ISSN: 0973-7510
E-ISSN: 2581-690X
Shimoga region of Western Ghats is one of the richest biodiversity hotspots in India, which receives an average rainfall of 2869 mm and provides congenial atmosphere for mushrooms. In the present study eleven mushrooms were collected from Siddapura, Theertha halli and Agumbe forests of Shimoga district during monsoon (June-September 2013). The samples were labeled as WGM-1 to WGM-11 (Western Ghats Mushroom). These mushrooms were identified by ITS (Internal Transcribed Spacer) region sequence homology available at National Centre for Biotechnological Information (NCBI) Gen Bank as Lentinus squrossulus (WGM-1), Pleurotus salmoneostramenius (WGM-2), Termitomyces sp.-1 (WGM-3), Termitomyces sp.-2 (WGM-4), Leucoagaricus purpureolilacinus (WGM-5), Tricholosporum porphyrophyllum (WGM-6), Agrocybe pediades (WGM-7), Leucocoprinus birnbaumii (WGM-8), Podoscypha petalodes (WGM-9), Xylaria sp. (WGM-10) and Antrodia serialis (WGM-11). Among the eleven species, L. squrossulus, P. salmoneostramenius, Termitomyces sp.1, Termitomyces sp.2, were edible mushrooms. This study unravels the abundance of the mushroom species in parts of Shimoga region of Western Ghats area.
Western Ghats, Mushroom, ITS region, Molecular identification.
Ethno mycology refers to the relationship between Man and fungi. Human association with mushroom fungi is from the time immemorial as wild mushrooms are picked up and eaten by man kind since antiquity. Thousands of years ago, fructifications of higher fungi have been used as a source of food due to their attractive flavor and taste (Rai, 1997). Mushrooms are efficient degraders of lingo-celluloses; hence they play a vital role in biodegradation. Some mushrooms have pharmaceutical value such as antimicrobial, anticancer, antioxidants etc., (Chadha and Sharma, 1995).
Mushrooms grow during rainy season when climatic conditions are cool and humid. Diversity of mushroom species varies according to the ecosystem and their habitat. Shimoga region is in the heart of Western Ghats, which is one of the hot-spots of biodiversity in India. This district comes under south-eastern transitional zone and receives an average annual rainfall of 2869 mm (Bhat, et al., 2012) making an ideal habitat for blooming variety of mushrooms. Defining the number and kinds of fungi on earth has been a point of discussion and several studies have focused on enumerating the world fungal diversity (Crous et al., 2006). Only a fraction of total fungal wealth has been subjected to scientific scrutiny and mycologists continue to unravel the unexplored and hidden wealth. One third of fungal diversity of the globe exists in India and of this only 50 % are characterized so far (Manoharachary et al., 2005).
Mushrooms are ephemeral and disappear within a day. Therefore, documentation of mushrooms needs constant survey during appropriate season. Mushrooms can be identified based on their morphological and molecular characters. The Phenotypic characters include the shape, size, texture, colour and odour of the fruiting body. Molecular tools such as 18S rRNA/ITS (Internal transcribed spacer) region can be used to identify mushrooms at any stage (Rajarathnam and Thiagarajan., 2012). Several wild fungi were documented elsewhere and identified using ITS sequence (Oyetayo., 2012). In this study, we report molecular characterization of 11 mushrooms documented from Shimoga region of Western Ghats.
Collection and documentation of mushrooms
Field survey was made to document the wild mushrooms in forest area of Shimoga district (Shimoga, Siddapura, Agumbe and Theertha halli) of Karnataka from June to September 2013. The survey was carried out with the help of information provided by tribal communities like Adivasis, Halakki vokkals and Siddis in the locality during the visits as they were familiar with mushroom types and season of their appearance. The mushroom samples were collected in paper bags and field notes like date, weather condition, abundance, habitat and phenotypic characters were recorded.
Molecular characterization
Genomic DNA Isolation
Total genomic DNA from cap tissue was extracted using CTAB method (Sambrook et al., 1989). The DNA obtained was stored in Tris-EDTA (10:1) buffer at -20°C. The DNA concentration was measured using nano drop (Eppendorff) and then PCR amplification was carried out in 40 µl reaction mixture containing 4.0µl of 10 X PCR Taq. Buffer, 4.0 µl of 10 mM dNTP’s mix, 2.0 µl of ITS primers (ITS1-5’TCCGTAGGTGAACCTGCGG3’and ITS4-5’ TCCTCCGCTTATTGATATGC 3’, 0.6 µl of Taq. DNA polymerase, 2.0 µl of Template DNA (~50 ng) and 27.4 µl of sterile distilled water.
PCR amplification and elution
The PCR reaction was carried out in a Thermal Cycler (Applied Biosystems). Programmed as initial denaturation at 96°C for 3min, 40 cycles of denaturation of 94°C for 1 min, annealing at 60°C for 30 sec and extension at 72°C for 1 min and final extension at 72°C for 10 min. The amplified products were separated by agarose gel electrophoresis. The gel was visualized under UV light and documented using Alpha Innotech Gel documentation unit. The amplified product was eluted using Gene JET™ Gel Extraction Kit (Thermo Scientific) following manufacturer protocol. The eluted product was cloned into pTZ57R/T cloning vector using Ins T/A clone PCR product cloning kit [MBI, Fermentas Life Sciences, USA (#K1214)] after determining the appropriate vector: insert ratios (Sambrook et al., 1989). The ligation reaction was performed in a 10µl reaction volume at 16°C overnight. The ligated product was transformed in to E. coli (DH5á) cells using heat shock method (Sambrook et al., 1989) and plated on Luria Berton (LB) agar medium containing antibiotic (ampicillin, 100 µg/ml). The recombinant clones were initially screened by blue white selection, followed by colony PCR using M13 forward and reverse primers (Sambrook et al., 1989). The transformed colony was multiplied in LB broth containing 100 µl ampicillin for overnight and the recombinant plasmid was isolated using GenEluteTM HP Plasmid MiniPrep Kit (Sigma, USA) following the manufactures protocol. The isolated plasmid was sequenced at Sci Genome Labs Private Ltd. Kerala, INDIA using M13 forward and reverses primers.
Sequence analysis and homology search
Sequence results were analysed with Vec Screen online software from NCBI for removing the vector contamination. Forward and reverse primer sequences were checked against each other by generating the reverse complement of the “reverse” sequence using Fast PCR Professional (Experimental test version 5. 0. 83) and aligning it with the “forward” sequence with the help of CLUSTAL W Multiple Sequence Alignment Programme using the online software SDSC Biology Workbench (San Diego Supercomputer Center). The full length gene homology search was performed with blast programme of National Centre for Biotechnology Information (NCBI) (http://www.ncbi.nlm.nih.gov/BLAST) (Altschul et al., 1990).
Mushrooms have been used as food and medicine by the ancient Egyptian, Greek, Roman and Chinese civilizations. The diversity of fungi and their natural beauty occupy prime place in the biological world and India has been a cradle for these species. To understand the occurrence, abundance, locality or habitat and edibility of the mushrooms, traditional knowledge of the tribal folks was very much essential, therefore, we sought villager’s knowledge and accompanied them during the survey for collection of mushrooms. Field information of the mushroom species was recorded during collection (Table 1). Then the samples were labeled as WGM-1, WGM-2, WGM-3, WGM-4, WGM-5, WGM-6, WGM-7, WGM-8, WGM-9, WGM-10 and WGM-11. The habitats were varied from soil to tree stump and leaf litter as it is a versatility of the forest ecosystem which provides diversified niche for different types of mushrooms under same umbrella. Srivastava et al. (2011) collected mushrooms belonging to four species of Termitomyces namely, Termitomyces heimii, Termitomyces clypeatus, Termitomyces mammiformis and Termitomyces microcarpus from Gorakhpur forest division and characterized by different morphological traits such as shape of stipe, pileus, margin of fruit body, colour of fruit body, gills, flesh, annulus, pseudorrhiza and spore print. Dwivedi et al., (2012) reported 52 mushroom species from Amarkantak Biosphere Reserve of Madhya Pradesh. Those mushrooms belonged to different genera out of which only 14 mushroom samples were identified up to species level.
Table (1):
Field characters of mushrooms collected from Western Ghats region of Karnataka.
Sl. No |
Mushroom collected |
Date of collection |
Place of collection |
Vernacular name |
Habitat |
|---|---|---|---|---|---|
1 |
WGM-1 |
22/09/2013 |
Siddapura, shimoga district. |
Mara anabe |
Tree |
2 |
WGM -2 |
18/09/2013 |
TheerthahalliShimoga district. |
Henneanabe |
Branch of tree |
3 |
WGM -3 |
17/09/2013 |
Siddapurashimogha district |
Beruanabe |
Soil |
4 |
WGM -4 |
22/09/2013 |
Agumbe forest of shimogha district |
Huthadaanabe |
Termite mound |
5 |
WGM -5 |
22/10/2013 |
Siddapura, of shimogha district |
Thuppadaanabe |
Soil |
6 |
WGM -6 |
22/10/2013 |
Agumbe forest of shimogha district |
Nayeeanabe |
Humus |
7 |
WGM -7 |
17/08/2013 |
Siddapurashimogha district |
Chikkaanabe |
Soil |
8 |
WGM -8 |
19/08/2013 |
Agumbeforest, shimogha district |
Huchaanabe |
Soil |
9 |
WGM -9 |
10/10/2013 |
Siddapura, shimogha district |
Mara anabe |
Black soil |
10 |
WGM -10 |
28/09/2013 |
Theerthahallishimogha district. |
Kaddianabe |
Humus soil |
11 |
WGM -11 |
26/08/2013 |
Agumbe forest shimogha district |
Mara anabe |
Decay wood |
WGM-Western Ghat Mushroom
Molecular tools provide more accurate methods for identification of both prokaryotes and eukaryotes. The eleven mushrooms were identified up to species level by using ITS region sequence. PCR amplification of genomic DNA of the 11 mushrooms in this study yielded amplified product sizes varying from 455 bp to 773 bp (Table, 2) which were corresponding to almost full length gene sequence of ITS. The sequence homology of the11 species ranged from 83-99% (Table, 2) when aligned with the sequences present in NCBI Gen Bank. WGM-1 has 98% homology with Lentinus squrossulus, WGM-2 has 99 % homology with Pleurotus salmoneostramenius (Fig.1). WGM-3 has 99% homology with Termitomyces sp.WGM-4 showed 97% homology with Termitomyces sp. WGM-5 showed 90% homology with Leucoagaricus purpureolilacinus, WGM-6 with 83% homology with Triholosporum porphyrophyllum, WGM-7 with 99% homology for Agrocybe pediades, WGM-8 with 99% homology for Leucocoprinus birnbaumii, WGM-9 showed 99% homology with Pedoscypha petalodes, WGM-10 had 89% homology with Xylaria sp. and WGM-11 had 99% homology with Antrodia serialis.
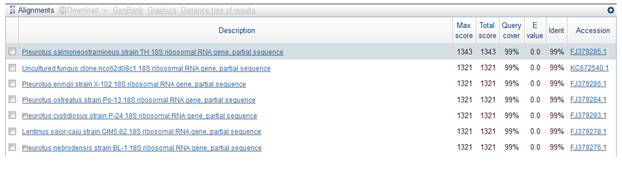
Fig-1, Full length sequence and homology search of Pleurotus salmoneostramenius
(The above figure is representative of molecular identification)
Table (2):
List of Mushroom species identified by ITS region sequence and their DNA amplicon size.
Sl. No |
Mushroom Species Designation |
Size of amplified DNA(bp) |
Mushroom species identified |
Blast search homology (%) |
|---|---|---|---|---|
1 |
WGM-1 |
684 |
Lentinus squrossulus |
97 |
2 |
WGM-2 |
737 |
Pleurotus salmoneostramenius |
99 |
3 |
WGM-3 |
735 |
Termitomyces sp. |
99 |
4 |
WGM-4 |
773 |
Termitomyces sp. |
99 |
5 |
WGM-5 |
716 |
Leucoagaricus purpureolilacinus |
94 |
6 |
WGM-6 |
719 |
Trycholos porumporpyrophyllum |
87 |
7 |
WGM-7 |
729 |
Agrocybepediades |
99 |
8 |
WGM-8 |
764 |
Leucocoprinousbimbaumii |
99 |
9 |
WGM-9 |
693 |
Podoscyphapetalodes |
99 |
10 |
WGM-10 |
455 |
Xylaria sp. |
89 |
11 |
WBG-11 |
735 |
Antrodiaserialis |
95 |
WGM-Western Ghat Mushroom
The ITS region/ 18S rRNA gene sequence are the most widely used techniques in molecular phylogenetics of mushroom as these sequences are conserved irrespective of life history and evolution (Rajaratnam and Thiagarajan, 2012). An edible mushroom from the Theertha halli forest area of Western Ghats of Shivamoga district of Karnataka was identified using ITS region of ribosomal DNA sequences as Termitomyces sp. (Earanna et al., 2013). Our study documented the abundance of the 11 mushroom flora from the Western ghats region (Shimoga) of Karnataka.

Fig-2: Mushrooms collected from Shimoga regions of Karnataka. a) L. squrossulus, b) P. salmoneostramenius, c) Termitomyces sp., d) Termitomyces sp. e) L. purpureolilacinus, f) T. porphyrophyllum, g) Agrocybe pediades, h) Leucocoprinous bimbaumii, i) Podoscypha petalodes, j) Antrodia serialis, k) Xylaria sp.
- Altschul, S. F., Gish, W., Miller, W., Myers, E. W. and Lipman, D.J 1990. Basic Local Alignment Search Tool. J. Mol. Biol., 215: 403-410.
- Bhat, S. P., Chandran S. M. D. and Ramachandra, T. V. Status of Forests in Shimoga, Central Western Ghats, Lake., 2012; 7: 1-10.
- Chadha, K. L. and Sharma, S. R., 1995. Mushroom research in India-History, Infrastructure and Achievements. In: KL Chadha (Eds.). Adv. Horticulture, MPH (Malhotra Publishing House): New Delhi: pp. 1-8.
- Crous, P. W. How many species of Fungi are there in tip of Africa. Stu. Mycology., 2006; 55:13-17.
- Dwivedi, S., Tiwari, M. K., Chauhan, U. K., and Pandey, A. K., Biodiversity of mushrooms of Amarkantak Biosphere Reserve forest of Central India. Int. J Pharm. Life Sci., 2012; 3(1): 1363-1367.
- Earanna, N., Nandini, K., Ganavi, M. C., Sajeevan, R. S., and Karaba N. Nataraja, Molecular and nutritional characterization of an edible mushroom from the Western Ghats of Karnataka, India- A new report. Mushroom Res., 2013; 22(1): 1-7.
- Manoharachary, C., Sridhar, K. R., Singh., Adholeya, T. S., Suryanarayanan, S., Rawat. and Johri, B. N., Fungal Biodiversity: Distribution, Conservation and Prospecting of Fungi from India. Curr. Sci., 2005; 89(1): 58-71.
- Oyetayo, V. O., Wild Termitomyces species collected from Ondo and Ekiti States are more related to African speciesas revealed by ITS region of rDNA. Sci. World J., 2012; 1:1-5.
- Rai, R. D., Medicinal mushroom: In: Advances in mushroom biology and production. Mushroom Society of India. NRCM, Solan, HP., 1997; 355-368.
- Rajaratnam, S. and Thiagarajan, T., Molecular characterization of wild mushroom. Eur. J Biol., 2012; 2(2): 369-373.
- Srivastava, B., Dwivedi, A. K. and Pandey, V. N., Ethnobotanical survey, distribution and utilization of Termitomyces species in Gorakhpur forest division. Plant Sci. Feed., 2011; 1(3): 28-33.
- Sambrook, J., Fritsch, F. F. and Maniatis, T., 1989. Molecular cloning: A laboratory manual, Cold Spring Harbour Laboratory Press, Cold Spring Harbor, NY, pp. 931-957.
© The Author(s) 2016. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.


