ISSN: 0973-7510
E-ISSN: 2581-690X
The non-hospital environment particularly poultry farms and abattoirs are fast becoming reservoir channels for the transmission of antibiotic resistant bacteria including those that produce metallo beta-lactamases (MBLs). Food-producing animal’s habouring multidrug resistant bacteria including those that produce metallo-beta-lactamases (MBLs) poses health risks to the human population. This study investigated the prevalence of blaIMP-1 and blaVIM-1 MBL genes in Pseudomonas aeruginosa isolates from food-producing animals by multiplex PCR technique. Anal swab samples (n=120) were bacteriologically analyzed on cetrimide selective agar for the selective isolation of P. aeruginosa isolates. Antibiogram was carried out as per the Clinical and Laboratory Standard Institute (CLSI) criteria. The production of MBLs was detected phenotypically and genotypically using the modified Hodges test method and multiplex PCR technique respectively. DNA products were run on 1.5 % agarose gel, and visualized using a UV transilluminator at 260 nm. Data was analyzed statistically using SPSS version 23.0. Out of the 120 anal swab samples, a total of 43 (35.8 %) isolates of P. aeruginosa was bacteriologically recovered. The P. aeruginosa isolates were found to be resistant to ampicillin (88.4 %), cefotaxime (81.4 %), gentamicin (79.1 %), sulphamethoxazole-trimethoprim (72.1 %), oxacillin (76.7 %), nitrofurantoin (76.7 %), meropenem (62.8 %), ofloxacin (67.4 %), imipenem (65.1 %) and cloxacillin (69.8 %). MBL was phenotypically detected in 15 (34.9 %) isolates of P. aeruginosa. However, the multiplex PCR technique significantly confirmed MBL production in only 12 (27.9) isolates of P. aeruginosa (p<0.001) that harboured the blaIMP-1 MBL genes. The blaVIM-1 MBL genes were not detected in the MBL positive P. aeruginosa phenotypes. The P. aeruginosa isolates that harboured the blaIMP-1 MBL genes were found to be multiply resistant. This study reported for the first time the prevalence of blaIMP-1 MBL genes from P. aeruginosa isolates from anal swab samples of food-producing animals in Abakaliki, Nigeria. The long-term exposure of food-producing animals to antibiotics could cause accumulation of antibiotic resistance determinants in the gut microbiota of these animals.
Pseudomonas aeruginosa, Abattoir, MBL genes, IMP-1, VIM-1, Multidrug resistance, Nigeria
Pseudomonas aeruginosa is a Gram-negative, oxidase-positive, and aerobic bacterium found in the family Pseudomonadaceae, and they are widespread in nature.1 It is a free-living, non-enteric bacterium, commonly found in soil and water1. P. aeruginosa has biofilm-forming capabilities, and they are commonly found in moist or wet environments such as bath tubs, sinks and as common contaminants in hospitals and abattoirs. P. aeruginosa is known for its resistance to antibiotics, and the bacterium is multidrug resistant in nature1,2. Metallo-b-lactamases (MBLs) are carbapenem-hydrolyzing enzymes that have the exceptional ability to hydrolyze the carbapenems including imipenem, meropenem, ertapenem3,4,5,6. MBLs are a type of class B carbapenemases which are usually found in members of the Enterobacteriaceae family, Acinetobacter species and P. aeruginosa isolates, where they mediate bacterial resistance to the carbapenems3,6. MBL-producing bacteria are of public health interest owing to the fact that bacteria that harbour genes for their production are notably resistant to a handful of antibiotics 6,7,8,9,10,11. They confer variable range of resistance to all beta-lactam antibiotics except the monobactams such as aztreonam. More so, the presence of MBL-producing bacteria put the use of the carbapenems under threat5,6,12. Gram negative bacteria including P. aeruginosa that produces MBLs has been previously reported from hospital and non-hospital environment here in Nigeria and elsewhere8,13,14,15,16, 17,18,19. The emergence and spread of carbapenem resistance significantly limits possible treatment options for treating life-threatening infections caused by these bacteria. Pathogenic bacteria that harbour genes for the production of carbapenemases such as MBLs are of public health importance since the carbapenems are often the last line of drugs for the treatment and management of multidrug resistant infections 3. Antibiotic resistance genes are crucial in the niche colonization of microorganisms since microbes need to combat antimicrobial compounds produced by other microorganisms and higher organisms in their immediate environment 20,21. MBL-producing bacteria are a public health menace, and this necessitates the need to detect by genotypic and phenotypic techniques the prevalence of these organisms in the community in order to forestall any health danger due to them. It is in view of this that this study investigated by multiplex PCR technique the prevalence of MBL genes, particularly blaIMP-1 and blaVIM-1 in P. aeruginosa isolates from the non-hospital milieu.
Ethical consideration
This research was conducted in line with the World Medical Association (WMA) declaration of Helsinki on the principles for medical research involving human subjects and identifiable human and animal material/data 22. There was no written ethical approval as the samples were collected at the point of slaughter.
Collection and processing of samples
One hundred and twenty (120) anal swab samples were used for this study; and each of the samples were analyzed using standard microbiology techniques. All samples were aseptically inoculated in nutrient broth (Oxoid, UK) and incubated overnight at 37oC 23. Bacterial growth was phenotypically confirmed by the presence of visible bacterial growth as evidenced by turbidity after incubation. All tubes showing turbidity were each sub-cultured onto freshly prepared plates of cetrimide selective agar (Oxoid, UK) for the selective isolation of P. aeruginosa isolates.
Pseudomonas aeruginosa isolation
Suspension of the turbid solution from the broth culture was aseptically plated onto cetrimide selective agar (CSA) plate(s) for the selective isolation of P. aeruginosa. The plates were incubated overnight at 37oC. P. aeruginosa isolates were subcultured onto freshly prepared CSA plates for the isolation of discrete colonies of P. aeruginosa; and the presence of P. aeruginosa on the culture plates was determined qualitatively and quantitatively based on colonial morphology or characteristics, microscopy, and biochemical testing 23.
Antibiotic susceptibility profiling
The susceptibility of the P. aeruginosa isolates to selected antibiotics was carried out by the Clinical and Laboratory Standard Institute (CLSI) guidelines using the modified Kirby-Bauer disk diffusion technique on Mueller-Hinton (MH) agar plates [Oxoid, UK] 24,25. Antibiotic disks in the single disk format including: imipenem (10 µg), meropenem (10 µg), ertapenem (10 µg), cefoxitin (30 µg), ceftazidime (30 µg), sulphamethoxazole-trimethoprim (25 µg), gentamicin (10 µg), cefotaxime (30 µg), ceftriaxone (30 µg), ciprofloxacin (10 µg), ofloxacin (10 µg), oxacillin (10 µg), ampicillin (10 µg), cefepime (30 µg), aztreonam (30 µg), nitrofurantoin (10 µg) and cloxacillin (10 µg) were used for susceptibility testing. All antibiotics were procured from Oxoid limited [Oxoid, UK]. Inhibition zone diameter was measured, recorded and interpreted based on the CLSI criteria 24,25.
Modified Hodges (Cloverleaf) Test
The presence of MBL in the P. aeruginosa isolates was phenotypically determined. Briefly, the susceptibility of the isolates to imipenem, meropenem, and ertapenem was evaluated as per the CLSI criteria 10,24,26. Isolates found to show reduced susceptibility to the carbapenems as per antibiotic breakpoints recommended by CLSI was phenotypically confirmed for MBL production. The modified Hodges or Cloverleaf test was performed by aseptically swabbing MH agar plates with Escherichia coli ATCC 25922 strain. The inoculated MH agar plates were allowed for about 5 min; and imipenem (10 µg) single disks were aseptically placed at the center of the MH agar plates. The test bacteria (adjusted to 0.5 McFarland turbidity standards) were heavily streaked in three different directions from the edge of the imipenem (10 µg) disk to the center of the MH agar plates. Susceptibility plates were incubated overnight at 30oC. The plates were macroscopically observed for indentation, and the growth of the test bacteria towards the imipenem (10 µg) susceptibility disk. Growth of test bacteria towards the carbapenem disk is indicative of metallo-b-lactamase production phenotypically 4,10,14,26.
Multiplex PCR characterization of MBL genes
Table 1 shows the gene sequence of forward and reverse primers used for the amplification of blaIMP-1 MBL genes and blaVIM-1 MBL genes. All isolates of P. aeruginosa positive for MBL production by the modified Hodges test technique was assessed for the presence of blaIMP-1 and blaVIM-1 MBL genes by multiplex PCR technique using specific primers supplied by Inqaba Biotech limited (Inqaba Biotechnical Industries Ltd, South Africa). The specific primers for the multiplex PCR was constructed as described in a previous report 25,27. Multiplex PCR amplification of the MBL genes in the test isolates was performed in a thermal cycler (Lumex instruments, Canada) with a final volume of 26.5 µl master mix comprising 0.2 µl of Taq polymerase enzyme U/µl, 2.5 µl of 10X PCR buffer along with 2.5 µl MgCl2, 1 µl of 10 pM from each of the forward and reverse primers, 2.5 µl of dNTPs MIX (2 mM), 3 µl of DNA template (from the test isolates), 14.8 µl of nuclease-free water. The initial denaturation temperature was at 95oC for 2 min, and this was followed by 25 cycles of DNA denaturation at 95oC for 30 sec. The primer annealing was carried out at 48oC for 30 sec, and primer extension was carried out at 72oC for 30 sec. After the last cycle, a final extension step was carried out at 72oC for 2 min.
Table (1):
Gene sequence of forward and reverse primers for multiplex PCR technique.
| Gene target(s) | Primer sequence (5ꞌ to 3ꞌ, as synthesized) |
Expected amplicon size (bp) |
|---|---|---|
| blaIMP-1 | F1 (5ꞌ-ACC GCA GCA GAG TCT TTG CC-3ꞌ) R1 (5ꞌ-ACA ACC AGT TTT GCC TTA CC-3ꞌ) |
587 |
| blaVIM-1 | F3 (5ꞌ-AGT GGT GAG TAT CCG ACA G-3ꞌ) R3 (5ꞌ-ATG AAA GTG CGT GGA GAC-3ꞌ) |
261 |
| F-forward primer, R-Reverse primer | ||
Gel electrophoresis
Agarose gel electrophoresis was carried out in order to separate the amplified DNA products according to their individual sizes when an electric field is applied across the gel 27. This was done using 1.5% agarose gel prepared by dissolving 3 g of the agarose powder in 0.5X TBE buffer. The agarose solution was heated in a microwave until all the agarose was dissolved. The molten agarose gel was cooled to about 50oC before pouring, and an aliquot of 1.0 µl of 10 mg/ml of ethidium bromide (EtBr) dye was added to the agarose gel using gloved hands. A comb was then placed in a sealed mould (gel casting chamber), and the molten agarose gel was poured into the sealed gel casting chamber. The gel was allowed to cool for 20 min before the seal and the comb was removed. Thereafter, the solidified gel was placed in the gel electrophoresis vessel or chamber that was filled with 0.5X TBE buffer. And the PCR products (i.e. the amplified genes in the Eppendorf tubes) was mixed with 1 µl 6X coloured loading buffer per 5 µl of the PCR product and then aseptically pipetted into the wells in the gel. An aliquot of 5 µl of the 100 bp DNA marker/ladder was also loaded on one of the gel well while the PCR reagent without DNA was loaded in another well as a negative control. The gel was run at 80 V for 2 h; and then visualized with UV transilluminator at 260 nm.
Table (2):
Isolation and characterization of P. aeruginosa isolates.
| Organism | Sample source | Oxidase test | Gram staining | Colonial features of P. aeruginosa on culture media | Isolation rate of P. aeruginosa n (%) |
|---|---|---|---|---|---|
| Pseudomonas aeruginosa
|
Anal swabs of cow | Positive | Negative | Colonies with greenish and bluish pigmentation | 43 (35.8) |
| Key: n-number of isolates; %-percentage | |||||
Statistical analysis
The Statistical Package for Social Sciences (SPSS) version 23.0 (SPSS, Chicago, IL, USA) was used for data analysis. The differences in data were considered statistically significant at p < 0.05.
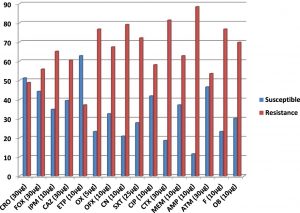
In this study, the prevalence of Pseudomonas aeruginosa isolates that harboured genes for the production of metallo-b-lactamase (MBL) enzymes was phenotypically and genotypically detected using modified Hodges technique and multiplex PCR technique respectively. Table 2 shows the frequency of isolation of P. aeruginosa isolates from the anal swabs of cow bacteriologically analyzed in this study. A total of 43 (35.8%) isolates of P. aeruginosa was bacteriologically analyzed from 120 anal swab samples used in this study. The P. aeruginosa isolates recovered produced colonies with greenish and bluish pigmentation on cetrimide selective agar, due to pyocyanin and pyoverdin pigments typical of Pseudomonas species (Table 2). Figure 1 shows the percentage susceptibility profile of the P. aeruginosa isolates to the tested antibiotics used in this study. The isolates of P. aeruginosa showed varied levels of resistance and susceptibility to the tested antibiotics. Interestingly, the P. aeruginosa isolates were found to be highly resistant to beta-lactam antibiotics in the penicillin family including ampicillin (88.4%), oxacillin (76.7%) and cloxacillin (69.8 %).
Fig. 1. Percentage susceptibility of 43 isolates of P. aeruginosa
Key: IPM = imipenem, MEM = meropenem, ETP = ertapenem, FOX = cefoxitin, CAZ = ceftazidime, SXT=sulphamethoxazole-trimethoprim, CN = gentamicin, CTX = cefotaxime, CRO = ceftriaxone, CIP = ciprofloxacin, OFX = ofloxacin, OX = oxacillin, AMP = ampicillin, ATM = aztreonam, F = nitrofurantoin, OB = cloxacillin
Table (3):
Phenotypic detection of MBL-producing P. aeruginosa isolates.
| Organism | Source | MBL positive n (%) | MBL negative n (%) |
|---|---|---|---|
| P. aeruginosa | Anal swabs of cow |
15 (34.9) | 28 (65.1) |
| Key: MBL-metallo-β-lactamase; n-number of isolates; %-percentage | |||
The next resistance profile of the P. aeruginosa isolates was resistance to antibiotics in the cephalosporin class specifically 2nd– and 3rd generation cephalosporins including cefotaxime (81.4 %), ceftazidime (60.5%), cefoxitin (55.8%) and ceftriaxone (48.8%). All the isolates of P. aeruginosa were also resistant to some non-beta-lactam antibiotics including gentamicin (79.1%), sulphamethoxazole-trimethoprim (72.1 %), nitrofurantoin (76.7%) and ofloxacin (67.4%). The P. aeruginosa isolates also showed reduced susceptibility to the carbapenems used in this study including ertapenem (37.2%), meropenem (62.8%) and imipenem (65.1%). The result of the prevalence of MBL phenotypes detected by modified Hodges (Cloverleaf) method is shown in Table 3. MBL production was phenotypically detected in a total of 15 (34.9%) isolates of P. aeruginosa out of the 43 isolates of P. aeruginosa that was phenotypically screened for MBL production by the modified Hodges test method. Table 4 shows the distribution of the blaIMP-1 and blaVIM-1 MBL genes investigated by multiplex PCR technique in the P. aeruginosa isolates analyzed in this study. The occurrence of blaIMP-1 MBL genes was significantly detected in a total of 12 (27.9%) isolates of P. aeruginosa (p<0.001) by multiplex PCR technique. The blaVIM-1 MBL genes was not detected in the P. aeruginosa isolates that was genotypically screened by multiplex PCR technique using specific primers (Table 4). The isolates of P. aeruginosa that produced MBL and was confirmed by multiplex PCR technique to harbour the blaIMP-1 MBL genes was found to be multiply resistant to antibiotics in the class of fluoroquinolones, carbapenems, cephalosporins and aminoglycosides.
Table (4):
Prevalence of MBL genes in the 43 isolates of P. aeruginosa from anal swab samples.
| MBL genes detected | No. of isolates (%) | Resistance profile |
|---|---|---|
| blaIMP-1 | 12 (27.9) | Fluoroquinolones, aminoglycosides, cephalosporins, carbapenems |
| blaVIM-1 | 0 (0) | |
| (p<0.001) |
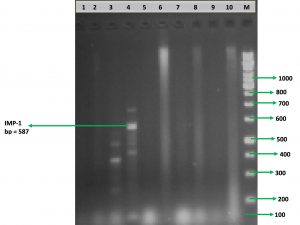
Fig. 2. Electrophoretic analysis and identification of blaIMP-1 MBL gene in the P. aeruginosa isolates. Lane M is the DNA marker/ladder which is a 100 bp marker. Lanes 2-10 shows the different DNA samples. Lane 4 shows amplified products of blaIMP-1 MBL gene with a base pair size of 587 bp. There was no amplified gene product in lanes 1,2,3,5-10. Lane 1 is the negative control which contains nuclease free water.
Antibiotics are invaluable for the fight and control of infectious diseases in both human and animal population. But this potent compound has over the years succumbed to the evolving nature of antibiotic resistant bacteria (including those that produce metallo-b-lactamases, MBLs). Antibiotic resistant bacteria are known to remain viable even in the face of potent antimicrobial onslaught. We asked if P. aeruginosa isolates from abattoir in Abakaliki metropolis, Nigeria harbour gene for the production of MBLs. To answer this question, we investigated the prevalence of MBL genes, particularly blaIMP-1 and blaVIM-1 MBL genes in P. aeruginosa isolates from food-producing animals using both genotypic and phenotypic techniques. The occurrence of P. aeruginosa in this study was high (n=43; 35.8%), which is similar to the work of Saderi et al.10 and Olutayo and Abimbola 28. The P. aeruginosa isolates recovered in this study also showed high level resistance to the carbapenems including imipenem, ertapenem and meropenem; and this was found to be in accordance with previous reports carried out by Franco et al.12 in Brazil. Reduced susceptibility of the P. aeruginosa isolates was also observed in cefoxitin, cefotaxime, ceftriaxone and ceftazidime used for treating infections caused by P. aeruginosa; and this report of ours is similar to the study conducted by Aibinu et al.14 and Olutayo and Abimbola 28 who reported similar levels of reduced susceptibility of P. aeruginosa isolates to antibiotics in southwest Nigeria. P. aeruginosa is notorious for its resistance to antibiotics due to the permeability barrier associated with the outer membrane (OM) layer of the organism 1,20,29. Bashir et al.30 and Akinduti et al.31 also reported in their study carried out in Kashmir and Abeokuta respectively that P. aeruginosa is a multidrug resistant organism that is notoriously resistant to several antibiotic classes. Antibiotics are usually added to the feed and water of food-producing animals including livestock and poultry birds to promote growth and limit bacterial infection in the animals. But such supplementation of antibiotics in the water and feed of food-producing animals poses health risk to the human population due to the possible development and dissemination of antibiotic resistance in the food chain.32 The production of metallo-b-lactamase (MBL) enzymes in the P. aeruginosa isolates recovered in this study was phenotypically confirmed in 15 (34.9%) isolates. However, multiplex PCR technique only confirmed MBL production in 12 (27.9%) isolates of P. aeruginosa that were found to harbour the blaIMP-1 MBL genes. In a related study carried out in Nigeria, Aibinu et al.(14) reported the phenotypic detection of MBL in isolates of P. aeruginosa in southwest Nigeria. Also, we had previously reported that isolates of P. aeruginosa are notorious in the production of MBLs, and this has contributed to the increasing resistance of the organism to the carbapenems and some non-beta-lactam antibiotics 4. Our report on the phenotypic detection of MBLs in P. aeruginosa isolates is not in agreement with those reported by Abd El-Baky et al.33 in which 31 isolates of P. aeruginosa were phenotypically detected to produce MBL enzymes in Asia. Akinduti et al.31 also reported a lower rate of MBL-positive P. aeruginosa isolates (3.3%) in their study carried out in southwest Nigeria. In Japan, Shibata et al.27 also reported a higher occurrence rate of MBL-producing P. aeruginosa isolates in their study in which 116 P. aeruginosa isolates were discovered phenotypically to produce MBL enzymes. The prevalence of MBL genes was significantly (p<0.001) detected by multiplex PCR technique in the P. aeruginosa isolates genotypically investigated for the presence of blaIMP-1 MBL genes and blaVIM-1 MBL genes. It was discovered that only the blaIMP-1 MBL genes were detected by multiplex PCR in the isolates of P. aeruginosa investigated in this study. The blaVIM-1 MBL genes were not detected by the multiplex PCR technique used. blaIMP-1 MBL genes have been previously reported to be a widely distributed MBL genes that mediate bacterial resistance to the carbapenems including imipenem.3 In a similar study conducted in Iraq, Anoar et al.34 reported that the prevalence of blaIMP-1 MBL gene in isolates of P. aeruginosa was between 4-5%. In Japan, Shibata et al.27 reported that out of a total of 357 isolates of Gram-negative bacteria carrying blaIMP-1 MBL genes, 116 isolates were confirmed to be P. aeruginosa. These result reported by Shibata et al.27 is not in agreement with ours where the number of bacterial isolates that were genotypically confirmed by multiplex PCR to harbour blaIMP-1 MBL gene was 12 P. aeruginosa isolates. The presence of blaIMP-1 MBL genes in P. aeruginosa isolates analyzed in this study may be due to several risk factors associated with the acquisition, development and spread of antibiotic resistance genes in the community of which the undue exposure and use of antibiotics in animal husbandry and in livestock production have been implicated as a key factor driving the development and spread of drug resistant bacteria in the community 32,35. Several studies have also shown that the blaIMP-1 MBL genes are the most prevalent MBL genes harboured by Gram-negative bacteria including E. coli, Klebsiella species and P. aeruginosa; and the blaIMP-1 MBL genes occur worldwide 3,9,36.
The result of our study is the first report on the prevalence of P. aeruginosa isolates of food-producing animals in Abakaliki, Nigeria that harbours the blaIMP-1 MBL genes. Antibiotic usage for the propagation of food-producing animals is one of the major factors through which resistant bacteria including those that produce MBLs emerge and spread in the community. The surveillance of antibiotic resistance and their genetic determinants in livestock and other agricultural practices in this part of the world should be improved upon through proper detection and awareness on proper antibiotic usage in order to contain the nefarious activities of drug resistant bacteria.
Conflict of Interest
The author(s) declare that there is no conflict of interest.
- Madigan M.T., Martinko J.M., Dunlap P.V and Clark D.P. Brock Biology of microorganisms. 12th edition. Pearson Benjamin Cummings Publishers. USA. 2009; Pp.795-796.
- Brooks G.F., Butel J.S and Morse S.A. Medical Microbiology, 23rd edition. McGraw Hill Publishers. USA. 2004; Pp. 248-260.
- Walsh T.R., Toleman M.A., Poirel L and Nordmann P. Metallo b – Lactamases: The Quiet Before the Storm? Clinical Microbiology Review. 2005; 18(2):306-325.
- Ejikeugwu PC, Ugwu CM., Iroha IR., Eze P, Gugu TH and Esimone CO. Phenotypic Detection of Metallo-b-Lactamase Enzyme in Enugu, Southeast Nigeria. American Journal of Biological, Chemical and Pharmaceutical Science. 2014; 2(2):1-6.
- Tortola M.T., Lavilla S., Miro E., Gonzalez J.J., Larrosa N., Sabate M., Navarro F and Prats G. First Detection of a Carbapenem – Hydrolyzing Metallo enzyme in Two Enterobacteriaceae Isolates in Spain. Antimicrobial Agents and Chemotherapy. 2005; 49(8):3492-3494.
- Thompson KS. Extended – spectrum b-Lactamase, AmpC, and Carbapenemase Issues. Journal of Clinical Microbiology. 2010; 48(4):1019-1025.
- Okazaki R, Hagiwara S, Kimura T, Tokue Y, Kambe M, Murata M, Aoki M, Kaneko M, Oshima K and Murakami M. Metallo-b-lactamase-producing Klebsiella pneumoniae infection in a non-hospital environment. Acute Medicine and Surgery. 2016; 3:32-35.
- Overturf GD. Carbapenemases: A Brief Review for Pediatric Infectious Disease Specialists. The Pediatric Infectious Disease Journal. 2010; 29(1):68-70.
- Rossolini G.M, Condemi M.A, Pantanella F, Docquier J.D, Amicosante G and Thaller M.C. Metallo-b-lactamase producers in environmental microbiota: new molecular class B enzyme in Janthinobcaterium lividum. Antimicrobial Agents and Chemotherapy. 2001; 45(3):837-844.
- Saderi H., Karimi Z., Owlia P., Bahar M.A., Rad S.M. Phenotypic detection of metallo – beta – lactamase producing Pseudomonas aeruginosa strains isolated from burned patients. Iranian Journal of Pathology. 2008; 3(1):20-24.
- Safari M, Nejad A.S.M, Bahador A, Jafari R and Alikhani M.Y. Prevalence of ESBL and MBL encoding genes in Acinetobacter baumannii strains isolated from patients of intensive care units (ICU). Saudi Journal of Biological Sciences. 2015; 22:424-429.
- Franco M.R.G., Caiaffa-Filho H.H., Burattini M.N and Rossi F. Metallo-beta-lactamases among imipenem-resistant Pseudomonas aeruginosa in a Brazilian University hospital. Clinics. 2010; 65(9):825-829.
- Ejikeugwu C, Esimone C, Iroha I, Ugwu C, Ezeador C, Duru C, Adikwu M. Phenotypic detection of AmpC beta-lactamase among anal Pseudomonas aeruginosa isolates in a Nigerian abattoir. Archives of Clinical Microbiology. 2016; 7(2):1-5.
- Aibinu I, Nwanneka T, Odugbemi T. Occurrence of ESBLs and MBL in Clinical Isolates of Pseudomonas aeruginosa from Lagos, Nigeria. Journal of American Science. 2007; 3(4):81-85.
- Carfi A., Pares S., Duéé E., Galleni M., Duez C., Frere J.M and Dideberg O. The 3 – D structure of a zinc metallo –b – lactamase from Bacillus cereus reveals a new type of protein fold. The Embo Journal. 1995; 14(20):4914-4921.
- Deshpande P., Rodrigues C., Shetty A., Kapadia F., Hedge A and Soman R. New Delhi Metallo-b-lactamase (NDM-1) in Enterobacteriaceae: Treatment options with Carbapenems Compromised. Japi. 2010; 58:147-149.
- Mahajan A, Tandon VR. New Delhi Metallo (Beta-lactamase 1): Fact or fiction. Journal of Medical Education and Research. 2010; 12(3):109.
- Lolans K., Queenan A.M., Bush K., Sahud A and Quinn J.P. First Nosocomial Outbreak of Pseudomonas aeruginosa Producing and Integron – Borne Metallo-b-Lactamase (VIM-2) in the United States. Antimicrobial Agents Chemotherapy. 2005; 49(8):3538-3540.
- Chakraborty D, Basu S, Das S. A study on infections caused by Metallo-Beta-Lactamase Producing Gram-negative Bacteria in Intensive Care Unit Patients. American Journal of infectious Diseases. 2010; 6(2):34-39.
- Fernandez M, Conde S, Torre J, Molina-Santiago C, Ramos J.L and Duque E. Mechanisms of resistance to chloramphenicol in Pseudomonas putida KT2440. Antimicrob. Agents Chemother. 2012; 56(2):1001-1009.
- Bush K, Jacoby GA, Medeiros AA. A Functional Classification of Scheme for Beta Lactamases and its Correlation to Molecular Structure. Antimicrob Chem other 1995; 39:1211-1233.
- World Medical Association (WMA) Declaration of Helsinki. Ethical Principles for Medical Research involving Human Subjects. Note of Clarification on Paragraph 30 added by the WMA General Assembly, Tokyo 2004. 9.10.2004.
- Cheesbrough M. District Laboratory Practice in Tropical Countries. 2nd edition. Cambridge University Press, UK. 2006; Pp. 178-187.
- Clinical Laboratory Standard Institute, CLSI. Performance standards for antimicrobial disk susceptibility test. Fifteenth informational supplement, CLSI document M100-S15. Wayne, PA. USA. 2011.
- Ejikeugwu C, Esimone C, Iroha I, Igwe D.O, Ugwu M, Ezeador C, Duru C, Adikwu M. Molecular Identification of MBL Genes blaIMP-1 and blaVIM-1 in Escherichia coli Strains Isolated from Abattoir by Multiplex PCR Technique. Research Journal of Microbiology. 2017; 12: 266-273.
- Varaiya A., Kulkarni N., Kulkarni M., Bhalekar P and Dogra J. Incidence of metallo beta – lactamase producing Pseudomonas aeruginosa in ICU patients. Indian J Med Res. 2008; 127:398-402.
- Shibata N, Doi Y, Yamane K, Yagi T, Kurokawa H, Shibayama K, Kato H, Kai K and Arakawa Y. PCR typing of genetic determinants for metallo-b-lactamases and integrases carried by Gram-negative bacteria isolated in Japan, with focus on the class 3 integron. Journal of Clinical Microbiology. 2003; 41(12):5407-5413.
- Olutayo IF, Abimbola OA. Antibiogram of Escherichia coli and Pseudomonas strains isolated from wastewater generated by an abattoir as it journeys into a receiving river. Advances in Microbiology. 2016; 6:303-309.
- Wegener HC. Antibiotics in animal feed and their role in resistance development. Current Opinion in Microbiology. 2003; 6:439-445.
- Bashir D., Thokar M.A., Fomda B.A., Bashir G., Zahoor D., Ahmad S and Toboli A.S. Detection of metallo – beta – lactamase (MBL) producing Pseudomonas aeruginosa at a tertiary care hospital in Kashmir. African Journal of Microbiology Research. 2011; 5(2):164-172.
- Akinduti P.A, Ejilude O, Motayo B.O and Adeyokinu A.F. Emerging multidrug resistant AmpC beta-lactamase and carbapenemase enteric isolates in Abeokuta, Nigeria. Nature and Science. 2012; 10(7): 70-74.
- Shea KM. Committee on Environmental Health and Committee on Infectious Disease: Non-therapeutic use of antimicrobial agents in animal agriculture: implications for pediatrics. Pediatrics. 2004; 114:862-868.
- Abd El-Baky RM, Abd El-Azeim NH, Gad GFM. Prevalence of extended spectrum beta lactamase, AmpC beta lactamase, and metallo beta lactamase among clinical isolates of Pseudomonas aeruginosa. Journal of Advanced Biotechnology and Bioengineering. 2013; 1:22-29.
- Anoar KA, Ali FA, Omer AA. Detection of metallo b-lactamase enzyme in some Gram-negative bacteria isolated from burn patients in Sulaimani city, Iraq. European Scientific Journal. 2014; 10(3):485-496.
- Usha PTA, Sabitha J, Nisha AR. Antimicrobial Drug Resistance: A Global Concern. Veterinary World. 2010; 3(3):138-139.
- Mansouri S, Neyestanaka D.K, Shokoohi M, Halimi S, Beigverdi R, Rezagholezadeh F and Hashemi A. Characterization of AmpC, CTX-M and MBLs types of b-lactamases in clinical isolates of Klebsiella pneumoniae and Escherichia coli producing extended spectrum beta lactamases in Kerman, Iran. Jundishapur J Microbiol. 2014; 7(2):e8756.
© The Author(s) 2018. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.