ISSN: 0973-7510
E-ISSN: 2581-690X
A thermophilic bacterial strain having the ability to produce L-tryptophan enzymatically was isolated and identified from a less explored hot spring of West Bengal. The isolate was identified using polyphasic taxonomic approach as a strain of Bacillus licheniformis. Initially, the 16S rRNA gene and later the whole genome of the isolate was sequenced and submitted to the NCBI Gene Bank for future reference. The isolate showed considerable tryptophan synthase activity and may be a potential candidate for mass production of L-tryptophan by enzymatic means.
L-tryptophan, Tryptophan Synthase, Bacillus licheniformis, Paniphala Hot Spring
L-tryptophan, an essential aromatic amino acid, is widely used in food, chemical and pharmaceutical industries. Demand for L-tryptophan is increasing day by day. It is effectively used in treating schizophrenia, insomnia, and alcoholism1-3; for the large-scale production of many small peptide antibiotics and a number of value-added products like serotonin,4 melatonin5 etc. In addition to that, immunomodulatory effects of tryptophan and tryptophan metabolites (both host and gut microflora derived) have been reported from various preclinical studies.6
Industrially L-tryptophan is produced either by microbial fermentation from cheap and readily available carbon and nitrogen sources or by enzymatic means using intermediates of the chorismate pathway like indole–glycerol phosphate or indole as substrate (see Figure 1). Tryptophan synthase, the key enzyme for enzymatic production of L-tryptophan, is a bifunctional tetrameric enzyme that catalyses the conversion of indole 3-glycerol phosphate and serine to tryptophan and water in the last two steps of the biosynthetic pathway in most of the producer organisms.7 In bacteria, the two reactions are catalysed by separate α and β-subunits, which combine to form a stable (αβ)2-heterotetramer.8,9 Tryptophan synthase is considered as a ‘Biocatalyst Extraordinaire’, due to its ability to synthesise L-tryptophan and a wide array of non-canonical amino acids.10 Apart from that, tryptophan synthase, the key biosynthetic enzyme of the tryptophan biosynthesis pathway is recently used as a drug target to develop specific drugs for controlling various pathogenic bacteria.11
Figure 1. General scheme of the tryptophan synthase reaction. Indole, which is formed from IGP by the α-subunits is channeled to β-subunits, which synthesise tryptophan from indole and serine (Kriechbaumer et al.9)
In the present study, we have isolated, purified, and identified a thermophilic bacterial strain from the Paniphala hot spring water which is situated near Baraboni area of Asansol, West Bengal, India (23°45′33″N, 86°58′54″E) (Figure 2).
Isolation of thermophilic tryptophan producing bacteria
Water samples (temperature- 60.2°C, pH-6.9) were collected from the hot springs of Paniphala, West Bengal in sterile screw capped bottles and were passed through sterile bacterial filter disc using syringe filters. The bacterial cells collected on the filter pad were added directly in sterile Davis & Mingioli’s (DM) broth medium containing KH2PO4 0.3 g, K2HPO4 0.7 g, tri sodium citrate 0.05 g, MgSO4.7H2O 0.02 g, (NH4)2SO4 0.1 g, glucose 1%, distilled water 100 mL, pH 7.2 and incubated at 55°C for visible growth. The culture broth was serial diluted and plated on DM agar medium and incubated at 55°C. After 24 hrs incubation, colonies with good growth were isolated and purified by four-way streaking on sterile DM agar medium.
Genomic DNA isolation and sequencing of 16s rRNA gene of isolate 3C1
A Genomic DNA kit (PureLink™ Genomic DNA Mini Kit Catalog number: K182001) was used for genomic DNA isolation of B. licheniformis 3C1. The purity of the isolated Genomic DNA samples was tested by agarose gel electrophoresis. The isolated genomic DNA sample was purified using the HiMedia Genomic DNA purification Kit and was sent to National Centre for Microbial Resource-NCCS, Sait Trinity, Pashan, Pune- 411021.
Preparation of cell lysate
B. licheniformis 3C1 cells were inoculated into 20-mL volumes of the modified M9 broth from a slant culture and incubated at 37°C and 140 rpm for 16 h. The broth culture then centrifuged at 10000g for 5 min at 4°C. The supernatant was kept in a separate tube and the cell pellet was suspended in 50 mM Tris-HCl buffer (pH 8.2), washed through centrifugation, and resuspended in the same buffer. The suspended cells are disrupted using an ultrasonic homogenizer. The lysates were clarified by centrifugation at 12000g for 10 min at 4 °C. The supernatants and clarified cell lysates were used to measure tryptophan synthase activity.
Determination of optimum temperature and pH of the growth medium
To determine the optimum temperature for maximum growth of the bacterial isolate 3C1 was grown in 20 ml portion of sterile DM medium in conical flask in triplicate set at various growth temperature ranging from 30 to 70°C in shaking (200 rpm) condition in an incubator shaker after inoculating the medium with 2% v/v of aqueous suspension of washed bacterial cells (5 x 107 cells/ml) from 24 hrs old slant culture.
To determine the optimum medium pH for maximum growth of the isolated bacterial strain, 20 ml portion of DM medium supplemented with indole and serine were taken in 100 ml conical flasks in triplicate set for each pH ranging from 5 to 9. Medium pH was adjusted using 1N HCl or 1N NaOH as required. After sterilization, each flasks was inoculated with 2% v/v of aqueous suspension of washed bacterial cells (5 x 107 cells/ml) from 24 hrs old slant culture and were incubated in an incubator shaker at 60°C Growth was measured after 24, 48 and 72 hrs of incubation in a UV-VIS spectrophotometer at 600 nm.
Assay of tryptophan synthase
Ability of the isolated thermophilic strain for the enzymatic production of L-tryptophan was tested in an assay mixture containing cell extract as a source of tryptophan synthase along with indole-3-glycerol phosphate (IGP) and serine as the precursor substrates in a buffered environment. For this purpose, cell extract of 3C1 (200 μL) was incubated for 30 mins, at 55°C in 80 mM potassium phosphate buffer, pH 8.0 containing 100 μM IGP, 60 mM L-serine, 50 μM pyridoxal phosphate (final concentration) in a total volume of 1 mL.12 After incubation, the reaction was stopped by adding 1 mL 10% TCA solution, and the formation of L-tryptophan as the end product was assayed following the method described below.
Spectrophotometric quantification of L-tryptophan
Tryptophan formed in the assay mixture was measured following the method developed by Ren et al.13 Diphenylamine sulphonate of the assay mixture oxidises to diphenylbenzidine sulphonic acid after reacting with sodium nitrite in the sulphuric acid medium. The unstable oxidation product reacts quickly with sodium nitrite to produce a diazotized intermediate. When the diazotized intermediate is coupled with tryptophan, a pink color product is developed, which is stable for at least 1 h at the ambient temperature. This colored product has an absorption maximum of 522 nm and the molar absorptivity is 0.89×104 L/mol.cm.
Isolation of thermophilic tryptophan producing bacterial strain
Out of 132 pure cultures, isolate from the water samples of the Paniphala hot spring, as described in the materials and methods section, 3C1 was finally selected on the basis of its tryptophanase negative character, maximum growth rate in DM broth medium, and tryptophan synthase activity for further study (data not shown).
Morphological, cultural, and physio-biochemical characteristics of the selected isolate 3C1 were studied following standard protocol available elsewhere, and the results obtained are presented in Table 1.
Table (1):
Showing Morpho-physio-biochemicals characters of isolate 3C1. ( *Methyl Red (MR) and Voges Proskauer Tests are done to know the type of glucose fermentation pathway an organism uses. If the organism ferments glucose by mixed acid fermentation, it shows a positive MR test, and if uses a butanediol fermentation pathway it shows positive for VP test.)
| Sample | 3C1 | |
|---|---|---|
| Colony Morphology on R2A agar | Shape | Irregular |
| Color | Off White (more yellowish) | |
| Opacity | Opaque | |
| Consistency | Sticky | |
| Margin | Irregular, undulating | |
| Elevation | slightly raised with an uneven surface | |
| Cellular morphology (Figure 3) | Shape | Large rods |
| Arrangement | Mostly filamentous | |
| Gram’s Nature | Positive | |
| Spore forming ability | Spore former | |
| motility | Motile | |
| Physio-Biochemical tests | Amylase Test | Positive |
| Nitrate reduction test | Positive | |
| Indole production | Negative | |
| Methyl Red test* | Positive | |
| Voges Proskauer test | Negative | |
| Citrate utilization | Negative | |
Determination of optimum temperature and pH of the selected growth medium for isolate 3C1
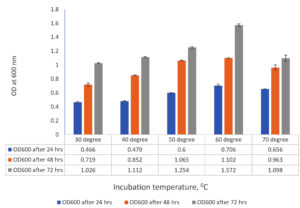
Growth of the selected isolated in varying initial medium pH of 7.2 and incubation temperature of 55°C were tested to find out the optimum temperature (Figure 4 ) and medium pH (Figure 5) for the growth of 3C1.
Figure 4. Optimization of incubation temperature for optimum growth of the isolate 3C1. Series 1= OD600 after 24 hrs growth, Series 2= OD600 after 48 hrs growth, Series 2= OD600 after 72 hrs growth. All data are average of three replicates
Figure 5. Optimization of medium pH for maximum growth of isolate 3C1. Series 1= OD600 after 24 hrs growth, Series 2= OD600 after 48 hrs growth, Series 2= OD600 after 72 hrs growth. All data are average of three replicates
Identification of the isolate 3C1 on the basis of its 16S rDNA sequence homology
Genomic DNA of the selected thermophilic bacterial strain viz 3C1 was isolated and the 16S rRNA gene was sequenced following the method described in the materials and methods section. The sequenced 16S rRNA gene of isolate 3C1 (Figure 6) was obtained from the NCMR-MCCS Pune in Fasta format and was compared the 16S rDNA sequences available in the NCBI database using BLAST. The similar sequences were analyzed using CLUSTALW and the phylogenetic tree of the sequences was generated (Figure 7) which revealed that the type strain of Bacillus licheniformis ATCC 14580(T) is the closest relative of isolate 3C1with a sequence homology of 99.67% (Table 2). The 16S rRNA gene sequence of isolate 3C1 is submitted in NCBI Gene Bank having accession number MN294568.
Table (2):
Microbial Identification Report for PRN B_APR_18_250 & 251, NCMR, NCCS, Pune
PRN |
Strain No. |
Closest neighbour |
Accession No. |
% Similarity |
|---|---|---|---|---|
B_APR_18_251 |
3C1 |
Bacillus licheniformis ATCC 14580(T) |
AE017333 |
99.67 |
Figure 7. Phylogenetic tree of the selected isolate having tryptophan synthase activity obtained from Paniphala hot spring, Barabani, Asansol, West Bengal, India
The isolated strain was identified on the basis of its morpho-physio-biochemical characteristics (Table 1) and 16S rRNA gene sequence homology (Figure 6 & 7) as a strain of Bacillus licheniformis.
Geothermal hot springs with a temperature range of 50-60°C or higher are excellent ecological niche for versatile thermophilic microorganisms. In the present work bio-prospecting of thermophilic microbes of a local, less explored hot spring, situated at Paniphala, near Barabani, Asansol, Burdwan West was done for enzymatic production of L-tryptophan. Water samples were collected from the hot spring, its physical parameters were recorded and culturable microbes were purified on suitable agar medium and tested for their tryptophan synthase activity. Finally, on the basis of tryptophan synthesis efficiency one isolate was selected for further study. Bioprospecting of thermophilic microbes isolated from several hot springs throughout the world by various workers are previously reported but reports on enzymatic tryptophan production by thermophilic microbes are limited.
The isolate was identified as a gram positive rod shaped bacteria, closest to Bacillus licheniformis with 99.62 % similarity of 16S rRNA gene sequence. The strain is positive for catalase, amylase and protease but negative for IMViC, nitrate reduction and indole test. It showed optimum growth temperature at 60°C and optimum pH at 8.0 with a wide range of growth temperature and pH which makes it suitable as industrial strain. Industrial strain having capacity to grown optimally in a wide range of temperature has added advantages. Wen et al.14 and Ferrer et al.15 reported production of L-tryptophan from its precursors by Corynebacterium glutamicum and E. coli at 30°C and 37°C, respectively. To improve L-tryptophan production the tryptophan synthase of E. coli was genetically engineered.16,17 so that it can act optimally at higher temperature and pH (40°C and pH 8.0 respectively) then its natural optimum growth temperature and pH of 30°C and 7.0 respectively. Several authors reviewed18,19 the use of various thermophilic bacterial strain for biotransformation of precursor metabolites into final products and concluded that thermophiles has many advantages over their mesophilic counterparts for the said purpose. Capacity of the isolated strain for enzymatic synthesis of L-tryptophan was assayed in vitro in terms of tryptophan formation from a mixture of IGP (indole glycerol phosphate), serine, pyridoxal phosphate and toluene treated permeabilised cells with respect of time. The added advantage is that the isolated strain is negative for tryptophanase (indole test negative). Though thermophilic bacteria having high tryptophanase activity are not yet reported in large number, similar activity by several mesophilic counterparts are already reported by several researchers.14-16, 18 In most of the reports the main bottleneck was the stability of the tryptophan synthase enzyme of the mesophilic tryptophan synthase at commercial production condition for a prolonged period.20-22 Being a thermophilic, tryptophan synthase from test strain is expected to show high thermal-stability for a prolonged period. From the initial study, it appears that the isolated Bacillus strains may be further exploited for enzymatic production of L-tryptophan.
ACKNOWLEDGMENTS
The author is thankful to the University Grants Commission, India, for providing the necessary funds for the research work. The co-operation of the administration of the MUC Women’s College, Burdwan, West Bengal, India, by providing the necessary infrastructure for the research work is thankfully acknowledged.
FUNDING
The research was funded by the University Grant Commission (UGC), vide Grant number PSW-034/15-16 (ERO) ID No.-WB1-048, S. No.- 300310, dated 04.07.2017.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
- Ano Y, Takaichi Y, Ohya R, Uchida K, Nakayama H, Takashima A. Tryptophan-tyrosine dipeptide improves tau-related symptoms in tauopathy mice. Nutr Neurosci. 2022;26(8):766-777.
Crossref - Nayak BN, Singh RB, Buttar H. Role of tryptophan in health and disease: systematic review of the anti-oxidant, anti-inflammation, and nutritional aspects of tryptophan and its metabolites. World Heart Journal. 2019;11(2):161-178.
- Badawy AA, Doughrty DM, Marsh-Richard DM, Steptoe A. Activation of liver tryptophan pyrrolase mediates the decrease in tryptophan availability to the brain after acute alcohol consumption by normal subjects. Alcohol and Alcoholism. 2009;44(3):267-271.
Crossref - Mateos SS, Sanchez CL, Paredes SD, Barriga C, Rodriguez AB. Circadian levels of serotonin in plasma and brain after oral administration of tryptophan in rats. Basic Clin Pharmacol Toxicol. 2009;104(1):52-59.
Crossref - Paredes SD, Marchena AM, Bejarano I, et al. Melatonin and tryptophan affect the activity-rest rhythm, core and peripheral temperatures, and interleukin levels in the ringdove: changes with age. J Gerontol A Biol Sci Med Sci. 2009;64(3):340-350.
Crossref - Zelante T, Iannitti RG, Cunha C, et al. Tryptophan catabolites from microbiota engage aryl hydrocarbon receptor and balance mucosal reactivity via interleukin-22. Immunity. 2013;39(2):372-385.
Crossref - Raboni S, Spyrakis F, Campanini B, et al. Pyridoxal 5-phosphate-dependent enzymes: catalysis, conformation, and genomics. Comprehensive Natural Products II Chemistry and Biology. 2010;7:273-350.
Crossref - Dierkers AT, Niks D, Schlichting I, Dunn MF. Tryptophan synthase: structure and function of the monovalent cation site. Biochemistry. 2009;48(46):10997-1010.
Crossref - Kriechbaumer V, Weigang L, Fießelmann A. Characterisation of the tryptophan synthase alpha subunit in maize. BMC Plant Biol. 2008;8:44.
Crossref - Watkins-Dulaney E, Straathof S, Arnold F. Tryptophan synthase: biocatalyst extraordinaire. Chem Bio Chem. 2021;22(1):5-16.
Crossref - Bosken YK, Ai R, Hilario E, et al. Discovery of antimicrobial agent targeting tryptophan synthase. Protein Sci. 2022;31(2):432-442.
Crossref - Darkoh C, Chappell C, Gonzales C, Okhuysen P. A rapid and specific method for the detection of indole in complex biological samples. Appl Environ Microbiol. 2015;81(23):8093-8097.
Crossref - Ren J, Zhao M, Wang J, Cui C, Yang B. Spectrophotometric method for determination of tryptophan in protein hydrolysates. Food Technol Biotechnol. 2007;45(4):360-366.
- Ishiwata K, Fukuhara N, Shimada M, Makiguchi N, Soda K. Enzymatic production of L-tryptophan from DL-serine and indole by a coupled reaction of tryptophan synthase and amino acid racemase. Biotechnol Appl Biochem. 1990;12(2):141-149.
Crossref - Ferrer L, Elsaraf M, Mindt M, Wendisch VF. l-serine biosensor-controlled fermentative production of l-tryptophan derivatives by Corynebacterium glutamicum. Biology. 2022;11(5):744.
Crossref - Xu L, Han F, Dong Z, Wei Z. Engineering improves enzymatic synthesis of L-tryptophan by tryptophan synthase from Escherichia coli. Microorganisms. 2020;8(4):519.
Crossref - Li Z, Ding D, Wang H, Liu L, Fang H, Chen T, Zhang D. Engineering Escherichia coli to improve tryptophan production via genetic manipulation of precursor and cofactor pathways. Synthetic and Systems Biotechnology. 2020;5(3):200-5.
Crossref - D’Este M, Alvarado-Morales M, Angelidaki I. Amino acids production focusing on fermentation technologies-A review. Biotechnol Adv. 2018;36(1):14-25.
Crossref - Taylor IN, Brown RC, Bycroft M, et al. Application of thermophilic enzymes in commercial biotransformation processes. Biochem Soc Trans. 2004;32(Pt 2):290-292.
Crossref - Mindt M, Walter T, Kugler P, Wendisch VF . Microbial engineering for production of N-functionalized amino acids and amines. Biotechnol J. 2020;15(7):e1900451.
Crossref - Wendisch VF. Metabolic engineering advances and prospects for amino acid production. Metab Eng. 2020;58:17-34
- Ferrer L, Mindt M, Suarez-Diez M, et al. Fermentative indole production via bacterial tryptophan synthase alpha subunit and plant indole-3-glycerol phosphate lyase enzymes. J Agric Food Chem. 2022;70(18):5634-45.
Crossref
© The Author(s) 2023. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.