ISSN: 0973-7510
E-ISSN: 2581-690X
Ectoine is a valuable active compound produced by halophilic bacteria with broad applications in the pharmaceutical and cosmetic industries. The increasing industrial demand underscores the need for enhanced production capacity and drives the discovery of effective and efficient ectoine-producing bacteria strains. Although Indonesia possesses numerous high-salinity habitats, such as traditional salt ponds, the potential for novel ectoine-producing strains remains underexplored. Therefore, this study aims to isolate ectoine-producing bacteria from traditional salt ponds in Buleleng Regency and optimize the yield of potential isolates. Halophilic bacteria were initially cultivated from saltwater and soil samples on MM63 medium. The ectoine-producing capability of these isolates was then evaluated using a batch culture approach incorporating an osmotic shock step. Ectoine quantification was performed through reversed-phase High-Performance Liquid Chromatography using a mobile phase of water/acetonitrile (95/5). To maximize production, a series of optimization experiments determined the ideal media composition, initial pH, incubation temperature, and incubation time. The results showed that seven ectoine-producing halophilic bacteria were successfully isolated, with one of the best isolates being Halomonas huangheensis LES5 AG4. The application of osmotic shock increased ectoine production by up to 3.6-fold, specifically from 94 mg/L to 335 mg/L. Furthermore, ectoine production was significantly increased to 646 mg/L after optimizing several key factors, including ammonium chloride (0.47% w/v), glycerol (1.42% w/v), NaCl (11.67% w/v), Initial pH (7.1), incubation temperature (30 °C), and incubation time (16 hours). These results show that Halomonas huangheensis LES5 AG4 holds significant potential for development as an industrial-scale ectoine producer.
Ectoine, Halophilic Bacteria, Salt Pond, Halomonas huangheensis
Ectoine, also called 1,4,5,6-tetrahydro-2-methyl-4-pyrimidine carboxylic acid, is a compatible organic molecule with significant potential as an active ingredient in pharmaceutical and cosmetic products. Numerous studies have shown the efficacy and safety for a variety of biomedical applications. These include treating respiratory diseases and hypersensitivity reactions,1 dry eye disease,2 as well as irritation and inflammation on the ocular surface.3 Furthermore, ectoine has been shown to enhance antibody production.4 Aside from biomedical uses, ectoine is also reported to have considerable potential as a skin protective agent. Several studies have reported the effectiveness in treating inflammatory skin diseases,5 acting as a skin-whitening agent,6 while also mitigating cytotoxicity and irritation.7 Ectoine functions as an anti-irritant, anti-UVB, and antibacterial agent.8
Given the safety and wide-ranging benefits, ectoine has been increasingly used as an active ingredient in various health and cosmetic products. This growing commercial availability of ectoine-based products has directly increased the market demand. As reported by a Business Research Institute, the global ectoine market was valued at approximately USD 0.07 billion in 2024 and is projected to reach USD 0.11 billion by 2033, representing an estimated compound annual growth rate (CAGR) of 5.4%.9 This strong market growth underscores ectoine potential to become a key component in future product formulations.
Ectoine is produced by halophilic and halotolerant bacteria as a cellular defense mechanism against high osmotic stress in saline environments. Several ectoine-producing halophilic bacteria have been identified, including Halomonas elongata, Brevibacterium epidermis, Halomonas salina, Corynebacterium glutamicum, and Halomonas hydrothermalis.10 Given Indonesia’s diverse ecosystems with high salinity, such as salt ponds, mangrove mud, and marine sediments, there is significant potential for discovering novel ectoine-producing bacteria. However, basic studies exploring this potential remain limited. A recent study by Parwata et al. reported the isolation of several ectoine-producing halophilic bacteria strains from saltwater and sediment samples collected from the Salt Pond of Pejarakan Village in Bali.11 However, the ectoine productivity of these specific strains was reportedly too low for industrial-scale applications. This underscores the urgent need for the discovery and characterization of new bacteria strains with high ectoine productivity.
Using the bacteria milking method, ectoine can be produced on an industrial scale. This method is executed by cultivating bacteria in a high-salt medium to stimulate ectoine biosynthesis, followed by a transfer into a low-salt medium to induce the release from the cells.12 However, a significant challenge with this method is that the high salt concentrations required for optimal ectoine production by halophilic microbes13,14 can cause accelerated bioreactor corrosion and inhibit bacteria cell growth, thereby limiting overall ectoine yield.15 Discovering new halophilic bacteria strains that can achieve high ectoine productivity at more moderate salt concentrations is essential for making the process both economically viable and sustainable.
To address the significant gap between potential microbial productivity and the negative environmental and economic impacts associated with high salinity cultivation, this study was designed to identify novel halophilic bacteria strains capable of achieving high ectoine yields under industrially favorable, moderate salinity conditions. Bacteria isolates were collected from traditional salt ponds located in the Buleleng Regency of Bali Province, Indonesia. This study then focused on systematically enhancing ectoine synthesis from the most productive isolates by optimizing several critical cultivation parameters, specifically media nutrient composition, pH, incubation temperature, and incubation time. The ultimate objective was to establish the selected microbe efficacy as a viable candidate for large-scale ectoine production, which was evaluated by implementing an osmotic shock technique to confirm the maximum production potential.
Chemicals
All chemicals, including KH2PO4, KOH, MgSO4.7H2O, FeSO4.7H2O, NaCl, (NH4)2SO4, (NH4)2Cl, and glycerol, were purchased from Merck (Germany) and were of pro analysis grade. Yeast extract, tryptone, peptone, Bacto agar, maltose, glucose, and L-aspartic acid were obtained from Himedia (India). Methanol, chloroform, and acetonitrile were purchased from Merck (Germany).
Collection of sample
Soil and saltwater samples were collected from three traditional solar salt ponds located within the Buleleng Regency, Bali Province, Indonesia. The specific sampling locations included Pejarakan Village, Les Village, and Tedjakula Village (Figure 1). Soil samples were systematically acquired from the basin of each of the three salt ponds. Saltwater samples were collected from the concentration vessels used in the salt production process at each of the three villages. Soil samples were collected from four distinct locations within each salt pond to represent a gradient of salinity concentrations. Concurrently, salt samples were obtained from two locations showing a salinity range of 25-29 °Bי. All samples were transferred into sterile vials and immediately transported to the laboratory in an icebox for subsequent analysis.
Figure 1. Traditional salt ponds in Buleleng Regency, Bali Province, Indonesia. (A) Salt pond in Pejarakan Village, (B) Salt pond in Les Village, (C) Salt pond in Tedjakula Village
Isolation of halophilic bacteria
Halophilic bacteria were grown and isolated using MM63 agar medium containing 1.36% (w/v) KH2PO4, 0.42% (w/v) KOH, 0.2% (w/v) (NH4)2SO4, 0.025% (w/v) MgSO4.7H2O, 0.00011% (w/v) FeSO4.7H2O, 0.5% (w/v) Glucose, 5% (w/v) NaCl, and 1.75% (w/v) bacto agar.16 Saltwater samples were filtered using sterile Whatman filter paper. Soil samples were treated with a 5% (w/v) NaCl solution, agitated, and subsequently filtered. Each resulting filtrate and saltwater sample was then inoculated onto MM63 agar media using the spread-plate method. The medium was incubated at 37 °C for 1-2 days. The growing bacteria colonies were then purified by repeatedly streaking onto fresh MM63 agar medium until a pure colony was obtained. The experiments were conducted without replication.
Ectoine production and determination of bacteria growth
Bacteria strains were initially inoculated into 4 mL of MM63 broth containing 5% w/v NaCl in a shaker for 12 hours at 37 °C. Subsequently, 0.5 mL of inoculum was transferred aseptically into 20 mL of fresh MM63 media. The medium was incubated in a shaker at 37 °C with an agitation of 180 rpm for 24 hours. After completion of the incubation period, the bacteria cells were harvested by centrifugation at 8,000 x g for 10 minutes. The resulting cell pellet was then used for the extraction of ectoine. Bacteria growth was investigated spectrophotometrically by measuring the cell optical density (OD) at 600 nm using a Shimadzu UV-1800 Spectrophotometer (Japan).
Ectoine was extracted from the bacteria cell following a modified procedure based on the Bligh and Dyer method.17 Initially, the collected cell pellets were dried in an oven at 70 °C for three hours. The dried material was then extracted using a biphasic solvent mixture composed of methanol/chloroform/water at a ratio of 10:5:4 (v/v/v). About 350 µL of this solvent blend was added to the dried pellets, and the extraction efficiency was maximized through ultrasonication for 30 minutes. Subsequently, the solvent partitioning was initiated by adding equal volumes (65 µL each) of pure chloroform and water to the mixture. The resulting solution was thoroughly mixed by shaking for 10 minutes. Phase separation was accelerated by centrifugation at 10,000 x g for 15 minutes. The upper aqueous phase, which contained the target compound ectoine, was carefully collected and reserved for quantitative analysis.
Determination of ectoine concentration
Ectoine concentration in the extracted samples was quantified in duplicate using a Shimadzu High-Performance Liquid Chromatography Model SPD-20A/20AV (Japan). A sample volume of 20 µL was injected into the system for analysis. Chromatographic separation was then achieved using a Silica C18 column (Sigma Aldrich, USA) maintained at a constant temperature of 28 °C. The separation of the compatible solutes was performed under isocratic conditions. The mobile phase consisted of a mixture of water and acetonitrile (95/5, v/v) delivered through the column at a constant flow rate of 1 mL/min. Detection of ectoine was performed by monitoring the characteristic absorbance at 210 nm using a UV detector.11 Quantification of ectoine was achieved by comparing the peak areas of the samples against a standard curve generated using an authentic standard purchased from Sigma Aldrich.
Identification of halophilic bacteria
The identity of the isolated bacteria was determined through sequencing of the 16S ribosomal RNA (rRNA) gene. Chromosomal DNA was first extracted from the bacteria cultures using the Presto Mini gDNA Bacteria Kit (Geneaid, Taipei, Taiwan). The extracted genomic DNA was used as a template for the amplification of the 16S rRNA gene through Polymerase Chain Reaction (PCR). The reaction used the universal primers Bact27F (order: AGAGTTTGATCATGGCTCAG) and Uni1492R (order: GGTTACCTTGTTACGACTT). The reaction was carried out following predenaturation at 94 °C for 4 min, denaturation at 94 °C for 30 seconds, annealing at 50 °C for 1 min, and extension/propagation at 72 °C for 2 min. The cycle was repeated 34 times, and the resulting 16S rRNA gene amplicons were subsequently purified and forwarded for Sanger sequencing. The raw sequence data obtained were assembled and edited using the DNA Baser Assembler software (version 5.21.0BT, Heracle BioSoft). For preliminary identification, the assembled sequence was aligned against those in public databases using the Basic Local Alignment Search Tool (BLAST). The acquired sequence, along with related references identified through BLAST, was then used to construct a phylogenetic tree. This evolutionary analysis was carried out using the Molecular Evolutionary Genetics Analysis (MEGA) software (version 12.0.14).18
Optimization of culture conditions for enhanced ectoine production
Optimization of culture conditions was conducted by systematically evaluating several key parameters, including the concentrations of the nitrogen source, carbon source, and NaCl, the initial medium pH, incubation temperature, and incubation time. The optimization of nutritional component concentrations (nitrogen source, carbon source, and NaCl) was performed using Response Surface Methodology (RSM). The experimental design used a Central Composite Design (CCD). Furthermore, the RSM analysis was conducted using Minitab version 19 (2020). The initial medium pH, incubation temperature, and incubation time were optimized using a one-variable-at-a-time (OVAT) approach. This methodology was selected based on the assumption that no significant interactions existed between the variables, allowing for the independent assessment of each parameter effect on ectoine production. All experiments were performed in duplicate.
Isolation of halophilic bacteria and primary screening for ectoine-producing isolates
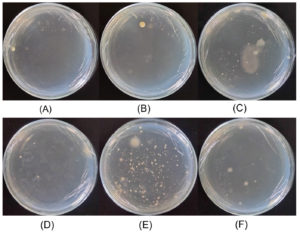
Figure 2 shows the growth results of halophilic bacteria cultured on MM63 agar containing 5% (w/v) NaCl. Bacteria growth was observed in all samples from the Pejarakan, Les, and Tedjakula salt ponds, with varying colony counts. The saltwater sample from the Pejarakan salt pond (Figure 2A) and the corresponding soil sample (Figure 2B) both yielded a low number of colonies. In contrast, the saltwater sample from the Les salt pond (Figure 2C) produced a higher number of colonies. However, the soil sample (Figure 2D) showed limited growth. The highest colony count was observed in the saltwater sample from the Tedjakula salt pond (Figure 2E). Meanwhile, the soil sample from the salt pond (Figure 2F) showed slightly more growth. These results show that, under the specified conditions, the MM63 medium with a 5% (w/v) NaCl content supported greater halophilic bacteria growth from the saltwater samples than from those collected at the three locations.
Figure 2. Bacteria colonies cultured on MM63 agar media supplemented with 5% (w/v) NaCl. The colonies were isolated from: (A) saltwater and (B) soil samples collected from Pejarakan Village Salt Pond; (C) saltwater and (D) soil samples collected from Les Village Salt Pond; and (E) saltwater and (F) soil samples collected from Tedjakula Village Salt Pond
A total of 12 bacteria colonies were randomly selected from each medium
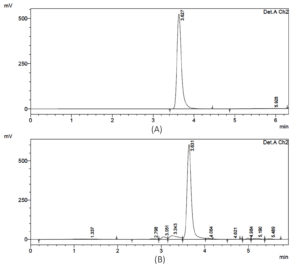
(Figure 2) and evaluated for the capacity to synthesize ectoine. Analytical results showed that seven isolates successfully produced ectoine, confirmed by a retention time of 3.6 minutes on the HPLC chromatogram. This peak corresponds precisely to the ectoine standard, as shown in Figure 3.
Figure 3. HPLC chromatograms of ectoine. (A) ectoine standard, (B) ectoine extracted from the halophilic bacteria culture
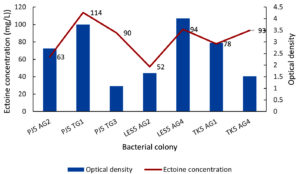
All seven isolates showed ectoine production, with concentrations ranging from 52 to 114 mg/L (Figure 4). Two isolates, LES5 AG2 and PJ5 AG2, showed the lowest ectoine yields, at 52 mg/L and 63 mg/L, respectively. Four other isolates produced ectoine at moderate levels, including TK5 AG1 (78 mg/L), PJ5 TG3 (90 mg/L), TK5 AG4 (93 mg/L), and LES5 AG4 (94 mg/L). The highest ectoine concentration was produced by isolate PJ5 TG1, which yielded 114 mg/L. Bacteria growth, measured by OD at 600 nm, did not correlate directly with ectoine production levels. Specifically, isolates with high growth rates, such as PJ5 AG2, LES5 AG4, and TK5 AG1, produced relatively low levels of ectoine. Conversely, isolates PJ5 TG3 and TK5 AG4 were remarkable for producing high levels of ectoine despite having comparatively low growth rates. This suggests that the strains possess efficient ectoine biosynthetic pathways, but the overall productivity could be enhanced by optimizing conditions to increase biomass production.
Figure 4. Bacteria growth (OD) and concentration of ectoine produced by halophilic bacteria in MM63 media. The bacteria were isolated from saltwater sample (AG) and soil sample (TG) collected at the salt pond of Pejarakan Village (PJ), Les Village (LES), and Tedjakula Village (TK)
The ectoine productivity of the three highest-producing bacteria isolates, namely PJ5 TG1, LES5 AG4, and TK5 AG4, was further investigated using a two-stage cultivation method with osmotic shock (OS). This method comprised an initial growth phase in media with a low NaCl concentration to maximize cell biomass, followed by a transfer to media with a higher NaCl concentration to induce ectoine production. Osmotic shock conditions were tested by shifting the NaCl concentration from 5%-10% (w/v). The treatments were performed without replication.
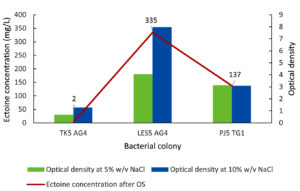
Figure 5 shows the ectoine production results following the osmotic shock from 5%-10% (w/v) NaCl. Isolate PJ5 TG1 showed a 1.2-fold increase in ectoine production, rising from 114 mg/L (Figure 4) to 137 mg/L (Figure 5). In contrast, ectoine production by isolate TK5 AG4 decreased significantly after the osmotic shock, falling from 93 mg/L to 2 mg/L. More importantly, isolate LES5 AG4 showed a substantial 3.6-fold increase in ectoine production, from 94 mg/L to 335 mg/L, after the application of the osmotic shock. Bacteria growth (OD) showed varying responses to osmotic shock. Specifically, isolates TK5 AG4 and LES5 AG4 showed a significant increase in growth (1.9-fold), while PJ5 TG1 had no growth acceleration following the shock. These results show that the enhanced growth of isolate LES5 AG4 is positively correlated with the observed increase in ectoine production by the bacteria.
Figure 5. Impact of osmotic shock (OS) on the growth (OD) and ectoine concentration produced by the halophilic bacteria
Ectoine is synthesized intracellularly from the precursor L-aspartate-β-semialdehyde (ASA) through a three-enzyme pathway catalyzed by L-2,4-diaminobutyrate transaminase (EctB), 2,4-diaminobutyrate acetyltransferase (EctA), and ectoine synthase (EctC).19 Given this relationship, increasing bacteria growth would theoretically lead to a higher biomass and a greater yield of ectoine.
Based on the superior ectoine yield achieved following two-stage cultivation and osmotic shock experiments, isolate LES5 AG4 was designated as the most promising producer. Consequently, the next phase focused on maximizing ectoine production from the bacteria through the optimization of the media composition, fermentation conditions, and method.
Identification of ectoine-producing halophilic bacteria isolate LES5 AG4
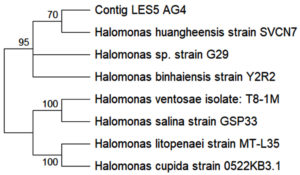
The identification of the ectoine-producing halophilic bacteria LES5 AG4 was performed using 16S rRNA gene sequence analysis (Figure 6). The resulting sequence analysis indicated that the LES5 AG4 isolate showed close phylogenetic homology with the species Halomonas huangheensis (GenBank accession number: PX904975). This is in line with previous studies in which similar bacteria, characterized as Gram-negative, rod-shaped, and moderately halophilic, were successfully isolated from saline-alkali soil samples in Shandong Province, China.20 Based on available literature, this is the first report documenting ectoine production by specific bacteria species.
Optimization of culture conditions for enhanced ectoine production
Screening of nitrogen and carbon sources
The optimization of ectoine production from the halophilic bacteria LES5 AG4 was initially focused on screening the nitrogen and carbon sources. Five distinct nitrogen sources were evaluated to determine the effect on ectoine yield, including tryptone, peptone, yeast extract, ammonium sulfate, and ammonium chloride. Similarly, five carbon sources were tested, namely maltose, glucose, aspartate, glycerol, and monosodium glutamate (MSG). All ectoine production experiments were conducted using MM63 minimal medium supplemented with a 5% (w/v) NaCl concentration. Among the components, only nitrogen and carbon were systematically varied.
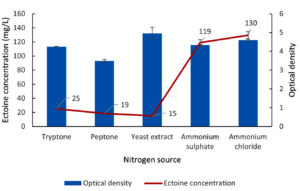
The results detailing the optimization of nitrogen sources for ectoine production from the LES5 AG4 bacteria isolate are shown in Figure 7. Yeast extract yielded the highest bacteria growth, evidenced by an OD value of 5.0. Conversely, peptone proved to be the least effective nitrogen source for cell proliferation, resulting in an OD of 3.5. Tryptone and ammonium sulfate supported comparable and relatively high cell densities, with OD values of 4.2 and 4.3, respectively. Ammonium chloride facilitated higher microbial growth (OD 4.6) than ammonium sulfate, though it remained lower than that observed with yeast extract.
Figure 7. Impact of various nitrogen sources on the growth (OD) and ectoine concentration produced by the halophilic bacteria isolate LES5 AG4
A crucial result was that ectoine production levels were not directly proportional to bacteria growth. Despite supporting the highest cell density, yeast extract yielded the lowest ectoine concentration at 15 mg/L. Similarly, peptone and tryptone produced only marginally higher ectoine levels of 19 mg/L and 25 mg/L, respectively. The inorganic nitrogen sources, ammonium sulfate and ammonium chloride, significantly enhanced ectoine biosynthesis, resulting in concentrations of 119 mg/L and 130 mg/L, respectively. These data suggest that ammonium sulfate and, more effectively, ammonium chloride serve as superior nitrogen sources for simultaneously supporting adequate bacteria growth and inducing high-level ectoine production in the LES5 AG4 isolate.
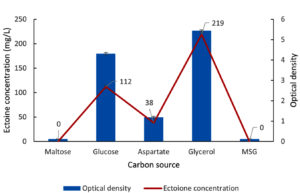
Figure 8. Impact of various carbon sources on the growth (OD) and ectoine concentration produced by the halophilic bacteria isolate LES5 AG4
Figure 8 shows the bacteria growth and ectoine concentration produced by the LES5 AG4 isolate across the tested carbon sources. The bacteria showed no observable growth in the MM63 medium when supplied with maltose or MSG as the sole carbon source, resulting in zero ectoine production. L-aspartate proved to be an inefficient carbon source for both bacteria proliferation and ectoine biosynthesis. Specifically, growth was minimal (OD of 1.2), yielding an ectoine concentration of only 38 mg/L. Glucose, on the other hand, supported a substantially higher ectoine yield of 112 mg/L, correlating with a high growth rate (OD of 4.3). Most importantly, glycerol facilitated the highest ectoine concentration recorded at 219 mg/L, accompanied by the maximum observed growth rate (OD of 5.4). These results conclusively show that glycerol is the superior carbon source for both maximizing cell growth and enhancing ectoine production in the LES5 AG4 isolate.
Optimization of nutrient level
The optimization of culture conditions, specifically the levels of the carbon source (glycerol), the nitrogen source (ammonium chloride), and sodium chloride (NaCl), was performed to enhance the biomass and ectoine production by the halophilic bacteria LES5 AG4. This optimization used a two-stage cultivation process. Stage I was dedicated to maximizing biomass accumulation, and Stage II was designed to stimulate and increase ectoine synthesis. The experimental design was generated using a CCD, as shown in Table 1.
Table (1):
CCD matrix for the optimization of biomass production (cultivation I) and ectoine production (cultivation II) by the halophilic bacteria isolate LES5 AG4
| Cultivation | Variable | Level | ||||
|---|---|---|---|---|---|---|
| I | Coded | -1.682 | -1 | 0 | 1 | 1.682 |
| Ammonium chloride (% w/v) | 0.05 | 0.15 | 0.30 | 0.45 | 0.55 | |
| Glycerol (% w/v) | 0.30 | 0.52 | 0.85 | 1.18 | 1.40 | |
| NaCl (% w/v) | 5.00 | 6.62 | 9.00 | 11.38 | 13.00 | |
| II | Coded | -1.682 | -1 | 0 | 1 | 1.682 |
| Ammonium chloride (% w/v) | 0.10 | 0.24 | 0.45 | 0.66 | 0.8 | |
| Glycerol (% w/v) | 0.60 | 1.05 | 1.70 | 2.35 | 2.80 | |
| NaCl (% w/v) | 7.00 | 9.23 | 12.50 | 15.77 | 18.00 | |
The RSM analysis of the data from Cultivation I yielded a model for cell dry weight (CDW) with a high coefficient of determination (R2 = 91.89%). This value shows a strong fit between the predicted model and the observed experimental data variance. The Analysis of Variance (ANOVA) results, detailed in Table 2, showed that only the linear term for glycerol concentration had a statistically significant effect on CDW (p < 0.05). Conversely, the linear terms for ammonium chloride and NaCl concentrations did not show a significant effect.
Table (2):
Summary of ANOVA for the CDW model (Cultivation I) and ectoine concentration model (Cultivation II)
| Source | P-value | |
|---|---|---|
| Cultivation I | Cultivation II | |
| Model | 0.000 | 0.003 |
| Linear | 0.000 | 0.062 |
| Ammonium chloride | 0.082 | 0.640 |
| NaCl | 0.951 | 0.012 |
| Glycerol | 0.000 | 0.505 |
| Square | 0.002 | 0.000 |
| Ammonium chloride* Ammonium chloride | 0.011 | 0.046 |
| NaCl*NaCl | 0.001 | 0.000 |
| Glycerol*Glycerol | 0.009 | 0.283 |
| 2-Way Interaction | 0.992 | 0.832 |
| Ammonium chloride* NaCl | 0.793 | 0.665 |
| Ammonium chloride* Glycerol | 0.884 | 0.811 |
| NaCl*Glycerol | 0.977 | 0.454 |
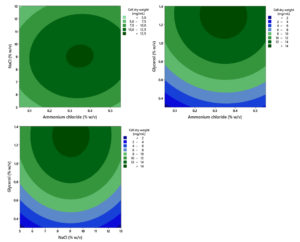
Increasing glycerol concentration within the tested range of 0.3%-1.3% (w/v) led to a corresponding rise in bacteria biomass production (CDW), as shown in Figure 9. However, in the square model, all three variables, namely ammonium chloride, NaCl, and glycerol, showed a statistically significant effect (p < 0.05). This significance in the quadratic terms strongly suggests that the optimal concentrations for all three factors were located within the defined experimental design space. Furthermore, the analysis showed no significant two-way interaction effects among all three variables (p > 0.05). Based on the established response model, the predicted optimal concentrations for maximizing biomass production (CDW) were determined to be 9.04% (w/v) NaCl, 1.30% (w/v) glycerol, and 0.34% (w/v) ammonium chloride, respectively.
Figure 9. Contour plot showing the combined effect of ammonium chloride, glycerol, and NaCl concentration on the growth (CDW) of the halophilic bacteria isolate LES5 AG4
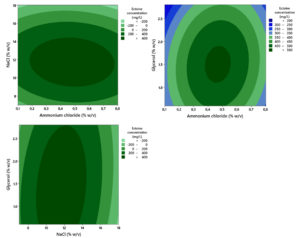
Ectoine production by the bacteria was maximized during Cultivation II. Bacteria cells were initially cultivated in MM63 medium containing the optimal nutrient concentrations determined in Cultivation I, and subsequently transferred aseptically into the MM63 medium designed for Stage II. The RSM analysis of the Cultivation II data yielded a response model for the ectoine concentration produced, with a coefficient of determination (R2) at 85.99%. The ANOVA results (Table 2) showed that, for the linear terms, only the NaCl concentration was statistically significant (p < 0.05). Conversely, the linear terms for ammonium chloride and glycerol concentrations were not statistically significant. In the tested concentration range of 11%-18% w/v, increasing the NaCl concentration resulted in a trend of decreasing ectoine concentration (Figure 10).
Figure 10. Contour plot showing the combined effect of ammonium chloride, glycerol, and NaCl concentration on the ectoine concentration produced by the halophilic bacteria isolate LES5 AG4
Regarding the quadratic terms, ammonium chloride and NaCl concentrations showed statistically significant effects (p < 0.05). The significance suggests that the optimal concentrations for these two variables were contained within the experimental design space. Although the quadratic term for glycerol concentration was statistically insignificant
(p > 0.05), inspection of the model contour plot confirmed that the optimum concentration was within the experimental range. The analysis also confirmed no significant two-way interaction effects among the three experimental variables (p > 0.05), showing that the variables exerted independent effects on ectoine production. Based on the established response model, the predicted optimal concentrations for maximizing ectoine production were determined to be 0.47% (w/v) ammonium chloride, 1.42% (w/v) glycerol, and 11.67% (w/v) NaCl.
Optimization of initial medium pH, incubation temperature, and incubation time
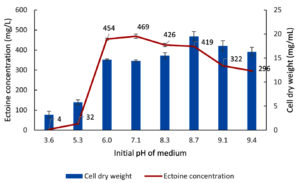
Initial medium pH, incubation temperature, and incubation time were subjected to subsequent optimization. As shown in Figure 11, optimization of the initial medium pH was conducted across a range from 3.6-9.4. The experimental results showed that at acidic pH (3.6 and 5.3), both biomass production (CDW) and ectoine production by the halophilic bacteria LES5 AG4 were significantly low. A subsequent increase in pH led to a concurrent rise in both bacteria growth and ectoine production. The highest ectoine yields were recorded at initial medium pH of 6.0 and 7.1, measuring 454 mg/L and 469 mg/L, respectively, with corresponding CDW values of 14.7 mg/mL and 14.4 mg/mL. At alkaline pH above 7.1, biomass production showed a slight increase, reaching the optimum value of 19.5 mg/mL at pH 8.7. However, ectoine levels showed a tendency to decrease under alkaline conditions, particularly at pH values above 8.7. These results show that increased biomass production is not consistently correlated with enhanced ectoine production, suggesting the initial medium pH exerts differential effects on bacteria growth and ectoine biosynthesis. Figure 11 shows that the optimum initial pH of media for ectoine production by bacteria was 7.1.
Figure 11. Impact of initial medium pH on the growth (CDW) and ectoine concentration produced by the halophilic bacteria isolate LES5 AG4
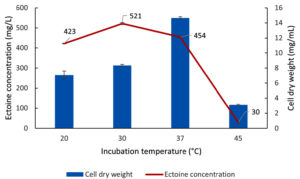
Optimization of the incubation temperature for biomass and ectoine production by the LES5 AG4 bacteria was conducted across a range of 20 to 45 °C (Figure 12). Incubation at a temperature of 37 °C yielded the highest biomass production (CDW) at 14.65 mg/mL. In contrast, a temperature of 30 °C resulted in the optimum ectoine production, reaching 521 mg/L, despite the biomass production being substantially lower at only 8.35 mg/mL compared to the 37 °C condition. These results further underscore that temperature exerts differential effects on bacteria growth and ectoine biosynthesis. At higher temperatures (45 °C), both biomass and ectoine production decreased drastically, suggesting that high temperatures inhibit both bacteria growth and ectoine biosynthesis pathways. Conversely, at low temperatures (20 °C), the bacteria maintained a relatively high ectoine yield (423 mg/L) with an estimated growth (CDW) of 7.1 mg/mL. Figure 12 confirms that the optimum incubation temperature for ectoine production by the halophilic bacteria LES5 AG4 was 30 °C.
Figure 12. Impact of incubation temperature on the growth (CDW) and ectoine concentration produced by the halophilic bacteria isolate LES5 AG4
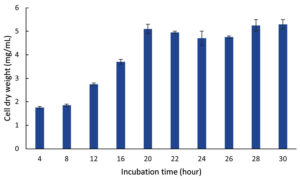
Optimization of the incubation time was performed for two stages, namely Cultivation I for biomass production and II for ectoine production. Figure 13 shows the biomass production (CDW) of the LES5 AG4 bacteria during Cultivation I over an incubation period ranging from 4-30 hours. During the initial incubation period (4 and 8 hours), the bacteria showed low growth, with CDW values of 1.8 mg/mL and 1.9 mg/mL, respectively. Increasing the incubation time led to a significant enhancement in bacteria growth, which peaked at 5.1 mg/mL after incubation for 20 hours. Bacteria growth was then stable between incubation times of 20 and 30 hours, showing the onset of the stationary phase. Based on these results, an incubation time of 20 hours is deemed suitable to achieve optimal biomass for transition to Cultivation II. Although a 30-hour incubation yielded a marginally higher CDW (5.3 mg/mL), at this point, the cells have been established in the stationary phase. Prolonged duration in the stationary phase can result in an accumulation of aged bacteria cells and potentially inhibitory secondary metabolites, which may negatively impact subsequent ectoine production and overall cellular metabolism.
Figure 13. Growth kinetics (CDW) of the halophilic bacteria isolate LES5 AG4 over various incubation times during Cultivation I
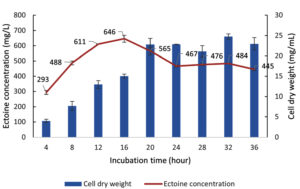
Data for biomass (CDW) and ectoine production by the halophilic bacteria LES5 AG4 during Cultivation II, with varying incubation times, are presented in Figure 14. Following the transfer to fresh media (Cultivation II), the bacteria showed significant growth, achieving a CDW of 22.8 mg/mL after incubation for 20 hours. This observation shows that the transfer to a nutrient-rich media induced a new exponential growth phase with a significantly greater growth rate compared to Cultivation I. Similar to the previous stage, after 20 hours of incubation, bacteria growth was stable and continued until 36 hours of incubation, marking the onset of the stationary phase. This suggests that the LES5 AG4 growth profile remains consistent, regardless of the differing nutrient levels in the MM63 media between the two cultivations.
Figure 14. Growth kinetics (CDW) and ectoine concentration produced by the halophilic bacteria isolate LES5 AG4 over various incubation times during Cultivation II
Incubation for only 4 hours in Cultivation II yielded an ectoine level of 293 mg/L. This initial quantity is attributed to the carryover ectoine synthesized during Cultivation I, which had an initial measured concentration of 285 mg/L. Therefore, the 4-hour incubation only contributed a marginal increase in ectoine concentration (8 mg/L). Further extending the incubation time resulted in a significant increase in ectoine production, reaching the optimum yield of 646 mg/L at the 16th hour. This result is particularly remarkable because the biomass production after incubation for 16 hours (15.0 mg/mL) was substantially lower than the maximum CDW achieved at the 20th hour (22.8 mg/mL).
The ectoine yield at 16 hours (646 mg/L) was demonstrably greater than the yield at 20 hours (565 mg/L). Incubation beyond 16 hours led to a significant reduction in ectoine levels, dropping to 467 mg/L by the 24th hour, before stabilizing until the 36th hour. This observation reinforces the principle that the maximum yield of a secondary metabolite (ectoine) often occurs during the late exponential or early stationary phase of growth and is not necessarily correlated with the point of maximum biomass accumulation.
This study successfully isolated seven ectoine-producing halophilic bacteria from soil and saltwater samples obtained from salt ponds in Buleleng Regency, Bali. The best ectoine-producing bacteria isolate LES5 AG4 was identified as Halomonas huangheensis. The adoption of a two-stage cultivation method coupled with osmotic shock successfully enhanced both biomass production and ectoine accumulation of the bacteria. This methodology is consistent with previous reports showing increased biomass and ectoine yields in halophilic bacteria, such as Halomonas elongata BK-AG2521 and Halomonas boliviensis.22 The elevated bacteria growth observed during the second cultivation stage is hypothesized to result from the dilution or removal of inhibitory byproducts generated during the initial cultivation.21 Furthermore, the increase in ectoine production during the second cultivation is directly attributable to the elevated NaCl concentration in the culture medium. Halophilic bacteria generally respond to an increase in environmental salinity by synthesizing compatible organic solutes, such as ectoine, which function to counteract high external osmotic pressure and protect cell integrity.13,14
The growth and ectoine production of the best isolate, Halomonas huangheensis LES5 AG4, were significantly affected by different nitrogen and carbon sources. In this study, ammonium chloride was identified as the best nitrogen source for ectoine production. This result stands in contrast to Chen et al, who showed that ammonium sulfate served as the optimal nitrogen source for ectoine biosynthesis in Halomonas salina BCRC17875. Conversely, Chen et al. reported that the use of NH4Cl resulted in a significantly lower ectoine yield for the specific strain.23 This difference suggests strain-specific variations in nitrogen assimilation pathways, as NH4Cl yielded the highest ectoine levels for Halomonas huangheensis LES5 AG4, marginally surpassing the yield achieved with ammonium sulfate. In line with this study, Chen et al. also reported that complex organic sources such as peptone and tryptone were suboptimal for ectoine production in H. salina BCRC17875.23
The best carbon source for ectoine production by Halomonas huangheensis LES5 AG4 was glycerol. These results contrast sharply with those of other studies reported. For instance, Chen et al. identified a combination of yeast extract and glutamate as the optimal carbon source for ectoine production in Halomonas salina BCRC17875.23 Similarly, MSG was reported in another study as the preferred carbon source for ectoine synthesis by Brevibacterium epidermis DSM20659.24 These reports stand in direct opposition to the current study, showing that MSG inhibited ectoine synthesis entirely in Halomonas huangheensis LES5 AG4. Although glucose was shown to prevent ectoine synthesis in H. salina BCRC17875,23 this carbon source induced relatively high levels of ectoine production in Halomonas huangheensis LES5 AG4. These stark differences underscore significant, strain-specific metabolic variability in the use of carbon substrates for ectoine biosynthesis.
This study found that sodium chloride (NaCl) is a critical factor influencing both the growth and ectoine production of the halophilic bacteria Halomonas huangheensis LES5 AG4. Optimal ectoine production was achieved at a higher salt (11.67% w/v) compared to the concentration supporting maximal biomass accumulation (9.04% w/v). This dissociation between optimal conditions for growth and for ectoine synthesis is consistent with reports from other halophiles. For instance, Halomonas elongata BK-AG25 showed optimal biomass production at an NaCl content of 8.9% (w/v), while ectoine production peaked at a considerably higher concentration of 18% (w/v).21 Halomonas cupida J9 was also reported to yield optimal ectoine at 15% NaCl, while optimal biomass production occurred at a lower range (6%-10% w/v).24 However, this pattern is not universal, as both ectoine and biomass production were simultaneously optimized in Halomonas salina BCRC17875 and Brevibacterium epidermis DSM20659, at the same high salt concentration, namely 11.7% w/v23 and 5.9% w/v25, respectively.
The optimum initial pH of media for ectoine production varied considerably among halophilic bacteria. Specifically, the optimal pH for ectoine production by Halomonas huangheensis LES5 AG4 in this study was neutral (pH 7,1). However, the optimal condition for the halophilic bacteria Halomonas sp. H02 was reported to be acidic (pH 5.9).26 Ectoine production from other halophiles, such as Brevibacterium epidermis DSM20659 and Alkalibacillus haloalkaliphilus, was found to be optimal under alkaline conditions (pH 7.8 for B. epidermis DSM2065925 and pH 9.0 for A. haloalkaliphilus27). In this study, optimum biomass production occurred at a more alkaline pH (8.7). This observation also contrasts with those reported for B. epidermis DSM20659 and A. haloalkaliphilus, where optimum biomass production was attained at a pH slightly more acidic than the pH for maximum ectoine production (pH 7.5 for B. epidermis and 8.8 for A. haloalkaliphilus). These comparative results clearly show that different bacteria species show distinct physiological responses to the initial medium pH, suggesting a high degree of strain-specific optimization is necessary for maximizing both ectoine and biomass yields.
The influence of temperature on both biomass and ectoine production in Halomonas huangheensis LES5 AG4 is comparable to other reports for the halophilic bacteria Halomonas elongata BK-AG25. Specifically, the optimal temperature for biomass accumulation was determined to be 37.4 °C, while the maximum ectoine yield was observed at 33 °C.21 Ectoine production by another halophilic bacteria, Brevibacterium epidermis DSM20659, was reported to be optimal at a lower temperature of 25 °C.25 In these bacteria, the maximum biomass yield coincided with the optimal temperature for ectoine production. These data suggest that ectoine biosynthesis across various halophilic bacteria is generally most effective within a mesophilic temperature range (25-35 °C). However, an exception was observed in Alkalibacillus haloalkaliphilus, where ectoine production peaked at a higher temperature of 40 °C, with the optimal temperature for biomass production being 45 °C.27
The maximum ectoine concentration achieved by Halomonas huangheensis LES5 AG4 under optimized conditions, including nutrient content, pH, temperature, and incubation time, was 646 mg/L. This concentration is significantly lower than the yields reported for several other Halomonas species, including Halomonas cupida J9 (4.12 g/L),24 Halomonas sp. H02 (1.84 g/L),26 Piscibacillus halophilus (1.49 g/L),28 Halomonas sp. NEC-1 (1.38 g/L),29 Halomonas smyrnensis IIIM VA-6 (6.5 g/L),30 and Halomonas salina BCRC17875 (13.96 g/L).23 However, a critical distinction is the significantly shorter incubation period required for ectoine production by Halomonas huangheensis LES5 AG4, which was only 16 hours. This is substantially faster compared to the aforementioned Halomonas strains, including H. cupida J9 (48 hours), Halomonas sp. (72 hours), H. salina BCRC17875 (44 hours), Piscibacillus halophilus (48 hours), and Halomonas smyrnensis IIIM VA-6 (72 hours). Furthermore, the ectoine yield of 646 mg/L achieved by Halomonas huangheensis LES5 AG4 in this study surpassed those of Alkalibacillus haloalkaliphilus (200 mg/L)27 and Virgibacillus salarius BHTA19 (274 mg/L).31 The incubation time for A. haloalkaliphilus was shorter (11 hours) than that of Halomonas huangheensis LES5 AG4 (16 hours).
This study represents the initial report of ectoine production using the halophilic bacteria Halomonas huangheensis. Previous genomic analyses of this species showed the presence of the UspA gene, which is hypothesized to facilitate ectoine synthesis in saline environments, particularly under osmotic stress conditions such as high salinity and low temperature.32 The results presented corroborate this genomic evidence by demonstrating that Halomonas huangheensis LES5 AG4 possesses the capacity to produce ectoine, with concentration showing a further increase following an osmotic shock process.
In conclusion, this study successfully achieved the primary objective by isolating and characterizing a novel ectoine-producing halophilic bacteria strain, identified as Halomonas huangheensis LES5 AG4, from traditional salt ponds in the Buleleng Regency, Bali Province, Indonesia. Crucially, this strain showed the capacity to produce ectoine under moderate salinity conditions, addressing a major limitation in industrial bioprocesses. A systematic optimization of critical cultivation parameters yielded the optimal values for maximal ectoine synthesis by the bacteria, including ammonium chloride (0.47% w/v), glycerol (1.42% w/v), NaCl (11.67% w/v), Initial pH (7.1), incubation temperature (30 °C), and incubation time (16 hours). Under these optimized conditions, the Halomonas huangheensis LES5 AG4 yielded 646 mg/L of ectoine, representing a 6,9-fold increase compared to those obtained under unoptimized parameters. This significant enhancement confirms the strain high productivity potential and the technical viability for industrial-scale production. Future studies should focus on upscaling ectoine production using large-scale bioreactors to evaluate the commercial viability and industrial applicability of Halomonas huangheensis LES5 AG4.
ACKNOWLEDGMENTS
The authors are grateful to the Department of Chemistry, Faculty of Mathematics and Natural Sciences, Universitas Pendidikan Ganesha, for providing the resources and facilities required for this study.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
IPP designed the study. IPP, SM and KSMJ carried out the laboratory work and analyzed the data. IPP wrote the manuscript. SM and KSMJ reviewed and revised the manuscript. All authors read and approved the final manuscript for publication.
FUNDING
This study was funded by the Directorate of Research and Community Service, Directorate General of Research and Development, Ministry of Higher Education, Science, and Technology, in accordance with the Implementation Contract for the State University Operational Assistance Program for Research Program Number: 100/C3/DT.05.00/PL/2025, dated May 28, 2025
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
Not applicable.
- Kadam P, Khisti M, Ravishankar V, et al. Recent advances in production and applications of ectoine, a compatible solute of industrial relevance. Bioresour Technol. 2024;393:130016.
Crossref - Chen X, Lin N, Li JM, et al. Ectoine, from a Natural Bacteria Protectant to a New Treatment of Dry Eye Disease. Pharmaceutics. 2024;16(2):236.
Crossref - Bilstein A, Heinrich A, Rybachuk A, Mosges R. Ectoine in the Treatment of irritations and Inflammations of the Eye Surface. Biomed Res Int. 2021;2021:8885032.
Crossref - Jarusintanakorn S, Mastrobattista E, Yamabhai M. Ectoine enhances recombinant antibody production in Chinese hamster ovary cells by promoting cell cycle arrest. N Biotechnol. 2024;83:56-65.
Crossref - Kauth M, Trusova OV. Topical Ectoine Application in Children and Adults to Treat Inflammatory Diseases Associated with an Impaired Skin Barrier: A Systematic Review. Dermatol Ther (Heidelb). 2022;12(2):295–313.
Crossref - Hseu YC, Chen XZ, Gowrisankar YV, Yen HR, Chuang JY, Yang HL. The Skin-Whitening Effects of Ectoine via the Suppression of a-MSH-Stimulated Melanogenesis and the Activation of Antioxidant Nrf2 Pathways in UVA-Irradiated Keratinocytes. Antioxidants. 2020;9(1):63.
Crossref - Bujak T, Zagorska-Dziok M, Niziol-Lukaszewska Z. Complexes of Ectoine with the Anionic Surfactants as Active Ingredients of Cleansing Cosmetics with Reduced Irritating Potential. Molecules. 2020;25(6):1433.
Crossref - Simbara AT, Habibah FF, Hertadi R. Rhamnolipid-Modified PHB–Ectoine Nanoparticles for Multifunctional Skin Protection Against UVB, Irritation, and Bacteria. ACS Omega. 2025;10(12):12200-12213.
Crossref - Ectoine Market Size, Share, Growth, and Industry Analysis, By Type (No less than 99%, Below 99%), By Application (Skin Care Products, Others), Regional Insights and Forecast From 2025 To 2033. https://www.businessresearchinsights.com/market-reports/ectoine-market-100579, accessed 21 March, 2025
- Liu M, Liu H, Shi M, Jiang M, Li L, Zheng Y. Microbial production of ectoine and hydroxyectoine as high value chemicals. Microb Cell Fact. 2021;20(1):76.
Crossref - Parwata IP, Maryam S, Tika IN. Ectoine and Hydroxyectoine from Halophilic Bacteria Isolated from Traditional Solar Saltern at Pejarakan Village, Buleleng Regency, Bali. Hayati. 2022;29(5):669-680.
Crossref - Sauer T, Galinski EA. Bacterial milking: A novel bioprocess for production of compatible solutes. Biotechnol Bioeng. 1998;57(3):306-313.
Crossref - Shivanand P, Mugeraya G. Halophilic bacteria and their compatible solutes–osmoregulation and potential applications. Curr Sci. 2011;100(10):1516-1521. https://www.jstor.org/stable/24076671
- Roberts MF. Organic compatible solutes of halotolerant and halophilic microorganisms. Aquat Biosyst. 2005;1:5.
Crossref - Zhang LH, Lang YJ, Nagata S. Efficient production of ectoine using ectoine-excreting strain. Extremophiles. 2009;13(4):717-724.
Crossref - Fatollahi P, Ghasemi M, Yazdian F, Sadeghi A. Ectoine production in bioreactor by Halomonas elongata DSM2581: Using MWCNT and Fe-nanoparticle. Biotechnol Prog. 2021;37(1):e3073.
Crossref - Van-Thuoc D, Hien TT, Sudesh K. Identification and characterization of ectoine-producing bacteria isolated from Can Gio mangrove soil in Vietnam. Ann Microbiol. 2019;69: 819-828.
Crossref - Kumar S, Stecher G, Suleski M, Sanderford M, Sharma S, Tamura K. Molecular Evolutionary Genetics Analysis Version 12 for adaptive and green computing. Mol Biol Evol. 2024;41(12):msae263.
Crossref - Zhang H, Liang Z, Zhao M, et al. Metabolic Engineering of Escherichia colifor Ectoine Production With a Fermentation Strategy of Supplementing the Amino Donor. Front Bioeng Biotechnol. 2022;10:824859. Published 2022 Jan 25.
Crossref - Miao C, Jia F, Wan Y, Zhang W, Lin M, Jin W. Halomonas huangheensis sp. nov., a moderately halophilic bacterium isolated from a saline-alkali soil. Int J Syst Evol Microbiol. 2014;64(Pt 3):915-920.
Crossref - Parwata IP, Wahyuningrum D, Suhandono S, Hertadi R. Ectoine Production by Halomonas elongata Bk-AG25: Two-Step Optimization using the Response Surface Methodology. Asian J Microbiol Biotechnol Environ Sci. 2020;22(1):181-188.
- Van-Thuoc D, Guzman H, Quillaguaman J, Hatti-Kaul R. High productivity of ectoines by Halomonas boliviensis using a combined two-step fed-batch culture and milking process. J Biotechnol. 2010;147(1):46-51.
Crossref - Chen WC, Hsu CC, Lan JC, Chang YK, Wang LF, Wei YH. Production and characterization of ectoine using a moderately halophilic strain Halomonas salina BCRC17875. J Biosci Bioeng. 2018;125(5):578-584.
Crossref - Chen Y, Liu Y, Meng Y, et al. Elucidating the salt-tolerant mechanism of Halomonas cupida J9 and unsterile ectoine production from lignocellulosic biomass. Microb Cell Fact. 2024;23(1):237.
Crossref - Onraedt AE, Walcarius BA, Soetaert WK, Vandamme EJ. Optimization of ectoine synthesis through fed-batch fermentation of Brevibacterium epidermis. Biotechnol Prog. 2005;21(4):1206-1212.
Crossref - Li TT, Qu A, Yuan XN, et al. Response surface method optimization of ectoine fermentation medium with moderate halophilic bacteria Halomonas sp. H02. IOP Conf. Ser.: Earth Environ. Sci. 2017;77:012019.
Crossref - Bergmann S, David F, Franco-Lara E, Wittmann C, Krull R. Ectoine production by Alkalibacillus haloalkaliphiles-bioprocess development using response surface methodology and model-driven strategies. Eng Life Sci. 2013;13(4):399-407.
Crossref - Orhan F, Akncoglu A, Ceyran E. Ectoine production from a novel bacterial strain and high-purity purification with a cost-effective and single-step method. J Biotechnol. 2024;388:24-34.
Crossref - Ma Y, Sun Z, Tashilhamo et al. Halomonassp. NEC-1, an ectoine-rich and wide pH tolerant halophilic bacterium isolated from a salt lake of Xizang, China. J Ocean Limnol. 2025;43:1636-1646.
Crossref - Radha A, Najar IN, Nekkala N et al. Efficient production of ectoine using isolated Halomonas smyrnensisIIIM VA-6 at the reduced salinity approach. FEMS Microbiol Lett. 2026;373:fnag001.
Crossref - Leszczewicz M, Broncel N, Fraczak O, Kapela T, Makowski K. Screening of Moderately Halophilic Bacteria Producing Ectoine Resulting in the Selection of Virgibacillus salarius BHTA19. Food Technol Biotechnol. 2025;63(3):310-319.
Crossref - Rekadwad BN, Li WJ, Rekha PD. The diversity of unique 1,4,5,6-Tetrahydro-2-methyl-4-pyrimidinecarboxylic acid coding common genes and Universal stress protein in Ectoine TRAP cluster (UspA) in 32 Halomonas species. BMC Res Notes. 2021;14(1):296.
Crossref
© The Author(s) 2026. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.