ISSN: 0973-7510
E-ISSN: 2581-690X
Actinomycetes are Gram-positive filamentous bacteria well known for the production of bioactive compounds. Recently, many halophilic habitats have been explored for isolation of actinomycetes that exhibit biotechnological potentials. In this investigation, a saline habitat of Rajasthan, Sambhar Salt Lake (SSL) was selected to study the actinomycetes population and Carboxy Methyl Cellulase (CMCase) production by native isolates. A total of sixteen actinomycete isolates, halotolerant and moderately halophilic, were obtained using culture-dependent methods and characterized morphologically and biochemically. They were identified as members of Streptomyces, Nocardiopsis, Pseudonocardia, Saccharospolyspora, and Microbispora. Streptomyces was the most dominating genus, followed by Nocardiopsis. Agar plate assay was used for screening the isolates for CMCase production. Thirteen were found to produce the enzyme, apparent by hydrolysis observed on media plates. The highest relative activity of 22.04 was shown by isolate SSL 14 identified as Nocardiopsis sp. by 16S rDNA sequencing studies and thus selected for further optimization studies. Maximum enzyme (1.08 ± 0.09 U/ml) was produced using medium containing Carboxy Methyl Cellulose (Carbon source) and yeast extract (nitrogen source) at 12% NaCl and pH 9.0, incubated at 30 °C for 96 h. Maximum CMCase production at high salt concentration and pH suggests that Nocardiopsis SSL 14 can be used for industrial processes that operate under excessive saline and alkaline conditions.
CMCase, Halophiles, International Streptomyces Project (ISP), Nocardiopsis, Saltern
Actinomycetes are the most economically and biotechnologically valuable prokaryotes used to produce a wide variety of bioactive compounds.1 They also contribute to the cycling of nutrient elements by producing extracellular hydrolytic enzymes. One such hydrolytic endoglucanase enzyme is CMCase, which acts on the internal amorphous region and randomly hydrolyses long cellulose chains. It acts synergistically with exo-(1,4)-β-D-glucanase to hydrolyse cellulose.2 CMCases are used for enhancing fabric feel, look, and colour in textile processing.3 They decrease pulp viscosity and improve pulp beatability of the paper.4 Besides this, they are also used in food,5 beverages,6 detergent,7 biofuel industries,8 drug delivery,9 and wound healing.10 The demand for CMCase functional at high pH, temperature and salt concentrations has increased gradually. This emphasizes the need for the exploration of CMCase producing actinomycetes from extreme habitats. In order to suffice this demand, a saline habitat of Rajasthan (India), Sambhar Salt Lake, was selected which forms a unique ecosystem possessing a wide range of salinity in its water and salt formation ponds. It is India’s largest inland saline lake that produces salt for domestic and industrial needs. The present study was conducted to isolate actinomycetes and investigate CMCase production from halophilic and alkaline habitats of Sambhar Salt Lake, Rajasthan.
Study Area: Sambhar Salt Lake
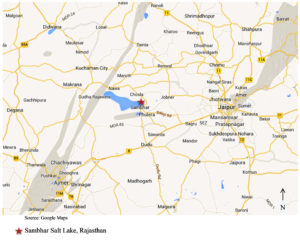
Sambhar Salt Lake is the India’s largest saline lake lying at a latitude and longitude of 26°55’12″N and 75°12’00″E, respectively. It is situated off National Highway No. 8, about 64 km northeast of Ajmer and 96 km southwest of Jaipur, extending up to Nagaur district in Rajasthan (Figure 1). The lake is surrounded by Aravali mountain ranges.
Sampling sites
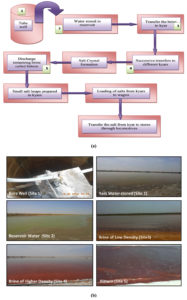
Eastern part of the lake is divided into solar salt pans (kyars) called salterns, where salt is produced for industrial and human consumption. Surface water samples were collected from five major stages (1 to 5) of salt production in various seasons, i.e., monsoon (A1-E1), winter (A2-E2), and summer (A3-E3) in the year 2012-13 (Figure 2a, b). The density of water in salterns was measured on-site by hydrometer. The pH and salinity of samples were measured using microprocessor water and soil analysis Kit Model 1160E.
Figure 2. Salt production process and sample collection sites (1 to 5) at Sambhar Salt Lake, Rajasthan (a) Flow chart of salt production, (b) Sampling Sites (1-5).
Isolation and Characterization of Actinomycetes
Actinomycetes were isolated on Actinomycetes Isolation Agar (AIA) medium containing antifungal agents nystatin and cycloheximide (50 µg/ml). For all first isolations, medium was prepared in sterile sample water and for subsequent inoculations medium was supplemented with NaCl (determined from salinity of samples). AIA plates were incubated at 28°C for 5-20 days for the appearance of actinomycete colonies and maintained on AIA at 4°C. Purified colonies were characterized by their morphological and physiological features as per the International Streptomyces Project (ISP).11 Micro morphological features were studied by the Coverslip culture method.12 The observed structures were compared with those described in Bergey’s manual13 to identify isolates up to the genus level. Various biochemical tests, including substrate degradation and utilization of different sugars as sole carbon source, were also studied.14,15
Screening for CMCase Production
All selected isolates were inoculated on carboxy methyl cellulose (CMC) agar medium and incubated at 28°C for 15 days, after which they were stained by flooding them with Congo red solution (1% w/v) for 15 min, followed by destaining with 1 M NaCl for 15 min.16
Selection of Potent Isolate for Optimization Studies
Relative enzyme activity of the isolates was measured from the screening assays using the formula D2– d2/d2, where d and D represent the widths of growth and hydrolysis, respectively.17 The isolate with maximum relative activity was selected for further optimization studies.
CMCase (Endoglucanase) Activity Assay
CMC containing broth (100 ml) was used for cellulase production. It was inoculated with 1% inoculum grown for 48 h at 30°C and 150 rpm. The culture was centrifuged at 10,000 rpm and 4°C for 10 min. The CMCase activity was determined in the crude supernatant in terms of reducing sugars released, which was quantified by dinitro salicylic acid (DNS) method.18 One unit (U) was defined as the amount of enzyme that released 1 mg of glucose per minute under experimental conditions.
Optimization of Culture Conditions for enzyme production
Culture conditions for CMCase production were optimized by following One Factor at a Time (OFAT) approach.19
Effect of Incubation Time
The production media was inoculated with selected actinomycete isolate (1%) and incubated at 30°C for different time intervals (24-144 h) at 150 rpm. CMCase production was estimated after every 24 h for 7 days by the method described by.18 Total biomass was measured and expressed as mg dry mass per 50 ml of culture medium.20
Effect of Incubation Temperature, pH, and salt concentration
The selected isolate was incubated at different temperatures (25-60°C); pH (5.0-10.0) and NaCl concentrations (1-13%) underdetermined optimum conditions.
Effect of Carbon and Nitrogen Sources
In order to maximize enzyme production effect of different carbon sources (1% w/v) and nitrogen sources (1% w/v) were studied by incorporating them into production media and incubating them under determined optimum conditions.
Identification of Selected Isolate by 16S rDNA Sequencing
Isolate showing maximum enzyme production was grown in nutrient broth medium and DNA was isolated in the form of a single discrete PCR amplicon band on agarose gel. 8F and 1492R primers were used for carrying out forward and reverse DNA sequencing reaction of PCR amplicon using BDT v3.1 Cycle sequencing kit on ABI 3730xl Genetic Analyzer. With the help of aligner software, a consensus sequence of 16S rDNA gene was generated, and the sequences which were similar to it were searched by BLAST. First ten sequences selected based on maximum identity score were then aligned using Clustal W. RDP database was used to generate distance matrix, the evolutionary history of the taxa was inferred by the Neighbour-Joining method,21 and MEGA 4 was used to construct phylogenetic trees.22
Statistical Analysis
The data for all experiments were expressed as mean ± standard deviation. The experiments for enzyme assays were set with three replicates, and three aliquots were taken from each replicate for all parameters, except for biomass determination. The data were analysed by one way analysis of variance (ANOVA) followed by LSD post hoc test. All the analysis was performed using the IBM SPSS software (Version 20).
The study was divided into three different seasons during which water samples were collected and analysed for their pH and salinity content. The results revealed that Sambhar Lake water has an alkaline pH ranging from 9.0-12.5 and high salinity from 30.959 -357.0 g/l (Table 1).
Table (1):
Physico-chemical analysis of water samples collected in different seasons.
| Rainy Season | |||||
|---|---|---|---|---|---|
| Water Samples | A1 | B1 | C1 | D1 | E1 |
| Salt Production Stage | Rain water stored | Reservoir | Brine | Brine | Brine |
| Density (°Be) | 1 | 13 | 20 | 26 | 17 |
| Salinity (g/l) | 30.959 ± 1.04 | 101.961± 1.906 | 186.640 ± 2.30 | 294.521 ± 2.355 | 205.563 ± 1.467 |
| pH | 12.0 ± 0.91 | 12.5 ± 0.264 | 11.2 ± 0.86 | 9.0 ± 0.14 | 11.0 ± 0.55 |
| Winter Season | |||||
| Water Samples | A2 | B2 | C2 | D2 | E2 |
| Salt Production Stage | Brine | Brine | Reservoir | Brine | Bittern |
| Density (°Be) | 25 | 23.5 | 6 | 15.5 | 28 |
| Salinity (g/l) | 293 ± 3.168 | 334 ± 2.0 | 82 ± 1.87 | 120 ± 1.896 | 354 ± 2.104 |
| pH | 10.2 ± 1.3 | 11.8 ±0.2 | 11.0 ±0.264 | 10.5 ± 0.40 | 10.8 ± 0.692 |
| Summer Season | |||||
| Water Samples | A3 | B3 | C3 | D3 | E3 |
| Salt Production Stage | Reservoir | Brine | Brine | Brine | Bittern |
| Density (°Be) | 6 | 7.6 | 25 | 27 | 29 |
| Salinity (g/l) | 82 ± 1.81 | 147 ± 1.63 | 299 ± 1.575 | 275 ± 2.68 | 357 ± 1.70 |
| pH | 11.0 ± 0.264 | 10.3 ± 0.346 | 10.0 ± 0.529 | 9.6 ± 0.346 | 10.9 ± 0.40 |
Isolation and Characterization of Actinomycetes Isolates
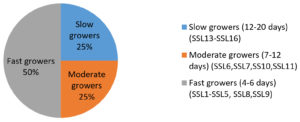
A total of sixteen isolates was recovered from Sambhar salterns and were designated by numbers 1 to 16 and prefixed with “SSL” as an abbreviation of Sambhar Salt Lake. They were classified based on incubation time required for complete growth on AIA medium as slow, moderate, and fast growers (Figure 3). Four isolates (SSL 6, SSL 7, SSL 10 and SSL 11) showed optimum growth at 8% NaCl and three isolates (SSL 14, SSL 15, SSL 16) at 12% NaCl hence classified as moderate halophiles. The remaining nine were classified as halotolerant as they grew well in the absence of salt and also tolerated salinity in the range of 5-8% NaCl. The colonies of thirteen isolates were tough, leathery, or powdery in appearance with the presence of aerial (AM) and substrate mycelium (SM), whereas three isolates were non-filamentous. All the isolates were Gram-positive. The growth was more rapid on ISP 2 and ISP 6 media, along with the production of diffusible pigment by many isolates. AM of many isolates was white coloured on different ISP media, and SM was of varied colours such as cream, yellow, orange, and brown (Table 2). The nature of AM and SM and their fragmentation in AIA medium helped in identifying the isolates up to the genus level (Table 3). Streptomyces was found to be the most dominant genus. Biochemical features of isolates are represented in Table 4.
Table (2):
Characteristics of Actinomycete Isolates on ISP Media.
| Isolate | ISP2 | ISP3 | ISP4 | ISP5 | ISP6 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| AM | SM | Diffusible pigment | AM | SM | AM | SM | AM | SM | AM | SM | Diffusible pigment | ||||||
| SSL1 | White | Bright orange | + | White | White | Light Cream | Light Cream | Creamish white | Creamish White | White | Dark orange | – | |||||
| SSL2 | Greyish white | Brown | + | White | Dark brown | White | Light brown | White | Light Cream | White | Orange | – | |||||
| SSL3 | White | Dark Yellow | – | White | Chocolate brown | White | Yellowish cream | White | Creamish White | White | Orange | – | |||||
| SSL4 | Cream | Cream | – | Greyish white | Greenish brown | White | Yellowish cream | White | Light Cream | White | Light orange | – | |||||
| SSL5 | Grey | Dark Yellow | – | Greyish white | Greenish brown | White | Light yellow | White | Light Cream | White | Orange | – | |||||
| SSL6 | White | Bright orange | + | White | Light Cream | Cream | Cream | No Growth | No Growth | White | Brownish orange | + | |||||
| SSL7 | White | Orange | + | cream | Light Yellow | Cream | Cream | no AM | Light Cream | White | Orange | + | |||||
| SSL8 | No AM & SM, orange-coloured colonies | – | No AM & SM, light cream growth | No Growth | No Growth | No AM & SM, Yellow colonies | – | ||||||||||
| SSL9 | No AM &SM, cream-coloured colonies | – | No growth | No Growth | No Growth | No AM &SM Cream colonies | – | ||||||||||
| SSL10 | White | Orange | + | Cream | Light Yellow | White | Light Yellow | No AM | White | White | Light orange | – | |||||
| SSL11 | White | Bright orange | + | Cream | Brown | Creamish white | Light brown | No Growth | No Growth | White | Orange | + | |||||
| SSL12 | No AM & SM, Yellowish cream colonies | – | Peach | No Growth | No Growth | No AM & SM, orange colonies | – | ||||||||||
| SSL13 | Light Grey | Light Brown | – | White | Light Cream | White | White | No Growth | No Growth | White | Orange | – | |||||
| SSL14 | White | Bright orange | + | White | Sandy Yellow | Light yellow | Light yellow | No AM | White | White | Dark orange | + | |||||
| SSL15 | White | Bright orange | + | White | Sandy Yellow | Yellow | Yellow | White | Light Yellow | White | Light orange | + | |||||
| SSL16 | White | Bright orange | + | Cream | Cream | Cream | Cream | White | Light Yellow | Brownish white | Light orange | – | |||||
(+) Diffusible pigment produced; (-) Diffusible pigment not produced
Table (3):
Genera of Actinomycetes of Sambhar Salt Lake Tentatively Identified on the Basis of Micro-morphological Features (Holt et al., 2000).
Isolate |
Characteristics of Mycelium |
Tentative Genera |
|---|---|---|
SSL 1 |
AM and SM present. AM fragments in to short filaments. Spore chain was of rectiflexibiles (RF) nature. |
Nocardiopsis |
SSL 2 |
SM extensively branched that does not fragment. The AM forms rectiflexibiles (RF) spore chains at maturity. |
Streptomyces |
SSL 3 |
Extensively branched non fragmenting SM and AM. AM carried spiral spore chain with coccoid shaped spores. |
Streptomyces |
SSL 4 |
Both mycelia were well-developed. AM carried long spore chain of Retinaculiaperti type and SM was stable. |
Streptomyces |
SSL 5 |
Extensively branched and stable SM and AM with long chains of spores (spirales type) on latter one. |
Streptomyces |
SSL 6 |
SM stable and branched. AM sparse, often produced in tufts in older parts of colony and produced only at optimum growth temperature. Spore chain was of straight rectus type. |
Saccharopolyspora |
SSL 7 |
Segmented AM and SM. Long chains of cylindrical shaped spores of unequal sizes are present on AM. Spore chain morphology was of rectiflexibiles type. |
Pseudonocardia |
SSL 8 |
AM and SM absent. Colonies were of cream colour. Rod shaped cells were present usually arranged in chain. |
Georgenia |
SSL 9 |
AM and SM absent. Cream colonies with brown tint present. Cocci shaped cells are usually arranged in singlets. |
Kokuria |
SSL 10 |
Extensively developed AM and SM absent. AM forms short chain of spores of rectiflexibiles type at maturity that fragments in rod shaped spores. |
Streptomyces |
SSL 11 |
Well-developed SM, hyphae are long and densely branched. Abundant AM that forms long chain of spores of rectiflexibiles type at maturity. |
Nocardiopsis |
SSL 12 |
AM and SM absent. Cream colonies with brown tint present. Cells are cocci shaped usually arranged in singlets. |
Kokuria |
SSL 13 |
Compact SM that grows into and on top of agar and does not bear spores. AM was stable and produces spores formed in longitudinal pairs present on short sporophores. The AM forms bud at the side and later the bud or occasionally the tip of the side branch swells up producing two conidia separated by cross walls. |
Microbispora |
SSL 14 |
Well-developed SM and AM, hyphae are long and densely branched. AM at maturity fragments to form long chain of spores. Spore chain was of rectiflexibiles type. |
Nocardiopsis |
SSL 15 |
Branched and stable SM. Aerial hyphae carried spore chain of rectiflexibiles type. Rod shaped spores were present in long chains. |
Streptomyces |
SSL 16 |
SM well developed and branched. AM sparse and had rectiflexibiles type of spore chain segmented into bead like chain of spores. |
Saccharopolyspora |
Table (4):
Biochemical Characterization of Actinomycete Isolates.
| Isolates | Catalase Test | Oxidase Test | Gelatin liquefaction Test | Casein Hydrolysis Test | Xanthine Hydrolysis Test | Hypoxanthine Hydrolysis Test | Tyrosine Hydrolysis Test | Carbohydrate Utilization Test | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Glucose | Maltose | Xylose | Lactose | Fructose | Sucrose | Mannitol | ||||||||
| SSL 1 | + | + | – | + | – | – | – | + | + | + | + | + | + | + |
| SSL 2 | + | + | – | ++ | + | ++ | – | + | + | + | + | + | + | + |
| SSL 3 | + | + | – | +++ | ++ | – | – | + | + | + | + | + | + | + |
| SSL 4 | + | + | + | +++ | – | +++ | – | + | + | + | + | + | + | + |
| SSL 5 | + | + | + | ++ | – | – | – | + | + | + | + | + | + | + |
| SSL 6 | + | + | – | + | ++ | + | ++ | + | + | +/- | + | + | + | + |
| SSL 7 | + | + | – | + | +++ | – | – | + | + | +/- | +/- | + | + | + |
| SSL 8 | + | + | – | – | + | – | – | + | +/- | +/- | + | – | – | + |
| SSL 9 | + | + | – | – | + | – | – | – | – | – | – | +/- | +/- | – |
| SSL 10 | + | + | – | + | + | – | ++ | + | + | +/- | + | + | + | + |
| SSL 11 | + | + | + | + | ++ | – | +++ | + | + | + | + | + | + | + |
| SSL 12 | + | + | – | – | – | +++ | – | – | +/- | – | – | +/- | +/- | – |
| SSL 13 | + | + | – | + | – | ++ | – | + | – | – | – | – | – | – |
| SSL 14 | + | + | – | + | +++ | +++ | + | + | + | +/- | + | + | + | + |
| SSL 15 | + | + | – | + | + | +++ | +++ | + | + | +/- | + | + | + | + |
| SSL 16 | + | + | – | + | – | – | – | + | + | +/- | + | + | + | + |
(+) Positive test; (-) Negative test; (+/-) Indecisive utilization of carbohydrates; (+ + +) Strong Hydrolytic Activity; (+ +) Moderate Hydrolytic Activity; (+) Weak Hydrolytic Activity
Screening for CMCase Production and Relative Enzyme Activity Determinations
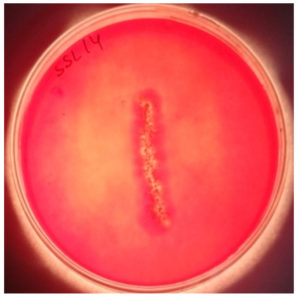
All filamentous isolates formed a clear zone of hydrolysis when CMC agar plates were flooded with Congo red and NaCl. Three non-filamentous actinomycetes isolates SSL 8, SSL 9 and SSL 12 did not show the production of CMCase. Relative enzyme activity was calculated for all positive isolates (Table 5), and highest was shown by SSL 14 (Figure 4), and thus, it was selected for enzyme optimization studies.
Table (5):
Relative Enzyme Activity of Actinomycete Isolates.
S.No. |
Isolates |
Relative CMCase activity |
|---|---|---|
1. |
SSL 1 |
16.488 |
2. |
SSL 2 |
19.25 |
3. |
SSL 3 |
11.96 |
4. |
SSL 4 |
6.249 |
5. |
SSL 5 |
12.69 |
6. |
SSL 6 |
6.41 |
7. |
SSL 7 |
4.861 |
8. |
SSL 10 |
7.028 |
9. |
SSL 11 |
20.496 |
10. |
SSL 13 |
10.56 |
11. |
SSL 14 |
22.04 |
12. |
SSL 15 |
14.391 |
13. |
SSL 16 |
17.063 |
Optimization of Culture Conditions
Effect of Incubation Time
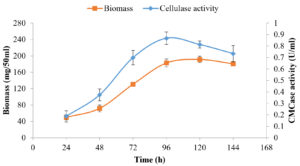
The majority of the actinomycetes are slow-growing. CMCase production by SSL14 was found to be maximum at 96 h of growth (0.868 ± 0.054 U/ml), and thus, the other parameters were optimized at the same incubation time. A significant difference in CMCase activity was observed between 96 and 120 h (p < 0.05). Growth analysis showed that enzyme was produced during log and stationary phase (Figure 5).
Figure 5. Fermentation Time Course for Cell Growth and Extracellular CMCase Produced by Isolate SSL 14
Effect of incubation temperature, pH, and salt concentration
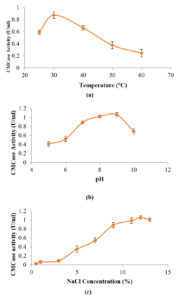
Maximum activity (0.877 ± 0.055 U/ml) was observed at 30°C (Figure 6a), which is also the optimum growth temperature for actinomycetes.23 As the temperature was increased from 40°C to 60°C, enzyme activity significantly declined. Among, the different pH tested, highest CMCase activity occurred at pH 9.0 (1.063 ± 0.045 U/ml) (Figure 6b) however no significant difference between pH 8.0 and 9.0 (p > 0.05) was observed. This observation revealed the alkaliphilic nature of CMCase produced.
Results presented in Figure 6c showed that CMCase production was least at 0.5% NaCl, and it increased gradually with the increasing salt concentration, with maximum occurring at 12% NaCl (1.055 ± 0.046 U/ml). These results suggested the moderately halophilic nature of the isolate.
Figure 6. Optimization of (a) Temperature, (b) pH, (c) NaCl for CMCase Production by Isolate SSL 14.
Effect of Carbon and Nitrogen Sources
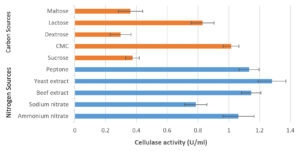
From the different carbon sources tested, the CMCase production was maximum (1.015 ± 0.051 U/ml) in the medium supplemented with CMC as a carbon source and minimum in the presence of dextrose (Figure 7). It was found that yeast extract gave maximum activity (1.08 ± 0.09 U/ml). Also, the enzyme activity was comparable when the medium was supplemented with beef extract, peptone, and ammonium nitrate (Figure 7).
Identification of isolate SSL 14 by 16S rDNA sequencing
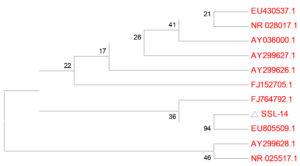
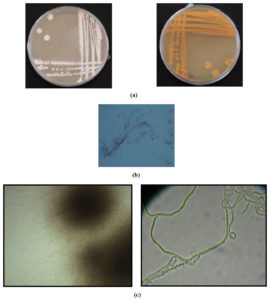
The phylogenetic analyses based on BLAST search using 16S rDNA gene sequence of the isolate SSL 14 revealed 100% similarity to Nocardiopsis sp. BCEAA 7603 (Figure 8), and thus, it was designated as Nocardiopsis sp. SSL 14 (GenBank Accession number KU860459). These results were also supported by its morphological features (Figure 9a-c) as described above.
Figure 8. Neighbor-joining tree showing the position of isolate Nocardiopsis SSL14 to first ten similar sequences. Values are the branch lengths reflecting the actual distances between the sequences. EU805509.1: Nocardiopsis sp. BCEAA 7603); FJ764792.1: Nocardiopsis sp. E-143; EU430537.1: Nocardiopsis exhalans strain VTT E-063001; FJ152705.1: Uncultured bacterium clone TX1A_153; AY299626.1: Streptomyces sp. YIM 80125; AY299627.1: Nocardiopsis sp. YIM 80129; AY036000.1: Nocardiopsis exhalans; NR_028017.1: Nocardiopsis exhalans strain ES10.1; AY299628.1: Nocardiopsis sp. YIM 80130; NR_025517.1: Nocardiopsis metallicus strain R2A
Salt lakes and multi-pond solar salterns are well-known hypersaline habitats having high salt concentration and alkaline pH.24 They possess enormous potential in terms of microbial diversity that can be exploited for many industrial applications. One such hypersaline habitat, Sambhar Salt Lake, India, was studied for its actinomycetes population and their bioactive potential. The analysis of pH and salinity of lake water in different seasons revealed its alkaline and saline nature. Many previous studies also reported the same nature of Sambhar Salt Lake.25,26 Similar values for alkalinity and salinity were also reported for other athalassohaline habitats like El- Djerid Salt Lake in southern Tunisia.27 Actinomycetes from such halophilic habitats are composed of heterogeneous physiological groups of different genera. In this study, a total of 16 actinomycetes isolates (SSL 1-SSL 16), including halotolerant and moderate halophiles, were recovered from solar salt pans of Sambhar Salt Lake, India. Other hypersaline environments studied for their presence were crystallizer ponds of solar salterns from Tuticorin in Bay of Bengal,28 and the saline desert of Kutch, India.29 Halotolerant and halophilic actinomycetes have been reported from other Salt Lake habitats.30,31 To prevent osmotic lysis of cells in the presence of salt, most of the halophiles and halotolerant microorganisms accumulate compatible solutes in high concentrations within the cytoplasm as a part of their adaptive strategy.32
Halophilic and halotolerant species have been reported for the genera Streptomyces, Pseudonocardia, Streptosporangium, Nocardiopsis, Salinispora, and Streptomonspora.33 The presence of Streptomyces as a predominant genus in halophilic environments of both marine and non-marine origin has been reported in many studies,28,34,35 and a similar pattern was noticed in the present study also. Nocardiopsis was the second most abundant actinomycete isolated in the present study, and its presence was also observed in Kovalam, Puthalum, and Thamaraikulam salt works, India.36 Member of a rare genus Pseudonocardia was also recovered in the present study. Its presence has also been noticed in Bay of Bengal’s hypersaline environments, India,37 and in the South China Sea.38 Rajagopal and Kannan39 recently reported isolation of Actinoalloteichus sp. from sediments of the Havelock Island of Andaman, India.
Sufficient studies were not available on actinomycetes population of Sambhar Lake water till 2012 but in 2013, Yadav et al.40 published reports on actinomycetes diversity from Sambhar Lake water which showed the presence of the same actinomycetes genera.
Actinomycetes are studied most extensively for antibiotic production, but less focus is on enzyme production, especially those isolated from halophilic habitats.41 In this study, 81% of the isolates were found to be positive for CMCase production. The Enzyme produced by isolate SSL 14 identified as Nocardiopsis sp. was optimized following the OFAT approach. Similar enzyme optimization studies using the OFAT approach were performed for cellulase production by Streptomyces sp. B-PNG23 and by Fomitopsis sp. RCK 2010.42,43
It has been reported that enzyme production by actinomycetes starts in the early log phase, increases in the late log phase, and continues till the onset of the stationary phase, after which it declines.44 The results of the present study were found to be in accordance with this theory. CMCase production by Nocardiopsis SSL 14 was highest at 96 h of incubation. A novel actinomycete, Nocardiopsis sp. KNU was found to produce maximum cellulase in the late log phase that continued up to the stationary phase,45 whereas non-growth associated CMCase production was observed in Streptomyces drozdowiczii.46 In another study on Streptomyces sp., maximum enzyme production was observed after five days of incubation.47 It has been reported in the literature that end product act as an inhibitor to hydrolysis of cellulose,48,49 and therefore is responsible for the decline in activity after a few days. The depletion of nutrients in the medium can also adversely affect enzyme production after a few days of incubation.50
The effect of temperature on the enzyme productions was related to growth temperature, maximally produced at 30°C. Enzymes are not stable at very high or very low pH values as they get, denatured and thus, these kinds of pH values adversely affect enzyme production by microorganisms.51 This fact is supported by the finding of the present study, where the production of CMCase was very low at lower pH values. However, maximum enzyme production at alkaline pH was observed and can be attributed to the origin of isolate and thereby its adaptation to saline habitat of Sambhar Lake. Different species of Streptomyces were reported to show maximum enzyme production at a broad range of pH from 5.5 to 8.0.52-55
Nocardiopsis SSL 14 showed the highest activity when the medium was supplemented with 12% NaCl. This adaptation of proteins in a media with high salt concentrations is attributed to the presence of a large number of acidic residues on protein surface supported by electrostatic interactions and an increased number of salt bridges.56
Synthesis of cellulase is subjected to induction and catabolite repression in both bacteria and fungi. Cellulose and its derivatives act as inducers, and soluble sugars like glucose, maltose, etc., act as repressors.57 Consistent with these theories, in the present study also, maximum enzyme production occurred in the presence of CMC and decreased significantly in the presence of glucose and maltose. Thus, CMC was easily metabolized by SSL 14 and also acted as an inducer for CMCase production. In a study on Streptomyces sp., it was found that CMC (1%) was better than glucose and other carbon sources for endoglucanase production.58
Yeast extract was found to be the best source for CMCase production by SSL 14, similar to that reported for Streptomyces sp. F2621,59 and Streptomyces drozdowiczii.46 Inorganic sources such as ammonium nitrate and peptone did not show a significant difference in enzyme activity. A similar finding has also been reported in Streptomyces sp. T3-1 by Jang and Chen.58 Wang, in 1982, reported that ammonium ions could be absorbed to the mycelium resulting in a lowering of the pH, thereby inhibiting cellulase production.60 This could be responsible for lowering of CMCase production in the presence of ammonium salt.
In a nutshell, the findings of the present investigation have shown that CMCase was produced by Nocardiopsis SSL 14 under high salt concentrations (12% NaCl) and alkaline pH 9.0. This property is advantageous in the bioremediation of cellulosic materials and the production of bio-energy. The enzyme can be used with ionic liquids (ILs) in the pre-treatment of lignocellulosic biomass to produce renewable products in an eco-friendly way.61-62 The enzyme can be further purified and characterized for its ability to function at high pH and salt concentrations and can be tested for its use in commercial detergent formulations. The production of protease by actinomycete isolates of Sambhar Lake and their application as a detergent additive has been studied.63 Thus, it is also suggested that these actinomycetes might also produce other enzymes and products that could prove beneficial for different biotechnological applications.
ACKNOWLEDGMENTS
None.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
CS conceptualized the study, acquired the funding, investigated and performed the experiments. PC performed the data curation and formal analysis. PC and PM visualized the study. CS and PC wrote the original draft. PB and NM performed the supervision. PB, NM and PM reviewed and edited the final manuscript for publication.
FUNDING
This work was supported by University Grants Commission (UGC), India with grant number MS-122/304071/07-08/CRO.
DATA AVAILABILITY
16S rDNA sequence of Nocardiopsis sp. SSL 14 is available in GenBank with Accession number KU860459. All other datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
- Lam LS. Discovery of novel metabolites from marine actinomycetes. Curr Opin Microbiol. 2006;9(3):245-251.
Crossref - Coughlan MP. Mechanisms of cellulose degradation by fungi and bacteria. Animal Feed Science and Technology. 1991;32(1-3):77-100.
Crossref - Bhat MK. Cellulases and related enzymes in biotechnology. Biotechnol Adv. 2000;18(5):355-383.
Crossref - Oksanen T, Pere J, Buchert J, Viikari L. The effect of T. reesei cellulases and hemicellulases on the paper technical properties of never-dried bleached kraft pulp. Cellulose. 1997;4:329-339.
Crossref - Othman NEA, Ismail F, Aziz AA, Wahab NA. Preparation and characterization of palm-based sodium carboxymethyl cellulose for application in food additive. Biointerface Res Appl Chem. 2021;11(5):13053-13063.
Crossref - Galante YM, De Conti A, Monteverdi R. Application of Trichoderma enzymes in textile industry. In: Harman GF, Kubicek CP (Eds.), Trichoderma & Gliocladium-Enzymes, biological control and commercial applications. Taylor & Francis, London, 1998:311-326.
- Abe TO, Lajide L, Owolabi BJ, Adebayo, AO, Ogunjobi JK, Oluwasina OO. Synthesis and application of carboxymethyl cellulose from Gliricidia sepium and Cola gigantea. BioRes. 2018;13(3):6077-6097.
- Kuhad RC, Gupta R, Singh, A. Microbial cellulases and their industrial applications. Enzyme Res. 2011;2011:280696.
Crossref - Inphonlek S, Sunintaboon P, Leonard M, Durand A. Chitosan/carboxymethylcellulose-stabilized poly(lactide-co-glycolide) particles as bio-based drug delivery carriers. Carbohydr Polym. 2020;242:116417.
Crossref - Koneru A, Dharmalingam K, Anandalaksh, R. Cellulose based nanocomposite hydrogel films consisting of sodium carboxymethylcellulose-grapefruit seed extract nanoparticles for potential wound healing applications. Int J Biol Macromol. 2020;148:833-842.
Crossref - Shirling EB, Gottlieb D. Methods for characterization of Streptomyces Species. Int J Syst Bacteriol. 1966;16(3):313-340.
Crossref - Williams ST, Cross T. Actinomycetes. In: Booth C, Methods in Microbiology, Academic Press. 1971;4:295-334.
Crossref - Holt JG, Krieg NR, Sneath PHA, Staley JT, Williams ST. Bergey’s Manual of Determinative Bacteriology, Lippincott Williams and Wilkins, USA, 2000:605-704.
- Gordon RE, Barnett, DA, Handerhan JE, Pang CHN. Nocardia coeliaca, Nocardia autotrophica, and the nocardin strain. Int J Syst Bacteriol. 1974;24(1):54-63.
Crossref - Pridham TG, Gottlieb D. The utilization of carbon compounds by some Actinomycetales as an aid for species determination. J Bacteriol. 1948;56(1):107-14.
Crossref - Carder JH. Detection and quantitation of cellulase by Congo red staining of substrate in a cup-plate diffusion assay. Anal Biochem. 1986;153(1):75-79.
Crossref - Bradner JR, Gillings M, Nevalainen KMH. Qualitative assessment of hydrolytic activities in Antarctic microfungi grown at different temperatures on solid media. World J Microbiol Biotechnol. 1999;15(1):131-132.
Crossref - Miller L. Use of dinitro salicylic acid reagent for determination of reducing sugar. Anal Chem. 1959;31(3):426-428.
Crossref - Hegde SV, Ramesha A, Srinvas C. Optimization of amylase production from an endophytic fungi Discosia sp. isolated from Calophyllum inophyllum. J Agric Technol. 2011;7(3):805-813.
- Dastager GS, Agasar D, Pandey A. Production and partial purification of a-amylase from a novel isolate Streptomyces gulbargensis. J Ind Microbiol Biotechnol. 2009;36(2):189-194.
Crossref - Saitou N, Nei M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987;4(4):406-425.
Crossref - Tamura K, Dudley J, Nei M, Kumar S. MEGA4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol. 2007;24(8):1596-1599.
Crossref - Goodfellow M, Williams ST. Ecology of actinomycetes. Annu Rev Microbiol. 1983;37:189-216.
Crossref - Zafrilla B, Martinez-Espinosa RM, Alonso MA, Bonete MJ. Biodiversity of Archaea and floral of two inland saltern ecosystems in the Alto Vinalopo Valley, Spain. Saline Systems. 2010;6:10.
Crossref - Jose PA, Jebakumar SRD. Phylogenetic appraisal of antagonistic, slow growing actinomycetes isolated from hypersaline inland solar salterns at Sambhar Salt Lake, India. Front Microbiol. 2013;4:190.
Crossref - Sinha R, Raymahashay BC. Evaporite mineralogy and geochemical evolution of the Sambhar Salt Lake, Rajasthan, India. Sediment Geol. 2004;166(1-2):59-71.
Crossref - Hedi A, Sadfi N, Fardeau ML, et al. Studies on the biodiversity of halophilic microorganisms isolated from El-Djerid Salt Lake (Tunisia) under aerobic conditions. Int J Microbiol. 2009;2009:731786.
Crossref - Jose PA, Jebakumar SRD. Phylogenetic diversity of actinomycetes cultured from coastal multi pond solar saltern in Tuticorin, India. Aquat Biosyst. 2012;8(1):23.
Crossref - Thumar JT, Dhulia, K, Singh, SP. Isolation and partial purification of an antimicrobial agent from halotolerant alkaliphilic Streptomyces aburaviensis strain Kut-8. World J Microbiol Biotechnol. 2010;26(11):2081-2087.
Crossref - Oren A. Diversity of halophilic microorganisms: environments, phylogeny, physiology and applications. J Ind Microbiol Biotechnol. 2002;28(1):56-63.
Crossref - Cai M, Tang SK, Chen YG, Li Y, Zhang YQ, Li WJ. Streptomonospora amylolytica sp. nov. and Streptomonospora flavalba sp. nov., two novel halophilic actinomycetes isolated from a salt lake. Int J Syst Evol Microbiol. 2009;59(10):2471-2475.
Crossref - Galinski EA. Compatible solutes of halophilic eubacteria: molecular principles, water-solute interactions, stress protection. Experientia. 1993;49:487-496.
Crossref - Hamedi J, Mohammadipanah F, Ventosa A. Systematic and biotechnological aspects of halophilic and halotolerant actinomycetes. Extremophiles. 2013;17(1):1-13.
Crossref - Jensen PR, Dwight R, Fenical W. Distribution of actinomycetes in near-shore tropical marine sediments. Appl Env Microbiol. 1991;57(4):1102-1108.
Crossref - Dharmaraj S. Marine Streptomyces as a novel source of bioactive substances. World J Microbiol Biotechnol. 2010;26(12):2123-2139.
Crossref - Jenifer JSA, Selva Donio MTB, Viji VT, Velmuruga S. Halo-alkaliphilic actinomycetes from solar salt works in India: Diversity and antimicrobial activity. Blue Biotechnology Journal. 2013;2(1):137-151.
- Peela S, Bapiraju K, Terli R. Studies on antagonistic marine actinomycetes from the Bay of Bengal. World J Microbiol Biotechnol. 2005;21:583-585.
Crossref - Tian XP, Long LJ, Li SM, Zhang J. Pseudonocardia antitumoralis sp. nov.,a new deoxynyboquinone- producing actinomycete isolated from a deep-sea sedimental sample in South China Sea. Int J Syst Evol Microbiol. 2013;63(3):893-899.
Crossref - Rajagopal G, Kannan S. Systematic characterization of potential cellulolytic marine actinobacteria Actinoalloteichus sp. MHA15. Biotechnol Rep (Amst). 2017;13:30-36.
Crossref - Yadav AK, Vardhan S, Kashyap S, Yandigeri M, Arora DK. Actinomycetes diversity among rRNA gene clones and cellular isolates from Sambhar Salt Lake, India. Scientific World Journal. 2013;2013:781301.
Crossref - Chakraborty S, Khopade A, Kokare C, Mahadik K, Chopade B. Isolation and characterization of novel a amylase from marine Streptomyces sp. D1. J Mol Catal B: Enzymatic. 2009;58:17-23.
Crossref - Azzeddine B, Abdelaziz M, Estelle C, Mouloud K. Optimization and partial characterization of endoglucanase produced by Streptomyces sp. B-PNG23. Arch Biol Sci Belgrade. 2013;65(2):549-558.
Crossref - Deswal D, Khasa YP, Kuhad RC. Optimization of cellulase production by a brown rot fungus Fomitopsis sp. RCK2010 under solid-state fermentation. Bioresour Technol. 2011;102(10):6065-6072.
Crossref - Chakraborty S, Khopade A, Biaob R, et al. Characterization and stability studies on surfactant, detergent and oxidant stable a -amylase from marine haloalkaliphilic Saccharopolyspora sp. A9. J Mol Catal B: Enzymatic. 2011;68(1):52-58.
Crossref - Saratale GD, Oh SE. Production of thermotolerant and alkalotolerant cellulolytic enzymes by isolated Nocardiopsis sp. KNU. Biodegradation. 2011;22(5):905-919.
Crossref - Lima ALG, Nascimento RP, Bon EPS, Coelho RRR. Streptomyces drozdowiczii cellulase production using agro-industrial by-products and its potential use in the detergent and textile industries. Enzyme Microb Tech. 2005;37(2):272-277.
Crossref - Alani F, Anderson WA, Moo-Young MN. New isolate of Streptomyces sp. with novel thermoalkalo tolerant cellulases. Biotechnol Lett. 2008;30(1):123-126.
Crossref - Van Dycke BH. Enzymatic hydrolysis of cellulose – a kinetic study. Ph.D. Thesis. Massachusetts Institute of Technology. Cambridge, U.S. 1972.
- Howell JA, Mangat M. Enzyme deactivation during cellulose hydrolysis. Biotechnol Bioeng. 1978;20(6):847-863.
Crossref - Nochure SV, Roberts MF, Demain AI. True cellulases production by Clostridium thermocellum grown on different carbon sources. Biotechnol Lett. 1993;15:641-646.
Crossref - Haltrich D, Nidetzky B, Kulbe KD, Steiner W, Zupancic S. Production of fungal xylanases. Bioresour Technol. 1996;58(2):137-161.
Crossref - Theberge M, Lacaze P, Shareck F, Morosoli R, Kluepfel D. Purification and characterization of an endoglucanase from Streptomyces lavidans 66 and DNA sequence of the gene. Appl Microbiol. 1992;58(3):815-820.
Crossref - Hmad IB, Gargouri A. Neutral and alkaline cellulases: Production, engineering and applications. J Basic Microbiol. 2017;57(8):653-658.
Crossref - Solingen VP, Meijer D, Kleij WA, et al. Cloning and expression of an endocellulase gene from a novel Streptomycete isolated from an East African soda lake. Extremophiles. 2001;5(5):333-341.
Crossref - Jaradat Z, Dawagreh A, Ababneh Q, Saadoun I. Influence of culture conditions on cellulase production by Streptomyces sp. (Strain J2). Jordan J Biol Sci. 2008;1:141-146.
- Danson MJ, Hough DW. The structural basis of protein halophilicity. Comp Biochem Physiol. 1997;117(3):307-312.
Crossref - Stoppok W, Rapp P, Wagner F. Formation, location, and regulation of endo-1,4-3-glucanases and 3-glucosidases from Celluilomnonas uda. Appl Environ Microbiol. 1982;44(1):44-53.
Crossref - Jang HD, Chen KS. Production and characterization of thermostable cellulases from Streptomyces transformant T3-1. World J Microbiol Biotechnol. 2003;19:263-268.
Crossref - Smruti P, Rao KK, Krishna M. The production of betaglucosidase by Aspergillus terreus. Curr Microbiol. 1995;30(5):255-258.
Crossref - Wang CW. Cellulolytic enzymes of Volvariella volvacea. In: Tropical Mushrooms Biological Nature and Cultivation Methods. Chang ST, Quimio TH (Eds.), The Chinese University Press, Hong Kong, 1982:167-186.
- Gunny AAN, Arbain D. Ionic liquids: green solvent for pre-treatment of lignocellulosic biomass. Adv Mater Res. 2013;701:399-402.
Crossref - Gunny AAN, Arbain D, Gumba RE, Jong BC, Jamal P. Potential halophilic cellulases for in situ enzymatic saccharification of ionic liquids pre-treated lignocelluloses. Bioresour Technol. 2014;155:177-181.
Crossref - Sharma C, Mehtani P. Application of protease produced from an actinomycete strain as a detergent additive. 2012;1:19-21.
© The Author(s) 2022. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.