ISSN: 0973-7510
E-ISSN: 2581-690X
Recent studies have documented an increase in the incidence of antifungal resistance in newly emerging species closely related to C. albicans, and the coexistence of genotypic variants. Hence, an application of PCR-based molecular typing is crucial in identifying these fungi. Our study used molecular methods to characterize the latest genotypic subgroups of C. albicans and analysed if there was a relationship between the genotypes and the antifungal resistance pattern. The study was conducted in JSS Hospital, Mysuru, Karnataka between July 2018 and December 2020. A total of 1427 Candida species were isolated from clinical samples. Candida albicans were isolated and confirmed using Germ tube test, ID VITEK 2 and PCR (ITS primer). DNA extraction was done using the Hi-Media Yeast DNA Extraction Kit. The amplified products were analysed using Agarose gel electrophoresis (2%). Among 1427 Candida species, 282 were Candida albicans. The following resistance was exhibited to major antifungals – Caspofungin (3.5%), Amphotericin B (1.4%), flucytosine (2.8%) Fluconazole (6%) Micafungin (2.8%) Voriconazole (3.1%) and all were sensitive to miconazole. ABC genotyping showed Genotype A (450 bp) predominant (87.58%) followed by genotype B (840bp) (9.92 %) and genotype C (450bp and 840 bp) (0.2%). Genotype D and E were not observed. Our study showed the growing antifungal resistance in clinical isolates. Genotype A was predominant in South Karnataka region followed by Genotype B and C. There was no correlation between genotyping and antifungal resistance. However, a study with greater number of samples from diverse geographical locations may give more insight.
Antifungal susceptibility, clinical isolates, Candida albicans, Genotype A, Genotype B, Genotype C, Genotyping, Karnataka, South Indian region
Candida albicans, a ubiquitous commensal organism and the most prevalent and pathogenic Candida species, is responsible for majority of oral and systemic candidiasis,1-3 community-onset and nosocomial candidiasis.4-6 Candida albicans are responsible for the high morbidity and mortality rates affecting the immunocompromised and immunocompetent patients.3,5,7 This is due to an increase in antimicrobial resistance and the restricted number of antifungal drugs.8
C. albicans can either grow as a yeast or in filamentous forms and is capable of forming true hyphae.9,10 C. dubliniensis is a recently recognized species that is phenotypically and closely related to C. albicans and is susceptible to the same range of antifungal agents.11 Recently documented studies have shown an increase in the incidence of newly emerging species closely related to C. albicans like C dubliniensis, as well as the coexistence of genotypic variants and antifungal resistance during infections.11
Molecular typing of an infectious agent helps in epidemiological studies and in designing suitable infection control strategies.12 PCR-based molecular typing methods are crucial in identifying these fungi13,14 as they have a high discriminative power and are reproducible.15 Molecular typing methods such as genome sequencing, multi-locus sequence typing, pulsed field gel electrophoresis, restriction fragment length polymorphism, and randomly amplified polymorphic DNA analysis have been developed for the characterization and deeper comprehension of C. albicans.16-18 Several studies have identified the ABC genotypes of C. albicans from clinical specimens.19,20 This method was developed based on the presence or absence of a transposable group I intron in the gene which encodes the 25S ribosomal RNA.21 The use of molecular typing in the identification of C. albicans have provided vital information in supplementing the growing knowledge of yeast epidemiology. Ribosomal DNA is commonly used for PCR-based molecular methods for the species level identification of Candida. Ribosomal sequences have been widely used for genetic typing and identification of several fungal pathogens.22,23
In India, since no studies have been performed on genotyping of C. albicans, we characterized the clinical isolates of C. albicans based on ABC genotyping and analysed its relation to the antifungal resistance. Identifying the different genotypes of Candida albicans, helps to track its relatedness with the emerging genetic variants. It also helps us identify antifungal susceptibility and virulence between different genotypes.
Sample collection
Samples were collected from patients attending tertiary healthcare (JSS Hospital) hospital in Mysuru, Karnataka between July 2018 to December 2020. Ethical clearance was obtained from the JSS Institutional Ethical Committee on 06/08/2018 (JSSMC/IEC/18/06/2018-19). Clinical samples like urine, blood, vaginal swab, pus, surgery wounds, and tracheal secretions were collected from the patients based on their clinical symptoms. A total of 73,878 samples were screened for Candida spp. and later confirmed as Candida albicans by phenotypic and genotypic methods. The samples were collected under sterile conditions and transported to the laboratory for analysis.
Isolation and identification of isolates
The samples were cultured on Sabarauds Dextrose Agar at 37°C for 48 hrs. The identification of the suspected isolates were done by Chlamydospore production on cornmeal agar, germ tube test and differential culturing on Candida Hichrome agar.
Differential culturing
The suspected isolates were sub-cultured onto CHROMagar Candida® (CHROM -agar Himedia Bangalore) and incubated at 37 °C for 72 hours to obtain pure cultures and were screened for green colonies.
DNA Extraction
The isolates were inoculated in 2ml of Yeast Extract Peptone Dextrose (YPD) and incubated at 30°C for 16 hrs in a gyratory shaker. DNA extraction was performed on the confirmed isolates using HiMedia Yeast DNA Extraction Kit following the instructions provided by the manufacturers. Following DNA extraction, the concentration and purity of the DNA was checked using a Nanodrop (Thermo Scientific; Wilmington, DE, USA).
Molecular confirmation
Molecular confirmation of the samples confirmed as Candida albicans were done by internal transcribed spacer (ITS) PCR. The ITS region and the primers used for the molecular confirmation of Candida albicans were ITS1 (5′- TCCGTAGGTGAACCTGCGG-3′) and ITS4 (5′-TCCTCCGCTTATTGATAT-3′). The cycling conditions followed for PCR are described below.
Initial denaturation: 95°C – 5min
Denaturation: 95°C – 1min
No. of cycles: 30
Primer annealing: 60°C – 1 min
Extension: 72°C – 1min
Final extension: 72°C – 5 min
Agarose gel electrophoresis was performed and the gel was observed under a Gel documentation system to view the different bands separated based on the size and molecular weight.
ABC Genotyping: ABC Genotyping of the isolates were performed as described by McCullough et al.12 The following primers were used for amplification CA-INT-L (5′-ATAAGGGAAGTCGGCAAAATAGATCCGTAA-3′), CA-INT-R(5′-CCTTGGCTGTGGTTTCGCTAGATAGTAGAT-3′).
Reaction components were assembled as described below. Thaw the reagents on ice. Take a 0.2ml PCR tube and add the reagents in the following order: water (upto 25 μL), buffer (2.5 μL), dNTP’s (0.2mM), MgCl2 (5 μL), forward primers (1.0 μL), reverse primers (1.0 μL), Taq Polymerase and template DNA (1.0 μL). Gently mix the tubes by tapping it. Briefly centrifuge the tubes. Prepare a positive control and negative control. PCR program was set as
Initial denaturation: 95°C – 3 min
Denaturation: 95°C – 1 min
No. of cycles: 30
Primer annealing: 60°C – 1 min
Extension: 72°C – 1 min
Final extension: 72°C – 5 min
The amplified products were subjected to agarose gel electrophoresis (2%) and viewed under a Gel Doc System (BioRad Inc).
Antifungal susceptibility testing by VITEK 2 (Biomerix Inc)
3ml sterile saline was transferred to a test tube. Using a straight wire, a homogenous culture of C. albicans was transferred to the saline tube. The suspension was adjusted to 2 Mc Farland standards. The prepared suspension was placed in the VITEK 2 YST cassette for identification and VITEK 2 YST08 for antifungal susceptibility. The results were then analysed.
Controls
ATCC 90028 and SC 5314 were used as external control strains for identification and typing of our isolates.
Storage
The isolates were then stored on YPD glycerol at-70°C.
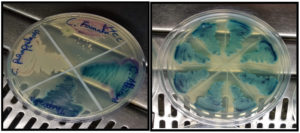
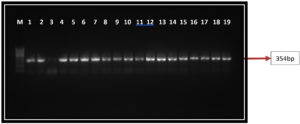
Among the 1427 Candida spp isolated from patients during the study period, 282 isolates were Candida albicans. The colonies of Candida albicans appeared as light green colonies on Hi CHROM-agar (Figure 1). Germ tubes were formed on incubation with human serum at 37°C (2.5-3 hrs). All the phenotypically identified Candida albicans were confirmed by PCR using the ITS primer PCR product of approximately 354bp, which was obtained as shown in Figure 2.
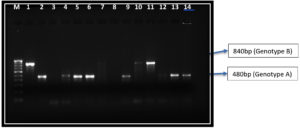
ATCC 90028 and ATCC 90029 were used as the reference strains for Candida albicans genotype A and genotype B and revealed band sizes of 450bp and 840bp respectively. CY 1123 produced two PCR products (450 and 840 bp) and was used as a reference strain for genotype C. Agarose gel electrophoresis revealed the different genotypes based on their band sizes (Table 1).
Table (1):
Different genotypes based on the band sizes.
No. |
Genotypes |
Basepair (bp) |
Ref. |
|---|---|---|---|
1. |
Genotype A |
450 |
McCullough, 1999 et al. |
2. |
Genotype B |
840 |
|
3. |
Genotype C |
450& 840 |
|
4. |
Genotype D |
1080 |
|
5. |
Genotype E |
1400 |
Tamura et al. |
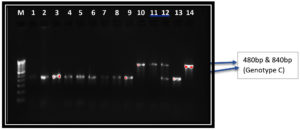
ABC genotyping of the 282 isolated strains of Candida albicans revealed genotypes A, B and C. However, we did not identify any Genotype D or E in our study. Genotype A, B and C accounted for 247 (87.58%), 28 (9.92%), 7(2.48%) respectively. Genotype A produced a band of 450bp while Genotype B produced a band size of 850bp (Figure 3). However, Genotype C revealed two bands, one of 450bp band size and the other 840bp. (Figure 4).
Antifungal susceptibility testing was performed on 282 isolates of C. albicans against the commonly used antifungals: fluconazole, voriconazole Micanazole, flucytosine, caspofungin micafungin and Amphotericin B. The results of the susceptibility pattern against the commonly used antifungals as well as the susceptibility exhibited by the different genotypes are summarised in Table 2. Miconazole was found to be the most effective antifungal being sensitive to all the antibiotics followed by Amphotericin B with the least resistance percentage of 1.4%. The samples showed the following resistance pattern to major antifungals like flucytosine and micafungin (2.8%) Voriconazole (3.1%) Caspofungin (3.5%) and fluconazole (6%).
Table (2):
AST pattern against the commonly used antifungal agents differentiated based on the genotypes.
Antifungal agent |
Overall, Candida albicans Antifungal resistance (%) |
Resistance of Genotype A (n=247) |
Resistance of Genotype B (n=28) |
Resistance of Genotype C (n=7) |
|---|---|---|---|---|
Caspofungin |
3.5 |
2.7 |
7.1 |
0 |
Amphotericin B |
1.4 |
7 |
7.1 |
0 |
Flucytosine |
2.8 |
1.9 |
3.5 |
0 |
Fluconazole |
6 |
3.9 |
7.1 |
0 |
Micafungin |
2.8 |
1.9 |
7.1 |
0 |
Voriconazole |
3.1 |
2.7 |
3.5 |
0 |
Miconazole |
0 |
0 |
0 |
0 |
Among all the identified A genotypes(n=247), 4%(n=10) were resistant to fluconazole, while 2.8 % (n=7) % were resistant to caspofungin and Voriconazole. Only 2 % (n=5) showed resistance to flucytosine and Micafungin. All were susceptible to Miconazole. Higher resistance was observed in Genotype B (n=28) compared to Genotype A, showing the most resistance 7.1 % (n=2) to Caspofungin, micafungin, fluconazole and Amphotericin B followed by 3.5 % (n=1) resistance to flucytosine and voriconazole. All were sensitive to Miconazole. Genotype C (n=7) were susceptible to all the antifungals.
Molecular typing has nowadays been widely used and have proved a very useful tool for epidemiological analysis and have a deeper insight on microbial pathogens, trace the origin of a particular infection, and helps to establish the relationship between different commensal and infectious organisms. It also helps to track the emergence of antibiotic resistance strains and the genetic relatedness between different organisms of the same species.24
Certain studies have used PCR techniques using primers that span highly variable sequences of ITS1 and ITS2, and conserved regions (18S, 5.8S and 28S rRNA genes) for differentiation of medically important Candida species.25-28 Our study has also used these ITS primer sequences for the molecular confirmation of Candida albicans. 27,28
An old and simple molecular method for the differentiation of C. albicans was analysing the RFLPs of cellular DNA, dividing them based on the position of a dimorphic band 12 and then subdividing them into types. It was a reliable and reproducible method and hence used for large epidemiological studies in US and UK.19
McCullough et al.12 reported that a PCR primer designed to span the 25S rRNA gene (rDNA) region classified C. albicans into four genotypes. Based on the length of the PCR product, the genotypes were inferred as genotype A (450bp), genotype B (840bp), genotype C (450 & 840bp), and genotype D (1,080bp). They also confirmed that C. albicans genotype D belonged to the same taxon as C. dubliniensis. This was the widely used method for typing of C. albicans isolates and also for differentiating C. albicans from C. dubliniensis.16,29,30 In our study, most of the Candida albicans were Genotype A (87.58%) followed by Genotype B (9.92%). Only a small percentage belonged to Genotype C (2.48%). Our study correlates with the studies done in other countries by Bii, C. et al. (Kenya),31; Emmanuel, N.N., et al.,( Nigeria),32 Watanabe et al. Japan16 Sawadago33 where, genotype A of C. albicans was the most predominant followed by Genotype B and Genotype C the least prevalent. This method was unique and reproducible and helped in the identification of the different genotypes of Candida albicans namely Genotype A, B, C, D and E.
Tamai IA et al. 34 isolated Candida albicans from HIV-infected patients and identified that 66% of the samples were genotype A.34 In two other studies in Turkey performed by Karahan35 among 190 Candida strains, nearly 51% were genotype A. Another study in Brazil, Chaves et al.36 also reported genotype A as the most common genotype. From all these studies it was concluded that Genotype A was the most common genotype in majority of the studies.
In contrast to all these studies, a study by Abdul Rahim37 from Dublin, Ireland and Moron 38 Malaysia found Genotype A (n=65) the predominant one, but Genotype C (n=20) was more prominent than Genotype B (n=15). This may be attributed to the geographical location. It was observed that the percentage of Genotype B and C were almost similar to each other in studies done in Iraq by Dalvand et al.38 and Samaka et al.39
Although resistance is rare in Candida albicans, long term use of antifungals during recurrent infections has led to emergence of resistance.40 Our study noted a growing resistance in Candida albicans against the commonly used antifungal agents. The highest resistance was exhibited against fluconazole being 6%, which is worrisome.
Our study showed that Genotype B showed more resistance compared to Genotype A & C. Although most of the Candida albicans were Genotype A (n=247), Genotype B (28) showed the most resistance. Genotype B showed a resistance of 7.1% each to the commonly used antifungals Amphotericin B, Fluconazole, caspofungin and Micafungin. However, Mercure et al.41 reported an association of Candida albicans genotype A with a high resistance to flucytosine. Studies by Dalvand,38 Samaka39 also analysed if there was an association between the antifungal resistance pattern and the Candida genotypes, however, found no association.
To conclude, our study showed a growing antifungal resistance in clinical isolates. ABC genotyping revealed that genotype A was predominant than B and C in the South Karnataka region. As genotyping techniques have high efficiency, in future, genotyping can be attempted with a greater sample size considering different geographical locations. This would help us analyse if there was any association between the genotype’s antifungal resistance and the pathogenesis. Our study showed Genotype B to have a higher resistance compared to Genotype A, inspite of most of the Candida albicans being Genotype A. All genotype C isolates were susceptible to the major antifungals. Most of the studies done on genotyping of Candida albicans focused on the isolates obtained from infected lesions of oral cavity, vaginal mucosa and stool samples. A genotyping method with high resolution would be useful for identifying the dissemination area, source and route of Candidiasis and for designing proper management strategies for Candidiasis.
ACKNOWLEDGMENTS
The authors would like to thank JSS Academy of Higher Education and research for providing all the facilities for conducting the study.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
None.
ETHICS STATEMENT
This study was approved by JSS Ethical Clearance Committee (Reference number JSSMC/IEC/18/06/2018-19)
AVAILABILITY OF DATA
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
- Pfaller MA, Messer SA, Moet GJ, Jones RN, Castanheira M. Candida bloodstream infections: Comparison of species distribution and resistance to echinocandin and azole antifungal agents in Intensive Care Unit (ICU) and non-ICU settings in the SENTRY Antimicrobial Surveillance Program (2008-2009). Int J Antimicrob Agents. 2011;38(1):65-69.
Crossref - Sardi JCO, Scorzoni L, Bernardi T, Fusco-Almeida AM, Mendes Giannini MJS. Candida species: Current epidemiology, pathogenicity, biofilm formation, natural antifungal products and new therapeutic options. J Med Microbiol. 2013;62(1):10-24.
Crossref - Pappas PG, Kauffman CA, Andes DR, et al. Clinical Practice Guideline for the Management of Candidiasis: 2016 Update by the Infectious Diseases Society of America. Clin Infect Dis. 2015;62(4):e1-e50.
Crossref - Chakrabarti A, Mohan B, Shrivastava SK, Marak RSK, Ghosh A, Ray P. Change in distribution & antifungal susceptibility of Candida species isolated from candidaemia cases in a tertiary care centre during 1996-2000. Indian J Med Res. 2002;116:5-12. http://www.ncbi.nlm.nih.gov/pubmed/12514972.
- Pfaller MA, Diekema DJ. Epidemiology of invasive candidiasis: A persistent public health problem. Clin Microbiol Rev. 2007;20(1):133-163.
Crossref - Kothari A, Sagar V. Epidemiology of Candida bloodstream infections in a tertiary care institute in India. Indian J Med Microbiol. 2009;27(2):171-172.
Crossref - Pfaller MA, Diekema DJ. Rare and Emerging Opportunistic Fungal Pathogens: Concern for Resistance beyond Candida albicans and Aspergillus fumigatus. J Clin Microbiol. 2004;42(10):4419-4431.
Crossref - Wiederhold NP. Antifungal resistance: current trends and future strategies to combat. Infect Drug Resist. 2017;10:249-259.
Crossref - Mukaremera L, Lee KK, Mora-Montes HM, Gow NAR. Candida albicans yeast, pseudohyphal, and hyphal morphogenesis differentially affects immune recognition. Front Immunol. 2017;8:1-12.
Crossref - Gilfillan GD, Sullivan DJ, Haynes K, Parkinson T, Coleman DC, Gow NAR. Candida dubliniensis: Phylogeny and putative virulence factors. Microbiology. 1998;144(4):829-838.
Crossref - Mancera E, Frazer C, Porman AM, Ruiz-Castro S, Johnson AD, Bennett RJ. Genetic modification of closely related Candida species. Front Microbiol. 2019;10:357.
Crossref - McCullough MJ, Clemons KV, Stevens DA. Molecular and phenotypic characterization of genotypic Candida albicans subgroups and comparison with Candida dubliniensis and Candida stellatoidea. J Clin Microbiol. 1999;37(2):417-421.
Crossref - Vrioni G, Matsiota-Bernard P. Molecular Typing of Candida Isolates from Patients Hospitalized in an Intensive Care Unit. J Infect. 2001;42(1):50-56.
Crossref - Shin JH, Nolte FS, Holloway BP, Morrison CJ. Rapid identification of up to three Candida species in a single reaction tube by a 5′ exonuclease assay using fluorescent DNA probes. J Clin Microbiol. 1999;37(1):165-170.
Crossref - Wilson MJ, Williams DW, Forbes MDL, Finlay IG, Lewis MAO. A molecular epidemiological study of sequential oral isolates of Candida albicans from terminally ill patients. J Oral Pathol Med. 2001;30(4):206-212.
Crossref - Tamura M, Watanabe K, Mikami Y, Yazawa K, Nishimura K. Molecular Characterization of New Clinical Isolates of Candida albicans and C. dubliniensis in Japan: Analysis Reveals a New Genotype of C. albicans with Group I Intron. J Clin Microbiol. 2001;39(12):4309-4315.
Crossref - Guiver M, Levi K, Oppenheim BA. Rapid identification of candida species by TaqMan PCR. J Clin Pathol. 2001;54(5):362-366.
Crossref - Luo G, Mitchell TG. Rapid Identification of Pathogenic Fungi Directly from Cultures by Using Multiplex PCR. J Clin Microbiol. 2002;40(8):2860.
Crossref - Dalle F, Franco N, Lopez J, et al. Comparative Genotyping of Candida albicans Bloodstream and Nonbloodstream Isolates at a Polymorphic Microsatellite Locus. J Clin Microbiol. 2000;38(12):4554.
Crossref - Zeng J, Zong L, Mao T, Huang Y, Xu Z. Distribution of Candida albican genotype and Candida species is associated with the severity of vulvovagianl candidiasis. Nan Fang Yi Ke Da Xue Xue Bao. 2011;31(10):1649-1653. PMID: 22027762
- Odds FC, Davidson AD, Jacobsen MD, et al. Candida albicans strain maintenance, replacement, and microvariation demonstrated by multilocus sequence typing. J Clin Microbiol. 2006;44(10):3647-3658.
Crossref - Mercure S, Montplaisir S, Lemay G. Correlation between the Presence of a Self-Splicing Intron in the 25S RDNA of C. Albicans and Strains Susceptibility to 5-Fluorocytosine. Nucleic Acids Research. 1993;21(25):6020–6027.
Crossref - Williams DW, Wilson MJ, Lewis MA, Potts AJ. Identification of Candida species by PCR and restriction fragment length polymorphism analysis of intergenic spacer regions of ribosomal DNA. J Clin Microbiol. 1995;33(9):2476-2479.
Crossref - McManus BA, Coleman DC. Molecular epidemiology, phylogeny and evolution of Candida albicans. Infect Genet Evol. 2014;21:166-178.
Crossref - Giri S, Kindo AJ. Evaluation of Five Phenotypic Tests in the Identification of Candida Species. Natl J Lab Med. 2015;4(4):13-18.
- Mirhendi H, Makimura K, Khoramizadeh M, Yamaguchi H. A one-enzyme PCR-RFLP assay for identification of six medically important Candida species. Nihon Ishinkin Gakkai Zasshi. 2006;47(3):225-229.
Crossref - Mandviwala T, Shinde R, Kalra A, Sobel JD, Akins RA. High-throughput identification and quantification of Candida species using high resolution derivative melt analysis of panfungal amplicons. J Mol Diagn. 2010;12(1):91-101.
Crossref - Ellepola ANB, Hurst SF, Elie CM, Morrison CJ. Rapid and unequivocal differentiation of Candida dubliniensis from other Candida species using species-specific DNA probes: comparison with phenotypic identification methods. Oral Microbiol Immunol. 2003;18(6):379-388.
Crossref - Wiebusch L, de Almeida-Apolonio AA, Rodrigues LMC, et al. Candida albicans isolated from urine: Phenotypic and molecular identification, virulence factors and antifungal susceptibility. Asian Pac J Trop Biomed. 2017;7(7):624-628.
Crossref - Saghrouni F, Ben Abdeljelil J, Boukadida J, Ben Said M. Molecular methods for strain typing of Candida albicans: a review. J Appl Microbiol. 2013;114(6):1559-1574.
Crossref - Bii CC, Kangogo MC, Revathi G, Wanyoike MW. Genotypes of Candida albicans from clinical sources in Nairobi Kenya. African J Microbiol Res. 2009;3(9):475-477.
- Emmanuel NN, Romeo O, Mebi AG, et al. Genotyping and fluconazole susceptibility of Candida albicans strains from patients with vulvovaginal candidiasis in Jos, Nigeria. Asian Pacific J Trop Dis. 2012;2(1):48-50.
Crossref - Sawadogo PM, Zida A, Soulama I, et al. Genotype analysis of clinical candida albicans isolates using pcrs targeting 25s rdna and alt repeat sequences of the rps and antifungal susceptibility in ouagadougou (Burkina Faso). Infect Drug Resist. 2019;12:3859-3866.
Crossref - Tamai IA, Salehi TZ, Sharifzadeh A, Shokri H, Khosravi AR. Repetitive sequences based on genotyping of candida albicans isolates obtained from iranian patients with human immunodeficiency virus. Iran J Basic Med Sci. 2014;17(11):831-835.
Crossref - Karahan ZC, Guriz H, Agirbasli H, et al. Genotype distribution of Candida albicans isolates by 25S intron analysis with regard to invasiveness. Mycoses. 2004;47(11-12):465-469.
Crossref - da Silva-Rocha WP, Lemos VLDB, Svidizisnki TIE, Milan EP, Chaves GM. Candida species distribution, genotyping and virulence factors of Candida albicans isolated from the oral cavity of kidney transplant recipients of two geographic regions of Brazil. BMC Oral Health. 2014;14(1):1-9.
Crossref - Abdulrahim MH, McManus BA, Flint SR, Coleman DC. Genotyping Candida albicans from Candida Leukoplakia and Non-Candida Leukoplakia Shows No Enrichment of Multilocus Sequence Typing Clades but Enrichment of ABC Genotype C in Candida Leukoplakia. PLoS One. 2013;8(9):1-11.
Crossref - Dalvand A, Katiraee F, Joozani RJ, Shokri H. Genotyping of Candida albicans isolated from animals using 25S ribosomal DNA and ALT repeats polymorphism in repetitive sequence. Curr Med Mycol. 2018;2018(4):12-19.
Crossref - Samaka HM, Al-Mohana AM, Al-Hamadani AH, Al-Charrakh AH. Genotyping and Antifungal Susceptibility Profile of Candida albicans Isolated from Cancer Patients. J Chem Pharm Sci. 2018;11(3):236-241.
Crossref - Arendrup MC, Patterson TF. Multidrug-Resistant Candida: Epidemiology, Molecular Mechanisms, and Treatment. J Infect Dis. 2017;216(suppl_3):S445-S451.
Crossref
© The Author(s) 2022. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.