATCC 7839, Pseudomonas aeruginosa ATCC 9027, Bacillus subtilis ATCC 6633, Staphylococcus aureus ATCC 6538, Aspergillus flavus, Fusarium moniliforme, Aspergillus niger and Candida albicans.18 Earlier reports of Hye-Won et al.,31 Anand and Kumaresh,38 and Liu et al.39 support our results reporting an antimicrobial activity of the actinomycetes, especially Streptomyces strains isolated from various sources.
Sanghvi et al.34 obtained by TLC, two fractions of an ethyl acetate extract of Streptomyces werraensis having antimicrobial activity. Previous publications have described the characterisation of major bioactive chemicals using a variety of techniques.34,40 The active antimicrobial compound was identified as methyl rosmarinate. It was the first isolated molecule from the plant Rabdosia serra (Lamiaceae). Further research showed the physicochemical properties of the purified antimicrobial compounds, quercetin 3-O-glucoside derivative, and a flavanone glycoside, extracted from Streptomyces antibiotics strain ess_amA8 isolated from the ant Tapinoma simrothi.18
According to the results, Streptomyces mediolani strain CF51DZ produces the antibiotic 51A, which may find use in the pharmaceutical sector. Additionally, there is evidence that insect bodies are the habitats of actinomycetes; symbioses occur between the two partners.
This work is the first study in Algeria investigating beetles-associated actinomycetes. The study has shown that Streptomyces spp. is a symbiont of P. morio, suggesting a connection between this Streptomyces species and the cork oak environment. This study has also provided a new research approach to establish the molecule 3-(3,4-dihydroxyphenyl)-1-methoxy-1-oxopropan-2-yl (E)-3-(3,4-dihydroxyphenyl)acrylate to be potential lead compound as an antibiotic for the drug discovery paradigms. The performed study highlights the significance of an advanced investigation aiming to track down new antibacterial metabolites from detected Streptomyces from underexplored habitats.
ACKNOWLEDGMENTS
The authors would like to express their gratitude to Dr G.N. for collecting the beetles and Laboratory of Aquatic and Terrestrial Ecosystems, University of Mohamed Cherif Messaadia, Souk-Ahras, Algeria for their valuable help during the completion of this work.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
This work was supported by the Algerian Ministry of Higher Education and Scientific Research for the Algerian-Tunisian cooperation project (Research – Development projects: Innovation and exploitation of the mixed Tunisian Algerian medicinal and aromatic flora for a controlled designation of origin of bioproducts – ACTIVPAM) with grant number PRD/TN/DZ/21/03 (AYARI-SEBAI).
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
- Djema A, Messaoudene M. The Algerian forest: Current situation and prospects. In Palahi M, Birot Y, Bravo F, Gorriz E (eds.), Modeling, valuing and managing Mediterranean forest ecosystems for non-timber goods and services. Proceedings of 57th European Forest Institute, Finland. 2009:17-28.
- Odegaard F. How manyspecies of arthropods? Erwin’s estimate revised. Biol J Linn Soc. 2000;71(4):583-597.
Crossref - Boukli HS, Abdellaoui-Hassaine K, Ponel P, Chaoui Boudghane-Bendiouis C, Bettioui R. Analyse de la structure fonctionnelle des peuplements de coleopteres dans le marais estuarien de la Tafna (Algerie). Bull Soc Zool Fr. 2014;139:5-19.
- Bukelskis D, Dabkeviciene D, Lukoseviciute-Bucelis A, Kriauciunas I, Lebedeva J, Kuisiene DN. Screening and transcriptional analysis of polyketide synthases and non-ribosomal peptide synthetases in bacterial strains from Krubera-Voronja Cave. Front Microbiol. 2019;10:2149.
Crossref - Cyske Z, Jaroszewicz W, Zabinska M, et al. Unexplored potential: biologically active compounds produced by microorganisms from hard-to-reach environments and their applications. Acta Biochim Polon. 2021;68(4):565-574.
Crossref - Ayari A, Morakchi H, Kirane-Gacemi D. Evaluation of antifungal activity of novel marine actinomycete, Streptomyces sp. AA13 isolated from sediments of Lake Oubeira (Algeria) against Candida albicans. Afr J Microbiol Res. 2016;10(6):156-171.
Crossref - Nahdhoit AR, Nihal DG. Screening of bioactive compounds for biomedical and industrial uses from Actinobacteria isolated from the Parsık Cave, Turkey, Identifying novel compounds from extreme environments. Johnson Matthey Technol Rev. 2023;67(2):159-170.
Crossref - Nait Marzoug A, Ayari A, Khaldi F, Guehria I, Gheid A. Effect of Peganum harmala L. extract supplemented ISP2 medium on growth and production of secondary metabolites of Streptomyces ayarius S115. Electron J Biotechnol. 2023;64(4):34-41.
Crossref - Sun J, Zhou XJ. Lignocellulolytic systems of insects and their potential for viable biofuels. In Sun J, Ding SY, Doran-Peterson J (eds.), Biological conversion of biomass for fuels and chemicals : exploration of natural utilization systems. Royal Society of Chemistry, Cambridge, UK. 2013;195-222.
Crossref - Schoenian I, Spiteller M, Ghaste M, Wirth R, Herz H, Spiteller D. Chemical basis of the synergism and antagonism in microbial communities in the nests of leaf-cuttingants. Proc Natl Acad Sci. 2011;108(5):1955-1960.
Crossref - Ramon I, Martinez-Carrasco AS, de la Nieta RS, et al. Characterization of actinomycetes strains isolated from the intestinal tract and feces of the larvae of the longhorn beetle Cerambyx welensii. Microorganisms. 2020;8(12):2013.
Crossref - Torres-Vila L, Mendiola-Diaz F, Sanchez-Gonzalez A. Dispersal differences of a pest and a protected Cerambyx species (Coleoptera: Cerambycidae) in oak open woodlands: A mark-recapture comparative study. Ecol Entomol. 2016;42(1):18-32.
Crossref - Human ZR, Slippers B, De Beer ZW, Wingfield MJ, Venter SN. Antifungal actinomycetes associated with the pine bark beetle, Orthotomicus erosus, in South Africa. S Afr J Sci. 2017;113(1/2):7.
Crossref - Shirling EB, Gottlieb D. Methods for characterization of Streptomyces species. Int J Syst Evol Microbiol. 1966;16(3):313-340.
Crossref - Gurtler V, Stanisich VA. New approches to typing and identification of bacteria using the 16S-23S rDNA spacer region. Microbiology. 1996;142(1):3-16.
Crossref - Ayari A, Morakchi H, Kirane D. Identification and antifungal activity of Streptomyces sp. S72 isolated from Lake Oubeira sediments in North-East of Algeria. Afr J Biotechnol. 2012;11(2):305-311.
Crossref - Ahmad I, Beg AZ.Antimicrobial and phytochemical studies on 45 Indian medicinal plants against multi drug resistant human pathogens. J Ethnopharmacol. 2001;74(2):113-123.
Crossref - Essam NS, Maged SA, Manal MY, Ashraf AM. Antimicrobial quercetin 3-O-glucoside derivative isolated from Streptomyces antibioticus strain ess_amA8. J King Saud Univ Sci. 2020;32(3):1838-1844.
Crossref - Lechevalier HA, Williams ST, Sharpe ME, Holt JG. Bergey’s manual of systematic bacteriology. Williams and Wilkins, Baltimore. 1989.
- Ravi RK, Vasantba JJ. Characterization and partial purification of an antibacterial agent from halophilic actinomycetes Kocuria sp. strain rsk4. BioImpacts. 2018;8(4):253-261.
Crossref - De Lima ProcoApio RE, Da Silva IR, Martins MK, De Azevedo JL, De ArauA Jo JM. Antibiotics produced by Streptomyces. Braz J Infect Dis. 2012;16(5):466-71.
Crossref - Singh V, Haque S, Khare S, et al. Isolation and purification of antibacterial compound from Streptomyces levis collected from soil sample of north India. PLoS ONE. 2018;13(7):e0200500.
Crossref - Hulcr J, Adams AS, Raffa KF, Hofstetter RW, Klepzig KD, Currie CR. Presence and diversity of Streptomyces in Dendroctonous and sympatric bark beetle galleries across North America. Microbial Ecol. 2011;61(4):759-768.
Crossref - Takizawa M, Colwell RR, Hill TR. Isolation and diversity of actinomycetes in the Chesapeake Bay. Appl Environ Microbiol. 1993;59(4):997-1002.
Crossref - Varalakshmi K, Subramanian G, Reddy MS. Fixed dose combination products of anti-hypertensive’s and anti-diabetic’s-Indian regulatory perspective against USA and EU. Intern J Pharm Pharm Sci. 2014;6:13-16.
- Barka A, Vatsa P, Sanchez L, Gaveau-vaillant N, Jacquard C, Klenk H. Taxonomy, physiology, and natural products of actinobacteria. Microbiol Mol Biol Rev. 2016;80(1):1-43.
Crossref - Holt G. Shorter Bergey’s manual of determinative bacteriology, Lippincott Williams and Wilkins, USA. 1994.
- Williams ST. Bergey’s manual of systematic bacteriology, 4th Ed. Baltimore, Hong Kong, London, Sydney. 1989.
- Almalki MA. Isolation and characterization of polyketide drug molecule from Streptomyces species with antimicrobial activity against clinical pathogens. J Infect Public Health. 2020;13(1):125-130.
Crossref - Firew E, Sudhamani M, Diriba M, Belachew T. Purification and characterization of bioactive metabolite from Streptomyces monomycini RVE129 derived from the Rift Valley soil of Hawassa, Ethiopia. Bio Med Res Int. 2022;141313.
Crossref - Hye-Won L, Jae-Hyung A, Minwook K, et al. Diversity and antimicrobial activity of actinomycetes from fecal sample of Rhinoceros Beetle Larvae. Kor J Microbiol. 2013;49(2):156-164.
Crossref - Nimaichand S, Zhu WY, Yang LL, et al. Streptomyces manipurensis sp. nov., a novel actinomycete isolated from a limestone deposit site in Manipur, India. Anton Leeuw Int J G. 2012;102:133-139.
Crossref - Hussain M, Rather M, Dangroo N, et al. Streptomyces puniceus strain AS13., production, characterization and evaluation of bioactive metabolites: a new face of dinactin as an antitumorantibiotic. Microbiol Res. 2018;207:196-202.
Crossref - Sanghvi G, Ghevariya D, Gosai S, et al. Isolation and partial purification of erythromycin from alkaliphilic Streptomyces werraensis isolated from Rajkot, India. Biotechnol Rep. 2014;1-2:2-7.
Crossref - Igbinosa EO, Beshiru A, Akporehe LU, Ogofure AG. Detection of methicillin-resistant staphylococci isolated from food producing animals: A public health implication. Vet Sci. 2016;3(3):14.
Crossref - Mousavi MNS, Mehramuz B, Sadeghi J, Alizadeh N, Oskouee MA, Kafil HS. The pathogenesis of Staphylococcus aureus in autoimmune diseases. Microb Pathog. 2017;111:503-507.
Crossref - Zgurskaya HI, Nikaido H. Multidrug resistance mechanisms: Drug efflux across two membrane. Mol Microbiol. 2000;37(2): 219-225.
Crossref - Anand C, Kumaresh SP. Influence of Titanium Carbide on the Three- Body Abrasive Wear Behaviour of Glass-Fabric Reinforced Epoxy Composites. Adv Materials. 2012;1(1):9-15.
Crossref - Liu C, Jiang Y, Wang X, et al. Diversity, antimicrobial activity, and biosynthetic potential of cultivable actinomycetes associated with lichen symbiosis. Microb Ecol. 2017;74(3):570-584.
Crossref - Thirumurugan D, Vijayakumar R. Characterization and structure elucidation of antibacterial compound of Streptomyces sp. ECR77 isolated from East coast of India. Curr Microbiol. 2015;70(5):745-755.
Crossref
© The Author(s) 2024. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.