ISSN: 0973-7510
E-ISSN: 2581-690X
The present work highlights a low-cost, ecofriendly and simple procedure for the synthesis of zinc oxide nanoparticles (ZnONPs), mediated by extracellular secondary metabolite pyoverdine – a siderophore produced by Pseudomonas aeruginosa. The in vitro bio-reduction of zinc nitrate into ZnONPs in the presence of siderophore pyoverdine was ascertained by UV–visible spectroscopy, X-ray diffraction (XRD), Fourier transform infrared spectroscopy and transmission electron microscopy (TEM) analysis. The synthesized ZnONPs were distinguished by a characteristic absorption peak at 416 nm in the UV–visible range. The TEM pictures of ZnONPs showed pseudospherical morphology, with particle size ranging between 50 and 100 nm. X-ray diffraction peaks obtained at 31.73°, 34.37°, 36.24°, 47.40°, 56.68°, 63.0° and 67.8° corresponding to the lattice plane of [100], [002], [101], [102], [110], [103] and [112] suggested the polycrystalline wurtzite structure of the nanoparticle. The ZnONPs were found to have an antibacterial effect against Gram-positive bacteria (Staphylococcus aureus and Bacillus sp.) and Gram-negative bacteria (Escherichia coli), and also showed a fungicidal effect against phytopathogenic fungi such as Rhizoctonia solani, Fusarium sp. and Penicillium sp. The present approach of using exogenously produced bacterial siderophore offers a unique in vitro green method of synthesizing ZnONPs, where a direct contact between the toxic metal and biological resource material can be avoided.
Pseudomonas aeruginosa, siderophore, XRD, FTIR, TEM, zinc oxide nanoparticles.
Pollution of water and soil by heavy metals is a matter of serious concern for all of us, as the persistent nature of heavy meals enables them to enter the food chain and cause long-term adverse effects on all forms of life1. The widespread application of metal containing pesticides, fertilizers and industrial sludge in agricultural fields, and the subsequent metal accumulation in plants, soil and the aquatic ecosystem, has posed serious environmental problems2. Though a few trace metals are essential for living organisms and play important physiological roles, their presence in higher concentrations can cause damage to cell membranes, DNA structure and enzyme activity, and disrupt cellular physiology3,4. Such large-scale application of metal-supplemented fertilizers and pesticides in agricultural fields over a longer period of time leads to excess metal loading of agricultural soil5 and disruption of the delicate ecological balance. Earlier reports have demonstrated that the application of Zn-containing sludge (90–250 mg kg”1 soil) in soil over a prolonged period completely diminished the bacterial population in the soil and severely reduced the crop yield. In such conditions, the major plant-growth-promoting activities of bacteria in the soil are greatly influenced by the presence of toxic metals such as lead, cadmium and zinc2. As a result, the overall agricultural yield is heavily dependent on the use of chemical fertilizers and pesticides.
Exploration of the environmental fate of accumulated toxic metals in the soil and the overall impact on the rhizospheric microbial population is urgently needed6. It has been suggested that excess metal loading of agricultural soil over a period of time not only leads to changes in the genetic makeup of the useful microbial communities7,8, but also has a negative impact on the plant-growth-promoting attributes of microorganisms, such as production of phytohormones, siderophore, hydrogen cyanide and indole acetic acid in the rhizosphere zone of plants9,10. Although, a high degree of adaptability in prokaryotic microorganisms enables them to withstand heavy metal stress, which ultimately leads to the development of heavy metal tolerance in microorganisms11. In this context, many microorganisms acquire the potential to transform the toxic metals into a less toxic form under both aerobic and anaerobic conditions12. The metabolic response of microorganisms against heavy metal stress, particularly in terms of changes in the production of metabolites and their role in the development of a metal tolerance strategy in microorganisms are highly interesting13.
Earlier reports have shown that bacterial strains with the ability to reduce nitrate are involved in the reduction of toxic silver ions into silver nanoparticles14,15. Biogenic synthesis of nanoparticles is a promising green technology that can be exploited for pharmaceuticals, cosmetics, agriculture and the environment, particularly in immobilization of potentially hazardous elements in soils16. It also offers an advantage over other traditional methods due to the small particle size, larger specific surface area and easy distribution in the soil. The nanomaterials are able to immobilize the labile fractions of potentially hazardous elements in the soil solution, resulting in reduced mobility and availability of toxic metals to plants in the metal-contaminated soils17. The antimicrobial activity of metal nanoparticles has received marked global attention as they specifically exhibit significant toxicity to microbes18, 19. The importance of the antimicrobial properties of metal nanoparticles is gaining ground due to the effective inhibitory action against a wide range of microbes20. Despite the broad scope of metal nanoparticles as antimicrobial agents, efforts are required to make this approach more cost effective and ecofriendly.
The siderophores present in the secondary metabolites of bacteria are known to play bioactive roles, including the chelation of metals21. Assuming that the chelation and transformation of Zn ions into Zn nanoparticles by bacterial siderophores might be contributing to the cellular defense strategy of the bacteria against Zn metal stress22, the present investigation was carried out to study the production of secondary metabolites by the rhizobacterium Pseudomonas aeruginosa in response to Zn metal stress. Subsequently, the role of secondary metabolite pyoverdine – a siderophore – was used to study the transformation of zinc ions into zinc oxide nanoparticles (ZnONPs) at room temperature in the absence of any reducing agent. Furthermore, the freshly synthesized ZnONPs were tested for their antibacterial and antifungal attributes against selected bacterial and phytopathogenic fungal strains.
Isolation and characterization of bacteria
Soil samples were collected from the rhizosphere of mustard (Brassica campestris) plants grown in soils irrigated by metal-contaminated wastewater near Lucknow city, India, located at 26° 55′ N and 80° 59′ E, and elevation of approximately 123 meters above sea level. A total of 10 bacterial isolates were obtained from the rhizosphere using standard microbiological techniques. The bacterial strains were first screened on the basis of their relative growth on nutrient agar plates supplemented with 1000 mg L-1 of zinc as zinc nitrate, Zn(NO3)2.6H2O. The metal-modified plates were inoculated and incubated at 28±2°C for 48 h. The highest concentration of zinc permitting bacterial growth on plates was considered as the maximum tolerance limit. Among the 10 bacterial isolates, only the strain DM1 showed maximum tolerance up to a 1000 mg L-1 concentration of zinc. Thus, the DM1 strain was selected for further studies.
The Zn (II) tolerant bacterial isolate DM1 was characterized with biochemical and molecular methods. For presumptive identification, bacterial isolate DM1 was characterized for physiological and biochemical characters according to Bergey’s Manual of Systematic Bacteriology23. Molecular characterization of the strain was carried out by using 16S rRNA sequences analysis (1.5 kb) carried out by Amnion Biosciences Pvt. Ltd. (Bengaluru, India). The sequence data were blasted using the National Center for Biotechnology Information online BLAST program to identify the selected bacterial isolate. The isolate was routinely maintained on nutrient agar slants at 4°C for further use.
Evaluation of the zinc-tolerance level
A log phase culture (OD610 = 0.1) of selected strain DM1 was seeded in nutrient broth modified with different concentrations (0–1200 mg L-1) of Zn(NO3)2.6H2O (zinc nitrate) and the optical density was measured at 610 nm by using a UV–visible spectrophotometer (1601 Shimadzu, Japan) up to stationary phase. The experiment was conducted in triplicate. The highest concentration of zinc permitting bacterial growth was considered as the maximum tolerance limit.
Growth conditions and extraction of siderophores
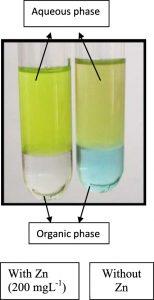
Extraction of siderophores from the secondary metabolites of cell-free supernatant was carried out by following the method of Meyer and Abdallah24. Briefly, the bacterial isolate P. aeruginosa DM1 was inoculated in sterile nutrient broth medium (pH 7.2). Bacteria were allowed to grow for 24 h at 35±2°C in a 500 mL Erlenmeyer flask with a working volume of 300 mL, under shaking conditions at 120 rpm on an orbital shaking incubator. Culture medium was then centrifuged at 5000 rpm to obtain cell-free supernatant25. The resultant supernatant was filter-sterilized. Furthermore, 5 mL of chloroform was mixed with 8 mL of supernatant and vortexed for 10 × 2 s. The chloroform layer with blue coloration was separated by funnel and a yellowish-green aqueous layer containing pyoverdine was scanned (320–500 nm) by double beam UV–visible spectroscopy.
Preparation and characterization of ZnONPs
To obtain ZnONPs, 1.0 mL pyoverdine extracted from the supernatant of exponential phase bacterial culture was mixed with 9.0 mL of Zn(NO3)2 solution (200 mg L-1)26, 27. The reaction mixture was incubated at room temperature. In the control set, Zn(NO3)2 solution was replaced by double distilled water and kept under a similar set of conditions. Experiments were carried out in triplicate. A visible change in the color of the Zn(NO3)2 + pyoverdine mixture, from light green to fluorescent green, was observed. The visual change in the color was indicative of synthesis of ZnONPs, which were characterized by using standard analytical techniques such as UV–visible spectroscopy, Fourier transform infrared spectroscopy (FTIR), X-ray diffraction (XRD) and transmission electron microscopy (TEM) analysis.
Monitoring of ZnONP synthesis by UV–visible spectroscopy-
UV–visible spectroscopy was used to monitor metal interaction with pyoverdine. The greenish aqueous layer of pyoverdine was separated and subjected to UV–visible spectrophotometric analysis. The change in absorbance of the resultant green colored pyoverdine in the presence of zinc metal was recorded between 320 and 500 nm by using a double beam UV–visible spectrophotometer (1601, Shimadzu, Japan).
Characterization of ZnONPs
FTIR analysis
FTIR (Nicolet 6700, Thermo Scientific, USA) was used to determine the nature of the interaction of pyoverdine with Zn metal. A dried powder of pyoverdine treated with or without Zn(NO3)2 solution was mixed with KBr (1:100 ratio) and the mixture was compressed into translucent sample disks by using manual hydraulic pressure. The infrared (IR) spectra were recorded in the range 4000–400 cm-1 28.
XRD analysis
XRD analysis was carried out using a PANalytical X’Pert Pro with Cu-Ka radiation (l=0.15406 nm) in the 2q range from 20° to 80°. The energetic X-rays penetrated deep into the materials to provide information about the bulk structure.
TEM
TEM analysis of biosynthesized ZnONPs was conducted at the Department of Materials Science and Engineering, Indian Institute of Technology, Kanpur (India) to determine the morphology and structure of the nanoparticles. Samples for microscopy were prepared by dropping 10µl of a ZnONP sample on a copper grid. The grids were dried before recording the micrographs.
Antimicrobial effect of ZnONPs
The synthesized ZnONPs were tested for their antimicrobial properties by the agar well-diffusion method against Gram-positive – Staphylococcus aureus, Bacillus sp. and Gram-negative – Escherichia coli – bacterial strains, and fungal phytopathogens Rhizoctonia solani, Fusarium sp. and Penicillium sp. For the antibacterial assay, an axenic culture of each bacterium grown in nutrient broth medium was taken and spread over individual agar plates by using a sterile spreader. Wells of 6 mm diameter were punched into nutrient agar plates using a borer. ZnONP (12.5, 25 and 50 µg mL-1) suspensions were poured into the wells of all the media plates. The plates were incubated at 28±2°C for 48 h and the level of the zone of inhibition of bacterial growth was measured on the third day of incubation18.
For the antifungal assay, an exponential phase culture of each fungus, sub-cultured in PDB (potato dextrose broth) medium, was uniformly spread over individual plates by using a sterile glass rod spreader. Wells of 6 mm diameter were punched into PDA (potato dextrose agar) plates using a borer. Suspensions of varying concentrations of ZnONPs (12.5, 25 and 50 µg mL-1) were poured into the wells of all the plates and they were incubated at 28±2°C for five days. The diameter of the zone of inhibition of fungal growth was measured on the fifth day of incubation29.
Characterization and molecular identification of the strain DM1
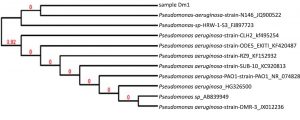
The selected bacterial strain from the rhizosphere of mustard plants was found to be a fluorescent, Gram-negative, motile, rod-shaped bacterium with a positive catalase test. Based on the morphological and biochemical study, the isolate was presumptively identified as a bacterial strain belonging to the genera Pseudomonas. Further identification of the isolated bacterial strain up to the species level was carried out using 16S rRNA sequence analysis and the BLAST program. The BLAST analysis of the isolate showed it shared a close relationship with the gene sequence of P. aeruginosa KF152932. (16S: 99% similarity with the reference strain KF152932). Thus, the bacterial isolate was identified as P. aeruginosa (GenBank accession number KF952773). A phylogenetic tree, based on a 16S rRNA partial gene sequence, is presented in Fig. 1.
Fig. 1. Phylogenetic tree of zinc-tolerant P. aeruginosa DM1 based on partial 16S rRNA sequence analysis.
Zinc tolerance
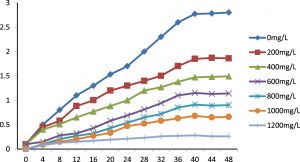
The bacterial strain P. aeruginosa DM1 that was isolated was grown in the presence of varying zinc nitrate concentrations (0, 200, 400, 600, 800, 1000 and 1200 mg L-1) to establish its zinc-tolerance properties. The bacterial growth was recorded at different time intervals (4, 8, 12, 16, 20, 24, 28, 32, 36, 40, 44 and 48 h). The results (Fig. 2) showed that the growth of bacteria gradually increased up to 40 h throughout the concentration range of zinc nitrate, and that was followed by sluggish growth beyond 40 h. The results for the concentration-dependent growth of the bacterial isolate exhibited an I-50 value (50% growth inhibitory concentration) at 400 mg L-1 of zinc nitrate. However, the bacterium could grow at a concentration of up to 1000 mg L-1 of zinc. The maximum tolerance limit value of the isolated DM1 strain against zinc was remarkably very high as compared to other bacterial isolates, showing Zn tolerance up to ≤ 500 mg L-1 (data not shown).
Pyoverdine-mediated synthesis of ZnONPs
UV–visible spectrophotometric analysis
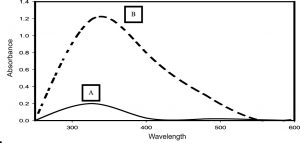
The siderophore pyoverdine extracted from the cell-free secondary metabolites of the P. aeruginosa DM1 strain was quantified using the absorbance maxima at 362 nm30, 15. Furthermore, Zn(NO3)2 solution (200 mg L-1) was mixed with aqueous solution of pyoverdine (20µM) and a change in the color (Fig. 3) of the reaction mixture from light green to fluorescent green was monitored for 24 h, indicating bio-reduction of Zn(NO3)2. The absorbance spectrum of the reaction mixture showed an increase in the characteristic absorbance maxima at 416 nm. These results together were indicative of the transformation of zinc nitrate into ZnONPs by the secondary metabolite pyoverdine (Fig. 4). Furthermore, it was interesting to see that the level of pyoverdine production by the bacterial isolate was stimulated in the presence of Zn metal (data not shown), when compared with the control (without Zn). On the contrary, production of another siderophore, pyocyanin, by the bacterial isolate declined (Fig. 5) due to the presence of zinc metal (200 mg L-1). An enhanced production of pyoverdine – a known metal chelator – in the presence of zinc could be a possible cellular defense strategy of the bacterial strain against Zn toxicity.
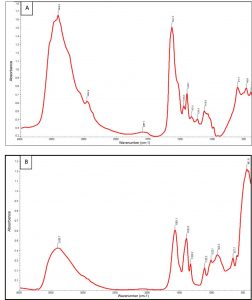
Fig. 4. UV–visible spectra (320–500 nm) of extracted pyoverdine from the secondary metabolites of P. aeruginosa DM1 grown without addition of zinc nitrate (A) and with addition of zinc nitrate (200 mg L-1).
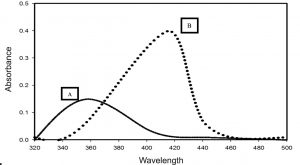
Fig. 5. UV–visible spectra (250–600 nm) of pyocyanin extracted from the secondary metabolites of a 4-day-old culture of P. aeruginosa DM1 grown with 200 mg L”1 of zinc nitrate (A) and without zinc nitrate (B), respectively.
FTIR analysis
FTIR measurement was performed to identify the possible interaction of pyoverdine with zinc nitrate in the range 400–4000 cm-1. The results showed a shift in certain characteristic IR absorbance peaks, as well as the emergence of a few new IR absorbance peaks due to the interaction of pyoverdine with zinc nitrate (Fig. 6 A, B).
The results of FTIR analysis showed that the strong IR peak at 3416 cm-1 shifted towards a lower frequency wavenumber (3408.7 cm-1), which is described as being mainly associated with conformational changes in the amine and hydroxyl groups31, 32. A sharp IR band at wavenumber 1641.8 cm-1, assigned to C=O stretching of the secondary amide (amide II), also shifted to a lower wavenumber (1581.1 cm-1). It clearly indicated the involvement of a protein/peptide moiety during the interaction of pyoverdine with zinc33. The IR peaks observed at 1402.9 cm-1 and 1339.2 cm”1 wavenumbers revealed the binding of one -C-H bond of functional group -CH2 and -N-O structure, indicative of the involvement of a hydroxamate type functional group, as reported elsewhere34, 35, 36. The results of the FTIR spectra revealed the involvement of a proteinaceous component of the pyoverdine in the reductive transformation of zinc ions into ZnONPs, as suggested by Jayaseelan et al.37 and Kumari et al.15. These results demonstrated a change in the conformational structure of the amino and hydroxamate groups of pyoverdine, which were perhaps involved in the interaction of siderophore with zinc metal.
Characterization of ZnONPs
XRD analysis
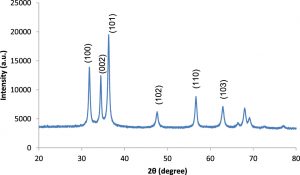
The results in Fig. 7 illustrate the XRD patterns of ZnONPs. The XRD analysis revealed seven sharp diffraction peaks in the 2q range from 20° to 80°=31.7, 34.3, 36.24, 47.46, 56.68, 63.0 and 67.8, which were found to be consistent with the reported standard Braggs’s diffraction of [100], [002], [101], [102], [110], [103] and [112] planes, characteristic of typical wurtzite type polycrystals38. Joint Committee on Powder Diffraction Standards (JCPDS card no. 36-1451) was used as a reference to assign the lattice planes according to the peaks obtained. Thus, the XRD pattern of ZnONPs clearly suggested that the nanoparticles were synthesized by the siderophore pyoverdine produced by P. aeruginosa DM1.
The selected area electron diffraction (SAED) pattern showed the crystalline nature of the particles. As depicted in Fig. 8B, the crystallinity of the particles was better due to the brighter spot.
TEM
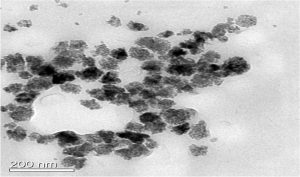
TEM was performed by FEI Tecnai G2 12 Twin TEM (120 kV) instrument. The mean size of ZnONPs was between 50 and 100 nm. Most of the nanoparticles were found to be in size range 20–100 nm. The bigger-sized particles in the range 100–200 nm were perhaps due to the agglomeration of smaller-sized particles (Fig. 8A).
Antimicrobial assay
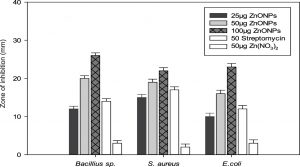
After characterization, the biologically synthesized ZnONPs were tested for their antibacterial and antifungal properties. Interestingly, different concentrations of the synthesized ZnONPs exhibited significant bactericidal activity against both Gram-negative (E. coli) and Gram-positive bacteria (S. aureus and Bacillus sp.) grown on nutrient agar medium. The fungicidal activity was tested against R. solani, Fusarium sp. and Penicillium sp. grown on PDA medium. The bactericidal and fungicidal activities were expressed in terms of the zone of inhibition (Fig. 9).
Generally, the antibacterial activity of ZnONPs increased considerably with a corresponding increase in the concentration of ZnONPs (ranging from 25 µg to 100 µg per well). Furthermore, the results showed that the zone of inhibition was formed by the ZnONPs against both the Gram-positive and Gram-negative bacterial strains (Fig. 9B). Upon loading of 100 µg ZnONPs into an 8 mm agar well, the maximum diameter of the zone of inhibition was found to be 28 mm, particularly against Bacillus sp., followed by E. coli (24 mm) and S. aureus (21 mm).
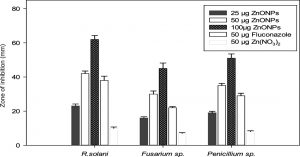
Fig. 9. (A) Evaluation of the antifungal activity of biogenic synthesized ZnONPs. (B) Evaluation of the bactericidal activity of biogenic synthesized ZnONPs.
Similarly, the antifungal activity of ZnONPs showed a concentration-dependent (25 µg to 100 µg per well) increase in the diameter of the zone of inhibition against all the phytopathogenic fungal strains. Furthermore, the results (Fig. 9A) revealed a maximum zone of inhibition against R. solani (65 mm), followed by Penicillium sp. (53 mm) and Fusarium sp. (48 mm) at 100 µg concentration. A comparison of the results on the antifungal effect of ZnONPs (50 µg per well) with established antifungal compound fluconazole (50 µg per well), as well as Zn(NO3)2 solution (50 µg per well), showed that ZnONPs were relatively more effective against all the phytopathogenic fungal strains. The bactericidal and fungicidal property of the ZnONPs suggested that these metal nanoparticles were diffusible through growth medium.
Since metal (Zn) supplemented fertilizers are commonly used in agricultural fields to enhance crop productivity, the long-term fate of zinc accumulation in the agriculture ecosystem can have significant adverse biological and ecological effects39. Bacterial transformation of toxic metals into less toxic forms have been reported elsewhere as a common cellular defense strategy of prokaryotic cells against the toxic effect of metals40. In the present investigation, the bacterial isolate DM1, identified as P. aeruginosa, with a significantly elevated level of zinc metal tolerance (1000 mg L-1), was used. Previously, workers had demonstrated that a lower concentration of Zn metal (200 µg mL-1) was toxic to bacteria41, 42. Under the influence of Zn metal (200 mg L-1), it was observed that production of pyoverdine siderophore was enhanced. On the contrary, cellular production of another siderophore, pyocyanin, was significantly reduced in the Zn-supplemented bacterial culture43. Thus, the selective production of metal-chelating siderophore by the bacterium P. aeruginosa suggested that Zn induced an enhanced production of a siderophore that could be a part of the cellular defense strategy of the bacterial isolate against the Zn metal stress 44.
In the UV–visible absorption spectrum, the characteristic absorbance peak of pyoverdine in aqueous solution was at 362 nm. The addition of zinc nitrate solution resulted in a new absorption peak at 416 nm, attributed to ZnONPs, as reported previously by other workers45. The FTIR analysis of a reaction mixture of bacterial siderophore pyoverdine and zinc nitrate solution exhibited strong IR absorption peaks at 3408.7 cm-1, 1581.1 cm-1, 1402 cm-1, 1339 cm-1 and 1120 cm-1 wavenumbers, indicating involvement of hydroxamate and amino groups in the reductive transformation of zinc nitrate into ZnONPs 46, 37. These observations were in agreement with previous findings that indicated that free amine groups, cysteine residues and carboxylate groups of enzymes or protein are involved in the transformation of zinc ions into zero-valent particles47, 48.
Furthermore, the data on the XRD pattern of synthesized ZnONPs in the 2q range and its comparison with standard Bragg’s diffraction peaks at 31.7, 34.3, 36.24, 47.46, 56.68, 63.0 and 67.8 theta values suggested typical wurtzite type polycrystals38, which were verified with the JCPDS card (no. 36-1451). The TEM images indicated a predominantly pseudospherical shape of the crystals, with smooth edges and the size range 50–100 nm. However, few undefined morphological aberrations in the particle shape might be due to agglomerations of the particles. The images observed in this study for ZnONPs through TEM were in agreement with those reported earlier49. A large number of reports are available on the biogenic synthesis of silver nanoparticles using various bacteria50, 51 and fungal strains19. Little effort has been made to study the biogenic synthesis of zinc nanoparticles37, 45, using the secondary metabolite siderophores. In the present study, it is reported that siderophore pyoverdine, a known iron chelator52, was able to interact with and transform zinc ions into zero-valent ZnONPs.
The antimicrobial impact of ZnONPs on microbial communities has been reported in the past53, 22. Earlier, it had also been indicated that the zero-valent metal nanoparticles can effectively penetrate the cell membrane of pathogenic microorganisms through the lipid bilayer, as they exhibit reduced hydrophobicity on account of the absence of surface charge54. In the light of the present investigation, it may be suggested that the antimicrobial characteristics of ZnONPs can be better exploited for the control of plant diseases, as reported by Khan and Rizvi55. However, the exact mechanism of the mode of action of ZnONPs against microorganisms still remains to be explored. The process of metal nanoparticle synthesis is more ecofriendly, as it does not involve the destruction of biological resource material and the method is free from the use of harsh chemicals. However, the use of siderophore pyoverdine for the synthesis of nanoparticles would require ample precaution as siderophores are reported to be cytotoxic56, 57. Besides the metal transformation potential of siderophores, efforts are required to evaluate the metal-induced changes in the cytotoxicity potential of these metabolites in the environment.
In the present investigation is the validation of pyoverdine (a siderophore from P. aeruginosa) mediated transformation of toxic doses of Zn ions into less toxic ZnONPs, which could be a cellular defense strategy of the Zn-tolerant bacterium P. aeruginosa against Zn metal stress. The chemical-free, in vitro biogenic synthesis of ZnONPs, without the destruction of biological resources, is an added advantage of this low-cost green technology. ZnONPs also offer a vast potential for use as a broad-spectrum antimicrobial agent to control microbial pathogens in plants.
Acknowledgements
We gratefully acknowledge the laboratory facilities provided by the Head, Department of Environmental Science, and the FTIR facility provided by the Director, University Science Instrumentation Centre (USIC), B. B. Ambedkar University, Lucknow, India. We also gratefully acknowledge the XRD and TEM facilities provided to us by the Director, Indian Institute of Technology, Kanpur (India). Manjari Barsainya is thankful to the University Grants Commission (UGC), New Delhi, India, for providing financial support in the form of research fellowship.
Conflict of Interest
The authors declares that there is no conflict of interests.
- Igwe J C, Ogunewe D N and Abia A A. Competitive adsorption of Zn (II) Cd (II) and Pb (II) ions from aqueous and non-aqueous solution by maize cob and husk. African Journal of Biotechnology, 2005; 4(10) 1113–1116.
- Stuczynski T I, McCarty G W and Siebielec G. Response of soil microbiological activities to cadmium lead and zinc salt amendments. Journal of Environmental Quality, 2003;32(4) 1346–1355.
- Teitzel G M, Geddie A, Susan K, Kirisits M J, Whiteley M and Parsek M R. Survival and growth in the presence of elevated copper: transcriptional profiling of copper-stressed Pseudomonas aeruginosa. Journal of Bacteriology, 2006; 188(20) 7242–7256.
- Chaudri A M, Allain C M, Barbosa-Jefferson V L, Nicholson F A, Chambers B J and McGrath S P. A study of the impacts of Zn and Cu on two rhizobial species in soils of a long-term field experiment.Plant and Soil, 2000; 221(2) 167–179.
- Huyer M and Page W J. Zn2+ increases siderophore production in Azotobacter vinelandii. Applied and Environmental Microbiology, 1988; 54(11) 2625–2631.
- Marques A P, Rangel A O and Castro P M. Remediation of heavy metal contaminated soils: phytoremediation as a potentially promising clean-up technology. Critical Reviews in Environmental Science and Technology, 2009; 39(8) 622–654.
- Sha R, Jiang L, Meng Q, Zhang G and Song Z. Producing cell free culture broth of rhamnolipids as a cost-effective fungicide against plant pathogens. Journal of Basic Microbiology, 2012;52(4) 458–466.
- Wightwick A and Allinson G. Pesticide residues in Victorian waterways: A review. Australasian Journal of Ecotoxicology, 2007; 13(3) 91–112.
- Rajkumar M and Freitas H. Influence of metal resistant-plant growth-promoting bacteria on the growth of Ricinus communis in soil contaminated with heavy metals.Chemosphere, 2008; 71(5) 834–842.
- Ahmad F, Ahmad I and Khan M S. Screening of free-living rhizospheric bacteria for their multiple plant growth promoting activities. Microbiological Research, 2008; 163(2) 173–181.
- Chai L, Huang S, Yang Z, Peng B, Huang Y and Chen Y. Cr (VI) remediation by indigenous bacteria in soils contaminated by chromium-containing slag. Journal of Hazardous Materials, 2009; 167(1) 516–522.
- Gericke M and Pinches A. Biological synthesis of metal nanoparticles. Hydrometallurgy, 2006; 83(1) 132–140.
- Rajapaksha R.M.C.P, Tobor-Kap³on M.A and Baath E. Metal toxicity affects fungal and bacterial activities in soil differently. Applied and Environmental Microbiology, 2004; 70(5) 2966–2973.
- Kalimuthu K, Babu R.S, Venkataraman D, Bilal M and Gurunathan S. Biosynthesis of silver nanocrystals by Bacillus licheniformis.Colloids and Surfaces B: Biointerfaces, 2008; 65(1) 150–153.
- Kumari R, Barsainya M, and Singh D.P. Biogenic synthesis of silver nanoparticle by using secondary metabolites from Pseudomonas aeruginosa DM1 and its anti-algal effect on Chlorella vulgaris and Chlorella pyrenoidosa. Environmental Science and Pollution Research, 2017; 24 4645–4654.
- Liu R and Lal R. Nanoenhanced materials for reclamation of mine lands and other degraded soils: a review. Journal of Nanotechnology, 2012; ID 461468.
- Arenas-Lago D, Rodríguez-Seijo A, Lago-Vila M, Couce L.A and Vega F.A. Using Ca3(PO4)2 nanoparticles to reduce metal mobility in shooting range soils. Science of the Total Environment, 2016; 571 1136–1146.
- Azam A, Ahmed A.S, Oves M, Khan M.S, Habib S.S and Memic A. Antimicrobial activity of metal oxide nanoparticles against gram-positive and gram-negative bacteria: a comparative study. International Journal of Nanomedicine, 2012; 7 6003–6009.
- Ottoni C.A, Simoes M.F, Fernandes S, dos Santos J.G da Silva E.S de Souza R.F.B et al. Screening of filamentous fungi for antimicrobial silver nanoparticles synthesis, 2017, AMB Express 7(1) 31.
- Dhas S P Shiny P J Khan S Mukherjee A and Chandrasekaran N. Toxic behavior of silver and zinc oxide nanoparticles on environmental microorganisms. Journal of Basic Microbiology, 2014; 54(9) 916–927.
- Dimkpa C.O, Svatoš A, Dabrowska P, Schmidt A, Boland W and Kothe E. Involvement of siderophores in the reduction of metal-induced inhibition of auxin synthesis in Streptomyces spp. Chemosphere, 2008; 74(1) 19–25.
- Shen Z, Chen Z, Hou Z, Li T and Lu X. Ecotoxicological effect of zinc oxide nanoparticles on soil microorganisms.Frontiers of Environmental Science and Engineering, 2015; 9(5) 912–918.
- Krieg N.R and J.G Holt. Bergey’s manual of systematic bacteriology, vol. 1, 1984; The Williams & Wilkins Co., Baltimore.
- Meyer J.A and Abdallah M.A. The fluorescent pigment of Pseudomonas fluorescens: biosynthesis purification and physicochemical properties. Microbiology, 1978; 107(2) 319–328.
- Wei X, Luo M, Li W, Yang L, Liang X, Xu L and Liu H. Synthesis of silver nanoparticles by solar irradiation of cell-free Bacillus amyloliquefaciens extracts and AgNO3.Bioresource Technology, 2012; 103(1) 273–278.
- Kumar C.G and Mamidyala S.K. Extracellular synthesis of silver nanoparticles using culture supernatant of Pseudomonas aeruginosa. Colloids and Surfaces B: Biointerfaces, 2011; 84(2) 462–466.
- Mishra M, Paliwal J.S, Singh S.K, Selvarajan E, Subathradevi C and Mohanasrinivasan V. Studies on the inhibitory activity of biologically synthesized and characterized zinc oxide nanoparticles using Lactobacillus sporogens against Staphylococcus aureus. Journal of Pure and Applied Microbiology, 2013;7(2) 1263–1268.
- Bryaskova R, Pencheva D, Nikolov S and Kantardjiev T. Synthesis and comparative study on the antimicrobial activity of hybrid materials based on silver nanoparticles (AgNps) stabilized by polyvinyl pyrrolidone (PVP). Journal of Chemical Biology, 2011; 4(4) 185–191.
- Jain S.K and Agrawal S.C. Fungitoxic effect of some organic volatile substances against fungi causing otomycosis.Mycoses, 1994; 37(7–8) 299–304.
- Buysens S, Heungens K, Poppe J and Hofte M. Involvement of pyochelin and pyoverdin in suppression of pythium-induced damping-off of tomato by Pseudomonas aeruginosa 7NSK2. Applied and Environmental Microbiology, 1996; 62(3) 865–871.
- Surewicz W.K, Mantsch H.H and Chapman D. Determination of protein secondary structure by Fourier transform infrared spectroscopy: a critical assessment. Biochemistry, 1993; 32(2) 389–394.
- Barsainya M and Singh D.P. Effect of secondary metabolites produced by Pseudomonas aeruginosa on root rot disease in mung bean [Vigna radiata (L) Wilczek] grown on salinized soil. Asian Journal of Microbiology Biotechnology and Environmental Sciences, 2016; 18(2) 503–512.
- Parikh R.Y, Singh S, Prasad B.L.V, Patole M.S, Sastry M and Shouche Y.S. Extracellular synthesis of crystalline silver nanoparticles and molecular evidence of silver resistance from Morganella sp: towards understanding biochemical synthesis mechanism. ChemBioChem, 2008; 9(9) 1415–1422.
- Yang J, Bremer P J, Lamont I L and McQuillan A J. Infrared spectroscopic studies of siderophore-related hydroxamic acid ligands adsorbed on titanium dioxide. Langmuir, 2006; 22(24) 10109–10117.
- Goloverda G, Jackson B, Kidd C and Kolesnichenko V. Synthesis of ultra small magnetic iron oxide nanoparticles and study of their colloid and surface chemistry.Journal of Magnetism and Magnetic Materials, 2009; 321(10) 1372–1376.
- Meraat R, Ziabari A.A, Issazadeh K, Shadan N and Jalali K.M. Synthesis and characterization of the antibacterial activity of zinc oxide nanoparticles against Salmonella typhi. Acta Metallurgica Sinica, 2016; 29(7) 601–608.
- Jayaseelan C, Rahuman A.A, Kirthi A.V, Marimuthu S, Santhosh Kumar T, Bagavan A and Rao K.B. Novel microbial route to synthesize ZnO nanoparticles using Aeromonas hydrophila and their activity against pathogenic bacteria and fungi. Spectrochimica Acta Part A: Molecular and Biomolecular Spectroscopy, 2012; 90 78–84.
- Shoeb M, Singh B.R, Khan J.A, Khan W, Singh B.N, Singh H.B et al. ROS-dependent anticandidal activity of zinc oxide nanoparticles synthesized by using egg albumen as a biotemplate. Advances in Natural Sciences: Nanoscience and Nanotechnology, 2013; 4(3) 035015.
- Singh R, Singh D.P, Kumar N, Bhargava S.K and Barman S.C. Accumulation and translocation of heavy metals in soil and plants from fly ash contaminated area. Journal of Environmental Biology, 2010; 31 421–430.
- Abhilash P.C, Powell J.R, Singh H.B and Singh B.K. Plant–microbe interactions: novel applications for exploitation in multipurpose remediation technologies. Trends in Biotechnology, 2012;30(8) 416–420.
- Babich H and Stotzky G. Toxicity of zinc to fungi bacteria and coliphages: influence of chloride ions. Applied and Environmental Microbiology, 1978; 36(6) 906–914.
- Nweke C.O, Alisi C.S, Okolo J.C and Nwanyanwu C.E. Toxicity of zinc to heterotrophic bacteria from a tropical river sediment. Applied Ecology and Environmental Research, 2007; 5(1) 123–132.
- Lee J.H, Kim Y.G, Cho M.H and Lee J. ZnO nanoparticles inhibit Pseudomonas aeruginosa biofilm formation and virulence factor production. Microbiological Research, 2014; 169(12) 888–896.
- Talam S, Karumuri S.R and Gunnam N. Synthesis characterization and spectroscopic properties of ZnO nanoparticles. ISRN Nanotechnology, 2012; 372505.
- Singh B.N, Rawat A.K.S, Khan W, Naqvi A.H and Singh B.R. Biosynthesis of stable antioxidant ZnO nanoparticles by Pseudomonas aeruginosa rhamnolipids. PloS One, 2014; 9(9) e106937.
- Osorio R, Osorio E, Medina-Castillo A.L and Toledano M. Polymer nanocarriers for dentin adhesion. Journal of Dental Research, 2014; 93(12) 1258–1263.
- Mukherjee P, Ahmad A, Mandal D, Senapati S, Sainkar S.R, Khan M.I et al. Bioreduction of AuCl4 ions by the fungus Verticillium sp and surface trapping of the gold nanoparticles formed. Angewandte Chemie International Edition, 2001;40(19) 3585–3588.
- Mashrai A, Khanam H and Aljawfi R.N. Biological synthesis of ZnO nanoparticles using C. albicans and studying their catalytic performance in the synthesis of steroidal pyrazolines. Arabian Journal of Chemistry, 2013; 10 S1530–S1536.
- Jain N, Bhargava A, Tarafdar J.C, Singh S.K & Panwar J. A biomimetic approach towards synthesis of zinc oxide nanoparticles. Applied microbiology and biotechnology, 2013; 97(2) 859-869.
- Thakkar K.N, Mhatre S.S and Parikh R.Y. Biological synthesis of metallic nanoparticles. Nanomedicine: Nanotechnology Biology and Medicine, 2010; 6(2) 257–262.
- Jeevan P, Ramya K and Rena A.E. Extracellular biosynthesis of silver nanoparticles by culture supernatant of Pseudomonas aeruginosa. Indian Journal of Biotechnology, 2012; 11(1) 72–76.
- Chen W.J, Kuo T.Y, Hsieh F.C, Chen P.Y, Wang C.S, Shih Y.L et al. Involvement of type VI secretion system in secretion of iron chelator pyoverdine in Pseudomonas taiwanensis. Scientific Reports, 2016;6 32950.
- Raghupathi K.R, Koodali R.T and Manna A.C. Size-dependent bacterial growth inhibition and mechanism of antibacterial activity of zinc oxide nanoparticles. Langmuir, 2011; 27(7) 4020–4028.
- Hulkoti N.I and Taranath T.C. Biosynthesis of nanoparticles using microbes – a review. Colloids and Surfaces B: Biointerfaces, 2014; 121 474–483.
- Khan M.R and Rizvi T.F. Nanotechnology: scope and application in plant disease management. Plant Pathology, 2014; 13 214–231.
- Pájaro M.C, Barberis I.L and Albesa I. Pseudomonas fluorescens: production of pyoverdine in human blood at 4 degrees C and cytotoxic effect of the pigment. Revista Latinoamericana de Microbiologia, 1994; 37(1) 1–6.
- Wagner V.E, Filiatrault M.J, Picardo K.F and Iglewski B.H. Pseudomonas aeruginosa virulence and pathogenesis in Pseudomonas: Genomics and Molecular Biology (Norfolk: Caister Academic Press), 2008; 129–158.
© The Author(s) 2018. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.