ISSN: 0973-7510
E-ISSN: 2581-690X
Antibiotic resistant and biofilm forming uropathogenic Enterobacteriaceae are rising. This study was conducted to evaluate the efflux pump and plasmid mediated efflux genes in ciprofloxacin (CIP) resistant hospital acquired uropathogenic Escherichia coli and Klebsiella pneumoniae. Also, to assess the anti-biofilm action of carbonyl cyanide m-chlorophenylhydrazone (CCCP). Uropathogenic E. coli and K. pneumoniae isolates were collected from Mansoura University Hospitals in Mansoura, Egypt. The effect of Sub- minimum inhibitory concentration (MIC) of CCCP on CIP MIC was evaluated and the MIC decrease factor (MDF) was calculated. The presence of oqxAB and qepA genes was detected by PCR. The effect CCCP on biofilm was detected in strong biofilm formers. 56 and 47 CIP-resistant uropathogenic E. coli and K. pneumoniae isolates respectively were detected. Significant MDF by CCCP was observed in 55.3% of these isolates. The qepA gene was only present in E. coli. However, oqxAB genes were found only in K. pneumoniae. Biofilm formation was detected in 58.9% and 72.3% of CIP-resistant E. coli and K. pneumoniae isolates, respectively. Biofilm formation was significantly decreased by CCCP. According to these findings, CIP resistance and plasmid-mediated efflux pumps in uropathogenic E. coli and K. pneumoniae are of rising concern. Efflux pump inhibitor CCCP represents a possible option to decrease the biofilm formation in these resistant urinary pathogens.
Ciprofloxacin Resistance, Biofilm, CCCP, Efflux Pump
Urinary tract infection (UTI) represents a predominant infection in both healthcare settings and community.1 The most common bacterial cause of UTI is Escherichia coli, but many other Enterobacteriaceae such as Klebsiella pneumoniae are also present.2
The Multidrug-resistant (MDR) Enterobacteriaceae is increasing as a cause of UTI.3 Biofilm formation plays a significant role in UTIs.4 When these uropathogens colonize the mucous membrane of the urinary tract, they tend to form biofilm, which favors their persistence for a long time. In addition, microbial biofilms are impermeable to many antibiotics that contribute to the development of MDR bacteria.5
Fluoroquinolones (FQs) represent one of first-line agents for UTI empirical therapy.6 Efflux pumps (EPs); QepA and oqxAB are important contributors of quinolones resistance in Enterobacteriaceae including E. coli, Klebsiella spp. These pumps actively extrude FQs from the cytoplasm. QepA is a proton-dependent transporter that belongs to the superfamily of major facilitators. QepA promotes resistance to quinolone,7 whereas OqxAB is a multidrug efflux pump with transmissible resistance-nodulation division that decreases bacterial susceptibility to ciprofloxacin (CIP).8
These EPs can be blocked by in vitro inhibitors such as carbonyl cyanide 3-chlorophenylhydrazone (CCCP). The effect of EP inhibitor CCCP against resistant Gram-negative bacteria has been investigated.9 This study was aimed at finding out the role of EP in CIP resistance of E. coli and K. pneumoniae causing hospital acquired UTI. In addition, the prevalence of plasmid-mediated EP genes, biofilm, and the anti-biofilm effect of CCCP against these isolates were investigated.
Bacterial Identification
A Prospective cross-sectional study was carried out for 18 months from March 2019 to August 2020 in Mansoura University hospitals (MUHs), Mansoura, Egypt. The Institutional Research Board at faculty of medicine, Mansoura University has approved this study (with a code number R.21.09.1463). Each patient included in this study signed an informed consent. Urine samples were collected from patients with suspected UTIs. Cystine–lactose–electrolyte-deficient (CLED) agar (Oxoid- UK) plates were used to culture collected urine samples according to method described before.10
Routine microbiological methods were used to identify bacterial isolates, biochemical reactions (Kligler iron agar, oxidase test, lysine iron agar, methyl red, Voges-Proskauer test, citrate tests, motility indole ornithine).10 API 20E (BioMerieux) was used to confirm the bacterial identification.
Antimicrobial Susceptibility Testing
The antibiotic susceptibilities of E. coli and K. pneumoniae isolates were determined by disc diffusion method according to guidelines of Clinical and Laboratory Standards Institute(CLSI).11 The tested antibiotics include: gentamicin, ciprofloxacin, cefuroxime, cefotaxime, ceftazidime, cefoxitin, amoxicillin/clavulanic, piperacillin/tazobactam, imipenem, trimethoprim/sulfamethoxazole, and aztreonam. Quality control was done using E. coli ATCC®25922TM strain. Criteria for multidrug-resistant (MDR) phenotypes were defined as described before.12
Determination of CIP and CCCP Minimum Inhibitory Concentrations
Broth micro dilution assay was done for determining CIP and CCCP minimum inhibitory concentrations (MICs) in E. coli and K. pneumoniae isolates. The procedure and break points interpretation were done according to CLSI recommendations.13 Pure powders of CIP and CCCP were acquired from Sigma Aldrich (St. Louis, MO, USA). E. coli ATCC®25922TM strain was used as Quality control.
Evaluating the Effect of EP Inhibitor CCCP on CIP Resistance
To detect the EP mechanism, the effect of EP inhibitor CCCP on CIP MIC was determined.14 CIP MIC in CIP resistant isolates was re-measured in the presence of a sub-MIC of CCCP (i.e. 0.5 × MIC) using serially rising concentrations of the antibiotics.
The MIC decrease factor (MDF) was calculated for each isolate using this formula; MDF = CIP MIC in the absence of CCCP/ CIP MIC in presence of CCCP.14 With MDF value of 4 or above, significant inhibition of EP by CCCP was considered (in the presence of CCCP, the CIP MIC drops fourfold or more).14
Plasmid Mediated EP Genes
Plasmid EP genes were assessed in CIP resistant isolates as previously described by two PCR: one PCR for oqxAB genes7 and the other PCR for the detection of qep A gene8 (Table 1).
Table (1):
Primers of plasmid EP genes.
Gene |
Sequence of primer |
Annealing temperature |
Product size |
Reference |
|---|---|---|---|---|
oqxA |
CTCGGCGCGATGATGCT CCACTCTTCACGGGAGACGA |
68°C |
392 |
7 |
oqxB |
TTCTCCCCCGGCGGGAAGTAC CTCGGCCATTTTGGCGCGTA |
64°C |
512 |
7 |
qepA |
GCA GGT CCA GCA GCG GGT AG CTT CCT GCC CGA GTA TCG TG |
60°C |
199 |
8 |
Detection of Biofilm
Biofilm formation assays were performed for CIP resistant isolates as previously described.15 Overnight culture of isolates was adjusted to turbidity 0.5 McFarland standard. Then for each isolate, 125 microliters of 1:100 diluted bacterial suspensions were put in sterile flat-bottomed 96-well microtiter plates and incubated at 37°C for 24 hours. The wells were washed thrice with 300 mL of sterile phosphate buffered saline (PH 7.3) and dried inverted at room temperature. For staining of the formed biofilm, 125 µL crystal violet of 0.1 percent was added for 10–15 minutes followed by rinsing with distilled water for three times. Uninoculated medium was used as control. The experiment was done in triplicates. A microtiter plate was read at the optical density (OD) 600 nm. The strains were then categorized using the following criteria: The cut-off OD (ODc)= mean OD of the negative control+ three standard deviations (SD). Biofilm were classified as strong biofilm producer when (OD more than 4×ODc); OD of non biofilm producer (OD equal to or less than ODc); weak biofilm producer (ODc< OD≤2x ODc); moderate biofilm producer (2 x ODc< OD≤4 x ODc).16
The Effect of Sub-MICs of CCCP on Biofilm Formation
The effect of CCCP on the formation of biofilm were assessed on strong biofilm producing CIP resistant isolates of K. pneumoniae and E. coli.
The biofilm assays were done as above but with the addition of sub MIC CCCP concentration of each strain compared to biofilm formation in absence of CCCP as control. The microtitre plate was read at the OD600nm after 24 h incubation at 37°C.
Statistical Analysis
After the KS test for normality, quantitative data will be described as mean (SD) or median (range). parametric tests were used for normally distributed data, while non parametric tests were used for non normally distributed data. A P value is considered statistically significant when it is < 0.05.
Bacterial Isolates
During the study period, 159 isolates (88 E. coli and 71 K. pneumoniae) were identified from a total of 279 cases of suspected UTIs. These urinary bacterial isolates were obtained from 89 (56%) females and 70 (44%) from males. The age of patients ranged from 19 to 77 years (48.4±13.3). Out of these 159 isolates, 56 isolates E. coli and 47 K. pneumoniae (64.8%) were CIP resistant (Table 2).
Table (2):
Antibiotic resistance pattern of uropathogenic E. coli and K. pneumoniae.
Antibiotic |
E. coli Total = 88 N (%) |
K. pneumoniae Total =71 N (%) |
|---|---|---|
Gentamicin |
24 (27.3) |
41 (57.7) |
Ciprofloxacin |
56 (63.6) |
47 (66.2) |
Cefuroxime |
76 (86.4) |
54 (76.1) |
Cefoxitin |
46 (52.3) |
36 (50.7) |
Cefotaxime |
66 (75) |
51 (71.8) |
Ceftazidime |
60 (68.2) |
55 (77.5) |
Amoxicillin / clavulanic acid |
67 (76.1) |
56 (78.9) |
Piperacillin-tazobactam |
30 (34.1) |
34 (47.9) |
Imipenem |
38 (43.2) |
55 (72.5) |
Trimethoprim-sulfamethoxazole |
54 (61.4) |
47 (66.2) |
Aztreonam |
46 (76.1) |
39 (54.9) |
Antibiotics Sensitivity Testing
The highest resistance among E. coli isolates was cefuroxime (86.4%) followed by amoxicillin / clavulanic acid (76.1%) and the least was gentamicin (27.3%). Regarding K. pneumoniae, the highest resistance was found in amoxicillin / clavulanic acid (78.9%) and cefuroxime (76.1%) and the least was in piperacillin-tazobactam (47.9%). CIP resistance was reported in 63.6% of E. coli and 66.2% of K. pneumoniae isolates (Table 2). 93% of the E. coli and 95% of K. pneumoniae were MDR.
Ciprofloxacin Resistance
Ciprofloxacin MIC values of isolated E. coli and K. pneumoniae ranged from 0.125 to 256 mg/L using the broth microdilution method. One hundred and three isolates were CIP resistant. CIP MIC estimates ranged from 1–256 mg/L. Sixty-four isolates (62.1%)of the isolates were high-level CIP resistance (MIC ≥ 32 mg/L), the MIC 50 and MIC90 were 32 mg/L and 256 mg/L, respectively.
Activity of EP
Reduction of fourfold or more CIP MIC in the presence of CCCP (MDF value of higher than 4) was detected in 57(55.3%) CIP resistant isolates.
Plasmid Mediated EP genes in Quinolone Resistant Isolates
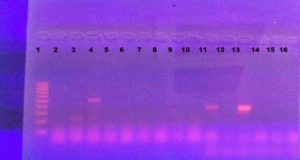
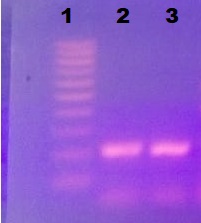
As regard resistant K. pneumoniae, both OqxA and OqxB genes were detected together in 8 isolates, while OqxA gene only was detected in 12 isolates. OqxA nor OqxB was not detected in any of E. coli isolates. qepA was detected in 12 isolates of E. coli (Figure 1, 2).
Figure 1. Detection of oqxAB genes by PCR to detect oqxA (392 bp) and oqxB (512bp). Lane 1 showing 100 pb DNA ladder Lane 4 showed oqxB. Lane 11 and 13 showed oqxA
Figure 2. Detection of qepA gene (199bp) by PCR. Lane 1 showing 100 pb DNA ladder and ladder 2, 3 showed qebA
Biofilm Formation in CIP Resistant Isolates
Among CIP resistant isolates, biofilm formation was detected in 33 (58.9%) of E. coli and 34(72.3%)of K. pneumoniae isolates. Six of E. coli in addition to 8 K. pneumoniae isolates expressed strong biofilm production capacity (Table 3). MICs of CCCP, CIP and resistance genes among strong biofilm producers were summarized in Table 4.
Table (3):
Biofilm formation in CIP resistant uropathogenic E.coli and K. pneumoniae.
E.coli 56 N (%) |
K. pneumonia 47 N (%) |
|
|---|---|---|
Strong biofilm formers |
6 (10.7) |
8(17) |
Moderate biofilm formers |
16(28.6) |
14(29.8) |
Weak biofilm formers |
11 (19.6) |
12(25.5) |
Non biofilm formers |
23 (41.1) |
13(27.7) |
Table (4):
MIC and plasmid mediated EP genes among strong biofilm producing CIP resistant E. coli and K. pneumoniae.
| Strain number | MIC | OqxA | OqxB | qepA | |||
|---|---|---|---|---|---|---|---|
| Species | CIP | CCCP | CIP+ subMIC CCCP (MDF) | ||||
| 7 | E. coli | 256.0 | 32.0 | 128 (2) | – | – | – |
| 67 | E. coli | 256.0 | 32.0 | 32 (8) | – | – | + |
| 70 | E. coli | 256.0 | 32.0 | 64 (4) | – | – | – |
| 9 | E. coli | 256.0 | 16.0 | 32 (8) | – | – | + |
| 89 | E. coli | 128.0 | 16.0 | 128 (0) | – | – | – |
| 81 | E. coli | 128.0 | 32.0 | 32 (4) | – | – | – |
| 942 | K. pneumoniae | 128.0 | 32.0 | 32 (4) | – | – | – |
| 959 | K. pneumoniae | 64.0 | 16.0 | 32 (2) | + | – | – |
| 824 | K. pneumoniae | 4.0 | 16.0 | 2 (2) | – | – | – |
| 6 | K. pneumoniae | 256.0 | 16.0 | 32 (8) | – | – | |
| 72 | K. pneumoniae | 4.0 | 32.0 | 2 (2) | – | – | – |
| 65 | K. pneumoniae | 4.0 | 64.0 | 1 (4) | – | – | – |
| 60 | K. pneumoniae | 256.0 | 32.0 | 128 (2) | + | – | – |
| 363 | K. pneumoniae | 256.0 | 32.0 | 32 (8) | + | + | – |
Effects of CCCP sub-MIC on biofilm formation
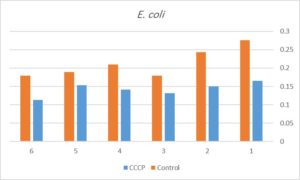
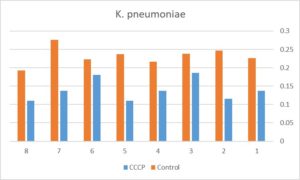
The effect of sub-MIC concentration of CCCP was evaluated on biofilm forming capacity of strong biofilm producing isolates (six E. coli and eight K. pneumoniae isolates). The results showed that sub-MIC of CCCP significantly inhibited the formation of E. coli and K. pneumoniae biofilms (P equal 0.002 and 0.0001 respectively) (Figure 3).
Treatment of MDR bacteria in UTI represents a challenge in healthcare settings.3 MDR organisms expressed simultaneous resistance to common antibiotics that are used in the treatment of UTI such as aminoglycosides, beta-lactams and FQs that are considered a great concern.17 Plasmid exchange of resistance aggravates the problem and represents major threat to health care system.18 In this study, 93% and 95% of isolated urinary E. coli and K. pneumoniae were MDR, respectively. Similar high percentage of MDR in uropathogenic E. coli in a study that was conducted in Egypt, 90.85% were MDR,19 but lower percentage of MDR E. coli was reported in previous studies in Saudi arabia20 and Australia.21
Resistance to broad spectrum 2nd generation (Cefuroxime) cephalosporin and amoxicillin-clavulanic acid was found to be the highest in uropathogenic E. coli representing (86.4%, 76.1%) and in K. pneumoniae representing (76.1%, 78.9%). Resistance to 3rd generation cephalosporin cefotaxime and ceftazidime in E. coli was 75%and 68.2%, respectively and in K. pneumoniae was 71.8% and 77.5%, respectively. Imipenem resistance was presented in about forty percentages (43.2%) of isolated uropathogenic E. coli and about seventy percentages (72.5%) of K. pneumoniae isolates.
Resistance to beta lactam antibiotics is a global phenomenon, but occurrence rates are variable. Matching with the result of this study, high resistance rate to 2nd and 3rd generations cephalosporin was found by Tandogdu22 in Klebsiella and by Esmaeel et al.23 in uropathogenic E. coli isolates in Egypt. Similarly, high prevalence of amoxicillin–clavulanic acid resistance was detected in K. pneumoniae,24 and in E. coli in Asian area.22 However, lower rate of resistance was detected in K. pneumoniae in Tunisia.25
About 64.8% of the isolates were CIP resistant including 62.1% of the isolates that showed high CIP resistance (MIC >32). This result consistent with previous report.23 High rates of CIP resistance 75% and 91.4% was reported in urinary isolates of E. coli in other studies in tertiary hospitals. High CIP resistance was described previously.26 Lower rate of CIP resistance (33%)was reported in urinary K. pneumoniae.24
This high rate of resistance in this study to beta-lactam antibiotics and FQ could be due to their wide use as an empirical therapy for community acquired UTIs and gastrointestinal infection representing a selective pressure for development of resistance in patient’s flora, which represents the main source of UTI infection. In addition, the use of these agents empirically to treat UTIs with the absence of antibiotic policy in the hospital may represent another factor for this high rate of resistance.27
Numerous mechanisms are responsible for FQ resistance for example decreases in membrane permeability, efflux systems, and topoisomerase mutations.28 EP genes that transfer quinolones outside the bacterial cytoplasm are involved in plasmid-mediated quinolone resistance. Two of them are well known; qepAB and oqxAB.29 By transporting antibiotics across the membrane, these EP systems OqxAB increased CIP and norfloxacin MICs 32- and 64-fold, respectively, above the control strain. Because OqxAB conferred CIP resistance on other Enterobacteriaceae, the spread of this FQ resistance mechanism is very likely.
The over expression of OqxAB and QepA mediates resistance to many antibiotics including quinolone. This plasmid-borne MDR-EP may represent a major problem, as it may pose resistance to many antimicrobial agents in addition to the spread of this resistance via horizontal transfer.30
The efflux systems as a cause of drug resistance in Enterobacteriaceae is growing.9 EP inhibitor CCCP was used to reveal the contribution of EPs in CIP resistance and MIC of CIP was assessed in the presence of sub-inhibitory concentrations of CCCP. Reduction in CIP MIC (MDF ≥ 4) was as a principle for significance, efflux activity was present in 57 (55.3%) of CIP resistant isolates. Likewise, the role drug EP was detected in 47.5% of CIP resistant isolates of K. pneumoniae.31 While the role of EP was at lower rate that range from 12.1% and 18.3% of isolates of Enterobacteriaceae in other study.14 Higher rate of EP inhibition by CCCP was previously reported with CIP MIC reduction ranged from 2-64 fold in 93% of MDR k. pneumoniae.32
As regard plasmid-mediated EPs, QepA– was found in E. coli , while oqxAB was detected in K. pneumoniae only. OqxA gene was the highest prevalence in K. pneumoniae (20 isolates) including eight isolates had both OqxA and OqxB genes. qepA was found in 12 isolates of E. coli. The same was reported in a study conducted in CIP resistant isolates in Egypt.14 Also efflux pump qepA was detected in 12.3% of CIP resistant bacteria isolated from UTI.33 Higher rate of oqxAB was detected in K. pneumoniae but it was not detected in E. coli. Moreover, QepA was not detected in any of Enterobacteriaceae species in a study conducted in South Africa.34 Also, QepA was not detected in other studies in India and South Africa.26,35
In this study, biofilm was produced in 60% and 75.5% of quinolone resistant E. coli and K. pneumoniae, respectively. A similar higher rate was reported in Egypt,19 Ethiopia36 and Iran,37 while, a lower rate of biofilm production Klebsiella and E. coli was also described.38,39
This study assessed the ability of EP inhibitor CCCP to inhibit biofilm formation in strong biofilm forming isolates with EP activity. Biofilm formation was inhibited by sub-MICs of CCCP in both E. coli and K. pneumoniae. The same was previously in a study showed inhibition of biofilm formation of Salmonella typhimurium by CCCP.40 Also, a study investigates K. pneumoniae biofilm showed CCCP inhibition of biofilms in a dose-dependent effect. With the effective inhibitory concentration of CCCP towards K. pneumoniae EP was 10 μg/mL.41
CIP resistance in hospital acquired uropathogenic E. coli and K. pneumoniae is of high prevalence and plasmid mediated EP is of rising concern. Biofilm development is a critical virulence determinant in these organisms. EP inhibitor CCCP represents a possible option to decrease the biofilm formation in these resistant urinary pathogens.
ACKNOWLEDGMENTS
None.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
RE designed the study. RE and GM conducted experiments. RE wrote the draft manuscript. RE and GM wrote and revised the manuscript. Both authors read and approved the final manuscript for publication.
FUNDING
None.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
The Institutional Research Board at faculty of Medicine, Mansoura University approved this study with code number R.21.09.1463.
- Erdem I, Ali RK, Ardic E, Omar SE, Mutlu R, Topkaya AE. Community-acquired Lower Urinary Tract Infections: Etiology, Antimicrobial Resistance, and Treatment Results in Female Patients. J Glob Infect Dis. 2018;10(3):129-132.
Crossref - Tandogdu Z, Wagenlehner FME. Global epidemiology of urinary tract infections. Curr Opin Infect Dis. 2016;29(1):73-79.
Crossref - Chowdhury N, Suhani S, Purkaystha A, et al. Identification of AcrAB-TolC Efflux Pump Genes and Detection of Mutation in Efflux Repressor AcrR from Omeprazole Responsive Multidrug-Resistant Escherichia coli Isolates Causing Urinary Tract Infections. Microbiol Insights. 2019;12:1178636119889629.
Crossref - De Souza GM, Neto ERDS, da Silva AM, et al. Comparative Study Of Genetic Diversity, Virulence Genotype, Biofilm Formation And Antimicrobial Resistance Of Uropathogenic Escherichia coli (UPEC) Isolated From Nosocomial And Community Acquired Urinary Tract Infections. Infect Drug Resist. 2019;12:3595-3606.
Crossref - Sharma S, Bhat GK, Shenoy S. Virulence factors and drug resistance in Escherichia coli isolated from extraintestinal infections. Indian J Med Microbiol. 2007;25(4):369-373.
Crossref - Mirzaii M, Jamshidi S, Zamanzadeh M, et al. Determination of gyrA and parC mutations and prevalence of plasmid-mediated quinolone resistance genes in Escherichia coli and Klebsiella pneumoniae isolated from patients with urinary tract infection in Iran. J Glob Antimicrob Resist. 2018;13:197-200.
Crossref - Kim HB, Wang M, Park CH, Kim E-C, Jacoby GA, Hooper DC. oqxAB encoding a multidrug efflux pump in human clinical isolates of Enterobacteriaceae. Antimicrob Agents Chemother. 2009;53(8):3582-3584.
Crossref - Yamane K, Wachino J-I, Suzuki S, et al. New plasmid-mediated fluoroquinolone efflux pump, QepA, found in an Escherichia coli clinical isolate. Antimicrob Agents Chemother. 2007;51(9):3354-3360.
Crossref - Poole K. Efflux-mediated resistance to fluoroquinolones in gram-negative bacteria. Antimicrob Agents Chemother. 2000;44(9):2233-2241.
Crossref - Forbes BA, Sahm DF, Weissfeld AS. Bailey & Scott’s Diagnostic Microbiology. 1999.
- Weinstein MP, Limbago B, Patel J, et al. M100 performance standards for antimicrobial susceptibility testing. Clin Lab Stand Inst. 2018;38:1-258.
- Magiorakos AP, Srinivasan A, Carey RB, et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect. 2012;18(3):268-281.
Crossref - Wikler MA. Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically: approved standard. CLSI. 2006;26:M7-A7.
- Hamed SM, Elkhatib WF, El-Mahallawy HA, Helmy MM, Ashour MS, Aboshanab KMA. Multiple mechanisms contributing to ciprofloxacin resistance among Gram negative bacteria causing infections to cancer patients. Sci Rep. 2018;8(1):12268.
Crossref - O’Toole GA. Microtiter dish biofilm formation assay. J Vis Exp. 2011;(47):2437.
Crossref - Stepanovic S, Vukovic D, Hola V, et al. Quantification of biofilm in microtiter plates: overview of testing conditions and practical recommendations for assessment of biofilm production by staphylococci. APMIS. 2007;115(8):891-899.
Crossref - Mishra MP, Debata NK, Padhy RN. Surveillance of multidrug resistant uropathogenic bacteria in hospitalized patients in Indian. Asian Pac J Trop Biomed. 2013;3(4):315-324.
Crossref - Mathers AJ, Peirano G, Pitout JDD. The role of epidemic resistance plasmids and international high-risk clones in the spread of multidrug-resistant Enterobacteriaceae. Clin Microbiol Rev. 2015;28(3):565-591.
Crossref - Gawad WE, Helmy OM, Tawakkol WM, Hashem AM. Antimicrobial Resistance, Biofilm Formation, and Phylogenetic Grouping of Uropathogenic Escherichia coli Isolates in Egypt: The Role of Efflux Pump-Mediated Resistance. Jundishapur J Microbiol. 2018;11(2).
Crossref - Alqasim A, Abu Jaffal A, Alyousef AA. Prevalence of Multidrug Resistance and Extended-Spectrum b-Lactamase Carriage of Clinical Uropathogenic Escherichia coli Isolates in Riyadh, Saudi Arabia. Int J Microbiol. 2018;2018:3026851.
Crossref - Fasugba O, Das A, Mnatzaganian G, Mitchell BG, Collignon P, Gardner A. Incidence of single-drug resistant, multidrug-resistant and extensively drug-resistant Escherichia coli urinary tract infections: An Australian laboratory-based retrospective study. J Glob Antimicrob Resist. 2019;16:254-259.
Crossref - Tandogdu Z, Cek M, Wagenlehner F, et al. Resistance patterns of nosocomial urinary tract infections in urology departments: 8-year results of the global prevalence of infections in urology study. World J Urol. 2014;32(3):791-801.
Crossref - Esmaeel NE, Gerges MA, Hosny TA, Ali AR, Gebriel MG. Detection of chromosomal and plasmid-mediated quinolone resistance among Escherichia coli Isolated from urinary tract infection cases; Zagazig University Hospitals, Egypt. Infect Drug Resist. 2020;13:413.
Crossref - El Bouamri MC, Arsalane L, El Kamouni Y, Zouhair S. Antimicrobial susceptibility of urinary Klebsiella pneumoniae and the emergence of carbapenem-resistant strains: A retrospective study from a university hospital in Morocco, North Africa. African J Urol. 2015;21(1):36-40.
Crossref - Khalifa ABH, Khedher M. Epidemiological study of Klebsiella spp. uropathogenic strains producing extended-spectrum β-lactamase in a Tunisian university hospital, 2009. Pathol Biol (Paris). 2012;60(2):e1-5.
- Piekarska K, Wołkowicz T, Zacharczuk K, et al. Co-existence of plasmid-mediated quinolone resistance determinants and mutations in gyrA and parC among fluoroquinolone-resistant clinical Enterobacteriaceae isolated in a tertiary hospital in Warsaw, Poland. Int J Antimicrob Agents. 2015;45(3):238-243.
Crossref - Baquero F, Negri M-C, Morosini M-I, Blazquez J. Antibiotic-selective environments. Clin Infect Dis. 1998;27(Suppl 1):S5-S11.
Crossref - Hooper DC, Jacoby GA. Topoisomerase inhibitors: fluoroquinolone mechanisms of action and resistance. Cold Spring Harb Perspect Med. 2016;6(9):a025320.
Crossref - Jacoby GA. Plasmid-Mediated Quinolone Resistance BT – Antimicrobial Drug Resistance: Mechanisms of Drug Resistance. In: Mayers DL, Sobel JD, Ouellette M, Kaye KS, Marchaim D, eds. Springer International Publishing; 2017;1:265-268.
Crossref - Li J, Zhang H, Ning J, et al. The nature and epidemiology of OqxAB, a multidrug efflux pump. Antimicrob Resist Infect Control. 2019;8:44.
Crossref - Pakzad I, Zayyen Karin M, Taherikalani M, Boustanshenas M, Lari AR. Contribution of AcrAB efflux pump to ciprofloxacin resistance in Klebsiella pneumoniae isolated from burn patients. GMS Hyg Infect Control. 2013;8(2):Doc15.
Crossref - Albarri O, AlMatar M, Ocal MM, Koksal F. Overexpression of Efflux Pumps AcrAB and OqxAB Contributes to Ciprofloxacin Resistance in Clinical Isolates of K. pneumonia. Curr Protein Pept Sci. 2022;23(5):356-368.
Crossref - Adekanmbi AO, Usidamen S, Akinlabi OC, Olaposi AV, Onilude AA. Carriage of plasmid-mediated qnr determinants and quinolone efflux pump (qepA) by ciprofloxacin-resistant bacteria recovered from Urinary Tract Infection (UTI) samples. Bull Natl Res Cent. 2022;46(1):1-7.
Crossref - Osei Sekyere J, Amoako DG. Genomic and phenotypic characterisation of fluoroquinolone resistance mechanisms in Enterobacteriaceae in Durban, South Africa. PLoS One. 2017;12(6):e0178888.
Crossref - Shetty SS, Deekshit VK, Jazeela K, et al. Plasmid-mediated fluoroquinolone resistance associated with extra-intestinal Escherichia coli isolates from hospital samples. Indian J Med Res. 2019;149(2):192-198.
Crossref - Awoke N, Kassa T, Teshager L. Magnitude of Biofilm Formation and Antimicrobial Resistance Pattern of Bacteria Isolated from Urinary Catheterized Inpatients of Jimma University Medical Center, Southwest Ethiopia. Int J Microbiol. 2019;2019:5729568.
Crossref - Karimi K, Zarei O, Sedighi P, Taheri M, Doosti-Irani A, Shokoohizadeh L. Investigation of Antibiotic Resistance and Biofilm Formation in Clinical Isolates of Klebsiella pneumoniae. Int J Microbiol. 2021;2021:5573388.
Crossref - Maharjan G, Khadka P, Siddhi Shilpakar G, Chapagain G, Dhungana GR. Catheter-Associated Urinary Tract Infection and Obstinate Biofilm Producers. Can J Infect Dis Med Microbiol. 2018;2018:7624857.
Crossref - Abdallah NMA, Elsayed SB, Mostafa MMY, El-gohary GM. Biofilm forming bacteria isolated from urinary tract infection, relation to catheterization and susceptibility to antibiotics. Int J Biotechnol Mol Biol Res. 2011;2(10):172-178.
- Baugh S, Ekanayaka AS, Piddock LJ V, Webber MA. Loss of or inhibition of all multidrug resistance efflux pumps of Salmonella enterica serovar Typhimurium results in impaired ability to form a biofilm. J Antimicrob Chemother. 2012;67(10):2409-2417.
Crossref - Tang M, Wei X, Wan X, Ding Z, Ding Y, Liu J. The role and relationship with efflux pump of biofilm formation in Klebsiella pneumoniae. Microb Pathog. 2020;147:104244.
Crossref
© The Author(s) 2022. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.