ISSN: 0973-7510
E-ISSN: 2581-690X
The present study was designed to compare the defense response of the host plant towards endophytic bacteria Pseudomonas aeruginosa PM389 and pathogenic bacteria Erwinia carotovora and to correlate the level of defense enzymes vis-a-vis bacterial colonization in the host. Wheat seedlings were treated with 107-108 cells ml-1 endophytic and pathogenic bacteria in the separate experimental set-up, and the level of plant defense enzyme was measured at various time intervals. Comparatively reduced level of most defense enzymes was produced in endophytic bacteria treated plants. While the endophytic bacterial population was almost constant after 24 HAI (hour after inoculation), the population of pathogenic bacteria kept fluctuating during the study period from 24 HAI. Unlike pathogenic bacteria, we observed attenuated defense response in challenged host plants towards endophytic bacteria, which helps endophytes establish inside plant. This study would be useful for understanding the mechanism of colonization and strategies of endophytes to fight against the host defense response.
Endophyte, Pseudomonas aeruginosa, defense enzymes, Erwinia carotovora
Plants harbor a diverse micro-flora of soil-borne bacteria and fungi that may have either beneficial or deleterious effects on the plant. Some of these bacteria, termed ‘endophytic bacteria’ colonize the interior of plant roots and shoots without harming their host plants.1 These endophytic bacteria can benefit host plants directly by providing nutrition (N, P, Fe etc.) and growth induction through phytohormone production, and indirectly through biocontrol and induced systemic tolerance.2 The majority of endophytic bacteria originate from the soil and colonize the plant interior. The exact mechanism of endophytic bacterial colonization is not well understood. However, various studies suggest that entry of endophytes in root is facilitated through wounds, cracks, and the points of lateral root emergence.3,4 The rhizosphere is the site of the complex interaction between plant and soil microflora, where plant exudates like organic acids, amino acids, and proteins etc., released from plants are responsible for initiating the colonization of bacterial endophytes in the plant interior.5 Since both endophytic (beneficial) and pathogenic (harmful) bacteria are capable of colonizing the host interior, successful establishment of endophytic bacteria inside the plant depends on genotypic compatibility, physiological status, and extent of host plant defense response.6 Thus, favored colonization of endophytes over pathogenic bacteria indicates interplay between plant and bacterial traits.
Plants respond to invading bacteria by eliciting defense responses which may be generalized or specific. The basal immune surveillance system of plant protects from microbial invasion through pattern triggered immunity (PTI) by identifying microbe-associated molecular pattern (PAMP/MAMP) like flagellin and lipopolysaccharides (LPS).7,8 However, some pathogenic bacteria dodge defense response by directly delivering the effector molecules inside the plants through a bacterial secretion system and cross the first line of defense.9 During co-evolution, plants have developed intracellular defense receptors (Resistance/R-gene-derived receptors) for the recognition of microbial effector molecules, i.e., effector-triggered immunity (ETI).10 ETI is highly specific for polymorphic effectors of different pathogens, which either releases certain hydrolytic enzymes like chitinase, β-1,3-glucanase or leads to hypersensitive responses and programmed cell death of infected cells.11,12 On microbial invasion, plants produce phytoalexin and defense enzymes responsible for lignin and callose deposition to make strong defense barriers against pathogens.13,14
Since both endophytic and pathogenic bacteria are alien to the host plant during the colonization process, endophytes initially trigger the host’s defense response similar to the pathogens. However, the endophytes somehow manage to combat the host defense later on to gain entry inside the plants, which requires further confirmation.15,16 Albeit beneficial, plants may respond towards endophytic bacteria as non-self body, which leads to induction of defense response in host plants. Despite the generation of plant defense response towards beneficial microbe, plants do not discourage interacting beneficial microbe. It suggests a well-coordinated and continuous molecular dialogue between the plant and the beneficial organism.17 To establish a mutualistic interaction with the plant, endophytes need to cope with host defense responses triggered locally in the roots upon endophytic MAMP perception. Thus, it is important to understand the strategies deployed by endophytic bacteria to deal with host defense responses and the role of plants in assisting endophytic bacteria in establishing inside the plants. To our knowledge, a few studies have been conducted on plant defense responses during endophytic bacterial colonization.18 Therefore, to address possible differences in intensities of defense response to endophytic and pathogenic bacteria, the present study aims to estimate the level of defense response by measuring the activity of defense enzymes produced after bacterial challenges and correlating them with successful colonization of given bacteria in-planta. We hypothesize that endophytes are evolved to suppress defense response, or they are not identified by plant factors and thus evade defense response.
The activity of four defense enzymes namely polyphenol oxidase (POD), peroxidase (P.O.), phenylalanine ammonia-lyase (PAL) and, β-1,3-glucanases was estimated.19 The present study has been carried out using endophytic bacteria Pseudomonas aeruginosa PM389 and a pathogenic strain Erwinia carotovora. PM389 was isolated from pearl millet and showed various plant growth promoting traits like nitrogen fixation, siderophore production, phosphate solubilization, and ability to cross-infect wheat plant.19 Erwinia carotovora is a broad range pathogen, and it causes secondary infection in wheat for enhancing the activity of major wheat pathogens like Fusarium spp. etc.20 We chose the wheat plant as host for defense response studies as it is a major cereal crop worldwide, and such studies on wheat can help increase crop yields in the future to feed the burgeoning population worldwide.
Bacterial Strains used
Plant growth-promoting endophytic bacteria used in the present study was Pseudomonas aeruginosa PM389 isolated from pearl millet.19 Erwinia carotovora, a broad host range plant pathogen, was procured from Indian Agriculture Research Institute, Pusa, India. Pure cultures were maintained on Luria Bertini (L.B.) agar media, and long-term storage was done at -70°C in glycerol (15%, v/v).
Test bacterial strains inoculation on wheat plant
Triticum aestivum variety GW322 seeds were surface sterilized with 70% ethanol for 2 min followed by 0.2% HgCl2 solution for 3 min. The seeds were washed thoroughly with several rounds of sterile distilled water to remove traces of sterilants. Finally, seeds were placed on moist sterile Whatman filter paper No.1 in Petridishes for germination and watered every alternate day for four days under dark conditions. Healthy seedlings were transferred on the 4th day to the tubes containing semi-solid Hoagland media (0.3% Agar). Composition of Hoagland media was (g l-1): KNO3, 202; Ca(NO3)2.4H2O, 118; Fe-EDTA, 15; MgSO4.7H2O, 493; NH4NO3, 80; H3BO3, 2.86; MnCl2.4H2O, 1.81; ZnSO4.7H2O, 0.22; CuSO4, 0.051; Na2MoO4.2H2O, 0.12; KH2PO4, 136 (pH to 6.0 with 3M KOH). Seedlings were grown hydroponically in Hoagland medium standard condition (14 h-light, 10 h-dark cycle at 25°C ± 2 and 15°C ± 2 respectively). On the 10th day of growth, plants were inoculated (separately) with107-108 cells ml-1 (in 1X PBS) endophyte and pathogen. Subsequently, plant samples were collected at 0, 6, and 12 hours post-inoculating bacterial strains and at a regular interval of 24 hours up to six days after inoculation (DAI). Collected plants were crushed in liquid nitrogen and stored at -70°C for further analysis of defense enzymes. Plants treated with 1X PBS served as control. All the treatments were carried out in four replicates (biological replicates), and each sample was further assayed in triplicates (technical replicates).
Plant defense enzymes estimation
The collected plant tissues were crushed in liquid nitrogen and stored at -70°C for the analysis of defense enzymes namely peroxidase, polyphenol oxidase, phenylalanine ammonia lyase and β-glucanase using standard methods described below. All the samples were homogenized in two times volume of respective buffer for the particular enzyme assay and centrifuged at 8000 g for 10 min at 4°C. The resulting supernatant was used for enzyme assays. Protein was estimated by the method as described by Lowry et al.21
β-glucanase activity
Plant samples were macerated in 0.05 mol l-1 sodium acetate buffer (pH 5. 0) by grinding at 4°C. A 10 μl crude enzyme extract was added to 10 μl of 4% laminarin and then incubated at 40°C for 10 min. The reaction was stopped by adding 375 μL of dinitrosalicylic reagent (0.8% NaOH, 0.25% dinitrosalicylic, 0.2% phenol, 0.05% Na2SO3) and 10 min heating on a boiling water bath followed by addition of 20 μl of 40% sodium potassium tartarate. The resultant colored solution was diluted three-fold with Milli Q water, vortexed and, absorbance was read at 575 nm in UV-Vis spectrophotometer (JASCO, USA). Various concentrations of glucose (0 to 1 mmol l-1) were used to obtain a calibration curve. One unit of enzyme activity is defined as the amount of enzyme that produced reducing sugar equivalent to 1 μmol of glucose equivalent per 10 min under the above conditions.22
Phenylalanine ammonia lyase (PAL) activity estimation
The plant sample was mixed in 50 mmol l-1 Tris, pH 8.8. To 40 μl of total enzyme extract, 24 μl of 10 mmol l-1 phenylalanine, and 176 μl Tris (pH 8.8) were added in successive order. The resultant mixture was incubated for 60 min at 30°C, and the reaction was stopped by adding 200 μl of 2 mol l-1 HCl and 300 μl toluene. The reaction mix was spun at 750 g for 5 min, and the upper layer was collected to read absorbance at 290 nm. Cinnamic acid (1-5 μg ml-1) was used to prepare a standard curve for the quantification.23
Peroxidase activity estimation
Plant tissue was extracted in 10 mmol l-1 phosphate buffer (pH 6.0). A 6 µl of enzyme extract was added to the reaction mix containing 24 µl 0.25% guaiacol dissolved in 0.01 mol l-1 potassium phosphate (pH 6.0) and 0.1 mol l-1 H2O2. Enzyme activity was measured by taking OD every 30 sec for 3 min at 470 nm and expressed as changes in absorbance min-1 mg-1 protein.24
Polyphenoloxidase activity estimation
Plant samples were homogenized in 0.1 mol l-1 sodium phosphate buffer (pH 6.5). The enzymatic assay was done using 20 µl of the enzyme extract, in which 320 µl of 0.1 mol l-1 sodium phosphate buffer (pH 6.5) was added. To start the reaction, 40 µl of 0.01 mol l-1 catechol was added to the reaction mix, and the activity was expressed as changes in absorbance (measured at every 30 sec for 1min) at 495 nm min-1 mg-1 protein.25
Evaluation of bacterial colonization in plants
Colonization studies were carried out using similar experimental set-up as described above for defense enzyme analysis of endophytic and pathogenic bacteria,. Their establishment in plants were evaluated by reisolation of inoculated bacteria at various time intervals followed by the confirmation of their identity by ERIC-PCR (Enterobacterial repetitive intergenic consensus sequence-PCR) to track the identity of bacterial inoculants.
Isolation of bacteria from plants
Healthy wheat plants were uprooted and washed thoroughly with sterilized tap water to remove media. Plant samples were surface sterilized by immersing in 70% ethanol for 2 minutes followed by 4% sodium hypochlorite for 12 minutes. All traces of sterilants were removed by rinsing the plant samples thoroughly with sterilized distilled water. A 100 µl of last wash was spread on Nutrient Agar (N.A.) media (HiMedia, India) to assess the sterilization efficiency of the protocol used. Surface sterilized plants were macerated separately in a sterile PBS solution. 100 µl of macerate was spread on NA plates and incubated at 37°C for two days. Bacterial colonies were counted individually for each treatment.
Enterobacterial repetitive intergenic consensus sequences-PCR (ERIC-PCR)
DNA template for PCR was prepared by “Boiling lysis method” as described by Misra et al.26 ERIC-PCR was performed using PTC thermal cycler (MJ Research, Inc., Walthon, MA, USA) following a standard protocol.27 The final volume of PCR reaction mixture was 50 µl, consisting of 1.5 units of Taq DNA polymerase, 125 µmol l-1 each dNTPs, 1.5 mmol l-1 MgCl2, 30 pmol of each primer, and 50 ng genomic DNA. The specific primers (Sigma-Aldrich, USA) used were 1R (5′-ATGTAAGCTCCTGGGGATTCAC-3′) and 2R (5′-AAGTAAGTGACTGGGGTGAGC G-3′). The thermal cycler program included an initial denaturation at 94°C for 5 min followed by 35 cycles of 1 min at 94°C, 55°C for 1 min and, 72°C for 1 min with a final extension of 5 min at 72°C. The amplified products were analyzed on 2% agarose gel using a gel documentation unit (Bio-Rad Laboratories, USA). The ERIC profiles were analyzed on the basis of the pattern of amplified DNA fragments separated in the gel. Isolates showing identical DNA band patterns were considered to be the same strain.
Statistical analysis
One-way ANOVA, standard deviation, and correlation analysis were done wherever applicable using Microsoft excel. Least Significant Difference (LSD)28 was done to identify significant differences in plant defense response towards endophytic and pathogenic bacteria. Duncan’s Multiple Range Test (DMRT) was used to compare the mean at different time intervals of the study period.
Comparative analysis of plant defense enzymes in response to endophytic and phytopathogenic bacteria
Peroxidase activity (PO)
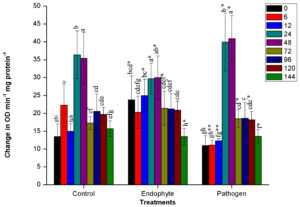
The PO activity was significantly (n=12, p<0.05) stimulated in plants immediately after inoculation of PM389 in plants treated with endophytic bacteria (from 0-12 HAI), whereas the appearance of PO activity in pathogen-treated plants took place after 24 HAI. The highest peroxidase activity was observed in plants collected at 24 and 48 HAI in all the treatments, and after 48 HAI activity was suddenly reduced in each treatment. During this period (24-48 HAI), pathogen treatment resulted in a significantly highest change in the activity of PO enzyme, while repressed enzyme activity was recorded in endophytic bacteria (PM389) treated plants in comparison to control treatment (Fig. 1a.).
Polyphenol oxidase (PPO)
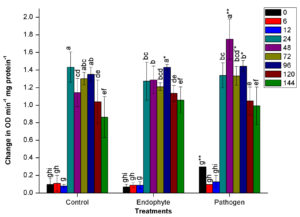
In all the treatments, a sudden increment in PPO activity was observed at 24 HAI. Maximum activity of PPO was observed at 48 HAI in pathogen-treated plants. There was a significant (n=12, p<0.05) increase with 26% higher PPO activity in pathogenic bacteria treated plants at 48 HAI than endophytic bacteria treated and un-inoculated control plants (Fig. 1b.). However, it followed a sharp decrease in the activity of PPO in all the treatments at 96 HAI. On the contrary, no significant deviation in PPO activity was observed in plants treated with endophytic bacteria from the PPO activity noted in control plants (Fig. 1b.).
β-glucanase
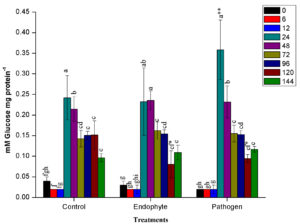
Starting from 24 HAI, the activity of β-glucanase in all plants remained higher in samples collected at various time intervals. At 24 HAI, β-glucanase activity was reported significantly higher in pathogen-treated plants than uninoculated control and endophyte-treated plants. Similar to results of PPO activity, plants treated with pathogenic bacteria showed enhanced activity of the β-glucanase enzyme. In contrast, endophytic bacteria PM389 did not elicit much higher glucanase activity than that of control (Fig. 1c). In all the treatments, glucanase activity was higher from 24-48 HAI and afterward started reducing drastically.
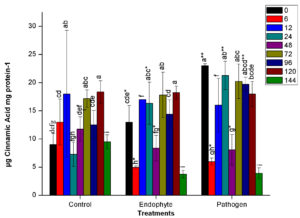
Fig. 1. Changes in the activity of different defense enzymes in wheat (Triticum aestivum var. GW322) plants at various time intervals in response to endophytic bacteria PM389 and phytopathogenic bacteria Erwinia carotovora. (a) Plant peroxidase, (b) Polyphenol oxidase (c) β-1,3-glucanase, and (d.) Phenylammonia lyase. Plants treated with phosphate buffer saline used as a control (PBS) treatment. Each value represents the mean of four biological and three technical replicates (12 replicates). Vertical bars represent standard deviation. The mean values were compared using Duncan’s multiple range test (DMRT) at p < 0.05. Values that are significantly different from each other are headed by different letters in each column of treatment.* represents a significant difference in the respective column from the column. ** represents a significant difference from both control and endophyte in the pathogen. *-/-* represents a significant difference in the pathogen from control/endophyte only (Part of this data has been published in another article in Press (Gupta et al. 2021 in Research Journal of Biotechnology))
Phenyl ammonia lyase (PAL)
Unlike other assays, no finite trend was observed in the PAL assay. There was alternate increase and decrease in enzyme activity at different time points in all treatments, including control plants. At 48 HAI, PAL activity decreased drastically for all the treatments. Further, the activity enhanced suddenly at 72 HAI. Similar to the results of other enzyme assays, pathogenic bacterial treatment led to the greatest change in enzyme activity in host plants. In pathogen-treated plants, significantly higher activity (n=12, p<0.05) was recorded at 0 HAI. At 6 HAI, activity was drastically reduced in both endophyte and pathogenic treatment followed an increase in activity until 24 HAI. At 24 HAI, a significant difference has been observed in pathogenic and endophytic treatment as shown in Fig. 1d. On the other hand, PM389 stimulated insignificantly lesser and constant defense response than E. carotovora from 12-24 HAI (Fig. 1d).
Endophytic and pathogenic colonization tracking in wheat plants
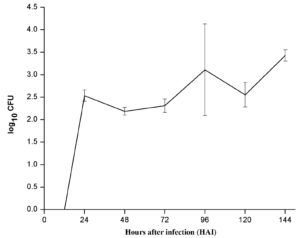
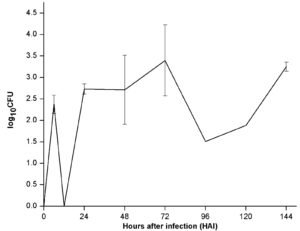
To estimate and compare colonization efficiency of endophyte (PM389), reisolation of inoculated bacteria from plants was done at various time intervals. The efficiency of colonization was assessed based on CFU count of each bacterial species inoculated separately. ERIC-PCR fingerprinting was used to confirm the identity of selected bacterial isolates. An identical ERIC-PCR profile of isolates recovered from inoculated plants to that of pure culture was used to confirm the colonization abilities of test strains. Recovery of endophytic bacteria from inoculated plants started from 24 HAI. The maximum population of endophytic bacteria was achieved after 144 HAI as depicted in Fig. 2a. The CFU count of endophytic isolates showed slight variation in plant samples collected at different time intervals. On the other hand, pathogenic bacteria failed to establish consistently during the entire study period. Unlike endophytic bacteria, pathogen recovery was obtained as early as 6 HAI with a sudden increase in population and followed a steep decline in population at 12 HAI and subsequent increase from 24 HAI. Further, a steep decline in population was noticed at 96 HAI, which again started increasing slowly in subsequent time points (Fig. 2b). For both pathogen and endophyte, population decrement was recorded with an increase in the level of either of the defense enzyme and vice-versa with few exceptions (Table 1). Thus, the population of pathogenic bacteria exhibited alternate rise, and decline in population during colonization.
Table (1):
Correlation analysis of changes in population count and levels of defence enzyme with time.
log10 CFU |
PO |
PPO |
β- glucanase |
PAL |
|---|---|---|---|---|
PM389 |
-0.23 |
0.94* |
0.70* |
0.02 |
Erwinia |
0.42 |
0.61 |
0.57 |
-0.42 |
*p<0.05, n =9
Fig. 2. Population dynamics of (a.) endophyte (PM389) and (b.) pathogen (Erwinia caratovora) on challenge inoculation in Triticum aestivum (var. GW322) at selected time points (0-144 HAI). Each value represents the mean of duplicates, and the experiment was repeated two times (4 replicates). Vertical bars represent standard deviation. The mean values were compared using Duncan’s multiple range test (DMRT) at p < 0.05. Values that are significantly different from each other are headed by different letters (Part of this data has been published in another article in Press (Gupta et al. 2021 in Research Journal of Biotechnology))
Plants synthesize several PR proteins and their homologues, including superoxide dismutase (SOD), peroxidase (POD), catalase (CAT) (oxidative enzymes), polyphenol oxidase (PPO), phenylalanine ammonia-lyase (PAL), β-1,3-glucanases (PR-2 family), and chitinases (PR-3 family) constitutively but the level of these proteins are mainly affected in response to microbial invasion.29,30 Therefore, in the present study, we compared activities of some of the above-mentioned defense enzymes in plants inoculated separately with endophytic bacteria Pseudomonas aeruginosa PM389 and pathogenic bacteria Erwinia carotovora. We also compared the kinetics of bacterial colonization in relation to immune response. To our knowledge, only a few works have demonstrated the differential behavior of host plants to beneficial and pathogenic bacteria.
Upsurge in peroxidase activity in plants following microbial challenge may lead to oxidative burst resulting from sudden changes in ion fluxes across the membrane.31,32 The appearance of peroxidase activity in the wheat plant immediately after inoculation of endophytic bacteria suggests rapid recognition of endophytic bacteria by plants. It might have led to the generation of ROS. Similar to our observation, increased ROS production (catalyzed by peroxidase) in response to microbial infection was noted as early as 2 min in an earlier study.33 Except for the initial sampling period, peroxidase activity remained decreased in response to PM389. This result is supported by results of Naffaa et al34 which suggested that the reduction in peroxidase level may be a possible mechanism to facilitate endophytic colonization of bacteria. On the contrary, peroxidase activity was not enhanced in response to pathogen up to 12 HAI indicating suppression of oxidative burst in the initial phase of infection. However, it was strongly induced from 24-48 HAI. Thus, the peroxidase activity was induced for longer period in pathogen-treated plants, which may restrict the entry of pathogen to certain extent.
Unlike the results of peroxidase activity, induction of glucanase appeared at 24 HAI. However, there are some studies that reported induction of different genes, including that for glucanase immediately after recognition of bacteria-derived effector molecules.35 We observed that inoculation with Erwinia showed significant induction of β-glucanase activity at 24 HAI, while no significant change was observed for above enzyme in plants inoculated with PM389. On the contrary, slightly repressed activity was observed in response to PM389 in comparison to control treatment. This finding is similar to the studies conducted on Sinorhizobium meliloti, which showed downregulation of β-glucanase (MtBGLU1) gene in the roots of Medicago truncatula after 24 h of inoculation.37 After 24 HAI, the β-glucanase expression was turned down for all the treatments. A decrease in activity in further sampling period may rely on the fact that β- glucanase gets intensified further only in the presence of glucan oligomers released during the lysis of the fungal cell wall. Since both treatments are bacterial in the present study, inoculated plants did not show further induction of β-glucanse activity due to the absence of elicitors.38
Similar to the results of β- glucanase activity, significant induction of PPO was observed in only pathogen treated plants at 48 HAI. It suggests that plants are also responsible for differentiating endophytes and pathogen during infection and help establish endophytic bacteria inside the plants. Recently, researchers reported two waves of transcriptional reprogramming in M. truncatula involving repression of defense-related genes followed by the activation of a nodule-specific transcriptome.39 Several collections of plant and bacterial mutants were used in the study, which established that the transcriptome switch depends upon a molecular dialogue between both partners. The fourth enzyme whose activity was assayed during plant-microbe interaction was phenyl ammonia lyase (PAL), the first key enzyme of phenylpropanoid pathway and flavanoid pathway generating precursors for lignin biosynthesis, phytoalexins, and other phenolic compounds that have direct effects on pathogens. Cinnamic acid is the product of PAL, which is directly linked to the cell lignification process.41,42 PAL activity was higher for pathogens at 0 HAI, suggesting the quick recognition by the plant’s phenylpropanoid pathway. Subsequently, PAL activity decreased in all treatments except in the samples collected from 72-120 HAI. Further, reduced PAL activity was recorded in endophyte treated plants, again supporting the notion that plants generate weak defense response against endophytes or beneficial bacteria.43
Attenuated defense response in endophyte treated plants than pathogen treated plants indicates that endophytes may gain entry into plants by suppressing or evading from plant defense response. It gains support from a few of earlier studies which report that various MAMPs like EPS (exopolysaccharide), LPS (lipopolysaccharide), and nod-factors of beneficial bacteria are capable of suppressing defense response of the plant to gain entry inside the plant.44 Downregulation of PR proteins in host plant on inoculation of endophytic bacteria Burkholderia phytofirmans has also been reported recently.18
To corroborate defense response to the success of colonization in plants, reisolation of PM389 and E. carotovora was made, and their extent of colonization was measured in terms of CFU count. Since ERIC sequences are distributed throughout the genome, and their distribution pattern and repeat number vary amongst different species/strains, bacterial inoculants’ identification was confirmed by ERIC-PCR based DNA fingerprinting approach.19,45 Based on the results of the reisolation study, it appears that colonization of PM389 started late compared to Erwinia, where the latter was recovered as early as 6 HAI showing infection severity. However, the extent of colonization of PM389 was consistent throughout the study period with a slight intermittent decrease in CFU count. The pattern of colonization by two types of bacteria corresponded to the pattern of defense response. Higher defense enzyme activities immediately after inoculation restricted bacterial colonization by endophytes, whereas pathogenic bacteria could colonize early when the defense enzyme activities were low. Similar to PM389, stress and defense-related genes were found to be upregulated initially and then, followed downregulation in Rhizobium inoculated plants.46 This phenomenon strengthens the fact that beneficial bacteria have evolved to establish in host plant through immuno-suppression of the host system. In contrast, Erwinia colonized earlier, but its population fluctuated at successive intervals. Successive increase and decrease of the pathogenic population at different time intervals evince constant fight of the host plant to ward off pathogen and pathogen further tries to pioneer in the plant by specific tactics. Overall, the population of both pathogenic and endophytic bacteria keeps dwindling though at different levels based on the corresponding levels of defense enzymes. Although the population of pathogenic and endophytic bacteria at 144 HAI is almost similar, which might be due to lower levels of defense enzyme at this time point or might arise from changed defence strategy of the pathogenic bacteria. It shows a similar pattern to that of the zig-zag model of the plant’s defense response against pathogenic establishment inside the plant.47
In conclusion, endophytes were compared with plant pathogen for induction of defense enzyme as well as their establishment in the host plant. It was observed that endophyte showed suppression of defense system of host, which could be a possible reason for successful colonization. It infers that a constant and positive molecular dialogue between host and endophytic bacteria might be involved for endophytic establishment, unlike pathogen establishment which shows constant fight occurring between pathogen and host plant.
Endophytes were compared with plant pathogen for induction of defense enzyme as well as their establishment in the host plant. It was observed that endophyte showed suppression of defense system of host, which could be a possible reason for successful colonization. It infers that a constant and positive molecular dialogue between host and endophytic bacteria might be involved for endophytic establishment, unlike pathogen establishment which shows constant fight occurring between pathogen and host plant.
ACKNOWLEDGMENTS
The authors are thankful to DST, New Delhi for providing research fellowship.
CONFLICT OF INTEREST
All authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
This research work was funded by Department of Science and Technology (DST), Govt. of India, New Delhi under SERC Fast Track Scheme Project (No.SR/FT/LS-0662/065/2008) Sanctioned to PNJ.
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the manuscript.
- Kandel SL, Joubert PM, Doty SL. Bacterial Endophyte Colonization and Distribution within Plants. Microorganisms. 2017; 5: 77.
Crossref - Bakker P, Pieterse CMJ, Loon L. Induced systemic resistance by fluorescent Pseudomonas spp. Phytopathol. 2007; 97: 239-243.
Crossref - Bais HP, Park S-W, Weir TL, Callaway RM, Vivanco JM. How plants communicate using the underground information superhighway. Trends Plant Sci. 2004; 9:26-32.
Crossef - Gupta G, Panwar J, Akhtar M, Jha P. Lichtfouse E (ed.), In Endophytic nitrogen-fixing bacteria as biofertilizer. Sustainable Agriculture Reviews, Springer. Netherlands. 2012:183-221.
Crossref - Kawasaki A, Donn S, Ryan PR, et al. Microbiome and exudates of the root and rhizosphere of brachypodium distachyon, a model for wheat. PLoS ONE. 2016;11:e0164533.
Crossref - Liu H, Carvalhais LC, Crawford M, et al. Inner Plant Values: Diversity, Colonization and Benefits from Endophytic Bacteria. Front Microbiol. 2017; 8:1-17.
Crossref - Boller T, Felix G. A renaissance of elicitors: perception of microbe-associated molecular patterns and danger signals by pattern-recognition receptors. Ann Rev Plant Biol. 2009;60:379-406.
Crossref - He P, Shan L, Lin N-C, et al. Specific bacterial suppressors of MAMP signaling upstream of MAPKKK in Arabidopsis innate immunity. Cell. 2006; 125:563-575.
Crossref - Lucke M, Correa MG, Levy A. The Role of Secretion Systems, Effectors, and Secondary Metabolites of Beneficial Rhizobacteria in Interactions With Plants and Microbes Front. Plant Sci. 2020;11:589416.
Crossref - Dangl JL, Jones JDG. Plant pathogens and integrated defence responses to infection. Nature 2001;411:826-833.
Crossref - Martin GB, Bogdanove AJ, Sessa G. Understanding the functions of plant disease resistance proteins. Ann Rev Plant Biol. 2003; 54:23-61.
Crossref - Nimchuk Z, Eulgem T, Holt Iii BF, Dangl JL. Recognition and response in the plant immune system. Ann Rev Gen. 2003; 37:579-609.
Crossref - Malinovsky FG, Fangel JU, Willats WGT. The role of the cell wall in plant immunity. Front Plant Sci. 2014;
Crossref - Glazebrook J. Contrasting mechanisms of defense against biotrophic and necrotrophic pathogens. Ann Rev Phytopathol. 2005; 43:205-227.
Crossref - Rabiey M, Hailey LE, Roy SR, et al. Endophytes vs tree pathogens and pests: can they be used as biological control agents to improve tree health?. Europ J Plant Pathol. 2019; 155:711–729.
Crossref - Hardoim PR, ‘V’ Overbeek LS, Elsas JDV. Properties of bacterial endophytes and their proposed role in plant growth. Trends Microbiol. 2008; 16:463-471.
Crossref - Van Wees SCM, Van ‘D’ Ent S, Pieterse CMJ. Plant immune responses triggered by beneficial microbes. Curr Opin Plant Biol. 2008; 11:443-448.
Crossref - Bordiec S, Paquis S, Lacroix H, et al. Comparative analysis of defence responses induced by the endophytic plant growth-promoting rhizobacterium Burkholderia phytofirmans strain PsJN and the non-host bacterium Pseudomonas syringae pv. pisi in grapevine cell suspensions. J Exp Bot. 2011; 62:595-603.
Crossref - Gupta G, Panwar J, Jha PN. Natural occurrence of Pseudomonas aeruginosa, a dominant cultivable diazotrophic endophytic bacterium colonizing Pennisetum glaucum (L.) R. Br. Appl Soil Ecol. 2013; 64:252-261.
Crossref - Newton AC, Toth IK, Neave P, Hyman LJ. Bacterial inoculum from a previous crop affects fungal disease development on subsequent nonhost crops. New phytol. 2004; 163:133-138.
Crossref - Lowry OH, Rosebrough NJ, Farr AL, Randall RJ. Protein measurement with the folin phenol reagent. J Biol Chem. 1951; 193:265-275.
Crossref - Pan S, Ye X, Kuc J. Association of ß-1,3-glucanase activity and isoform pattern with systemic resistance to blue mould in tobacco induced by stem injection with Peronospora tabacina or leaf inoculation with tobacco mosaic virus. Physiol Mol Plant Pathol, 1991; 39: 25-39.
Crossref - Ramamoorthy V, Raguchander T, Samiyappan R. Induction of defense-related proteins in tomato roots treated with Pseudomonas fluorescens Pf1 and Fusarium oxysporum f. sp. lycopersici. Plant Soil. 2002; 239:55-68.
Crossref - Hammerschmidt R, Nuckles EM, Kuć J. Association of enhanced peroxidase activity with induced systemic resistance of cucumber to Colletotrichum lagenarium. Physiol Plant Pathol. 1982; 20:73-82.
Crossref - Mayer AM, Harel E, Ben-Shaul R. Assay of catechol oxidase—a critical comparison of methods.Phytochem.1966; 5:783-789.
Crossref - Misra N, Gupta G, Jha PN. Assessment of mineral phosphate-solubilizing properties and molecular characterization of zinc-tolerant bacteria. J Basic Microbiol. 2012; 52:549-558.
Crossref - Versalovic J, Koeuth T, Lupski R. Distribution of repetitive DNA sequences in eubacteria and application to finerpriting of bacterial enomes.Nuc Acids Res. 1991; 19:6823-6831.
Crossref - Quinn GP, Keough MJ. Experimental design and data analysis for biologists. Cambridge university press, Cambridge. 2002.
Crossref - Jain S, Vaishnav A, Kumari S, Varma A, Tuteja N, Choudhary DK. Chitinolytic Bacillus-mediated induction of jasmonic acid and defense-related proteins in soybean (Glycine max, L. Merrill) plant against Rhizoctonia solani, and Fusarium oxysporum. J Plant Growth Regul.2017; 36, 200–214.
Crossref - Gomez-Vasquez R, Day R, Buschmann H, Randles S, Beeching JR, Cooper RM. Phenylpropanoids, phenylalanine ammonia lyase and peroxidases in elicitor-challenged cassava (Manihot esculenta) suspension cells and leaves. Ann Bot. 2004; 94:87-97.
Crossref - Kawano T. Roles of the reactive oxygen species-generating peroxidase reactions in plant defense and growth induction. Plant Cell Rep. 2003; 21:829-837.
Crossref - Survila M, Davidsson PR, Pennanen V, et al. Peroxidase-Generated Apoplastic ROS Impair Cuticle Integrity and Contribute to DAMP-Elicited Defenses. Front Plant Sci. 2016;7:1945.
Crossref - Chinchilla D, Zipfel C, Robatzek S, et al. flagellin-induced complex of the receptor FLS2 and BAK1 initiates plant defence. Nature. 2007; 448:497-500.
Crossref - Naffaa W, Ravel C, Boyer N, Guillaumin J-J. Peroxidase activity of perennial ryegrass and tall fescue seedlings artificially infected with endophytes. Agronomie.1999; 19:611-619.
Crossref - Ferrari S, Galletti R, Denoux C, ‘D’ Lorenzo G, Ausubel FM, Dewdney J. Resistance to Botrytis cinerea induced in Arabidopsis by elicitors is independent of salicylic acid, ethylene, or jasmonate signaling but requires PHYTOALEXIN DEFICIENT3. Plant Physiol. 2007;144:367-379.
Crossref - Mitra RM, Long SR. Plant and bacterial symbiotic mutants define three transcriptionally distinct stages in the development of the Medicago truncatula/Sinorhizobium meliloti symbiosis. Plant Physiol. 2004; 134:595-604.
Crossref - Sanchez L, Weidmann S, Arnould C, Bernard AR, Gianinazzi S, Gianinazzi-Pearson V. Pseudomonas fluorescens and Glomus mosseae trigger DMI3-dependent activation of genes related to a signal transduction pathway in roots of Medicago truncatula.Plant Physiol. 2005; 139:1065-1077.
Crossref - Fridlender M, Inbar J, Chet I. Biological control of soilborne plant pathogens by a β-1, 3 glucanase-producing Pseudomonas cepacia. Soil Biol Biochem.1993; 25:1211-1221.
Crossref - Lopez-Gomez M, Sandal N, Stougaard J, Boller T. Interplay of flg22-induced defence responses and nodulation in Lotus japonicus. J Exp Bot 2012; 63:393-401.
Crossref - Maunoury N, Redondo-Nieto M, Bourcy M, et al. Differentiation of symbiotic cells and endosymbionts in Medicago truncatula nodulation are coupled to two transcriptome-switches. PLoS ONE. 2010; 5(3):e9519.
Crossref - Zdor RE, Anderson A. Influence of root colonizing bacteria on the defense responses of bean. Plant Soil. 1992; 140:99-107.
Crossref - Shirsat S, Nair P. The mode of inhibition of the biosynthesis of phenylalanine ammonia lyase by its product cinnamic acid in aging potato parenchyma tissue. J Biosci.1986; 10:393-402.
Crossref - Liu J, Maldonado-Mendoza I, Lopez-Meyer M, Cheung F, Town CD, Harrison MJ. Arbuscular mycorrhizal symbiosis is accompanied by local and systemic alterations in gene expression and an increase in disease resistance in the shoots. The Plant J. 2007; 50:529-544.
Crossref - Zamioudis C, Pieterse CMJ. Modulation of host immunity by beneficial microbes. Mol Plant Microbe Interact. 2011; 25:139-150.
Crossref - Li W, Raoult D, Fournier P-E. Bacterial strain typing in the genomic era. FEMS Microbiol Rev. 2009; 33:892-916.
Crossref - Moreau S, Verdenaud M, Ott T, et al. Transcription reprogramming during root nodule development in Medicago truncatula. PLoS ONE. 2011; 6(1):e16463.
Crossref - Jones JDG, Dangl JL. The plant immune system. Nature. 2006; 444:323-329.
Crossref
© The Author(s) 2021. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.