ISSN: 0973-7510
E-ISSN: 2581-690X
We report in this study for the first time the prevalence of multiple resistant Staphylococcus haemolyticus in clinical settings in Saudi Arabia. A total of 1060 clinical specimens of hospitalized patients were screened for the presence of S. haemolyticus in the period between September and December 2020. Primary identification of the isolates was carried out by colonial characteristics on mannitol salt agar and clumping factor test, confirmation of presumptive isolates and antimicrobial susceptibility testing was performed by Vitek® 2, while PCR was employed to detect mecA and vanA genes. A total of 20 S. haemolyticus isolates were recovered from 20 samples (blood cultures, urine, nasal swab, wound swab, groin swab, and axilla swab), 90% (P <0.001, x2) of the isolates were multiple resistant to three antimicrobial agents and more. Resistance to oxacillin was exhibited in 95% of the isolates, while none of the isolates were resistant to vancomycin and linezolid, yet resistance to rifampicin was observed in 30 % of the isolates. The findings of this study highlights the emerging trends of Staphylococcus haemolyticus as potential drug resistant pathogen in hospital settings in Saudi Arabia, which requires in depth investigation on molecular understanding on antimicrobial resistance and virulence traits of the strains.
Staphylococcus haemolyticus, nosocomial, antimicrobial resistance, mecA, vanA, Saudi Arabia
Staphylococcus haemolyticus is a coagulase-negative common inhabitant of the human skin microbiota that has been recognized as an emerging important opportunistic pathogen1. Staphylococcus haemolyticus has been commonly and frequently reported in various nosocomial infections, particularly catheter-associated bacteremia2, urinary tract infection3, diabetic foot ulcer4, device-associated meningitis5 and wound infections6.
What makes Staphylococcus haemolyticus frequently encountered in clinical settings is perhaps its ability to from biofilm and more importantly its acquisition of multiple resistance to wide range of antimicrobial drug classes including glycopeptides1. Resistance to methicillin/oxacillin has been commonly reported in Staphylococcus haemolyticus of clinical origin, and this suggests the bacterium is able to acquire and re-transfer (reservoir) the SCCmec cassette to other staphylococci7, 6. Resistance to two or more classes of antimicrobial agents such as penicillins, tetracyclines, aminoglycosides, cephalosporins, quinolones, macrolides and glycopeptides among Staphylococcus haemolyticus of clinical importance have been frequently reported in different parts of the world8, 9, 10. However, the prevalence of Staphylococcus haemolyticus in Saudi Arabia is largely unknown, only two reports of Staphylococcus haemolyticus in Saudi Arabia has been published so far. In a survey of the causative agents associated with urinary tract infection (UTI) in pregnant women in southern Saudi Arabia, Staphylococcus haemolyticus was isolated from one (5.26%) UTI case out of the 151 cases examined; the antimicrobial susceptibility patterns of that isolated were not thoroughly investigated11. Asfour et al.12 reported a case of a premature baby with an endocardaitis-associated Staphylococcus haemolyticus that has been successfully treated with daptomycin in Riyadh. Thus, to the best of our knowledge, this is the first report to highlight the prevalence and antimicrobial susceptibility profiles of Staphylococcus haemolyticus in clinical settings in Saudi Arabia, particularly in Madinah province (north western Saudi Arabia).
Isolation and identification of Staphylococcus haemolyticus
A total of 1060 clinical samples of hospitalized patients at Ohud Hospital, Madinah, northwest Saudi Arabia, comprised of nasal swabs (320), groin swabs (230), axilla swabs (320) wound swabs (30), blood cultures (20), ear swabs (20), eye swabs (10) and urine samples (20) were examined for the presence of Staphylococcus haemolyticus in the period from September to December 2020. Samples were cultured on mannitol salt agar (Oxoid, Basingstoke, UK) and Columbia agar base (Oxoid) supplemented with 5% sheep blood (Oxoid), plates were incubated under aerobic atmosphere at 37 °C for 24-48 hrs3.
Presumptive Staphylococcus haemolyticus colonies on mannitol salt agar and blood agar were initially identified by means of clumping factor test, using the Maststaph™ kit (Mast Group Ltd, Liverpool, UK), further identification and confirmation was achieved by Vitek® 2 system (BioMerieux, Marcy-I’Etoile, France)4 to distinguish between Staphylococcus hameolyticus and other coagulase-negative staphylococci of clinical relevance (e.g. S. epidermidis, S. hominis and S. lugdunensis). Antimicrobial susceptibility in terms of minimum inhibitory concentration (MIC) of confirmed Staphylococcus haemolyticus was carried out by means of Vitek® 2 system (BioMerieux), antimicrobial agents belonging to nine different classes were as follows: benzylpenicillin, oxacillin (penicillins), levofloxacin, moxifloxacin (fluoroquinolones), gentamicin, tobramycin (aminoglycosides), vancomycin, teicoplanin (glycopeptides), erythromycin (macrolides), clindamycin (lincosamides), tetracycline, tigecycline (tetracyclines), linezolid (oxazolidinones), nitrofurantoin, fuscidic acid, rifampicin and trimethoprim/sulfamethoxazole (miscellaneous agents).
PCR based detection of resistance genes (mecA and vanA)
Molecular detection of mecA and vanA genes, using primer (F 5’-AAAATCGATGGTAAAGGTTGGC-3’ / R 5’-AGTTCTGGAGTACCGGATTTGC-3’( and primer (F 5’-ATGAATAGAATAAAAGTTGCAATAC-3’ / R 5’-CCCCTTTAACGCTAATACGAT-3’) respectively was achieved by PCR, as described earlier by Abulreesh et al.13. Briefly, the bacterial DNA was extracted by the Total RNA kit according to the manufacturer’s instructions (Geneaid Biotech Ltd, New Taipei City, Taiwan). The PCR reaction was prepared by adding 1 µl of primers (mecA F, mecA R) and (vanA F, vanA R) (100 pM pH8), 1 µl of template DNA, 18 µl of dH2O and 5 µl of Ultr-Pure Taq PCR master mix (Geneaid Biotech Ltd, New Taipei City, Taiwaný). Thermal cycling was performed on Veriti 96-Well Thermal Cycler (Applied Biosystems, Massachusetts, USA) with an initial denaturation at 94 °C for 2 min, then followed by 45 cycles of denaturation at 94 °C for 20 seconds, annealing at 57 °C and 54 °C for 30 seconds for mecA and vanA respectively and elongation at 72 °C for 1 minute. A final elongation step was utilized at 72 °C for 7 min before running the samples on 1.5% gel using the M12 Complete Electrophoresis Package (Edvotek Inc, Washington D.C., USA) for 40 min at 90 voltage. The amplification bands were visualized under UV light using the ChemiDoc-It2 Imaging System (Analytik Jena GmbH, Jena, Germany).
Of the 1060 samples collected during the period of September and December 2020, from hospitalized individuals at Ohud Hospital in Madinah, only 20 samples (1.9 %) were positive for Staphylococcus haemolyticus (Table 1), this was on the basis of colonial morphology on mannitol salt agar (colonies with reddish zones), negative reaction of clumping factor test and final confirmation by Viteck® 2 system. Staphylococcus haemolyticus were more prevalent in groin swabs (45 %), followed by urine samples (20 %) (Table 1). All 20 Staphylococcus haemolyticus isolates exhibited resistance to one or more antimicrobial agent class (Table 2). Multiple resistance (resistance to three antimicrobial classes or more) was observed in 18 isolates, while one isolate from urine was resistant to benzylpenicillin and showed intermediate resistance to erythromycin, likewise one isolate obtained from groin swabs exhibited resistance to benzylpenicillin, oxacillin and tetracycline only (Table 2). Therefore, Chi-Squared test (x2) showed that the number of Staphylococcus haemolyticus exhibiting multiple resistance are significantly (P <0.001) higher than non-multiple resistant isolates (Table 2).
Table (1):
Prevalence and antimicrobial resistance patterns of Staphylococcus haemolyticus of clinical origin.
| Sample type | N/P (%) | No resistant | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| BP | Ox | G | To | Le | Mo | E | Cl | Li | Te | Va | Tet | Ti | Nit | FA | Rif | TS | ||
| Nasal swabs | 320 / 2 | 2 | 1 | 1 | 1 | 1 | 1 | 2 | 1 | 0 | 0 | 0 | 1 | 0 | 0 | 2 | 0 | 1 |
| Groin swab | 320 / 9 | 9 | 9 | 6 | 6 | 5 | 4 | 8 | 5 | 0 | 0 | 0 | 8 | 0 | 0 | 7 | 4 | 1 |
| Axilla swabs | 320 / 1 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 1 | 0 | 0 | 1 | 0 | 0 |
| Wound swabs | 30 2 | 2 | 2 | 1 | 1 | 1 | 1 | 2 | 1 | 0 | 0 | 0 | 1 | 0 | 0 | 2 | 0 | 1 |
| Blood cultures | 20 / 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 0 | 0 | 0 | 0 | 2 | 0 | 0 | 2 | 1 | 0 |
| Urine samples | 10 / 4 | 4 | 4 | 2 | 2 | 2 | 2 | 3 | 2 | 0 | 0 | 0 | 2 | 0 | 0 | 3 | 1 | 1 |
| Ear swabs | 20/0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Eye swabs | 20/0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Total | 1060/ 20 (1.9) | 20 | 19 | 12 | 12 | 11 | 10 | 17 | 10 | 0 | 0 | 0 | 15 | 0 | 0 | 17 | 6 | 4 |
N = total number of samples, P = number of samples positive for S. haemolyticus, BP = benzylpenicillin, Ox = oxacillin, G = gentamicin, To = tobramycin, Le = levofloxacin , Mo = moxifloxacin, E = erythromycin, Cl = clindamycin, Li = linezolid, Te = teicoplanin, Va = vancomycin, Tet = tetracycline, Ti = tigecycline, Nit = nitrofurantoin, FA = fucidic acid , Rif = rifampicin, TS = trimethoprim / sulfamethoxazole
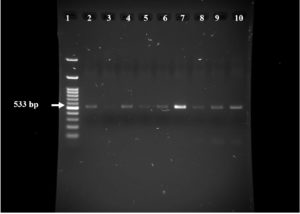
Resistance to benzylpenicillin was exhibited by all 20 (100%) isolates, while oxacillin resistance was found in 19 (95%) isolates, this was confirmed by cefoxitin screening where all isolates resistant to oxacillin were positive for cefoxitin screening test, further confirmation of oxacillin resistance was shown by the detection of mecA gene (533 bp product) (Table 2, Fig. 1) in all 19 isolates exhibiting MIC (0.5 µg or above) for oxacillin (Tables 2, 3)
Table (2):
Antibiotic resistance patterns, presence of mecA and vanA genes by PCR among 20 isolates of Staphylococcus haemolyticus of clinical origin.
Isolate number |
Origin |
Resistance profile |
No of classes |
Cefoxitin screen‡ |
Inducible clindamycin resistance‡‡ |
mecA |
vanA |
|---|---|---|---|---|---|---|---|
SHB1 |
Blood |
BP, Ox, G, To, Le, Mo, E, Tet, FA, Rif |
7 |
+ |
– |
+ |
– |
SHB2 |
Blood |
BP, Ox, G, To, Le, Mo, E, Tet, FA, Rif |
7 |
+ |
– |
+ |
– |
SHU1 |
Urine |
BP, Ox, G, To, Loe, Mo, E, Cl, Tet, FA, Rif, TS |
9 |
+ |
– |
+ |
– |
SHU2 |
Urine |
BP, Ox, G, To, Le, Mo, E, Cl, FA |
6 |
+ |
+ |
+ |
– |
SHU3 |
Urine |
BP, E (I)* |
1 |
– |
– |
– |
– |
SHU4 |
Urine |
BP, Ox, E, Tet, FA |
4 |
+ |
– |
+ |
– |
SHW1 |
Wound |
BP, Ox, G, To, Le, Mo, E, Cl, Tet, FA, TS |
8 |
+ |
– |
+ |
– |
SHW2 |
Wound |
BP, Ox, E, FA |
3 |
+ |
– |
+ |
– |
SHA1 |
Axilla |
BP, Ox, E, Cl, Tet, FA |
5 |
+ |
– |
+ |
– |
SHN1 |
Nasal |
BP, Ox, G, To, Le, Mo, E, Cl, FA, TS |
7 |
+ |
– |
+ |
– |
SHN2 |
Nasal |
BP, Ox, Le, Mo (I)*, E, Tet, FA |
5 |
+ |
– |
+ |
– |
SHG1 |
Groin |
BP, Ox, G, To, Le, Mo, E, Cl, Tet, FA, TS |
9 |
+ |
– |
+ |
– |
SHG2 |
Groin |
BP, Ox, E, Tet, FA |
4 |
+ |
– |
+ |
– |
SHG3 |
Groin |
BP, Ox, G, To, Le, Mo, E, Cl, Tet, FA, Rif, TS |
9 |
+ |
– |
+ |
– |
SHG4 |
Groin |
BP, Ox, G, To, Le, Mo, E, Cl, Tet, FA, TS |
8 |
+ |
– |
+ |
– |
SHG5 |
Groin |
BP. Ox, G, To, E, FA |
4 |
+ |
– |
+ |
– |
SHG6 |
Groin |
BP, Ox, G, To, Le, Mo (I)*, E, Cl, FA |
6 |
+ |
+ |
+ |
– |
SHG7 |
Groin |
BP, Ox, E, Tet, FA |
4 |
+ |
– |
+ |
– |
SHG8 |
Groin |
BP, Ox, G, To, Le, Mo, E, Cl, Tet, FA, Rif, TA |
9 |
+ |
– |
+ |
– |
SHG9 |
Groin |
BP, Ox, Tet |
2 |
+ |
– |
+ |
– |
Total |
P
| 19 |
2 |
19 |
0 |
†Chi-Squared (x2) to test the null hypothesis that none of the Staphylococcus haemolyticus isolates were exhibiting multiple-resistance patterns to antimicrobial agents,
BP = benzylpenicillin, Ox = oxacillin, G = gentamicin, To = tobramycin, Le = levofloxacin , Mo = moxifloxacin, E = erythromycin, Cl = clindamycin, Li = linezolid, Tet = tetracycline, Nit = nitrofurantoin, FA = fucidic acid , Rif = rifampicin, TS = trimethoprim / sulfamethoxazole; Cefoxitin screen‡ = Screened by Vitek® 2 system; Inducible clindamycin resistance‡‡ = screened by Vitek® 2 system
Fig. 1. Agarose gel electrophoresis (1% agarose) for mecA gene amplified size 533 bp as compared with 1 kbp ladder lane 1. Positive results are in lanes 2 (SHB1), 4 (SHB2), 5 (SHU2), 6 (SHW1), 7 (SHA1), 8 (SHN1), 9 (SHG) and 10 (SHG), negative result in lane 3 (SHU3).
Resistance to erythromycin and clindamycin was exhibited by 17 (85%) and 10 (50%) isolates respectively. Only 10% of the isolates exhibited positive to inducible clindamycin resistance screening (Table 2), these isolates were resistant to both clindamycin (linecosamides) and erythromycin (macrolides) (Table 2). The test was negative in about eight (40%) of the isolates exhibiting resistance to both erythromycin and clindamycin (Table 2).
None of the isolates (100%, n = 20) exhibited resistance to any of the antimicrobial agents belonging to glycopeptides (vancomycin and teicoplanin), oxazolidinones (linezolid), and nitrofurantoin (miscellaneous agents) however resistance to rifampicin (miscellaneous agents) was observed in four isolates only (20%) originating from blood cultures (two isolates), and one isolate from groin swabs and urine culture respectively (Tables 1, 2). Susceptibility to vancomycin was further confirmed by the absence of vanA gene from all 20 isolates (Table 2).
Resistance to gentamicin, tobramycin, levofloxacin, moxifloxacin, fusidic acid and trimethoprim/sulfamethoxazole were also observed in Staphylococcus haemolyticus reported in this study (Tables 1, 2). The MIC values of Staphylococcus haemolyticus isolates exhibiting resistance to more than six antimicrobial classes are shown in Table 3. As shown in the table, the majority of the isolates exhibited MIC of >= 4 µg for oxacillin, with only one isolate from exhibited an MIC of 0.5 µg and another one with MIC of 1µg, the MIC for oxacillin resistant staphylococci is >0.25 µg.
Table (3):
MIC of antimicrobial susceptibility patterns of Staphylococcus haemolyticus resistant to five and more antimicrobial classes.
| MIC of resistance profile | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Isolate number | Origin | PB | Ox | G | To | Le | Mo | E | Cl | Tet | FA | Rif | TS |
| SHB1 | Blood | ≥0.5 | ≥4 | ≥16 | 8 | ≥8 | 4 | ≥8 | ≤0.25 (S) | 2 | 8 | ≥32 | ≤10 (S) |
| SHB2 | Blood | ≥0.5 | ≥4 | ≥16 | ≥16 | ≥8 | 4 | ≥8 | ≤0.25 (S) | 2 | 8 | ≥32 | ≤10 (S) |
| SHW1 | Wound | ≥0.5 | ≥4 | ≥16 | ≥16 | ≥8 | 4 | ≥8 | ≥8 | 2 | 8 | ≤0.5 (S) | ≥320 |
| SHU1 | Urine | ≥0.5 | ≥4 | 8 | 2 | ≥8 | 4 | ≥8 | ≥8 | 2 | ≥32 | ≥32 | ≥320 |
| SHU2 | Urine | ≥0.5 | ≥4 | 4 | 2 | ≥8 | 4 | ≥8 | ≤0.25 | £1 (S) | ≥32 | ≤0.5 (S) | ≤10 (S) |
| SHN1 | Nasal | ≥0.5 | ≥4 | ≥16 | 8 | ≥8 | 4 | ≥8 | ≥8 | ≤1 (S) | 8 | ≤0.5 (S) | ≥320 |
| SHN2 | Nasal | ≥0.5 | 0.5 | ≤0.5 (S) | ≤1 (S) | 4 | 1 (I) | ≥8 | ≤0.25 (S) | ≥16 | ≥32 | ≤0.5 (S) | ≤10 (S) |
| SHA1 | Axella | ≥0.5 | ≥4 | ≤0.5 (S) | ≤1 (S) | ≤0.12 (S) | ≤0.25 (S) | ≥8 | ≥8 | ≥16 | 4 | ≤0.5 (S) | 20 (S) |
| SHG1 | Groin | ≥0.5 | ≥4 | ≥16 | 8 | ≥8 | 4 | ≥8 | ≥8 | 2 | 8 | ≤0.5 (S) | ≥320 |
| SHG3 | Groin | ≥0.5 | ≥4 | 8 | 2 | ≥8 | 4 | ≥8 | ≥8 | 2 | ≥32 | ≥32 | ≥320 |
| SHG4 | Groin | ≥0.5 | ≥4 | ≥16 | 8 | ≥8 | 2 | ≥8 | ≥8 | 2 | 16 | ≤0.5 (S) | ≥320 |
| SHG6 | Groin | ≥0.5 | 1 | 2 | ≤1 | 4 | 1 (I) | ≥8 | ≤0.25 | ≥16 | ≤0.5 (S) | ≤0.5 (S) | ≤10 (S) |
| SHG8 | Groin | ≥0.5 | ≥4 | 4 | 2 | ≥8 | 4 | ≥8 | ≥8 | 2 | ≥32 | ≥32 | ≥320 |
S = susceptible, I = intermediate, BP = benzylpenicillin, Ox = oxacillin, G = gentamicin, To = tobramycin, Le = levofloxacin , Mo = moxifloxacin, E = erythromycin, Cl = clindamycin, Tet = tetracycline, FA = fucidic acid , Rif = rifampicin, TS = trimethoprim / sulfamethoxazole
Epidemiological studies showed that Staphylococcus haemolyticus comes second to Staphylococcus epidemidis as the most frequently encountered coagulase-negative staphylococci associated with nosocomial infections1. Emerging Staphylococcus haemolyticus has been associated with a wide range of nosocomial infections such as bacteremia, UTI, and wound infection6, 14, 15, 16, the results obtained in this study report for the first time in Saudi Arabia the incidence of highly resistant Staphylococcus haemolyticus in blood cultures, UTI, wound infections among hospitalized patients, despite the low incidence of Staphylococcus haemolyticus in this study, the majority of the isolates exhibited multiple resistance to three and more antimicrobial classes. Although Staphylococcus haemolyticus is part of the human resident microbiota, the highly resistant isolates found in the nasal cavity and on the skin (groin) of hospitalized patient warrant attention in the handling of these patients given that they may be a source for the dissemination of these multiple resistant strains within the hospital environment which may be problematic since Staphylococcus haemolyticus is highly adaptable to hospital environments particularly on clinical devices1, 17.
In this study 90% of the Staphylococcus hameolyticus isolates exhibited multiple resistance (to three and more antimicrobial classes) (P <0.001, x2). Multiple resistant Staphylococcus haemolyticus of clinical origin are increasingly encountered worldwide3, 6, 8, 14, 15, 16, 18. We observed that 100% and 95% of the isolates were resistant to benzylpenicillin and oxacillin respectively, this result was also supported by cefoxitin screening, which is widely accepted as a surrogate for the detection methicillin resistance in staphylococci19, as well as the detection of mecA gene by PCR in these isolates. Similar results were reported elsewhere8, the high prevalence in pinicillins resistance in Staphylococcus haemolyticus is perhaps explained by the high diversity of staphylococcal cassette chromosome mec (SCCmec) element that is carried by Staphylococcus haemolyticus strains7, 8, 20, this high diversity of the Staphylococcus haemolyticus SSCmec genes suggest that the bacteria is an important reservoir for the dissemination of these genes among other staphylococci within health care settings20, 21.
Resistance to levofloxacin, moxifloxacin (Fluoruquinolones) gentamicin, tobramycin (aminoglycosides), erythromycin (macrolides), clindamycin (lincosamides) and tetracycline (tetracyclines) was observed in 50% or more of the Staphylococcus haemolyticus isolates. Similar observations were reported from India14, Brazil8, Taiwan6, Indonesia16, Jordan18, Iraq3, Thailand15 and Poland22. Resistance to fluoroquinolones, aminoglycoside, macrolides, lincosamides and tetracyclines in Staphylococcus haemolyticus is not surprising given the remarkable ability of the bacteria to receive the genes gyrA and parC (fluoroquinolones), aacA and aphD (aminoglycosides), ermA and ermC (macrolides), linA and lnuA (lincosamides) and tetK and tetM (tetracyckines) that mediate resistance to these antimicrobials17, 22. In addition, Staphylococcus hameolyticus strains can also carry fusB, drfC and drfG that mediate resistance to fusidic acid and trimethoprim respectively17, resistance to fusidic acid and trimethoprim/sulfamethoxazole were also observed in our isolates. Inducible clindamycin resistance was detected in two isolates only, detection of macrolides-lincozamides-streptogramin B (MLSB) phenotypes is important as it suggest the ability of these phenotypes to develop resistance to clindamycin during the therapy of the patients23. The ability of Staphylococcus haemolyticus to possess various resistance genes, together with their ability to have a diverse SSCmec genes render them to be remarkable in developing multiple resistant phenotypes that could be a source of disseminating resistance genes of different antimicrobials to other staphylococci via horizontal gene transfer in biofilm formations within hospital environment and that may lead to increase in nosocomial infections that are difficult to handle.
Multiple-resistance was also noted in Staphylococcus haemolyticus recovered from the skin, i.e. groin (95%, n = 9), axial (100%, n =1) and nasal cavity (1—%, n = 2) of hospitalized patients, some of the these isolates were resistant to about seven or eight antimicrobial classes, this is alarming given the possibility that these patients may act as a source of infection to health care personnel or source of contamination of hospital environment with these multiple resistant strains. The carriage of drug resistant Staphylococcus haemolyticus in the nares and skin of healthy as well as hospitalized individuals has been reported24.
Currently, vancomycin, linezolid and rifampicin are the treatment of choice for Staphylococcus haemolyticus infections14, 21, our results showed that resistance to vancomycin and linezolid do not exist, particularly with the absence of vanA gene (0% in this study) which in agreement with what reported worldwide, however, resistance to rifampicin exhibited by 30% of the isolates reported in this study is alarming, emerging Staphylococcus haemolyticus resistant to rifampicin have been reported7, Resistance to glycopeptides (vancomycin and teicoplanin) remains rarely detected in Staphylococcus haemolyticus3, 6, 14, 16, 18. Despite the total absence of vancomycin/teicoplanin, linezolid resistant isolates and the apparently low prevalence of rifampicin resistance among Staphylococcus haemolyticus reported in this study, wide surveillance of susceptibility patterns of these drugs among Staphylococcus haemolyticus of clinical origin is mandatory, since the development of resistance to these drugs may exist at low levels17.
In conclusion the results reported in this study shows for the first time the prevalence of multiple resistant Staphylococcus haemolyticus within clinical sittings in Saudi Arabia. The observed prevalence is still low, nonetheless, the majority of the isolates were multiple resistant and this may pose significant health threats. Staphylococcus haemolyticus adapt well to hospital environment and their ability to serve as recipient and/or donor of genes encoding for antimicrobial resistance may promote spreading of antimicrobial resistance to other nosocomial staphylococci, as well as a source of infection to hospital personnel. Therefore, this study on Staphylococcus haemolyticus prevalence highlighted the need for further investigation about the molecular basis of virulence properties and the pathogenesis of Staphylococcus haemolyticus of clinical origin in Saudi Arabia and their linkage to resistance genes.
ACKNOWLEDGMENTS
None.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
None.
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the manuscript.
- Czekaj T, Ciszewski M, Szewczyke EM. Staphylococcus haemolyticus – An emerging threat in the twilight of the antibiotics age. Microbiol. 2015;161:2061-2068.
Crossref - Argemi X, Hansmann Y, Prola K, Prevost G. Coagulase-negative staphylococci pathogenomics. Int. J. Molec. Sci. 2019;20:1215.
Crossref - Alhusain MT, Jaafar FN, Hassan AAA. Isolation and characterization of Staphylococcus haemolyticus from urinary tract infection. EurAsian J. Biosci. 2020;14: 1909-1913.
- Eltwisy HO, Abdel-Fattah M, Elsisi AM, Omar MM, Abdelmoteleb AA, El-Mokhtar MA. Pathogenesis of Staphylococcus haemolyticus on primary human skin fibroblast cells. Virulence. 2020;11:1142-1157.
Crossref - Vijayan P, Srinivas D, Siddaiah N, Bahubali VKH. Device-associated meningitis by linezolid-resistant Staphylococcus haemolyticus in a vancomycin-hypersensitive patient. J. Neurosci. Rural Pract. 2019;10:718-720.
Crossref - Chang PH, Liu TP, Hunag PY, et al. Clinical features, outcomes, and molecular characteristics of an outbreak of Staphylococcus haemolyticus infection, among a mass-burn casualty patients group, in a tertiary center in northern Taiwan. J. Microbiol. Immunol. Infect. 2018;51:847-855.
Crossref - Hosseinkhani F, Buirs MT, Jabalameli F, Emaneini M, van Leeuwen WB. High diversity in SCCmec elements among multidrug-resistant Staphylococcus haemolyticus strains originating from paediatric patients; characterization of a new composite island. J. Med. Microbiol. 2018;67:915-921.
Crossref - Barros EM, Ceotto H, Bastos MCF, dos Santos KRN, Giambiagi-deMarval M. Staphylococcus haemolyticus as an important hospital pathogen and carrier of methicillin resistance genes. J. Clin. Microbiol. 2011;50:166-168.
Crossref - Silva PV, Cruz RS, Keim LS, et al. The antimicrobial susceptibility, biofilm formation and genotypic profiles of Staphylococcus haemolyticus from bloodstream infections. Mem Inst Oswaldo Cruz Rio de Janeiro. 2013;108:812-816.
Crossref - Manoharan M, Sistla S, Ray P. Prevalence and molecular determinants of antimicrobial resistance in clinical isolates of Staphylococcus haemolyticus from India. Microb. Drug Resist. 2021;27:501-508.
Crossref - Alshabi AM, Alsharrani MS, ALkahtani SA, Akhtar MS. Prevalence of urinary tract infection and antibiotic resistance pattern in pregnant woen, Najran region, Saudi Arabia. Afri. J. Microbiol. Res. 2019;13: 407-413.
Crossref - Asfour SS, Aljobair F, Hammami I, Abdelrahim A, Al-Mouqdad MM. Successful treatment of Staphylococcus haemolyticus endocarditis inextermely premature baby with daptomycin. J. Pediatr. Neonat. Individ. Med. 2018;8:e080109.
Crossref - Abulreesh HH, Organji SR, Osman GEH, Elbanna K, Almalki MHK, Ahmad I. Prevalence of antibiotic resistance and virulence factors encoding genes in clinical Staphylococcus aureus isolates in Saudi Arabia. Clin. Epidemiol. Glob. Health. 2017;5: 196-202.
Crossref - Chaudhury A, Kumar AG. In vitro activity of antimicrobial agents against oxacillin resistant staphylococci with special reference to Staphylococcus haemolyticus. Ind. J. Med. Microbiol. 2007;25:50-52.
Crossref - Teeraputon S, Santanirand P, Wongchai T, et al. Prevalence of methicillin resistance and macrolide-lincosamide-streptogramin B resistance in Staphylococcus haemolyticus among clinical strains at a tertiary-care hospital in Thailand. New Microbe New Infect. 2017;19:28-33.
Crossref - Suhartono S, Hayati Z, Mahmuda M. Distribution of Staphylococcus haemolyticus as the most dominant species among staphylococcal infections at the Zainoel Abidin Hospital in Aceh, Indonesia. Biodiversitas. 2019; 20: 2076-2080.
Crossref - Pain M, Hjerde E, Klingenberg C, Cavanagh JP. Coparative genome analysis of Staphylococcus haemolyticus reveals key to hospital adaptation and pathogenicity. Front. Microbiol. 2019;10:2096.
Crossref - Al-Tamimi M, Abu-Raideh J, Himsawi N, Khasawneh A, Hawamdeh H. Methicillin and vancomycin resistance in coagulase-negative staphylococci isolated from the nostrils of hospitalized patients. J. Infect. Develop. Countries. 2020; 14: 28-35.
Crossref - Fernandes CJ, Fernandes LA, Collingnon P. Cefoxitin resistance as a surrogate marker for the detection of methicillin-resistant Staphylococcus aureus. J. Antimicrob. Chemother. 2005;55: 506-510:
Crossref - Szczuka E, Krajewska M, Lijewska D, Bosacka K, Kaznowski A. Diversity of staphylococcal cassette chromosome mec elements in nosocomial multiresistant Staphylococcus haemolyticus isolates. J. Appl. Genet. 2016;57:543-547.
Crossref - Azimi T, Mirzadeh M, Sabour S, Nasser A, Fallah F, Pourmand MR. Coagulase-negative staphylococci (CoNS) meningitis: a narrative review of the literature from 2000 to 2020. New Microbe New Infect. 2020;37: 100755.
Crossref - Szemraj M, Czekaj T, Kalisz J, Szewczyk EM. Differences in distribution of MLS antibiotics resistance genes in clinical isolates of staphylococci belonging to species: S. epidermidis, S. hominis, S. haemolyticus, S. simulans and S. warneri. BMC Microbiol. 2019;19: 124.
Crossref - Lewis II JS, Jorgensen JH. Inducible clindamycine resistance in staphylococci: should clinicians and microbiologists be concerned? Clin. Infect. Dis. 2005;40:280-285.
Crossref - Kateete DP, Asiimwe BB, Mayanja R, Najjuka CF, Rutebemberwa E. Species and drug susceptibility profiles of staphylococci isolated from healthy children in Eastern Uganda. PLoS One. 2020;15: e0229026.
Crossref
© The Author(s) 2021. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.