In December 2019, a novel coronavirus had emerged in Wuhan city, China that led to an outbreak resulting in a global pandemic, taking thousands of lives. The infectious virus was later classified as severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). Individuals infected by this novel virus initially exhibit nonspecific symptoms such as dry cough, fever, dizziness and many more bodily complications. From the “public health emergency of international concern” declaration by the World Health Organisation (WHO), several countries have taken steps in controlling the transmission and many researchers share their knowledge on the SARS-COV-2 characteristics and viral life cycle, that may aid in pharmaceutical and biopharmaceutical companies to develop SARS-CoV-2 vaccine and antiviral drugs that interfere with the viral life cycle. In this literature review the origin, classification, aetiology, life cycle, clinical manifestations, laboratory diagnosis and treatment are all reviewed.
Coronaviruses, COVID-19, Mutation, Spike Protein, Main Protease
Classification and origin of SARS-CoV-2
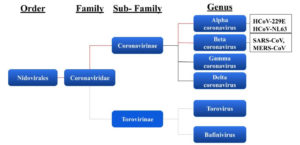
SARS-CoV-2 belongs to the order Nidovirales which is part of the Coronviranae subfamily that comes under the Coronaviridae family. Through antigenic and genetic relationships, Coronavirnae is categorised into 4 genera which include alphacoronavirus (α-CoV), betacoronavirus (β-CoV), gammacoronavirus (g-CoV) and deltacoronavirus (δ-CoV), where α-CoV and β-CoV are reported to infect mammals, and g-CoV and δ-CoV are reported to mainly infect birds and certain types of mammals.1 To date, the human coronavirus 229E (HCoV-229E) and NetherLand 63 (NL63) members from α-CoV genera, and the Organ Culture 43 (OC43) and Hong Kong University 1 (HKU1) members of β-CoV genera are the common types of viruses that infect humans that leads to the common cold.2 The genera of coronavirus can be further classified into the subgenera of Hibecovirus, Nobecovirus, Merbecovirus, Embecovirus and Sarbecovirus. Through phylogenetic analysis, SARS-CoV-2 is classified under the subgenus Sarbecovirus that is part of the genus β-CoV. In the past two decades, β-CoVs such as the severe acute respiratory syndrome (SARS-CoV) and the Middle Eastern respiratory syndrome coronavirus (MERS-CoV) and SARS-CoV-2 itself are seen to be highly pathogenic to humans. With the aid of next-gen sequencing, SARS-CoV-2 shows 50% similarity to MERS-CoV and 79% similarity to SARS-CoV.3-4
The natural host of coronaviruses (CoVs) are observed to be mostly from bats with the involvement of intermediary animal hosts that leads to the spillover event to humans. In the case of SARS-CoV and MERS-CoV, domestic civets and camels are found to be the intermediate host, respectively. For SARS-CoV-2, studies have shown and are ongoing to identify its intermediate host. From a study performed by Zhou et al., the RaTG13 sampled from a Rhinolophus affinis was clustered with SARS-CoV-2 with a genome sequence identity of 96% and concluded that there is a potential chance of SARS-CoV-2 to be originated from bats.5 Other than Rhinolophus affinis being a probable intermediate host, through metagenomic sequencing that was carried out by Lam et al., shows that Manis javanica (Malayan pangolin), which were seized in anti-smuggling operations in southern China, have two sublineages of SARS-CoV-2 related CoVs where one of the lineages is reported to portray strong similarity in the receptor-binding domain (RBD) to SARS-CoV-2 and to have a genome sequence identity of 92.22% to SARS-CoV-2 which deduces that pangolins to be one of the plausible intermediate hosts.5-6
Epidemiology of SARS-CoV-2
In November 2019, the world encountered a new emergence of unidentified acute respiratory tract infectious disease in the city of Wuhan, Hubei Province in China.7-8 Through similar traces in cases, it was found to have originated from a local fish market called Hunan South China Seafood Market in Wuhan and this was also later confirmed by the World Health Organization.9 With the onset, an epidemiological and etiological investigation was initiated by the China Centre for Disease Control and Prevention (China CDC).7,10 China reported the cluster of pneumonia-like new cases to WHO on 31st of December 2019, which led it to be identified as a novel coronavirus.11 WHO declared the novel coronavirus as a Public Health Emergency of International concern on 30th of January 2020. Up until 31st of January 2020, the novel coronavirus had spread to 19 different nations with 11, 791 infected cases and 213 deaths7. Ever since the onset, SARS-CoV-2 has caused a worldwide pandemic owing to its highly infectious nature. The highest spike in cases was witnessed in Iran, South Korea and Italy and has also spread to 150 nations.8 The rapid spread of SARS-CoV-2 also called for researchers to gather data and information that would aid in the development of cures and vaccines. The pioneering research was conducted by Prof. Yong Zhang, who delivered the genome of COVID-19 on 10th of January 2020.7
Table (1):
The non-structural proteins of coronaviruses and its respective functions52,43.
No. |
Type of nsp* (s) |
Function (s) |
|---|---|---|
1 |
nsp1 |
Blocks the host’s immune response by inhibiting type I (IFN)** expression and blocks the host’s mRNA translation |
2 |
nsp2 |
Interacts with prohibitin protein 1, 2 |
3 |
nsp3 |
Involves in polyprotein processing, aids in the formation of RTC. Coexpression with nsp 4 and nsp 6 results in the formation of DMV*** |
4 |
nsp4 |
Membrane rearrangement |
5 |
nsp5 |
Mediates processing at the 11 cleavage sites and can block inhibit IFN signalling |
6 |
nsp6 |
Activates autophagosomes formation from the host’s endoplasmic reticulum and contributes to DMVs formation |
7 |
nsp7 |
Cofactor of nsp12 |
8 |
nsp8 |
Cofactor of nsp12 |
9 |
nsp9 |
Associates in viral RNA production (RNA binding protein) |
10 |
nsp10 |
Cofactor of nsp14 and nsp16 |
11 |
nsp12 |
Performs viral RNA synthesising activity, with aid of nsp7 and nsp8 complex |
12 |
nsp13 |
By cooperative translocation, unwinds duplex RNA |
13 |
nsp14 |
Harbours guanine-N7-methyltransferase and ExoN activity |
14 |
nsp15 |
Induces endonuclease activity, limits the exposure of viral double-stranded RNA to host sensors |
15 |
nsp16 |
Catalyses the methylation of the penultimate nucleotide of the viral RNA, limit the detection by the host innate immunity |
*Non structural protein
**Interferon
***Double membrane vesicle
As of 15th of November 2020, WHO recorded 53,766,728 million cases and 1,308,975 million deaths in accordance with the reports received from various countries. From 9th November to 15th November the highest number of cases were reported by the United States of America with 1 million new cases, India with 306 cases, Italy with 242,00 cases, France with 203,000 cases and Brazil with 179,000. The highest recorded cumulative cases were in the United States of America with 22,960,102 cases, Europe with 15,047,248 cases and South- East Asia with 10,015,731. Focusing on South-East Asia, from the latter half of September to October, there has been a decline in the number of cases from 690,00 to 373,786 and deaths from 9,300, as reported in the week of 14th September to 4,534 in the week of 9th November. The countries reporting the highest number of cases in South-East Asia in the past week were Nepal, Maldives and India.12
As of 20th of November 2020, Malaysia has reported 52,638 confirmed cases with 329 deaths and 39,088 recovered cases. The first case in Malaysia appeared on 25th of January 2020 with the source of infection being traced to three Chinese nationals who arrived in Malaysia on the 24th of January 2020. They had been infected due to close contact with a Singapore national who had contracted SARS-CoV-2. The treatment was given in Sungai Buloh Hospital, Selangor, Malaysia.13,14 With the onset in Malaysia, the Ministry of Health (MOH) developed standard guidelines for managing the COVID-19. MOH also assigned 34 Hospitals and screening centres in each state. The first Malaysian case was detected on 4th of February 2020, involving a 41-year-old man who returned to Malaysia from Singapore. The individual was observed to have symptoms of fever and cough. He was kept under quarantine at Sungai Buloh Hospital, Selangor.15 This case also led to the first local transmission case on the 6th of February 2020, involving a 40-year-old Malaysian female, who was the sister of the 41-year-old man. The symptoms encountered were fever accompanied by a sore throat on the 1st of February 2020 and cough on the following day. The patient was kept under isolation at Sultanah Bahiyah Hospital, Alor Setar on the 3rd of February 2020. The first recovered and discharged case was on 4th of February 2020, involving a 4-year-old Chinese national girl who was kept under isolation at the Sultanah Maliha Hospital, Langkawi on 29th of January 2020.16 The first sporadic COVID-19 case was detected on 12th of March 2020 where the contracted individual did not have any travel history to COVID-19 affected area or had any contact with an infected person.17
The spike in cases was caused by a religious gathering in Sri Petaling, Kuala Lumpur. This also resulted in delivering the highest number of positive cases in Malaysia and also in Southeast Asia. The gathering consisted of 1500 individuals from several countries, which led to the outspread of the neighbouring countries namely Brunei, Indonesia, the Philippines, Thailand, Vietnam and Cambodia. The first case from the gathering was reported in Brunei, involving a 53-year-old man who was tested positive on return to Brunei. He was reported to show symptoms on 7th of March 2020 and was treated under isolation at the National Isolation centre in Tutong, Brunei. The Prime Minister of Malaysia implemented the Movement Control Order (MCO) on 16th of March 2020, due to the rise of cases above 553. Various safety measures were enforced such as social distancing, restricted interstate travel and limits on the number of family members allowed to travel, to contain the spread of COVID-19.16,18
The first deaths due to COVID-19 were reported on 17th of March 2020, involving a 60-year-old man with a history of chronic disease from Kuching, Sarawak and a 34-year-old man from Johor Bahru, who had contracted the virus through attending the religious gathering in Sri Petaling, Kuala Lumpur. On 28th of April 2020, the government then enforced Enhanced Movement Controlled Order (EMCO) for specific areas with high origins of positive cases. EMCO enforced more stringent rules such as restriction on residents from leaving their houses, having visitors or visiting any COVID- 19 infected areas.16
Transmission of SARS-CoV-2
Studies have shown that transmission of SARS-CoV-2 can also be zoonotic in nature. It was initially by this zoonotic transmission that humans came into contact with the virus. SARS-CoV-2 susceptibility to mutation, it led to an increased rate of transmission. SARS-CoV-2 infected individuals can be asymptomatic and symptomatic. Transmission of the virus from an individual to another is through contact with bodily fluid such as saliva and/or respiratory droplets and transmitted within 1-meter in distance.7 Transmission over 1-meter distance is also possible through airborne transmission as the droplet nuclei can prolong their presence in the air due to their size. SARS-CoV-2 can also be transmitted through indirect contact with objects that were used by the infected individual (eg: doorknob, handles of the stairwell).19-20 Touching or warmly greeting an individual who has the infection may pass the infection between other individuals. Making contact with a surface or item that has been infected and afterwards directly contacting the individual’s nose, eyes, or mouth also ease the transmission of the virus. The National Institutes of Health (NIH) proposes that infants, children, individuals who are 65 years or older, pregnant women and individuals with low immune systems are at increased risk of acquiring severe medical complications, due to COVID-19 infection.21 Moreover, inadequate implementation of infection control in private/public sectors that are in outbreaks22 and lacking admittance to PPE has been distinguished as one of the major risk factors for transmission.23 Through systematic literature and meta-analysis, it has been deduced that social distancing and the use of eye protection and face mask, reduced the risk of transmission.24
A recent study exhibited a relationship between ABO blood classifications and COVID-19 risk. Utilizing the information from Shenzhen and Wuhan, Zhao showed that blood group A individuals are more prone to be infected with SARS-CoV-2 whereas individuals who are blood group O to be least susceptible.25 This was also supported by a meta-analysis from Italy and Spain that found a higher risk of COVID-19 among blood group A and a lower risk among blood group O.26 In another study that was performed in Canada with 95 patients who were infected with SARS-CoV-2, found that 84% of patients who were either blood group A or AB required mechanical ventilation support whereas only 61% of the patients who were either blood group O or B required mechanical ventilation support. Additionally, patients with blood group A or AB had a more prolonged stay in the intensive care unit of 13.5 days, on average, compared with the patients with blood group O or B who stayed for an average of 9 days.27
Genomic structure of SARS-CoV-2
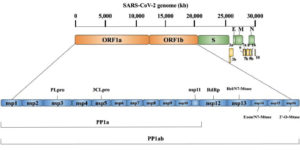
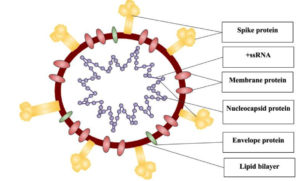
SARS-CoV-2 is pleomorphic or spherical in structure which consists of single-stranded positive-sense RNA (+ssRNA) that is bound to a nucleoprotein (NP) which is present inside a capsid, compromised with matrix proteins. With a viral genome size of 29.8kb-30kb, that encodes for approximately 9,860 amino acids, the genome comprises 38% of G + C contents and 11 protein-coding genes, with 12 expressed proteins.28 This large size genome is maintained by RNA processing enzymes such as the 3′-5′ exoribonuclease of nsp14 (nsp14-ExoN) which functions as a proofreader on the replication/transcription complexes (RTC), preventing lethal mutagenesis.29 SARS-CoV-2 is observed to be polycistronic mRNA that follows the 5′-cap and 3′-poly-A tail arrangement which comprises of the open reading frame (ORF) 1a and 1b and structural proteins namely spike (S) protein, envelope (E) protein, membrane (M) protein, nucleocapsid (N) protein, that is flanked with 5′ and 3′ untranslated regions (UTRs) which are 265 and 358 nucleotides in length.30
ORF 1a that encodes for 16 non-structural proteins (nsps), occupies nearly 67% of the genome, are cleaved by the polypeptide protein phosphatase 1a (pp1a) and protein phosphatase 1b (pp1b). The remaining ORFs encode for accessory proteins such as ORF 3, 6, 7a, 7b, 8 and 9b and structural proteins. Some of the nsps expressed include helicase (nsp13), RNA-dependent RNA polymerase (nsp12), two viral cysteine protease namely chymotrypsin-like or main protease (nsp5) and papain-like protease (nsp3) and several others that are associated with the replication and transcription process of SARS-CoV-2.30 The four major structural proteins namely nucleocapsid protein (N), envelope (E), spike surface glycoprotein (S) and accessory proteins which include ORF3a, ORF6, ORF7a/b, ORF8, and ORF9b is encoded by ORFs.31 The S protein functions as viral adhesion and fusion protein and consists of two subunits, S1 and S2. The S1 subunit contains a signal peptide, receptor-binding domain (RBD) and N-terminal domain (NTD), and E and M protein are involved in budding and assembly of viral particles that aids in the viral morphogenesis.31,32
Through structural analysis, the S1 subunit is observed to have a similar sequence identity of 70% with human SARS-CoV and bat SARS-like CoVs, with the core domain of RBD to be highly conserved. The S2 subunit of SARS-Cov-2 is reported to be highly conserved and has a sequence identity of 99% to human SARS-CoV and two bats SARS-like CoVs namely SL-CoV ZC45 and ZX C21. The 5′-UTR of SARS-CoV-2 consists of several stem-loops (SL) structures which include SL1-SL4, S5, SL5A, SL5B, and SL5C which are observed to bear resemblance with the human SARS-CoV and SL-CoV ZC45. Moreover, the novel coronavirus was seen to have the same SL6, SL7, and SL8 structures as the SARS-CoV, however with an extra SL. Furthermore, part of the S5 from SARS-CoV-2 was observed to be present in ORF 1a and ORF 1b that was identical in SARS-CoV, although not present in SL-CoV ZC45. It was also observed that SL-CoV ZC45 did have a SARS-CoV SL6-like stem-loop. From these analyses, the author was able to conclude that the 5′-UTR of SARS-CoV-2 share more similarities than the SL-CoV ZC45. The 3′-UTR was conserved among SARS-related CoVs including SARS-CoV, SAR-CoV-2 and human SARS-CoV.33
In RNA viruses, the rate of mutation is very high that leads to an increase in its evolvability and virulence. While most of the RNA viruses are susceptible to random mutation due to the absence of proofreading activity by exonuclease (ExoN), Nidoviruses mutation occurs due to dynamic equilibrium between the random host cell defence mechanism, viral survival and random perturbation.34-36 Mutations within the strains of SARS-CoV-2 are pivotal in affecting the rate of viral mortality and spread, which varies between countries. For CoVs, the mutation rate is estimated to be 4×10-4 nucleotide substitutions/site/year.36 Since the first SARS-CoV-2 genome was published in early January 2020, there were several new mutations detected all around the world, on different residue positions of the S protein, especially at the RBD region. Through a computational study performed by Chen et al., were able to identify 11,904 single mutations in six distinct clusters, when comparing the 24,715 complete SARS-CoV-2 genome samples with the first genome sequence of SARS-CoV-2 that was obtained on the 5th of January 2020. The six distinct clusters were to be found in different geographical locations which include the United States of America, United Kingdom, Australia, Canada, Iceland, Russia, Spain, Italy Germany, Belgium, France, China, India, Turkey, Saudi Arabia, Japan and Korea. Furthermore, the authors were able to detect 725 existing non-degenerated mutations on SARS-CoV-2 S protein, where 89 of those mutations were observed to be in the RBD region. Moreover, out of 89 mutations, 59 of the mutations are observed to be on the receptor-binding motif (RBM). The authors were able to conclude that the novel coronavirus has become comparatively more contagious and some of the mutated strains may tend to further mutate and become even more infectious.37
Another mutation analysis that was performed on all of the major proteins of 13 SARS-CoV-2 sequences from different countries namely China, India, Japan, Finland, Vietnam, Sweden, South Korea, Brazil, Taiwan, Australia, Nepal and Italy, showed that the Indian isolate portrayed mutation within the spike glycoprotein at 408 positions (Arginine replaced by Isoleucine) and replicase polyprotein mutations, with one mutation on the RNA-dependent RNA polymerase (RdRp) (A408V). A mutation on the main protease (Mpro) (R60C) was only detected in the Vietnam isolate.36
In Malaysia, the first mutation of SARS-CoV-2, announced by Health director-general Tan Sri Dr Noor Hisham Abdullah, was detected in two undocumented migrants that were from the Benteng LD (Lahad Datu) cluster.38 The D614G mutation resulted in the changes in the amino acid location at position 614, from aspartic acid (D) to glycine (G).39 Due to this change, the G614 variant was observed to have a higher viral load and produced a significantly higher infectious titer (2.6- to 9.3-fold increase) than of the D614 variant. Even Though the viral load is higher in the mutated variant, no differences were observed in the hospitalization outcomes.40 The existence of several kinds of S protein mutations represents various SARS-CoV-2 strains with different infectivities, but the real threat is in the increased mutation rate at the RBD that have led to highly infectious SARS-CoV-2 variants.37
Life cycle of SARS coronaviruses
Once the CoV enters the human lungs, the virus majorly affects the alveolar epithelial type 2 (AT2) cells.41 The virus uses human angiotensin-converting enzyme 2 (ACE2) a type 1 membrane protein as their receptor to gain entry to the host cell. Once bound to the receptor, the proteolytic cleavage of S protein by type 2 transmembrane serine protease (TMPRSS2) occurs resulting in the invasion of the host’s cytosol.42 The cleavage for the S2 protein section occurs in two regions. Separation of the RBD and fusion domains of the S protein occurs during the first cleavage and the following cleavage reveals the fusion peptide leading to the insertion into the plasma membrane.43 After the second cleavage, two heptad repeats in S2 bind together and form an antiparallel six-helix bundle. This results in the merging of viral and cellular membranes and ultimately fusing with the host cell membrane, allowing the viral genomic RNA to enter the cytosol.44
The viral genome RNA consists of a replicase gene that encodes for two sizable ORFs (rep1a and rep1b), that undergoes translation producing two large overlapping polyproteins namely pp1a and pp1ab.45 The expression of the two-terminal polyproteins was achieved by utilizing a slippery sequence (UUUAAAC) and an RNA pseudoknot which leads to a ribosomal frameshifting from rep1a reading frame to rep1b ORF.46 The cleavage between pp1a and pp1ab into nsps 1–11 and nsps 1–16, respectively, occurs in the presence of protease enzymes such as the papain-like proteases (PLpro), encoded within nsp 3, and a serine-type Mpro (chymotrypsin-like protease (3CLpro)) protease, encoded in nsp 59.47-49 Most of the nsps assemble for the formation of RTC in double-membrane vesicles (DMVs) which aids in the viral genomic and subgenomic synthesis.50 The genomic and subgenomic RNAs are produced with the help of negative-strand intermediates. Furthermore, with the involvement of RdRp, the negative-strand intermediate products are transcribed to positive-sense mRNAs.51
Following the viral genomic and subgenomic synthesis, the viral structural proteins migrate into the endoplasmic reticulum (ER) after undergoing translation. These proteins then arrive at the endoplasmic reticulum–Golgi intermediate compartment (ERGIC) through the secretory pathway, where the encapsidated viral genomes bud into the ERGIC’s membrane. This event leads to the maturation of the virion.53 The M protein manages the protein-protein interactions that are required during the virus assembly in the host cell. When the M and E proteins are expressed, virus-like particles (VLPs) are developed which results in the production of CoV envelope.54 The N protein enhances the VLP formation during the merging of the encapsidated viral genome into the membranes of the ERGIC.55 Furthermore, the M protein associates with the nucleocapsid, completing the process of the virion assembly.56 The viral release from the host cell occurs primarily by exocytosis, budding or cell death.57 Fully matured viruses are exocytosed from the endoplasmic reticulum or Golgi apparatus whereas premature virion particles undergo budding before the viruses are being released.58 During the budding process, the N protein binds to the region of the cell membrane where the E protein was inserted, giving access for appropriate orientation of the virion at the perinuclear, nuclear, endosomal or plasma membranes that result in the release of the virus. In certain situations, the SARS-CoV-2 affects the system of the host cell causing cell death and the release of the lysosomes along with the virus particles. The released viruses from the infected host cell may be released to the open environment through the said mode of transmission or may infect other healthy cells in the host’s body.43
Host’s immune response against SARS-CoV-2
The host’s innate and adaptive immune systems are triggered, through antigen-presenting cells (APC) (eg. macrophages and dendritic cells), upon viral entry into the host cell. These APCs are attached with pattern recognition receptors (PRR) namely NOD-like receptors (NLRs), toll-like receptors (TLRs) and RIG-I-like receptors (RLRs) that are situated at several regions in the host cells which include in the cytosol, lysosomes, plasma membranes and endosomal membrane. PRRs detect specific evolutionarily conserved structures on pathogens called pathogen-associated molecular patterns (PAMP) which consists of glycoproteins, nucleic acids, lipoproteins and carbohydrate moieties that are to be located in the structural components of viruses. Once detected by PRR, it stimulates the production of type I interferons (IFN) with antiviral functions, causing interference in viral’s replication ability.59-61
During the host’s adaptive immunity, the major histocompatibility complex (MHC) presents the viral antigen on the surface of the infected cells, facilitating recognition by virus-specific cytotoxic T lymphocytes (CTLs). In most scenarios, MHC class I molecules present the viral antigen, however, there are certain reports of MHC class II to present the viral antigen to CD4+ T cells.62 Furthermore, B cells are activated by CD4+ T cells which generate virus-specific antibodies namely immunoglobulin M (IgM) and immunoglobulin G (IgG).63 During antigen presentation by MHC class I molecules, it leads to the release of cytokines and activation of CD8+ T cells, causing the infected cells to undergo apoptosis.61 Additionally, T helper cells secrete pro-inflammatory cytokines that aid other immune cells and enhance antigen presentation. Activated T cells secrete chemokines which recruit more adaptive and innate cells to control the viral infectivity.63
Immune evasion of SARS-CoV-2
SARS-CoV-2 is reported to utilize several tactics that aids in its survival against the host immune system and most of its survival mechanism is similar to those of SARS-CoV and MERS-CoV. The immature dendritic cells there are present in the bronchioles and alveoli present antigen constantly to naive T lymphocytes during pathogenic stimuli resulting in a larger response through the production of cytokines.65 During SARS-CoV infection, the virus decreases the expression of costimulatory cell surface molecules such as CD80 and CD86 and human leukocyte antigen (HLA) class 2 molecules on immature DCs, obstructing viral antigenic presentation.64
SARS-CoV-2 expresses endonuclease, nsp1, nsp8 and DMV which affects the IFN α/β production. Endonuclease inhibits the detection and stimulation of dsRNA sensors namely melanoma differentiation-associated protein 5 (MDA5) and protein kinase R (PKR) that blocks the IFN α/β production and also suppress the host cell and RNase- L translation in the cells that are infected. Additionally, DMV also blocks the detection by MDA5 and PKR. To achieve interferon-stimulated genes (ISG) expression, IFN-α/β needs to bind to interferon-alpha/beta receptor (IFNAR) 1 and 2 receptor which then phosphorylates and activates signal transducer and activator of transcription (STAT) 1 and 2, leading to the expression.66,67 When nsp1 binds to STAT 1, it prevents phosphorylation68 and translocation, obstructing the release of antiviral proteins. In addition, nsp1 is also reported to directly cause RNase-host degradation.69
The CoVs also inhibit the host immune defence mechanism by suppressing the T cells functions by direct apoptosis and also cause functional exhaustion in CTLs and natural killers (NK) cells.64,70 When natural killer cell receptor NKG2A interacts with HLA E, induces inhibitory signals to NK cells resulting in inhibition of their activation further causing an inability to eliminate the viral infection.71 This event is seen to occur in SARS-CoV-2 infection, however, the process that drives NK cells exhaustion is poorly understood. Although, there are some potential strategies that include increased expression of NKG2A on NK cells or certain HLA-I molecules that cause an upregulated expression of HLA-I derived signal peptides leading to rising in NKG2A inhibitory signals in SARS-CoV-2 infected patients.70
Clinical features
General clinical manifestations
The initial onset symptoms begin approximately 14 days after the exposure, with almost all of the cases with an average time between 4-5 days from the day of exposure to the occurrence of first symptoms from the infected patients. In a study, it was revealed that 97.5% of individuals infected with COVID-19 show the symptoms within 11.5 days right after the initial exposure.72 Individuals infected with SARS-CoV-2 are usually seen with nonspecific symptoms such as dry cough, fever and dizziness. The other bodily systems that may also be affected include the respiratory tract system (clinical conditions: dry cough, winded, sore throat, rhinorrhoea, chest torment and haemoptysis), gastrointestinal system (clinical conditions: looseness of the bowels, vomiting, and nausea), musculoskeletal system (clinical conditions: muscle throb), and neurological system (clinical conditions: cerebral pain or disarray/confusion). Other common general indications observed in individuals are such as shortness of breath, fever and cough.73,74 Patients with serious clinical indications, experience intense type pneumonia, acute respiratory distress syndrome (ARDS), extrapulmonary signs and systemic confusions, for example, septic shock, and sepsis. The clinical course of the illness appears to foresee an ideal pattern in most patients.75
The author of the Chinese CDC reported stages of clinical manifestation of COVID-19 are divided according to the different severity of the infection. Under the mild infection which is 81% of the cases, patients showed mild pneumonia and non-pneumonia symptoms. In severe cases which is 14 % of the patients presented symptoms such as an increase in the frequency of respiration, dyspnoea and changes in the blood oxygen saturation level along with changes in the rate of oxygen delivery to patients within 24 to 48 hours are observed. As for the 5% of critical infections, patients with other indications such as experiencing septic shock, respiratory failure, few organs dysfunction or failure are persistently observed. As for all cases, the very initial symptoms observed are the presents of pneumonia in patients which are characterized by coughing, dyspnoea, fever and bilateral infiltrates indications are seen on chest imaging.76
Individuals who are more prone to have a serious infection from COVID-19 are individuals who have existing constant sickness likewise have a high risk of obtaining the infection. Certain types of diseases that trigger the risk of COVID-19 infection include chronic heart problems, for example, malignant growth, cardiovascular disease, coronary artery infection or cardiomyopathy, persistent obstructive pneumonic infection (COPD), diabetes type 2, smoking, obesity, persistent kidney infection, sickle cell disease, asthma, liver failure, persistent lung infections, for example, pneumonic fibrosis or cystic fibrosis, cerebrum and sensory system conditions and hypertension.77
Clinical manifestation in paediatric patients
SARS-CoV-2 is also reported to infect infants and young children, causing symptomatic and asymptomatic conditions, however, the paediatric manifestation is constantly expanding. In addition, children with pre-existing medical conditions are more susceptible to COVID-19 infection. Most of the child cases have mild indications, with the absence of fever or pneumonia, and the incubation period is assessed to be 1-14 days after the beginning of the illness. SARS-CoV-2 presents a longer average incubation period in children with 6.5 days compared to adults of 5.4 days.78 The common clinical signs in children are fever and cough; along with occasional manifestations such as exhaustion, myalgia, nasal clog, runny nose, wheezing, sore throat, migraine, spewing, unsteadiness, or stomach torment.79,80 There are additional occurrences of asymptomatic paediatric patients or individuals who just display cough or looseness of the bowels. Contaminated babies of children may introduce run of the mild side effects, for example, gastrointestinal appearances, asthma, or shortness of breath. However, in severe stages, it may lead to the distress of acute respiratory, dysfunction of coagulation, septic shock and refractory metabolic acidosis.81
Laboratory diagnosis
Specimen types, processing and transport
Adherence to sufficient standard working procedures (SOPs) is pivotal during the collection, assortment, packaging, storage and transportation of samples. The clinical physician should have the knowledge and practice the preventive extra precautionary measures during sample treatment. All examinations of clinical samples collected from the suspected individual should be performed in equipped laboratories instruments with specialised laboratory staff.82
Common rules on biosafety in the research facility should be carefully followed, and all the methodology that is to be attempted are based on a risk evaluation. Biosafety level 3 facilities are the minimum requirement to perform culturing of viral samples. The throat swab is the primary test that should be taken to examine a suspected individual. These materials include superior respiratory tract samples such as oropharyngeal and nasopharyngeal swab and inferior respiratory samples such as sputum and endotracheal or bronchoalveolar lavage suction in individuals showcasing severe respiratory ailments.82 As there has been evidence of SARS- CoV-2 in blood and stool these can be also collected as clinical samples to examine any suspected individual.83
After samples are packed in an appropriate container, it is transported to the laboratory expeditiously for the analysis of SARS-CoV-2. Viral samples are transported under the cold chain temperature of 2-8°C. In the instance of delay in transport to the laboratory, viral transport mediums are used such as solidification of samples at -20°C or preferably -70°C and are sent on a dry ice pack. It is essential to maintain a strategic distance for the process of defrosting/thawing and freezing of samples. Transportation within the national borders should follow relevant principles and guidelines. In the event of global transportation of samples, the United Nations model regulations should be tracked with different guidelines for the method of transport.84,85
Direct diagnosis
From several viral infection studies, the common analysis of SARS-CoV-2 is depending on the immediate identification of viral RNA (from nasopharyngeal swabs or on inferior respiratory specimens), antigens or the indirect recognizable specific antibody responses. The gold standard techniques are the main direct analysis method used for an active viral infection, while the identification of specified SARS-CoV-2 antibodies is the foundation for the past contact with the virus infection, both for analytic and epidemiological points. In clinical practice, the commonly utilized method is the gold standard method, while tests on inferior respiratory regions are performed based on different types of cases.82 In individuals with a decent result, viral RNA is recognized for 20 days or longer after the beginning of manifestations, and a bounce-back of the viral burden, after imperceptible with PCR, is possible.86
Real-time polymerase chain reaction (rt-PCR)
rt-PCR is one of the commonly utilized techniques for the detection of the SARS-CoV-2 genome where the primers act against the E protein (first-line screening) and RdRp regions (confirmation testing).87 Moreover, a one-step rt-PCR diagnosis has also been developed that recognises the viral N protein (first-line screening) and ORF1b (confirmatory testing).88 This rt-PCR can be conducted in two different methods which are multiplexed and single format tests. In the singleplex technique, identification of an individual’s RNase P (RP) primer is performed by utilizing each one of the probes and a primer set whereas the identification of the N gene in an individual is performed at three regions by using triple primers and a probe. The multiplexed format, N gene of SARS-CoV-2 is identified by using two sets of probe and primer and for the identification of the RP region in SARS-CoV-2, a single set of probe and primer is utilized. The sample collected from the superior and inferior regions of the respiratory system, isolation of RNA is performed which then undergoes reverse transcription to form cDNA.87 The differences in the sequences of a viral RNA are majorly affected due to the utilization of primers in various types of genes detected in SARS-CoV-2. From multiple studies conducted by different researchers, they indicated that drastic evolution and diversity of the gene results in false negatives in rt-PCR tests. Even when precise primers and probe sets are designed accordingly for rt-PCR assays, these false-negative results may be exhibited due to the rapid mutation of the SAR-CoV-2 strains that have to lead to misidentification among the primers and probe and the sequences, thus affecting the quality of the rt-PCR assay. Therefore, many different types of target genes should be amplified to avoid unworthy results.89,90
Reverse-Transcription-Loop-Mediated isothermal Amplification (RT-LAMP)
In modern years, the RT-LAMP technique has been used worldwide in the identification of the specific gene of viruses that involves the amplification of nucleic acid of a sequence at standard temperature. This is performed in a highly specific and sensitive assay compared to the rt-PCR technique. LAMP’s technique is mainly based on the formation of target DNA at a standard temperature of 60–65°C and by using the specialized designated DNA polymerase and primers, which is by emphasizing the strand DNA activity compared to the heat denaturation process in different PCR techniques.91 In an hour duration of time, more than 109 copies of sequences can be amplified that builds cauliflower moulded structures that similarly compromised DNA with numerous inverted repeats. LAMP is easy to use a procedure that gives sensitive, specific and reliable outcomes in less time when contrasted with other regular strategies.92 The combination method of LAMP along with reverse transcription provides a valuable discovery of the COVID-19 RNA strands. The advantages of this method are, it requires very minimal turnaround time and provides high sensitivity results.93
Antigen-Antibody based test
In this technique, the identification of COVID-19 antibodies is performed by utilizing enzyme-linked immunoassay known as a sandwich method whereby the double-antibody assay used, recognises NP by a microplate which is pre-filled with specific antibodies against the N protein and horseradish peroxidase (HRP)- marked secondary antibody against a similar protein. This immediate strategy is basic, quick, and does not require any special training, however a costly research facility instrument.94,95
Indirect diagnosis
Identification of specific IgG or IgM antibodies of COVID-19 infection can be also performed using indirect method. Moreover, WHO had emphasized the need for serological IgG and IgM testing which should be designed rapidly that ease a physician to perform the test under a low-cost budget and minimal cost range. Few studies performed by different authors have assessed that samples obtained from posterior nasopharyngeal demonstrated similar viral and antibody kinetics serological testing which is complementary to the rt-PCR test; this is because the responses of IgG and IgM towards the internal viral NP and S protein receptor-binding domain (RBD) started between 10 days after initial symptoms which were positively linked together in the neutralized antibodies titres; whereby the antibodies opposing RBD demonstrated an early increase comparing to other against NP, an increase in both the serological test was observed simultaneously. Moreover, the study also has demonstrated a longer duration of rt-PCR positivity in individuals who are experiencing a lack of prognosis and individuals who develop early antibodies.95
Enzyme-linked Immunoassay (ELISA)
A technique that recognizes the viral IgG and IgM reactions by recognising the antibodies against the S protein and NP, yet lacking characterized guidelines causes a hindrance. A meta-analysis was conducted in Brazil, to examine the accuracy of available diagnostic techniques to identify SARS-CoV-2 infection, and it was found that the sensitivity and particularity for IgG were 97% and 98%, respectively, and for the IgM antibodies, the sensitivity and specificity were 82% and 97%, respectively.94,95
Chemiluminescent Immunoassay (CLIA)
CLIA provides advanced critical preferences to overcome the traditional assay detection techniques, particularly in quantifying specific antibodies. When a substrate is added together with the Avidin-Horseradish Peroxidase chemical this results in a chemical reaction that produces light reactions. Thus, it helps in the assessment of the IgG and IgM titers by measuring the amount of absorbent light emitted out as a signal. In a study conducted by Padoan, the energy product which is kinetic energy between IgM and IgG antibodies demonstrated a quick increment in IgG and IgM between 6–7 days from initial symptoms. There was an absence of IgG and IgM in patients from day 1 to day 5 and out of the six patients being examined between day 6 and 7, three were identified as positive for IgM and four patients for IgG; between day 12 and 13 from the nine patients under observation, seven were positive for IgM and IgG were detected 100% in all the patients.95-96
Computed tomography (CT) scan
CT scan has high sensitivity and because of which many researchers suggest its utilization as one of the essential auxiliary demonstrative techniques for SARS-CoV-2. Furthermore, the results are obtained even before clinical side effects show up. There are some common features of COVID-19 patients that are observed in CT scans such as bilateral multilobar ground-glass opacification with diversely distributed in posteriors and peripheral,97 thickened lobular septa with alveolar filling, sub-pleura ascendance and amalgamation.98 Results with higher sensitivity and accuracy can be obtained by utilizing the combination of both gold standards diagnostic tests such as RT-qPCR and CT-scan in future. In addition, the high-resolution CT images of the chest are additionally demonstrated as a basic apparatus for recognition of SARS-CoV-2 which could be done during the initial phase of the infection to undertake rapid therapeutic measures.99 Prior likewise the ordinary images of the CT from the patients infected with SARS-CoV and MERS-CoV indicated comparable signs as in SARS-CoV-2.99
Animal models for COVID-19 research
Animal models are a vital study tool that helps researchers in gaining new insights for both pathogenesis and pre-clinical evaluations of antiviral drugs and vaccines against pathogenic diseases. The pathogenies of COVID-19 are mainly targeted to ACE2 and immunopathology. Hence, an effective animal model is an absolute necessity to understand its further mechanism. To date, COVID-19 has been observed in several animal models which include mouse, hamster, ferret, bat, macaque and cat.100,101 Laboratory mice are one of the most preferred animal models by immunologists and infectious disease researchers, however, due to the differences in human and murine ACE2 and TMPRSS2, mice are observed to be poorly infected with SARS-CoV-2. Transgenic mouse models that can express human ACE2 are available; nonetheless, most of the models still shows mild symptoms and limited infectibility, except for the K18-hACE2 transgenic mouse. This type of transgenic mouse was observed to develop several severe COVID-19 characteristics but lacks the expression of human FcgRT. Humanised FcgRT facilitates finer modelling of human antibody clearance in mice which will be useful during antibody-drug development.102
Scientific and technological advancements to combat COVID-19
Computational approach in drug discovery
At the current pandemic situation, de novo drug design is least to be considered as a therapeutic option as it is a tremendously time-consuming method. Alternatively, drug repositioning works as a preferred method of choice in drug discovery that establishes new therapeutic applications from existing drugs with known safety profiles and pharmacokinetics, thus decreasing the drug development duration and risk of adverse side effects. In recent years, there has been a great number of computational studies employed that predicts novel drug targets or drug repurposing candidates (synthetic and natural occurring). In silico approaches enables researchers to filter thousands of compounds at a lower cost and with a shorter duration spent in obtaining results, making it a vital tool in prioritizing potential drug candidates for further experimental validation.104
There are two approaches in computational studies to identify promising repurposable drugs and targets for COVID-19, namely virus-targeting and host-targeting approaches. In viral targeting approaches, it involves structure-based and deep-learning (DL) based drug screening methodologies which uses the viral target protein’s three-dimensional structure in predicting the binding affinity values, when bound to a known ligand. On the other hand, host-targeting approaches involve signature and network-based methodologies in identifying promising drugs candidates which impede the host mechanism that contributes to the viral pathogenesis. For instance, one of the clinical characteristics of COVID-19 is cytokine storm that is due to the excess release of pro-inflammatory cytokines and chemokines. Therefore, drugs that target specific dysregulated pathways may be beneficial in treating severe COVID-19 patients.105
Nanotechnology
Over the years, nanoparticles have shown greater importance in research due to their unique features such as being smaller in size, multifunctional and improved solubility, which facilitates the development of safer drugs/vaccines, early diagnostic devices. Various approaches in nanotechnology can be used in combating SARS-CoV-2 such as by designing and developing nano-based sensors, nano-antiviral drugs and nano-based vaccines.
Table (2):
Animal models for SARS-CoV-2 research100,103.
Animal species |
Scientific name |
Natural infection susceptibility |
Experimental infection susceptibility |
Immune Response |
Clinical signs |
Suitable study |
|---|---|---|---|---|---|---|
Mouse |
Mus musculus |
High |
High |
Yes |
Yes |
Pathogenesis, immune response and vaccine development study |
Hamster |
Mesocricetus auratus |
None |
High |
Yes |
Yes |
Transmission, pathogenesis, immune response and therapeutic study. |
Ferret |
Mustela putorius furo |
None |
High |
Yes |
Yes |
Transmission, pathogenesis and therapeutic study |
Bat |
Rousettus aegyptiacus |
High |
High |
Yes |
No |
Transmission study |
Monkey |
Chlorocebus aethiops |
None |
High |
Yes |
Yes |
Transmission, pathogenesis, immune response and vaccine development study |
Mink |
Neovison vison |
High |
Not done |
Yes |
Yes |
Transmission, pathogenesis and therapeutic study |
Various nanomaterials namely Metallica, polymeric, silica nanoparticles, quantum dots and carbon nanotubes have shown effective application on viral detection. The diagnostic system is developed by modifying the surface of the nanoparticle with biomolecules that are extracted from the virus, for example, viral RNA. Nanomaterials with high surface and volume ratios enhance the interactions between the sensor and the analyte which leads to the increase in detection limit and decrease in detection duration. In this framework, probes that use nanomaterials are widely utilised for the production of biosensors. Furthermore, nanotechnology-based biosensors provide better sensor’s responses by obtaining optical, electrical and catalytical properties, resulting in substantial analytical sensitivity. Among many nanoparticles for viral detection, gold nanoparticles (AuNPs) are highly preferred due to their electrical, photonic and catalytic characteristics106. From a study performed by Kim et al. to design a colourimetric assay to detect MERS-CoV, gold nanoparticles were functionalised with thiol modified probes, which hybridise with the target, inhibiting the aggregation of AuNPs by salt with subsequent colour change. This concept of colourimetric assay can be indisputably applied and adjusted as a diagnostic tool for other diseases, including COVID-19.107
Table (3):
Potential drugs that affect SARS-CoV-2 activity.
No. |
Drug (s) |
Class (chemical or pharmacological) |
Mechanism of action |
References |
|---|---|---|---|---|
1 |
Camostat |
Serine protease inhibitor |
Blocks the virus-activating host cell protease TMPRSS2* |
120 |
2 |
Teicoplanin |
Antibiotic |
Inhibits the cathepsin L activity, blocking the virus to enter the host cell |
121 |
3 |
Umifenovir (arbidol) |
Antiviral |
Inhibits the viral envelope’s membrane fusion by interacting between viral S-proteins and ACE2** receptors |
122 |
4 |
Kaletra (Ritonavir + Lopinavir) |
Protease inhibitor |
Inhibits 3Cl protease*** activity |
123 |
5 |
Remdesivir |
Antiviral |
Affects viral RdRp**** activity, inhibiting the formation of negative-sense RNA strands. Binds to Mpro***** |
124 |
6 |
Dexamethasone |
Corticosteroid |
Decreases gene transcription of various pro-inflammatory cytokines, chemokines and activates trans-expression |
125
|
7 |
Sevoflurane |
Anaesthetic agent |
Improves oxygenation in patients under ventilator |
126 |
* Transmembrane protease serine 2
** Angiotensin-converting enzyme 2
*** 3-chymotrypsin-like cysteine protease
**** RNA-dependent RNA polymerase
***** Main protease
An effective design of nanoparticle-antiviral drug combination is achievable and functions by interfering with the viral action through several approaches, such as by obstructing the viral entry into the host cells, interact with the viral particles and controlled release of antiviral agent for prolonged duration.108 Loczechin et al. performed a study to evaluate the antiviral activity of carbon quantum dots on HCoV-229E infection. In this study, the functional group of boronic acid was chemically integrated onto the carbon quantum dots and the interaction with the virus and its S protein was observed. Based on the results, the functional group on the carbon quantum dots were seen to obstruct viral entry by interacting with the S protein and entry receptors. Moreover, the carbon quantum dots were observed to inhibit the viral replication process, depicting the beneficial application of carbon quantum dots in the inhibition of the virus at different stages of infection.109
Table (4):
TCM derived compounds or extracts with anti-human coronavirus activity.
No. |
TCM compound (s) |
Mechanism of action |
References |
|---|---|---|---|
1 |
Quercetin and TSL-1 from Toona sinensis Roem |
Inhibits the entry of SARS-CoV into the host cell |
130
|
2 |
Emodin obtained from the genus Rheum and Polygonum |
Inhibits the SARS-CoV S protein interaction with ACE2 receptor |
127 |
3 |
Saikosaponins |
Prevents the early stages of human coronavirus 229E infection, causing inhibitory effects on viral attachment and penetration into the host’s cell |
128-129 |
4 |
Phenolic compounds derived from plants and root extract of Isatis indigotica |
Inhibits the SARS-3CLpro cleavage activity |
131 |
5 |
Houttuynia cordata water extract |
Inhibits SARS-3CLpro and RdRp’s activity, immunomodulation |
132 |
6 |
Flavonoids derived from rhoifolin, litchi seeds and quercetin |
Inhibits SARS-3CLpro activity |
133 |
7 |
Myricetin and scutellarein |
Affect the ATPase activity, Inhibiting SARS-CoV helicase protein |
134 |
In recent years, nanotechnology is increasingly used in the development of new generation vaccines, since the nanoparticle functions as an antigen carrier and also acts as an adjuvant.110 A wide range of nanoparticles can be used to construct nano-based vaccines which include peptide self-assembled virus-like particles (VLPs), organic-based nanoparticles (e.g. lipid nanoparticles) and inorganic based nanoparticles (e.g. AuNPs). Nano-based vaccines can prevent premature degradation of antigens and provide better antigen stability with controlled release, leading to the increase in the duration of the antigen exposure in the system, facilitating uptake by APCs. In the case of SARS-CoV-2, several mRNA-based nano-vaccines developed by Moderna and Pfizer, respectively, use lipid-based nanoparticles that function as nano carries to protect it from premature degradation by extracellular RNases and to provide efficient delivery of the mRNA to the immune cells. In addition, due to the controlled release of mRNA from the lipid-based nanoparticles, it causes constant protein translation, resulting in increased production of antibody titers with better B and T cell responses.111,112
Bacterial artificial chromosomes
The use of reverse genetics methodologies in generating recombinant SARS-CoV-2 are vital tools in understanding the deeper concepts of SARS-CoV-2 pathogenesis, leading to novel strategies to combat the virus. Furthermore, reverse genetics promotes the generation of reporter genes from recombinant viruses that are applied in in vivo models of infection and cell-based screening assays for prompt identification of therapeutic and preventive measures.113 Bacterial artificial chromosomes (BACs) are large genomic insert clones that are maintained as a single copy in Escherichia coli and are said to reverse the genetics system for large RNA viruses, including CoVs.114 Several studies have shown successful cloning of viral genomes and generation of reverse genetics systems, nevertheless, due to the toxicity and instability within the viral genome, the assembly of full-length complementary DNA of the virus in E. coli is being technically challenged. Two recent studies have shown to overcome the problem and able to generate the full-length SARS-CoV-2 genome through in vitro ligation and homologous recombination in Saccharomyces cerevisiae. However, in both of the studies in vitro transcription, a method that poses technical difficulties is required for the production of the full-length viral genome RNA. From another study by Ye et al., they were able to develop a reliable and convenient recombinant SARS-CoV-2 based on the use of a single BAC. Furthermore, rSAR-CoV-2 and SARS-CoV2 isolate from the study showed similar levels of fitness in Vero E6 cells and replication is validated golden Syrian hamster model.113
Treatment for SARS-CoV-2
Repurposing drugs
Currently, there are no specific antiviral drugs for the treatment of SARS-CoV-2.115 The COVID-19 pandemic has led to many clinical trials and ongoing trials where researchers evaluate a wide range of drugs and investigational agents for their efficiency as a therapeutic agent against the virus.116 The clinical trials that are performed and ongoing are based on the viral life cycle, leading to multiple targets for potential therapeutic intervention and the use of combination drugs that affect more than one target is seen to provide better results.117,118 Through molecular docking analysis on selected repurposed drugs against multiple proteins from SARS-CoV-2 namely S protein, 3Clpro, nsp10/16, nsp 9 and viral helicase showed that ritonavir, lopinavir and remdesivir give high binding efficiency to the respective viral protein targets. Based on the results obtained, the author concluded that ritonavir, lopinavir and remdesivir are to be potential therapeutic candidates against SARS-CoV-2.119
Table (5):
Various SARS-CoV-2 vaccine candidates that are in the development stage, manufactured by several companies137-138.
No. |
Company |
Potential vaccine Candidate |
Type of vaccine |
Mechanism of action |
|---|---|---|---|---|
1 |
University of Oxford |
Chimpanzee adenovirus vaccine vector |
Viral vectored vaccine |
Triggers predominant CTL* response |
2 |
Sinovac |
Formalin‐inactivated and alum‐adjuvanted candidate vaccine |
Inactivated vaccine |
Induces protective B cell response, however, requires booster dose. |
3 |
Moderna |
mRNA‐1273 |
mRNA based vaccine |
Activate predominant CTL response |
4 |
Inovio |
INO‐4800 |
DNA based vaccine |
Induces predominant CTL response |
5 |
Novavax |
NVX-CoV2373 |
Subunit vaccine (Recombinant nanoparticle vaccine) |
With the aid of S1 or RBD** subunit, triggers a humoral response |
* Cytotoxic T lymphocytes
** Receptor binding domain
Chinese medicinal treatment
The use of traditional Chinese medicine (TCM) aids in treating SARS-CoV-2 where the application was built from the previous treatment knowledge on SARS-CoV outbreak that occurred in late 2002 at Guangdong Province of China. From several studies, case reports and case series that explores the efficiency of TCM on SARS-CoV showed results that support the beneficial effect of TCM in the treatment and prevention of SARS-CoV.127 According to Lau et al, during the SARS-CoV outbreak, 1063 volunteers from high-risk virus laboratories were administered with TCM herbal extract, namely Sang Ju Yin plus Yu Ping Feng San. From the results obtained, to check the efficiency of the TCM herbal extract on SARS-CoV, it showed that none of the volunteers that consumed the herbal extract contracted the infection compared to 0.4% among the non-consumers (control group).128 Another study performed by Chen and Nakamura showed statistical evidence for the effectiveness of TCM as a line of treatment for SARS-CoV. From the study, the fatality rate of Singapore and Hong Kong was approximately 18% compared to Beijing with 52% until May 2003. However, a gradual decrease in Beijing’s mortality rate of 4%-1% was observed from May 2003 onwards and this effect was due to the use of TCM as conventional therapy.129
Vaccine
The global pandemic has caused a race in several countries to come up with the SARS-CoV-2 vaccine. Since the virus is highly similar to SAS-CoV features, the development and production of the SARS-CoV-2 vaccine are at a faster rate than ever before.135,136 Among various protein targets in SARS-CoV-2, the S protein is seen as the major target for the development of the vaccine, since the antibodies that affect S protein, (specifically at the RBD domain and other regions of the protein) neutralises the virus. Most of the potential vaccine candidates associate with the S antigen by viral vectored vaccines, inactivated vaccines, nucleic acid-based DNA/mRNA vaccines or by subunit vaccines.137,138
In conclusion, SARS-CoV-2 is a highly infectious CoV that causes dry cough, fever and fatigue as initial symptoms and leads to extreme pneumonia, ARDS, extrapulmonary signs and systemic confusions or death if left untreated. The origin of this virus is from bats with Manis javanica being the plausible intermediate host. The airborne virus can transmit between individuals through droplets that include sneezing and coughing and also by contact with contaminated objects. More studies are required on the pathogenic mechanism of SARS-CoV-2 in the host body to identify vulnerable targets for therapeutic intervention. With the aid of computational studies, it will facilitate the drug/vaccine design and development that affects the specific targeted viral protein resulting in decreased mortality and morbidity of SARS-CoV-2 from the human population.
ACKNOWLEDGMENTS
The author would like to thank Ms Christina Anthonette Nivetha Sethu Raman Vairavan for her support.
CONFLICT OF INTEREST
The authors declare that there is no conflicts of interest.
AUTHORS’ CONTRIBUTION
CASV designed the format for the literature review and edited the manuscript. CASV and DR analysed various texts and drafted the manuscript under the supervision of NS and GS. All the authors reviewed and approved the final version of the manuscript.
FUNDING
None.
ETHICS STATEMENT
Not applicable.
AVAILABILITY OF DATA
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
- Decaro N, Lorusso A. Novel human coronavirus (SARS-CoV-2): A lesson from animal coronaviruses. Vet Microbiol. 2020;244:108693.
Crossref - Neerukonda S, Katneni U. A Review on SARS-CoV-2 Virology, Pathophysiology, Animal Models, and Anti-Viral Interventions. Pathogens. 2020;9(6):426.
Crossref - Wassenaar TM, Zou Y. 2019_nCoV/SARS-CoV-2: rapid classification of beta coronaviruses and identification of Traditional Chinese Medicine as potential origin of zoonotic coronaviruses. Lett Appl Microbiol. 2020;70(5):342-348.
Crossref - Zhu Z, Lian X, Su X, Wu W, Marraro GA, Zeng Y. From SARS and MERS to COVID-19: a brief summary and comparison of severe acute respiratory infections caused by three highly pathogenic human coronaviruses. Respir Res. 2020;21(1):224.
Crossref - Zhou P, Yang X-L, Wang X-G, et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020;579(7798):270-273.
Crossref - Lam TT-L, Jia N, Zhang Y-W, et al. Identifying SARS-CoV-2-related coronaviruses in Malayan pangolins. Nature. 2020;583(7815):282-285.
Crossref - Adhikari SP, Meng S, Wu Y-J, et al. Epidemiology, causes, clinical manifestation and diagnosis, prevention and control of coronavirus disease (COVID-19) during the early outbreak period: a scoping review. Infect Dis Poverty. 2020;9(1):29.
Crossref - Callaway E, Cyranoski D, Mallapaty S, Stoye E, Tollefson J. The coronavirus pandemic in five powerful charts. Nature. 2020;579(7800):482-483.
Crossref - Mizumoto K, Kagaya K, Chowell G. Effect of a wet market on coronavirus disease (COVID-19) transmission dynamics in China, 2019-2020. Int J Infect Dis. 2020;97:96-101.
Crossref - Sun K, Chen J, Viboud C. Early epidemiological analysis of the coronavirus disease 2019 outbreak based on crowdsourced data: a population-level observational study. The Lancet Digital Health. 2020;2(4):e201-e208.
Crossref - Coronavirus Disease (COVID-19) – events as they happen. Who.int. https://www.who.int/emergencies/diseases/novel-coronavirus-2019/events-as-they-happen. 2020. Accessed October 27, 2020.
- Weekly epidemiological update – 17 November 2020. Who.int. https://www.who.int/publications/m/item/weekly-epidemiological-update—17-november-2020. 2020. Accessed November 22, 2020.
- [Breaking] 3 coronavirus cases confirmed in Johor Baru. New Straits Times. 2020. https://www.nst.com.my/news/nation/2020/01/559563/breaking-3-coronavirus-cases-confirmed-joh or-baru. Accessed October 27, 2020.
- First coronavirus cases in Malaysia: 3 Chinese nationals confirmed infected, quarantined in Sungai Buloh Hospital. Borneo Post Online. 2020. https://www.theborneopost.com/2020/01/25/first-coronavirus-cases-in-malaysia-3-chinese-nationals -confirmed-infected-quarantined-in-sungai-buloh-hospital/. Accessed October 27, 2020.
- First case of Malaysian positive for coronavirus. BERNAMA. https://www.bernama.com/en/general/news_covid-19.php?id=1811373. 2020. Accessed October 27, 2020.
- Elengoe A. COVID-19 Outbreak in Malaysia. Osong Public Health and Research Perspectives, 2020;11(3):93-100.
Crossref - COVID-19 Malaysia Updates. COVID-19 Malaysia. http://covid-19.moh.gov.my/terkini. 2020. Accessed October 27, 2020.
- Jakarta Globe. https://jakartaglobe.id/news/thirteen-indonesians-contract-covid19-at-malaysias-tabligh-islamic-gathering. Accessed October 27, 2020.
- Lin H, Liu W, Gao H, Nie J, Fan Q. Trends in Transmissibility of 2019 Novel Coronavirus-Infected Pneumonia in Wuhan and 29 Provinces in China. SSRN Electronic Journal. 2020.
Crossref - Jarvis M. Aerosol Transmission of SARS-CoV-2: Physical Principles and Implications. Front Public Health. 2020;8.
Crossref - Mediawati AS, Susanto R, Nurahmah. The Routes Of Covid-19 Transmission: A Literature Review. J Crit Rev. 2020;7(06):722-724.
Crossref - Baker MG. Nonrelocatable Occupations at Increased Risk During Pandemics: United States, 2018. Am J Public Health. 2020;110(8):1126-1132.
Crossref - Zhang Z, Liu S, Xiang M, et al. Protecting healthcare personnel from 2019-nCoV infection risks: lessons and suggestions. Front Med. 2020;14(2):229-231.
Crossref - COVID-19 clusters and outbreaks in occupational settings in the EU/EEA and the UK. European Centre for Disease Prevention and Control. 2020. https://www.ecdc.europa.eu/en/publications-data/covid-19-clusters-and-outbreaks-occupational-settings-eueea-and-uk. Accessed October 20, 2020.
- Zhao J, Yang Y, Huang H, et al. Relationship between the ABO Blood Group and the COVID-19 Susceptibility. medRxiv. 2020.
Crossref - The Severe Covid-19 GWAS Group. Genomewide Association Study of Severe Covid-19 with Respiratory Failure. N Eng J Med. 2020;383(16):1522-1534.
Crossref - Hunt K, Jacqueline HC. Blood type could be linked to Covid-19 risk and severity, new research suggests. CNN. 2020. https://edition.cnn.com/2020/10/14/health/blood-group-covid-19-scn-wellness/index.html. Accessed October 20, 2020.
- Mousavizadeh L, Ghasemi S. Genotype and phenotype of COVID-19: Their roles in pathogenesis. J Microbiol Immunol Infect. 2021;54(2):159-163.
Crossref - Ma Y, Wu L, Shaw N, et al. Structural basis and functional analysis of the SARS coronavirus nsp14-nsp10 complex. Proc Natl Acad Sci U S A. 2015; 112(30):9436-9441.
Crossref - Kim J-S, Jang J-H, Kim J-M, Chung Y-S, Yoo C-K, Han M-G. Genome-Wide Identification and Characterization of Point Mutations in the SARS-CoV-2 Genome. Osong Public Health Res Perspect. 2020;11(3):101-111.
Crossref - Walls A, Park Y, Tortorici M, Wall A, McGuire AT, Veesler D. Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein. Cell. 2020;181(2):281-292.e6.
Crossref - Schoeman D, Fielding BC. Coronavirus envelope protein: current knowledge. Virol J. 2019;16(1):69.
Crossref - Chan J-F-W, Kok K-H, Zhu Z et al. Genomic characterization of the 2019 novel human-pathogenic coronavirus isolated from a patient with atypical pneumonia after visiting Wuhan. Emerg Microbes Infect. 2020;9(1):221-236.
Crossref - Duffy S. Why are RNA virus mutation rates so damn high? PLoS Biol. 2018;16(8):e3000003.
Crossref - Wang R, Hozumi Y, Yin C, Wei G-W. Decoding SARS-CoV-2 Transmission and Evolution and Ramifications for COVID-19 Diagnosis, Vaccine, and Medicine. J Chem Inf Model. 2020;60(12):5853-5865.
Crossref - Khan M, Khan Z, Baig MH, et al. Comparative genome analysis of novel coronavirus (SARS-CoV-2) from different geographical locations and the effect of mutations on major target proteins: An in silico insight. PLoS One. 2020;15(9):e0238344.
Crossref - Chen J, Wang R, Wang M, Wei G-W. Mutations Strengthened SARS-CoV-2 Infectivity. J Mol Biol. 2020;432(19):5212-5226.
Crossref - Teng C. Mutated strain more infectious. The Star Online. 2020. https://www.thestar.com.my/news/nation/2020/10/03/mutated-strain-more-infectious. Accessed October 22, 2020.
- Eaaswarkhanth M, Al Madhoun A, Al-Mulla F. Could the D614G substitution in the SARS-CoV-2 spike (S) protein be associated with higher COVID-19 mortality? Int J Infect Dis. 2020;96:459-460.
Crossref - Korber B, Fischer WM, Gnanakaran S, et al. Tracking changes in SARS-CoV-2 Spike: Evidence that D614G increases infectivity of the COVID-19 virus. Cell. 2020;182(4):812-827.e19.
Crossref - Qian Z, Travanty EA, Oko L, et al. Innate immune response of human alveolar type II cells infected with severe acute respiratory syndrome-coronavirus. Am J Respir Cell Mol Biol. 2013;48(6):742-748.
Crossref - Xu J, Xu X, Jiang L, Dua K, Hansbro PM, Liu G. SARS-CoV-2 induces transcriptional signatures in human lung epithelial cells that promote lung fibrosis. Respir Res. 2020;21(1):182.
Crossref - Fehr AR, Perlman S. Coronaviruses: An Overview of Their Replication and Pathogenesis. Coronaviruses. 2015;1282:1-23.
Crossref - Celigoy J, McReynolds S, Caffrey M. The SARS-CoV heptad repeat 2 exhibits pH-induced helix formation. Biochem Biophys Res Commun. 2011;412(3):483-486.
Crossref - Mirzaei R, Karampoor S, Sholeh M, Moradi P, Ranjbar R, Ghasemi F. A contemporary review on pathogenesis and immunity of COVID-19 infection. Mol Biol Rep. 2020;47(7):5365-5376.
Crossref - Baranov P, Henderson C, Anderson C, Gesteland RF, Atkins JF, Howard MT. Programmed ribosomal frameshifting in decoding the SARS-CoV genome. Virology. 2005;332(2):498-510.
Crossref - Harcourt B, Jukneliene D, Kanjanahaluethai A, et al. Identification of Severe Acute Respiratory Syndrome Coronavirus Replicase Products and Characterization of Papain-Like Protease Activity. J Virol. 2004;78(24):13600-13612.
Crossref - Barretto N, Jukneliene D, Ratia K, Chen Z, Mesecar AD, Baker SC. The Papain-Like Protease of Severe Acute Respiratory Syndrome Coronavirus Has Deubiquitinating Activity. J Virol. 2005;79(24):15189-15198.
Crossref - Wu C, Liu Y, Yang Y, et al. Analysis of therapeutic targets for SARS-CoV-2 and discovery of potential drugs by computational methods. Acta Pharm Sin B. 2020;10(5):766-788.
Crossref - Santerre M, Arjona S, Allen CN, Shcherbik N, Sawaya BE. Why do SARS-CoV-2 NSPs rush to the ER?. J Neurol. 2021;268:2013-2022.
Crossref - Sola I, Almazan F, Zuniga S, Enjuanes L. Continuous and Discontinuous RNA Synthesis in Coronaviruses. Annu Rev Virol. 2015;2(1):265-288.
Crossref - Astuti I, Ysrafil. Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2): An overview of viral structure and host response. Diabetes Metab Syndr. 2020;14(4):407-412.
Crossref - Artika I, Dewantari AK, Wiyatno A. Molecular biology of coronaviruses: current knowledge. Heliyon. 2020;6(8):e04743.
Crossref - Xu R, Shi M, Li J, Song P, Li N. Construction of SARS-CoV-2 Virus-Like Particles by Mammalian Expression System. Front Bioeng Biotechnol. 2020;8:862.
Crossref - Siu YL, Teoh KT, Lo J, et al. The M, E, and N Structural Proteins of the Severe Acute Respiratory Syndrome Coronavirus Are Required for Efficient Assembly, Trafficking, and Release of Virus-Like Particles. J Virol. 2008;82(22):11318-11330.
Crossref - Bhowmik D, Nandi R, Jagadeesan R, Kumar N, Prakash A, Kumara D. Identification of potential inhibitors against SARS-CoV-2 by targeting proteins responsible for envelope formation and virion assembly using docking based virtual screening, and pharmacokinetics approaches. Infection, Genetics and Evolution. 2020;84:104451.
Crossref - Zhu N, Wang W, Liu Z, et al. Morphogenesis and cytopathic effect of SARS-CoV-2 infection in human airway epithelial cells. Nat Commun. 2020;11(1):3910.
Crossref - Fung TS, Liu D. Coronavirus infection, ER stress, apoptosis and innate immunity. Front Microbiol. 2014;5:296.
Crossref - Rabi FA, Al Zoubi MS, Kasasbeh GA, Salameh DM, Al-Nasser AD. SARS-CoV-2 and Coronavirus Disease 2019: What We Know So Far. Pathogens. 2020;9(3):231.
Crossref - Li G, Fan Y, Lai Y, et al. Coronavirus infections and immune responses. J Med Virol. 2020;92(4):424-432.
Crossref - Chang F-Y, Chen H-C, Chen P-J, et al. Immunologic aspects of characteristics, diagnosis, and treatment of coronavirus disease 2019 (COVID-19). J Biomed Sci. 2020;27(1):72.
Crossref - Kumar S, Nyodu R, Maurya VK, Saxena SK. Host Immune Response and Immunobiology of Human SARS-CoV-2 Infection. Medical Virology: From Pathogenesis to Disease Control. 2020;43-53.
Crossref - Mortaz E, Tabarsi P, Varahram M, et al. The Immune Response and Immunopathology of COVID-19. Front Immunol. 2020;11:2037.
Crossref - Law HKW, Cheung CY, Ng HY, et al. Chemokine up-regulation in SARS-coronavirus-infected, monocyte-derived human dendritic cells. Blood. 2005;106(7):2366-2374.
Crossref - Kim TS, Sun J, Legge KL, Braciale TJ. T cell responses during acute respiratory virus infection. Encyclopedia of Immunobiology. Elsevier. 2016;4:324-331.
Crossref - Bouayad A. Innate immune evasion by SARS-CoV -2: Comparison with SARS-CoV. Rev Med Virol. 2020;30(6):1-9.
Crossref - Ivashkiv LB, Donlin LT. Regulation of type I interferon responses. Nat Rev Immunol. 2014;14(1):36-49.
Crossref - Wathelet MG, Orr M, Frieman MB, Baric RS. Severe acute respiratory syndrome coronavirus evades antiviral signaling: role of nsp1 and rational design of an attenuated strain. J Virol. 2007;81(21):11620-11633.
Crossref - Kindler E, Gil-Cruz C, Spanier J, et al. Early endonuclease-mediated evasion of RNA sensing ensures efficient coronavirus replication. PLoS Pathog. 2017;13(2):e1006195.
Crossref - Zheng M, Gao Y, Wang G, et al. Functional exhaustion of antiviral lymphocytes in COVID-19 patients. Cell Mol Immunol. 2020;17(5):533-535.
Crossref - Lauterbach N, Wieten L, Popeijus HE, Voorter CEM, Tilanus MGJ. HLA-E regulates NKG2C+ natural killer cell function through presentation of a restricted peptide repertoire. Hum Immunol. 2015;76(8):578-586.
Crossref - Li J, Gong X, Wang Z, et al. Clinical features of familial clustering in patients infected with 2019 novel coronavirus in Wuhan, China. Virus Res. 2020;286:198043.
Crossref - Chen N, Zhou M, Dong X, et al. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet. 2020;395(10223):507-513.
Crossref - Wu Y-C, Chen C-S, Chan Y-J. The outbreak of COVID-19. J Chin Med Assoc. 2020;83(3):217-220.
Crossref - Kogan A, Segel MJ, Ram E, et al. Acute respiratory distress syndrome following cardiac surgery: Comparison of the American-European Consensus Conference definition versus the Berlin definition. Respiration. 2019;97(6):518-524.
Crossref - Wu Z, McGoogan JM. Characteristics of and Important Lessons From the Coronavirus Disease 2019 (COVID-19) Outbreak in China. The Journal of the American Medical Association. 2020;323(13):1239-1242.
Crossref - Kaur G, Lungarella G, Rahman I. SARS-CoV-2 COVID-19 susceptibility and lung inflammatory storm by smoking and vaping. J Inflamm (Lond). 2020;17(1):21.
Crossref - Abdelmaksoud A, Kroumpouzos G, Jafferany M, Lotti T, Sadoughifar R, Goldust M. COVID-19 in the pediatric population. Dermatol Ther. 2020;33(4):e13339.
Crossref - Zimmermann P, Curtis N. Coronavirus Infections in Children Including COVID-19 : An Overview of the Epidemiology, Clinical Features, Diagnosis, Treatment and Prevention Options in Children. Pediatr Infect Dis J. 2020;39(5):355-368.
Crossref - Zhang C, Gu J, Chen Q, et al. Clinical characteristics of 34 children with Coronavirus disease-2019 in the west of China: A multiple-center case series. bioRxiv. 2020
Crossref - Choi S-H, Kim HW, Kang J-M, Kim DY, Cho EY. Epidemiology and clinical features of coronavirus disease 2019 in children. Clin Exp Pediatr. 2020;63(4):125-132.
Crossref - Laboratory testing for 2019 novel coronavirus (2019-nCoV) in suspected human cases. 2020. https://www.who.int/publications/i/item/laboratory-testing-for-2019-novel-coronavirus-in-suspected-human-cases-20200117. Accessed October 22, 2020.
- Wu Y, Guo C, Tang L, et al. Prolonged presence of SARS-CoV-2 viral RNA in faecal samples. Lancet Gastroenterol Hepatol. 2020;5(5):434-435.
Crossref - Information for Laboratories about Coronavirus (COVID-19). Centers for Disease Control and Prevention. 2020. https://www.cdc.gov/coronavirus/2019-ncov/lab/guidelines-clinical-specimens.html. Accessed October 22, 2020.
- WHO | Guidelines for the safe transport of infectious substances and diagnostic specimens. 2020. https://www.who.int/csr/resources/publications/biosafety/WHO_EMC_97_3_EN/en/. 2020. Accessed October 22, 2020.
- To KK-W, Tsang OT-Y, Leung W-S, et al. Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: an observational cohort study. Lancet Infect Dis. 2020;20(5):565-574.
Crossref - Corman VM, Landt O, Kaiser M, et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Euro Surveill. 2020;25(3):2000045.
Crossref - Chu DKW, Pan Y, Cheng SMS, et al. Molecular Diagnosis of a Novel Coronavirus (2019-nCoV) Causing an Outbreak of Pneumonia. Clin Chem. 2020;66(4):549-555.
Crossref - Tahamtan A, Ardebili A. Real-time RT-PCR in COVID-19 detection: issues affecting the results. Expert Rev Mol Diagn. 2020;20(5):453-454.
Crossref - Phan T. Genetic diversity and evolution of SARS-CoV-2. Infect Genet Evol. 2020;81:104260.
Crossref - Nagamine K, Hase T, Notomi T. Accelerated reaction by loop-mediated isothermal amplification using loop primers. Mol Cell Probes. 2002;16(3):223-229.
Crossref - Galvez LC, Barbosa CFC, Koh RBL, Aquino VM. Loop-mediated isothermal amplification (LAMP) assays for the detection of abaca bunchy top virus and banana bunchy top virus in abaca. Crop Prot. 2020;131:105101.
Crossref - Huang WE, Lim B, Hsu C-C, et al. RT-LAMP for rapid diagnosis of coronavirus SARS-CoV-2. Microb Biotechnol. 2020;13(4):950-961.
Crossref - Castro R, Luz P, Wakimoto M, et al. COVID-19: a meta-analysis of diagnostic test accuracy of commercial assays registered in Brazil. Braz J Infect Dis. 2020;24(2):180-187.
Crossref - Russo A, Minichini C, Starace M, et al. Current Status of Laboratory Diagnosis for COVID-19: A Narrative Review. Infect Drug Resist. 2020;13:2657-2665.
Crossref - Padoan A, Cosma C, Sciacovelli L, Faggian D, Plebani M. Analytical performances of a chemiluminescence immunoassay for SARS-CoV-2 IgM/IgG and antibody kinetics. Clin Chem Lab Med. 2020;58(7):1081-1088.
Crossref - Pan Y, Guan H, Zhou S, et al. Initial CT findings and temporal changes in patients with the novel coronavirus pneumonia (2019-nCoV): a study of 63 patients in Wuhan, China. Eur Radiol. 2020;30(6):3306-3309.
Crossref - Shi H, Han X, Zheng C. Evolution of CT Manifestations in a Patient Recovered from 2019 Novel Coronavirus (2019-nCoV) Pneumonia in Wuhan, China. Radiology. 2020; 295(1):20.
Crossref - Chung M, Bernheim A, Mei X, et al. CT Imaging Features of 2019 Novel Coronavirus (2019-nCoV). Radiology. 2020;295(1):202-207.
Crossref - Parolin C, Virtuoso S, Giovanetti M, Angeletti S, Ciccozzi M, Borsetti A. Animal hosts and experimental models of sars-cov-2 infection. Chemotherapy. 2021;66(1-2):8-16.
Crossref - Jia W, Wang J, Sun B, Zhou J, Shi Y, Zhou Z. The mechanisms and animal models OF SARS-CoV-2 Infection. Front Cell Dev Biol. 2021;9:578825.
Crossref - Jarnagin K, Alvarez O, Shresta S, Webb DR. Animal models for sars-cov2/covid19 research-a commentary. Biochemical Pharmacology. 2021;188:114543.
Crossref - Munoz-Fontela C, Dowling WE, Funnell SG, et al. Animal models for covid-19. Nature. 2020;586(7830):509-515.
Crossref - Mohamed K, Yazdanpanah N, Saghazadeh A, Rezaei N. Computational drug discovery and repurposing for the treatment of COVID-19: A systematic review. Bioorg Chem. 2021;106:104490.
Crossref - Galindez G, Matschinske J, Rose T, et al. Lessons from the COVID-19 pandemic for advancing computational drug repurposing strategies. Nature Computational Science. 2021;1(1):33-41.
Crossref - Campos EV, Pereira AE, de Oliveira JL, et al. How can NANOTECHNOLOGY help to combat COVID-19? Opportunities and urgent need. J Nanobiotechnol. 2020;18(1):125.
Crossref - Kim H, Park M, Hwang J, et al. Development of Label-Free Colorimetric assay for mers-cov using gold nanoparticles. ACS Sensors. 2019;4(5):1306-1312.
Crossref - Abo-zeid Y, Urbanowicz RA, Thomson BJ, Irving WL, Tarr AW, Garnett MC. Enhanced nanoparticle uptake into virus infected cells: Could nanoparticles be useful in antiviral therapy? Int J Pharm. 2018;547(1-2):572-581.
Crossref - Loczechin A, Seron K, Barras A, et al. Functional carbon quantum dots as medical countermeasures to human coronavirus. ACS Appl Mater Interfaces. 2019;11(46):42964-42974.
Crossref - Zhu M, Wang R, Nie G. Applications of nanomaterials as vaccine adjuvants. Hum Vaccin Immunother. 2014;10(9):2761-2774.
Crossref - Li Y, Xiao Y, Chen Y, Huang K. Nano-based approaches in the development of antiviral agents and vaccines. Life Sciences. 2021;265:118761.
Crossref - Nanomedicine and the COVID-19 VACCINES. Nat Nanotechnol. 2020;15(12):963.
Crossref - Ye C, Chiem K, Park J-G, et al. Rescue of sars-cov-2 from a single bacterial artificial chromosome. mBio. 2020;11(5):168-220.
Crossref - Lai C. BAC Transgenesis: Cell-type specific expression in the NERVOUS System. Reference Module in Biomedical Sciences. 2016.
Crossref - Choudhary S, Malik Y, Tomar S. Identification of SARS-CoV-2 Cell Entry Inhibitors by Drug Repurposing Using in silico Structure-Based Virtual Screening Approach. Front Immunol. 2020;11:1664.
Crossref - Kalil AC. Treating COVID-19-Off-Label Drug Use, Compassionate Use, and Randomized Clinical Trials During Pandemics. The Journal of the American Medical Association. 2020;323(19):1897-1898.
Crossref - Al-Horani R, Kar S, Aliter K. Potential Anti-COVID-19 Therapeutics that Block the Early Stage of the Viral Life Cycle: Structures, Mechanisms, and Clinical Trials. Int J Mol Sci. 2020;21(15):5224.
Crossref - Cavasotto C, Di Filippo J. In silico Drug Repurposing for COVID-19: Targeting SARS-CoV-2 Proteins through Docking and Consensus Ranking. Mol Inform. 2021;40(1):2000115.
Crossref - Deshpande R, Tiwari A, Nyayanit N, Mdak M. In silico molecular docking analysis for repurposing therapeutics against multiple proteins from SARS-CoV-2. Eur J Pharmacol. 2020;886:173430.
Crossref - Bittmann S, Luchter E, Alieva EM, Villalon G, Weissenstein A. COVID 19: Camostat and The Role of Serine Protease Entry Inhibitor TMPRSS2. Journal of Regenerative Biology and Medicine. 2020.
Crossref - Baron SA, Devaux C, Colson P, Raoult D, Rolain J. Teicoplanin: an alternative drug for the treatment of COVID-19? Int J Antimicrob Agents. 2020;55(4):105944.
Crossref - Poduri R, Joshi G, Jagadeesh G. Drugs targeting various stages of the SARS-CoV-2 life cycle: Exploring promising drugs for the treatment of Covid-19. Cell Signal. 2020;74:109721.
Crossref - Yin W, Mao C, Luan X et al. Structural basis for inhibition of the RNA-dependent RNA polymerase from SARS-CoV-2 by remdesivir. Science 2020;368(6498):1499-1504.
Crossref - Ton A, Gentile F, Hsing M, Ban F, Cherkasov A. Rapid Identification of Potential Inhibitors of SARS-CoV-2 Main Protease by Deep Docking of 1.3 Billion Compounds. Mol Inform. 2020;39(8):2000028.
Crossref - Patel SK, Saikumar G, Rana J, et al. Dexamethasone: A boon for critically ill COVID-19 patients? Travel Med Infect Dis. 2020;37:101844.
Crossref - Orser BA, Wang D-S, Lu W-Y. Sedating ventilated COVID-19 patients with inhalational anesthetic drugs. EBioMedicine. 2020;55:102770.
Crossref - Ho TY, Wu S, Chen JC, Li CC, Hsiang CY. Emodin blocks the SARS coronavirus spike protein and angiotensin-converting enzyme 2 interaction. Antiviral Res. 2007;74(2):92-101.
Crossref - Cheng P-W, Ng L-T, Chiang L-C, Lin C-C. Antiviral effects of saikosaponins on human coronavirus 229E in vitro. Clin Exp Pharmacol Physiol. 2006;33(7):612-616.
Crossref - Bahbah E, Negida A, Nabet M. Purposing Saikosaponins for the treatment of COVID-19. Med Hypotheses. 2020;140:109782.
Crossref - Chen C-J, Michaelis M, Hsu H-K, et al. Toona sinensis Roem tender leaf extract inhibits SARS coronavirus replication. J Ethnopharmacol. 2008;120(1):108-111.
Crossref - Lin C-W, Tsai F-J, Tsai C-H, et al. Anti-SARS coronavirus 3C-like protease effects of Isatis indigotica root and plant-derived phenolic compounds. Antiviral Res. 2005;68(1):36-42.
Crossref - Lau K-M, Lee K-M, Koon C-M, et al. Immunomodulatory and anti-SARS activities of Houttuynia cordata. J Ethnopharmacol. 2008;118(1):79-85.
Crossref - Ng YL, Salim CK, Chu JJH. Flavonoids with inhibitory activity against SARS-CoV-2 3CLpro. J Enzyme Inhib Med Chem. 2020;35(1):1539-1544.
Crossref - Yu M-S, Lee J, Lee JM, et al. Identification of myricetin and scutellarein as novel chemical inhibitors of the SARS coronavirus helicase, nsP13. Bioorg Med Chem Lett. 2012;22(12):4049-4054.
Crossref - Chen W-H, Strych U, Hotez PJ, Bottazzi. The SARS-CoV-2 Vaccine Pipeline: an Overview. Curr Trop Med Rep. 2020;7(2):61-64.
Crossref - Krammer F. SARS-CoV-2 vaccines in development. Nature. 2020;586(7830):516-527.
Crossref - Wu S-C. Progress and Concept for COVID-19 Vaccine Development. Biotechnol J. 2020;15(6):2000147.
Crossref - Iacob S, Iacob D. SARS-CoV-2 Treatment Approaches: Numerous Options, No Certainty for a Versatile Virus. Front Pharmacol. 2020;11:1224.
Crossref
© The Author(s) 2021. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.