ISSN: 0973-7510
E-ISSN: 2581-690X
Denitrification based on immobilized microbial cellulose may offer an economical replacement for conventional treatment for nitrate removal. The environmental and bacterial biomass may influence the rate of biological denitrification processes. This study aimed to investigate the factors that affect denitrification rates, including carbon sources, pH, and bacterial inoculum. Different inoculum biomass of Pseudomonas aeruginosa and various carbon sources of glucose, sucrose, and cellulose with different concentrations were tested to assimilate 100 mg/L of KNO3 as nitrate source. Additionally, five additional inoculations, five different incubation time, and seven different pH levels were studied. The Pseudomonas aeruginosa isolates used different mineral media with three carbon sources, glucose, sucrose, and cellulose, with different concentrations at different rates to denitrify nitrate. The highest denitrification rate was with glucose after 18 hrs and was after 24 hrs when sucrose and cellulose were used, respectively. The bacterial biomass denitrification level was the highest, between 0.8% and 1% of OD600=1. Nitrate removal by Pseudomonas aeruginosa was the highest at pH 7, 8, and 9. This report suggests that when glucose is used as a carbon source, at neutral to alkaline pH, and 1% of denitrifying bacterial biomass, the highest level of biological denitrification process may be achieved.
Bacterial Biomass, Nitrification, Denitrification, Pseudomonas aeruginosa, carbon sources, Nitrate removal
Bacteria contribute to the nitrogen cycle, including nitrogen fixation, nitrification, and denitrification. The nitrogen removal by biological procedure involves a two-step process; nitrification and denitrification. The reduction of nitrate (NO3) to nitrogen (N2) by facultative bacteria such as Pseudomonas aeruginosa is known as a denitrification process. This process requires optimal conditions to occur naturally. A primary organic compound as a carbon source for energy, anoxic conditions so that the bacteria can use nitrate as a final electron acceptor to complete the denitrification process1. It was suggested that different organic carbon sources might give a different level of dentification and, consequently, the different mole fraction of nitrogen gas2.
The genus Pseudomonas is the most isolated and used denitrifying bacteria from soil and water samples. Another denitrifying Pseudomonas may include P. denitrificans, and P. stutzeri. The denitrifying bacteria rely primarily on the nir gene, which encodes for the periplasmic Nitrate Reductase for the denitrification process1,3,4.
Many studies have investigated the most efficient carbon sources for the biological denitrification processes. Carbon sources like wood sources, wheat straw, green waste, and maize cobs were explored for their efficacy in nitrate removal levels. Among all, Maize cobs were superior and had shown a high level of denitrification when it was used as a carbon source substrate due to its high glucose content. Wood sources were ideal for some bacteria with lesser extent by-products like total organic carbon and Nitrous oxide. Furthermore, the microbial composition may play a role in the denitrification process. It was shown that the nitrate level might influence the bacterial composition. Also, it could enhance the methane emissions rate in the denitrification process that depends on wood sources as a sole carbon source for the biodenitrification5.
The interaction between medium conditions and their contents to isolate denitrifying microbes was studied. It was shown that medium with a nitrogen concentration of 3 mM and neutral pH level with added vitamin with no riboflavin or sodium chloride was more appropriate to isolate denitrifying bacteria. Carbon sources such as succinate or ethyl alcohol were also supportive6. The environmental and bacterial biomass may influence the rate of biological denitrification processes. This study aimed to investigate the factors that affect denitrification rates, which may include carbon sources, pH, and bacterial inoculum. Different inoculum biomass of Pseudomonas aeruginosa and different carbon sources of glucose, sucrose, and cellulose with different concentrations were tested to assimilate 100 mg/L of KNO3 as nitrate source. Our primary study’s motivation was to design a cheap biological denitrification system that can aid in removing nitrate from water due to their scarce resource in Saudi Arabia. Designing biodenitrification system may require knowledge of the bacterial biomass and the nature of carbon sources used as electron donor for the denitrification process.
Isolation of Pseudomonas aeruginosa from soil
Soil samples were collected from different cities and regions in Saudi Arabia. At least 10 grams of soil samples were used to isolate denitrifying Pseudomonas aeruginosa. Approximately 90 ml of ddH2O was added to all soil samples to make a suspension for plating serially diluting bacteria on nutrient agar and Luria-Bertani Agar Medium medium at the level of 10-1 and 10-6. The plates were incubated at 30°C for 48 hours. The grown bacterial colonies on the agar were distinguished by colors, elevation, edge, and form. Individual well-spaced bacterial isolates were obtained by multiple subcultures on fresh agar plates 7,8. The identity of the isolates was confirmed by molecular techniques.
Storage of samples
The different isolated strains were incubated overnight in LB medium at 30°C or the temperatures representing their optimum growth. An aliquot from each was preserved by adding 50% sterile glycerol (v/v). Each bacterial isolate was preserved at -80°C for further use or by lyophilization for storage of the cells for a more extended period 9.
Molecular analysis of 16S rRNA, and nir Gene Sequencing
All bacterial isolates were confirmed by 16S rRNA, and nir gene was checked by nir gene primers. The PCR reactions were performed to amplify the 16S rRNA gene for microbial identification and to amplify the nir gene from bacterial isolates. The final volume of PCR reaction mixture was 25μl. The reaction mixture containing beads (beads contain buffer salts, nucleotides and Taq polymerase in a lyophilized powder), 3μl of DNA or lysate, 10 pmol of forward and reverse specific primers for nir gene F (TAY CAC CCS GAR CCG C), and nir gene R (CCG ACC GGT AAG TTC AAC GTG). Degenerate letters denote D (AGT), R (AG), M (AC), S (GC), B (GCT), N (AGCT), V (AGC), Y (CT), H (ACT), W (AT). The primers used for 16s rRNA amplification were as followed forward primer 8FLP (5’GCG GAT CCG CGG CTG CAG AGT TTG ATC CTG GCT CAG3’) and reverse primer 806R (5’GCG GAT CCG CGG CCG CGG ACT ACC AGG GTA TCT AAT3’) and ddH2O to a final volume of 25μl. The PCR was performed with the following cycling conditions to amplify the 16S rRNA gene or nir gene; 94°C for 5 min, then 35 cycles of denaturation at 94°C for 30 s, annealing at 57°C for 30 s, and extension at 73°C for 1 min followed by a final extension at 73°C for 10 min. The PCR amplification was analyzed by 1% agarose gel electrophoresis in 1X TBE buffer after it was stained with Ethidium Bromide. The gel image was captured by UV light at 254 nm in a UV transilluminator10,11. DNA Sequencing was performed to evaluate the data obtained by PCR using BigDye sequencing kit according to the manufacturer’s instructions (Thermofisher Scientific). The forward primer of each gene under investigation was used for cycle sequencing to generate sequencing data in a sequencer 3730XL DNA analyzer (Thermofisher Scientific). Sequencing data analysis was completed in BLAST algorithm available in the NCBI Genbank server (https://blast.ncbi.nlm.nih.gov/Blast.cgi). The 16S rRNA gene sequence identity used a criterion of 99-95% to speciate the bacteria or to identify the corresponding nir gene.
Optimization of nitrate denitrification by Pseudomonas aeruginosa
Pseudomonas Isolation was one of the isolates obtained from the field collections and has given promising preliminary denitrification results. Subsequently, it was selected for further analysis. The strain was grown in 100 ml Mineral medium broth in 250 ml conical flasks. Optimization of denitrification was performed under several condition, three different carbon sources were used (glucose, sucrose and cellulose) with different concentration (0.1%, 0.3%, 0.5%, 0.7% and 0.9%). The concentration of 100 mg/L of KNO3 was used. Five-time course incubation (2 h, 4 h, 8 h, 16 h, and 24 h) has also experimented. Finally, five different inoculations (200 ml, 400 ml, 600 ml, 800 ml, and 1000 ml), and seven different pH levels (3, 4, 5, 6, 7, 8, and 9) were investigated.
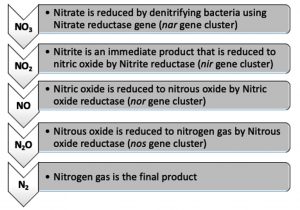
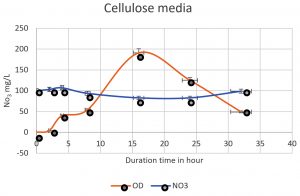
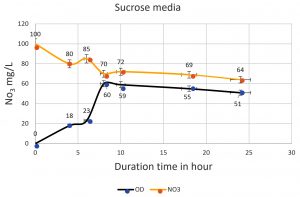
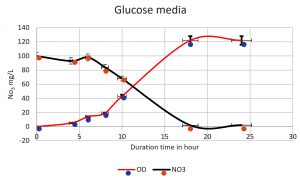
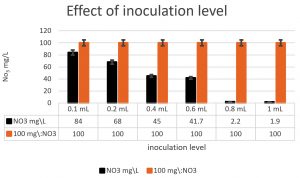
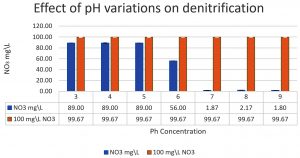
The denitrifying bacteria Pseudomonas aeruginosa from our isolates was identified and confirmed by molecular techniques (data not shown). Pseudomonas aeruginosa is able to denitrify the nitrate on mineral media with different sugar carbon sources due to their nir gene cluster through several chemical reactions to convert Nitrate to Nitrogen gas (Fig. 1). No major denitrification was observed with the use of cellulose as a carbon source even after 24 hours (Fig. 2). When sucrose was used as a carbon source, the highest reduction of nitrate was reached after 24 h (64 mg/L or 64%) by Pseudomonas aeruginosa (Fig. 3). The highest denitrification level was achieved after 18 hours with glucose as a carbon source (1.9 mg/L or 1.9%) (Fig. 4). The bacterial inoculum or biomass was investigated to assess how much bacteria is needed to give an optimum denitrification process. At least 100mg/L of nitrate was used as a fixed starting point. The maximum denitrification rate was achieved after 18h when the glucose media was inoculated with 0.8 % and 1% inoculum of OD600=1 (Fig. 5). The effect of pH variables on the denitrification processes was investigated. It has been shown that neutral to alkaline pH 7, 8, and 9 were suitable for the denitrification process by Pseudomonas aeruginosa on glucose media (Fig.6).
Fig.1. The denitrification pathway by denitrifying bacteria. it is a reduction process where the NO3 is reduced and converted to nitrogen gas through series of chemical reactions.
Fig. 2. The nitrate was denitrified by Pseudomonas aeruginosa on mineral media with cellulose. The highest denitrification level of nitrate was achieved after 16h (64 mg/L) off 100mg/L.
Fig. 3. The nitrate was reduced by Pseudomonas aeruginosa on mineral media with sucrose. The higher reduction of nitrate was reached after 24 h (64 mg/L). The bottom line shows the bacterial growth phase, and the top line shows the denitrification process starting from 100mg/L. The inverse relationship shows the duration of time from 0 to10 h. The direct relationship was started from 10h when bacteria began in the death phase.
Fig. 4. The nitrate was reduced on mineral media with glucose by Pseudomonas aeruginosa. The higher reduction of nitrate was reached at 18 h (1.9mg/L). An inverse relationship was shown in a time course from 0 to 18 h, Nitrate reduction was observed in log phase (the more bacterial growth, the higher the denitrification process).
Fig. 5. The bacterial inoculum was investigated to assess how much bacteria will give an optimum denitrification process. At least 100mg/L nitrate was used as a fixed starting point. The maximum denitrification was achieved after 18h when the glucose media was inoculated with 0.8 % and 1% of Pseudomonas aeruginosa inoculum of OD600=1.
The biological process of denitrification by denitrifying bacteria reduces the harmful effect on the environment. The biological conversion of nitrate to nitrogen gas by denitrifying bacteria may provide a low cost and efficient bioremediation treatment of contaminated groundwater. The biological treatment is controlled by anaerobic conditions, heterotrophic, and autotrophic denitrifying microbes12,13. The process of denitrification is initiated by microbes that can reduce nitrate to nitrogen gas through several chemical reactions where many reductase enzymes are used under optimal conditions such as pH, temperature, and perfect carbon source for energy. The denitrifying bacteria will have to find the ideal environmental conditions to process efficiently. These conditions may include good carbon sources, pH, and bacterial biomass. In this report, we aimed to investigate the factors that may affect the denitrification rates, which may include carbon sources, pH, and bacterial inoculum.
It appears that the source of carbon plays a significant role in the denitrification process by the bacteria by providing energy. Almost all bacterial species can metabolize glucose (C6H12O6) when used as a carbon source. The glucose is quickly metabolized by the bacteria because it is a simple sugar molecule that can enter the bacterial cell more efficiently than disaccharide or polysaccharide molecules. This substrate is degraded rapidly by the cell enzymatic machinery, and energy is produced14. Poor substrates such as sucrose or cellulose are not favorably absorbed or catalyzed by bacterial cells. Hydrolysis of substrate absorption is controlled by enzymes encoded by the genes in the bacterial chromosomes. Pseudomonas aeruginosa may slowly catalyze sucrose or cellulose molecules due to their enzymatic processivity level or reduced cellular absorbance to this kind of sugar9,15. The microbial production of cellulose is mainly dependent on carbon sources. It has been shown that the process by which the bacteria produce the cellulose is during Their active growth phase. Unlike plant cellulose, bacterial cellulose consists of linear β-1-4-glucan chains, a unique form found of cellulose found only in bacteria16.
Consistent with our findings, studies have shown that the rate of denitrification is superior between neutral to alkaline pH 7-9. Acidic environmental conditions have a negative effect on the level of denitrification by denitrifying bacteria. Studies have demonstrated that the denitrification potential of synthetic wastewater was inhibited below the neutral pH seven but was restored between pH7.5-9 17.
Interestingly, the pH level, along with the appropriate carbon source, may accelerate the nitrate removal and improve the denitrification process. The denitrification process is pH-dependent. It was observed that the pH 7, 8, 9 were the most efficient in the denitrification. This is because most denitrifying bacteria have an ideal pH level between 7-9. The denitrification progression may be halted or substantially declined If pH levels shift towards acidity (less than 6) or alkalinity (more than 9).
Additionally, the process of denitrification by denitrifying bacteria increase alkalinity and pH level18-20. Bacterial biomass may play an important factor in speeding the denitrification process. It was shown that suspended bacterial biomass might be superior over immobilized microorganisms in the denitrification process21,22. Our report showed that using an inoculum of 0.8% and 1% of OD600=1 was sufficient to accelerate the bacterial biomass to denitrify nitrate to 1.9% after 18 hours. It is thought that increasing the bacterial biomass of certain bacterial species and their enzymes may accelerate the process of denitrification19.
In this study, we aimed to investigate the effect of bacterial biomass level and the nature of carbon source for efficient denitrification process. The use of biological nitrate removal by denitrifying bacteria may offer a cost-effective approach. Optimizing the conditions for such treatment, such as defining the carbon source, working under ideal pH levels, and selecting the most appropriate bacterial biomass are considered essentials for the success of biological nitrate removal. The results of this report confirm that carbon source from glucose provides the best conditions for nitrate removal by the denitrifying bacteria. Additionally, suitable bacterial biomass at the neutral to alkaline pH levels may contribute to the highest level of biological denitrification.
ACKNOWLEDGMENTS
All authors would like to thank KACST and KSU for their full support during this project.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors designed the experiments. EA and RB performed the experiments. EA, RB, AB, SA analyzed and interpreted the data. EA, RB gathered information from the literature and drafted the manuscript. AB, SA reviewed the final manuscript. All authors have made a substantial and intellectual contribution to this manuscript’s work and approved it for publication.
FUNDING
This manuscript contains the results and findings of a research project funded by King Abdulaziz City for Science and Technology (KACST) (Grant No. ARP.30.34).
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the manuscript.
- Bernat K, Wojnowska-Baryla I. Carbon source in aerobic denitrification. Biochemical Engineering Journal. 2007;36(2):116-122.
Crossref - Knowles R. Denitrification. Microbiol Rev. 1982;46(1):43-70.
Crossref - Chen Q, Ni J. Heterotrophic nitrification-aerobic denitrification by novel isolated bacteria. J Ind Microbiol Biotechnol. 2011;38(9):1305-1310.
Crossref - Chen D, Yang K, Wang H, Lv B, Ma F. Characteristics of nitrate removal in a bio-ceramsite reactor by aerobic denitrification. Environmental Technology. 2015; 36(11):1457-1463.
Crossref - GrieBmeier V, Gescher J. Influence of the potential carbon sources for field denitrification beds on their microbial diversity and the fate of carbon and nitrate. Front Microbiol. 2018;9:1313.
Crossref - Heylen K, Vanparys B, Wittebolle L, Verstraete W, Boon N, De Vos P. Cultivation of denitrifying bacteria: optimization of isolation conditions and diversity study. Appl Environ Microbiol. 2006;72(4):2637-2643.
Crossref - Kross BC, Hallberg GR, Bruner DR, Cherryholmes K, Johnson JK. The nitrate contamination of private well water in Iowa. Am J Public Health. 1993;83(2):270-272.
Crossref - Shaaban MT, Attia M, Turky AS, Mowafy EI. Production of some biopolymers by some selective Egyptian soil bacterial isolates. Journal of Applied Sciences Research. 2012:94-105.
- Blaszczyk M. Effect of Medium Composition on the Denitrification of Nitrate by Paracoccus denitrificans. Appl Environ Microbiol. 1993;59(11):3951-3953.
Crossref - Braker G, Fesefeldt A, Witzel K-P. Development of PCR primer systems for amplification of nitrite reductase genes (nirK and nirS) to detect denitrifying bacteria in environmental samples. Appl Environ Microbiol. 1998;64(10):3769-3775.
Crossref - Tindall BJ, Sikorski J, Smibert RA, Krieg NR. Phenotypic characterization and the principles of comparative systematics. Methods for General and Molecular Microbiology, Third Edition. American Society of Microbiology. 2007:330-393.
Crossref - Koren D, Gould W, Bedard P. Biological removal of ammonia and nitrate from simulated mine and mill effluents. Hydrometallurgy. 2000;56(2):127-144.
Crossref - Soares M. Biological denitrification of groundwater. Water, Air, and Soil Pollution. 2000;123(1-4):183-193.
Crossref - Bren A, Park JO, Towbin BD, Dekel E, Rabinowitz JD, Alon U. Glucose becomes one of the worst carbon sources for E. coli on poor nitrogen sources due to suboptimal levels of cAMP. Scientific Reports. 2016;6(1):1-10.
Crossref - Peng Y-z, Yong M, Wang S-y. Denitrification potential enhancement by addition of external carbon sources in a pre-denitrification process. Journal of Environmental Sciences. 2007;19(3):284-289.
Crossref - Lynd LR, Weimer PJ, van Zyl WH, Pretorius IS. Microbial Cellulose Utilization: Fundamentals and Biotechnology. Microbiol Mol Biol Rev. 2002;66(3):506-577.
https://doi.org/10.1128/MMBR.66.3.506-577.2002 - Skiba U. Denitrification. 2008.
- Denitrifying Bacteria. In: Wastewater Bacteria.91-101.
- Gerardi M. » Bacterial Growth «. Wastewater Bacteria John Wiley & Sons, Inc. 2006:65-73.
Crossref - Rezaee A, Godini H, Dehestani S, Yazdanbakhsh AR, Mosavi G, Kazemnejad A. Biological denitrification by Pseudomonas stutzeri immobilized on microbial cellulose. World J Microbiol Biotechnol. 2008;24(11):2397-2402.
Crossref - Costa DD, Gomes AA, Fernandes M, da Costa Bortoluzzi RL, Magalhaes MdLB, Skoronski E. Using natural biomass microorganisms for drinking water denitrification. J Environ Manage. 2018;217:520-530.
Crossref - Drury CF, McKenney DJ, Findlay WI. Relationships between denitrification, microbial biomass and indigenous soil properties. Soil Biology and Biochemistry. 1991;23(8):751-755.
Crossref
© The Author(s) 2020. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.