ISSN: 0973-7510
E-ISSN: 2581-690X
The world is seeing a continuous rise in the levels of antibiotic resistance1. Organisms develop new resistance mechanisms, emerge, and spread the resistance worldwide, making it challenging to treat common infectious diseases. In the current study, clinical isolates received between the years 2017 to 2020 were cultured and the isolated organisms were screened for antibiotic resistance; isolates with multiple drug resistance were further subjected to confirmatory screening through Combined Disc Test (CDT) and Modified Hodge Test (M.H.T.), and molecular characterization to be finally tested for gene expression analysis. Molecular characterization involved screening of genes blaVIM-2, blaKPC-3, blaNDM-1, and blaIMP-11 responsible for imparting carbapenem drug resistance2. From the laboratories of tertiary care hospitals, a total of 1452 clinical isolates were collected and identified. The organisms were subjected to antibiotic susceptibility screening and carbapenem resistance screening. The isolates found positive in the screenings were subjected to molecular characterization for genes, blaVIM-2, blaKPC-3, blaNDM-1, and blaIMP-11, responsible for imparting carbapenem drug resistance. Most of the isolates were resistant variably to aminoglycosides but were found to be resistant to fluoroquinolones and β-lactams group of antibiotics. Carbapenem activity was detected in twelve percent of total isolates and 27 percent among multidrug-resistant isolates. blaNDM-1 gene was found present in 77% isolates, and five organisms among the total number of organisms showed pan drug resistance.
Carbapenem resistance, antibiotic resistance, blaVIM-2, blaKPC-3, blaNDM-1, blaIMP-11, Carbapenem-Resistant Enterobacteriaceae, Methicillin-Resistant Staphylococcus aureus
Patients’ more prolonged stay at healthcare setups increases the chance of acquiring secondary infections. Antibiotic-resistant organisms aggravate the worsened situation. Each year, in the US, >2.8 million antibiotic resistant infections occur1.According to the National Ambulatory Medical Care Survey 2016, conducted by the Center for Disease Control and Prevention (C.D.C.) in 2016, infectious and parasitic diseases accounted for 15.5 million doctor visits and 3.7 million emergency room visits in the U.S.A.3. As a result, more than 35,000 people die, making antibiotic-resistant infections a public health threat, according to the report published in the year 2019 by the C.D.C4. As per the annual report compiled by the Indian Council of Medical Research (ICMR) under the name of Antibiotic-Resistant Surveillance and Research Network in the year 2019, isolates from 107, 387 samples were studied taken/collected from various body parts5.
As per the annual report compiled by the Indian Council of Medical Research (ICMR) under the name of Antibiotic-Resistant Surveillance and Research Network in the year 2019, a total of 32,672 significant clinical isolates belonging to various genera and species of family Enterobacteriaceae were screened for antibiotic resistance. 45% of the isolates were found to be resistant toimipenem, 35% were found to be resistant to meropenem, and 40% were found to be resistant to ertapenem antibiotic4.One in every thirty-one hospital patients has at least one hospital-associated infection on any given day in the U.S.A.6; similar situations are observed across the world, and attempts are being made by healthcare workers to prevent them. Extended-spectrum β-lactamases (ESβLs), Vancomycin-Resistant Enterococci (VRE), and Carbapenem-Resistant Enterobacteriaceae (C.R.E.), and Methicillin-Resistant Staphylococcus aureus (MRSA), are the most frequently isolated organisms responsible for Hospital Acquired Infections (HAIs)6.
Antibiotic resistance is an ancient and dynamic mounting problem7. The resistance poses a global threat to the health of humans, animals and environment which is due to the emergence and existence of superbugs across animal, human and environmental due to excessive use of antibiotics in animals and humans. Antibiotic resistance has been declared as a global public health concern by many important organizations like Centers for Disease Control and Prevention (CDC), Infectious Disease society of America, World Economic Forum, and the World Health Organization8,9. Terrible complications may occur if effective global plans are not adopted soon. In this study we have tried to portray how resistance is spreading along with the intensity of the antibiotic resistance among different clinical specimen10.
The study does not involve samples of human origin, the strains used for processing were submitted by associate laboratories of hospitals after isolation; hence, Helsinki declaration is not required. During the study, no humans were prescribed medicines or placebo; further, biological vitals such as height, sex, weight etc. were not measured. The resistance screening performed on the antibiotics against established formulas and at the concentrations prescribed.
Selection of Bacterial Strains
A total of 1452 non-repetitive clinical isolates, obtained from laboratories of tertiary care hospitals of Ahmedabad between the period of 2017 to 2020, were identified using Vitek-2 compact. In percentage, identified organisms were E. coli (45%), Klebsiella spp. (18%), Enterobacter spp. (11%) and Proteus spp. (3%), of Enterobacteriaceae group, Acinetobacter spp. (2%), Burkholderia spp. (4%), Haemophilus spp. (5%) and Pseudomonas spp. (13%), of the non-Enterobacteriaceae group, which were subjected to susceptibility testing.
Antibiotic Susceptibility Testing
The isolates were subjected to antibiotic susceptibility testing, performed using the protocol described by Kirby-Bauer disk diffusion susceptibility test protocol, and the results were interpreted according to the clinical breakpoints recommended by the Clinical and Laboratory Standards Institute (CLSI) guideline11,12. According to the definition of C.D.C., in order not to miss any possible carbapenem producer, organisms showing resistance towards at least one carbapenem agent were considered for carbapenem confirmatory testing11. All consumable, used in the study, were procured from Hi-media Laboratories (Mumbai). Antibiotic discs taken are as following: Cefotaxime-clavulanic acid (30/10µg), aztreonam (30µg), cefepime (30µg), ceftriaxone (30µg), cefoxitin (30µg), cefotetan (30µg), meropenem (10µg), imipenem (10µg), ertapenem (10µg), cefotaxime (30µg), ceftazidime + clavulanic acid (30/10µg), ceftazidime (30µg)11. To differentiate among isolated cultures and to derive the antibiotic-resistance pattern of the organisms among extended-spectrum beta-lactamases (ESβLs), AmpC β-lactamases (AmpCβL) or Metallo-β-lactamases (MβL),the study used method suggested by Parul Sinha et al.13
Confirmation of Carbapenem Resistance
Modified Hodge Test
Using the ATCC E. coli 25922 strain a 5 ml Solution of 0.5 McF dilution was prepared. Prepared solution was diluted to 1:10 dilution factor for analysis. The prepared inoculum was lawn streaked on Muller Hinton agar plate and allowed to dry. At the center of the prepared plate meropenem (10µg) susceptibility disc was placed, and the test organism was streaked edge to edge on the plate. The results of the tests are considered positive when there is development of clover leaf-type indentation at the intersection of the test organism and ATCC strain within the zone of inhibition of the carbapenem susceptibility disc; results for the isolates developed otherwise were considered negative for carbapenem resistance14.
Combined Disc Test (CDT) / Ethylene Diamine Tetraacetic Acid Test
Muller Hinton Agar plates, pre-inoculated with the sample of the test isolate (0.5 McF standard) were processed with two sets of antibiotic discs, each one containing imipenem (10µg), meropenem (10 µg) or ertapenem (10 µg). 10 µl 0.1M EDTA solution was added to one of the two antibiotic discs Immediately after the discs were placed onto the agar. Plates were incubated at 37°C for 16 to 20 hours; zone diameters were recorded at the end of the incubation period. Zone diameter difference of ≥5mm between the APBA-free and APBA-containing discs or between the EDTA-free and EDTA-containing discs were considered indicative of Class A and B carbapenemase production, respectively15.
The organismspositive for at least one tests out of the two confirmatory tests, Combined Disc test (CDT) and Modified Hodge test, were selected; the remaining organisms were ruled out from the further analysis.
Molecular Characterization of Isolates
Using column extraction, genomic DNA of the isolates was extracted to perform Polymerase Chain Reaction (PCR) on a 7500 Real-Time RT-PCR machine using primers of Taqman chemistry for blaVIM-2, blaKPC-3, blaNDM-1, and blaIMP-11 carbapenem genes while maintaining the conditions described by (Van der zee et al., 2014). 5.0µl sample DNA was mixed with 25 µl of master mix and 2.50µl of assay mix topped up by 17.50µl of H2O ultimately making the final reaction volume to 50.0µl. RT-PCR reaction was set for single-plex PCR, despite primers have been showing positive reactions in multiplex conditions16.
Data analysis
Data generated during the study were processed by conventional statistical tools and had a P value of <0.05 for critical testing; 7500-software provided by the manufacturer was used to analyze the data obtained from RT-PCR.
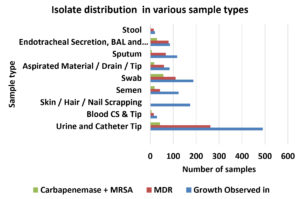
The prevalence of multi-drug resistant and carbapenem-resistant organisms in healthcare setup is a challenge for healthcare workers; in Egypt prevalence of carbapenem-resistant organisms among Klebsiella pneumoniae has been recorded as high as 44.3%17. Similar trends were observed in this study. Out of a total number of isolates (n=1452) screened for antibiotic resistance, n=651 isolates (45%) were found to be multi-drug resistant using current breakpoints recommended by CLSI (Table-1/Fig. 1)18.
Table (1):
Isolate wise distribution of multidrug resistant strains from various clinical specimen.
Specimen Type |
Growth Observed in (n) |
MDR (n) |
Carbapenemase + MRSA (n) |
|---|---|---|---|
Urine and Catheter Tip |
490 |
262 |
42 |
Blood CS & Tip |
29 |
15 |
5 |
Skin / Hair / Nail Scrapping |
174 |
NA |
NA |
Semen |
123 |
42 |
19 |
Swab |
188 |
110 |
57 |
Aspirated Material / Drain / Tip |
84 |
60 |
17 |
Sputum |
117 |
67 |
6 |
Endotracheal Secretion, BAL and Tip |
86 |
80 |
29 |
Stool |
20 |
15 |
2 |
total |
1311 |
651 |
177 |
MDR – Multidrug resistant, CS – Culture and sensitivity, BAL – Broncho Alveolar Lavage
Fig. 1. Isolate wise distribution from various specimen types with MDR and carbapenem isolation pattern
Most of the isolates collected during this study were resistant variably to aminoglycosides but were found to be resistant to fluoroquinolones and β-lactams group of antibiotics, which contradicts the finding of Endimiani et al19. Antibiotic susceptibility results of twelve isolates are worrisome, as the isolates showed pan-drug resistance, making it mandatory to develop new antimicrobials for the isolates along with a message for healthcare workers to maintain strict infection control measures to prevent cross-infection per see.
Carbapenem activity was detected in twelve percent of total isolates and 27 percent among multidrug-resistant isolates (n=177). CDT was positive for 25 percent of isolates (n=45), and M.H.T. was positive for 44 percent of isolates (n=77); whereas 31 percent of isolates (n=55) were found to be positive for both the tests (Table 2).
Table (2):
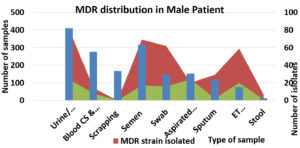
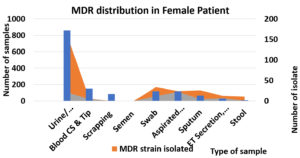
Gender wise distribution of MDR organisms.
| No. | Total (n) |
Gender (n) | Type of Specimen (n) | Growth observed (n) | MDR strain isolated (n) | Carbapenem Strain Isolated (n) | ||||
|---|---|---|---|---|---|---|---|---|---|---|
| M | F | M | F | M | F | M | F | |||
| 1 | 3113 | 1676 | 1437 | Urine and Catheter Tip | 410 | 860 | 81 | 162 | 24 | 21 |
| 1270 | ||||||||||
| Blood CS & Tip | 276 | 149 | 14 | 6 | 8 | 5 | ||||
| 425 | ||||||||||
| Skin / Hair / Nail Scrapping 253 | 168 | 85 | 0 | 0 | 0 | 0 | ||||
| Semen 317 | 317 | 0 | 69 | 0 | 17 | 0 | ||||
| Swab 266 | 147 | 119 | 62 | 34 | 16 | 12 | ||||
| Aspirated Material / Drain / Tip 270 | 153 | 117 | 20 | 24 | 24 | 22 | ||||
| Sputum 184 | 118 | 66 | 29 | 26 | 2 | 5 | ||||
| Endotracheal Secretion, BAL and Tip 104 | 75 | 29 | 59 | 13 | 20 | 7 | ||||
| Stool 24 | 13 | 11 | 7 | 11 | 1 | 1 | ||||
MDR – Multidrug resistant, CS – Culture and sensitivity, BAL – Broncho Alveolar Lavage
Gene blaNDM-1, contradicting the findings of researchers from Turkey, where it emerged, consequently finding its reservoir to the Middle East and Africa, and expanding into India21,22, agreeing to above research’s prevalence of the gene was found as high as 77% among multidrug-resistant strains along with blaKPC-3 gene (Table 3) making Indian population vulnerable to carbapenem-resistant organism infections21.
Table (3):
Results after phenotypic and automated isolation and deviation between both the techniques.
Organisms identified as |
Total isolates (n) |
Identified as the same organism after phenotypic characterization (n) |
Organism identified as after phenotypic characterization (n) |
Isolates showing carbepenem resistance (n) |
|---|---|---|---|---|
Acinetobacter spp. |
3 |
2 |
01 (E. coli) |
3 |
E. coli |
17 |
14 |
03 (Klebsiella spp.) |
17 |
Enterobacter spp. |
7 |
7 |
0 |
7 |
Klebsiella spp. |
33 |
30 |
03 (E. coli) |
33 |
Proteus spp. |
2 |
2 |
0 |
2 |
Pseudomonas spp. |
19 |
19 |
0 |
19 |
Total |
81 |
74 |
7 |
81 |
* Identification carried out using Vitek-2 compact
Table (4):
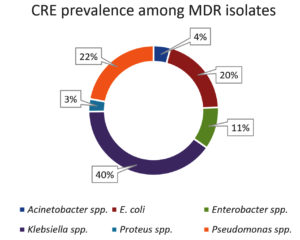
Prevalence of carbapenem resistance among multidrug resistant isolates.
| No. | Organism | Total Isolates (n, %) |
|---|---|---|
| 1 | Acinetobacter spp. | 4, 4.21% |
| 2 | E. coli | 19, 20.00% |
| 3 | Enterobacter spp. | 10, 10.53% |
| 4 | Klebsiella spp. | 34, 40.00% |
| 5 | Proteus spp. | 3, 3.16% |
| 6 | Pseudomonas spp. | 21, 22.11% |
| Total | 95, 100% | |
Furthermore, the presence of multiple blaKPC-3 and blaNDM-1 carbapenem resistance genes was observed, which explains why a total of five organisms showed pan-drug resistance.
Forty organisms that were positive during confirmatory screening, but showed negative results in RT-PCR, could be because of the combination of ESBLs and/or changes in an outer membrane protein22,23.
Table (5):
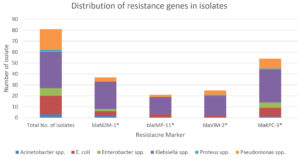
Molecular characterization and Carbapenem gene distribution in clinical isolates.
Organism |
Total No. of isolates (n) |
blaNDM-1* (n) |
blaIMP-11* (n) |
blaVIM-2* (n) |
blaKPC-3* (n) |
|---|---|---|---|---|---|
Acinetobacter spp. |
3 |
2 |
0 |
0 |
1 |
E. coli |
17 |
4 |
3 |
2 |
8 |
Enterobacter spp. |
7 |
2 |
0 |
0 |
5 |
Klebsiella spp. |
33 |
25 |
16 |
18 |
30 |
Proteus spp. |
2 |
0 |
0 |
1 |
1 |
Pseudomonas spp. |
19 |
4 |
2 |
4 |
9 |
It is evident, by looking at the findings of this study, that a high prevalence ratio of carbapenem-resistant strains is observed in clinical specimen. It is also clear that multiple genes are harbored by organisms, and such trends will continue until novel remedies are not developed for alternate therapeutic regimes. Hence, it seems to be the need of an hour to strictly implement antibiotic stewardship programs and stop over the counter sales of antibiotics; rigorous efforts are also to be made in finding drugs that has higher efficacy on the superbugs.
ACKNOWLEDGMENTS
We thank Biocare Research (I) Pvt. Ltd. and its entire staff members for their continuous support in sample pulling from their associate laboratories and providing laboratory support while carrying out this whole research.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
None.
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the manuscript.
- Antibiotic resistance. World Health Organization. https://www.who.int/news-room/fact-sheets/detail/antibiotic-resistance. 2020. Accessed May 27, 2021.
- Netikul T, Kiratisin P. Genetic Characterization of Carbapenem-Resistant Enterobacteriaceae and the Spread of Carbapenem-Resistant Klebsiella pneumonia ST340 at a University Hospital in Thailand. PLoS One. 2015;10(9):e0139116.
Crossref - Hedegaard H, Minino AM, Warner M. Drug overdose deaths in the United States, 1999-2019. NCHS Data Brief, No. 394. Hyattsville, MD: National Center for Health Statistics. 2020.
- Biggest threats and data – https://www.cdc.gov/drugresistance/biggest-threats.html, Accessed on- March 2, 2021.
- Walia K, Madhumathi J, Veeraraghavan B, et al. Establishing Antimicrobial Resistance Surveillance & Research Network in India: Journey so far. Indian J Med Res. 2019;149(2):164-179.
Crossref - Olaisen RH, Rossen LM, Warner M, Anderson RN. Unintentional injury death rates in rural and urban areas: United States, 1999-2017 pdf icon. NCHS Data Brief, No. 343.
- Singer AC, Shaw H, Rhodes V, Hart A. Review of Antimicrobial Resistance in the Environment and Its Relevance to Environmental Regulators. Front Microbiol. 2016;7:1728.
Crossref - Michael CA, Dominey-Howes D, Labbate M. The antimicrobial resistance crisis: causes, consequences, and management. Front Public Health. 2014;2:145.
Crossref - Spellberg B, Srinivasan A, Chambers HF. New societal approaches to empowering antibiotic stewardship. Journal of the American Medical Association. 2016;315(12):1229-1230.
Crossref - Laxminarayan R, Duse A, Wattal C, et al. Antibiotic resistance-the need for global solutions. Lancet Infect Dis. 2013;13(12):1057-1098.
Crossref - Aslam B, Wang W, Arshad MI, et al. Antibiotic resistance: a rundown of a global crisis. Infect Drug Resist. 2018;11:1645-1658.
Crossref - Hudzicki J. Kirby-Bauer Disk Diffusion Susceptibility Test Protocol. [ebook] American Society for Microbiology. 2016. https://asm.org/getattachment/2594ce26-bd44-47f6-8287-0657aa9185ad/Kirby-Bauer-Disk-Diffusion-Susceptibility-Test-Protocol-pdf.pdf.
- Parul S, Pallavi G, Rajni S, ArunaV, Maheshwari RK. Evaluation of a 12 Disc Test for Phenotypic Detection of β- lactamases Resistance in Gram-Negative Bacilli. Int J Curr Microbiol App Sci. 2016;5(6):105-114.
Crossref - Amjad A, Mirza IA, Abbasi SA, Farwa U, Malik N, Zia F. Modified Hodge test: A simple and effective test for detection of carbapenemase production. Iran J Microbiol. 2011;3(4):189-193.
- Omair M, Usman J, Kaleem F, Hassan A, Khalid A, Fahim Q. Evaluation of combined disc method for the detection of metallo-β-lactamase producing Gram-negative bacilli. Malays J Microbiol. 2012;8(1):21-25.
- Zee AVD, Roorda L, Bosman G, et al. Multicenter evaluation of Real time multiplex PCR for detection of Carbapenemase genes OXA-48, VIM, IMP, NDM and KPC. BMC Infectious Diseases. 2014;14:27.
Crossref - Weiner-Well Y, Rudensky B, Yinnon A, et al. Carriage rate of carbapenem-resistant Klebsiella pneumoniae in hospitalized patients during a national outbreak. J Hosp Infect. 2010;74(4):344-349.
Crossref - Moemen D, Masallat DT. Prevalence and characterization of carbapenem-resistant Klebsiella pneumoniae isolated from intensive care units of Mansoura University hospitals. Egyptian Journal of Basic and Applied Sciences. 2017;4(1):37-41.
Crossref - Endimiani A, Hujer AM, Perez F, et al. Characterization of blaKPC-containing Klebsiella pneumoniae isolates detected in different institutions in the Eastern U.S.A. J Antimicrob Chemother. 2009;63(3):427-437.
Crossref - Poirel L, Bonnin RA, Nordmann P. Genetic features of the widespread plasmid coding for the carbapenemase OXA-48. Antimicrob Agent Chemother. 2012;56(1):559-562.
Crossref - ZA Memish, A Assiri, M Almasri, et al. Molecular characterization of carbapenemase production among gram-negative bacteria in Saudi Arabia. Microbial Drug Resistance. 2015;21(3):307-314.
Crossref - Landman D, Salamera J, Singh M, Quale J. Accuracy of Carbapenem Nonsusceptibility for Identification of KPC-Possessing Enterobacteriaceae by Use of the Revised CLSI Break-points. J Clin Microbiol. 2011;49(11):3931-3933.
Crossref - Ganesh KS, Adithan C, Harish BN, Sujatha GS, Roy G, Malini A. Antimicrobial resistance in India: A review. J Nat Sci Biol Med. 2013;4(2):286-291.
Crossref
© The Author(s) 2021. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.