ISSN: 0973-7510
E-ISSN: 2581-690X
Resistance to antimycobacterial agents consistently remains a major obstacle to end TB in India. Geographical prevalence data regarding drug-resistant evolutionary genetics of M. tuberculosis (MTB) remains sparse in India. Our objective was to determine the genotypic drug resistance mutation pattern for Rifampicin and Isoniazid of MTB isolates to gain an understanding of the prevailing molecular epidemiology of drug-resistant tuberculosis. In this study 2528 M. tuberculosis DNA isolates from presumptive DRTB suspects received at the nodal TB reference laboratory in Kerala were tested for Rifampicin and Isoniazid resistance by sequence-based diagnostic Line Probe assay (LPA). Geographical prevalence and associations of rpoB, katG, inhA resistance codons was analyzed from January 2019 to March 2020. Among the 2528 DNA samples subjected for Rifampicin and Isoniazid resistance determination by LPA, 146 (5.8%) isolates were resistant to both drugs. Isoniazid mono-resistance was found in 164 (6.5%) and Rifampicin mono-resistance in 38 (1.5%) isolates. The most frequent rpoB mutation was S531L (60.32%) followed by S531W/L533P mutations seen in 8.15% of the isolates. S315T1 KatG mutation was seen in 97.33% of Isoniazid resistant isolates. 84.68% isolates with rpoB S531L mutation were found to be multidrug-resistant. 82.9% of isolates with rpoB S531L mutation showed katG S315T1 mutation. Mono isoniazid-resistant isolates were significantly higher compared to mono rifampicin-resistant isolates among the DNA isolates studied in our region. The molecular epidemiological pattern most frequently associated with multidrug resistance was rpoB S531L which was significantly associated with the co-presence of S315T1 mutation.
Mycobacterium tuberculosis, Multi-drug resistance, rpoB, katG, inhA mutations
Every year tuberculosis (TB) affects approximately 10 million individuals worldwide.1 Drug resistant tuberculosis patients are likely to remain infectious for a long period, resulting in reverberating public health consequences undermining tuberculosis control.2 India accounts for 27% global share of drug resistant TB cases.1 Multiple converging factors have ushered the explosive emergence of Multi Drug Resistant tuberculosis (MDR-TB) in India, paralyzing the regimen applicability of Isoniazid and Rifampicin, the potent first-line anti-TB drugs.
Chromosomal mutations entrapping anti-mycobacterial drug targets or the mycobacterial enzymes that activate pro-drugs exclusively mediate drug resistance in tuberculosis Rifampicin targets mycobacterial RNA polymerase and inhibits transcription. Rifampicin resistance determining mutations are predominantly (>96%) found in a 27 codon central region (81-bp RIF resistance-determining region), of the RNA polymerase subunit β (rpoB) encoding gene.3 Isoniazid (H) is activated by catalase peroxidase enzyme. It acts by formation of reactive radicals capable of damaging mycobacterial cell wall. H resistant isolates delineate mutations in the catalase peroxidase coding gene (katG) or the enoyl acyl reductase encoding promoter region (inh A) required for fatty acid biosynthesis and survival of mycobacterium species.4,5
Conventional phenotypic TB drug susceptibility testing (DST) is hitherto considered highly accurate gold standard for diagnosing anti-TB drug susceptibility. However these conventional methods are slow and cumbersome, requiring sequential procedures for isolation, identification, and DST testing of the MTB isolates. The delay in diagnosis may meanwhile result in spread and amplification of drug resistant strains due to inappropriate treatment.6 Sequence-based diagnostic assays which include INNO-LiPATB test7 and the GenoType MTBDRplus8 are WHO approved accessories for swift MDR-TB diagnosis by detection of specific drug resistant mutations.6 The drawback of the sequence based tests lies in the fact that they primarily depend on the demonstration of pre-set, internally incorporated mutations. The clinical utility of these diagnostics vary on the basis on geographical distribution of mutant strains.
The prevalent mutation sequences associated with Mycobacterial strain specific first line antitubercular drug resistance has been reported from North & Eastern India.9-11 Data of the prevalent first line drug resistant mutations from south India is sparse. The present study was undertaken at the Intermediate Reference Tuberculosis Laboratory, Kerala with the intention to put forth the scenario of genotypic-resistance of MTB isolates to Rifampicin and Isoniazid from Kerala. An attempt has been made to analyze the frequency, association and geographical generality of gene mutations in rpoB, katG and inhA genes by application of sequence based MTBDR plus assay.
A retrospective, cross-sectional study was undertaken in the Intermediate Reference Laboratory (IRL) for TB, Kerala. The Laboratory complies with the requirements of the national certification guidelines of the Central TB Division (CTD) MOH, GOI and is authorized by the CTD for Line probe assay testing as well as for Culture and Drug Sensitivity Testing (DST).
A total of 2528 MTB DNA specimens isolated from presumptive DRTB suspects received at the IRL between January 2019- March 2020 were included in the study. Follow up samples and multiple samples received from a single individual for testing were excluded from the study.
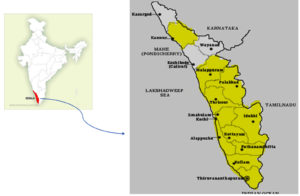
All samples were processed in a level 3 biosafety facility designed for TB containment.12 Sputum and extra-pulmonary samples received within 72 hours of collection in a triple layer packaging13 through a local courier facility for routine diagnosis and DST at the IRL laboratory from 11 districts of Kerala; Alappuzha, Ernakulam, Idukki, Kollam, Kannur, Kottayam, Malappuram, Palakkad, Pathanamthitta, Thrissur and Thiruvananthapuram (Fig. 1) were included in the present study.
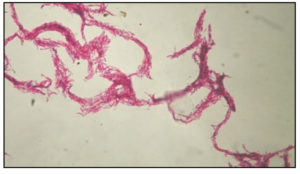
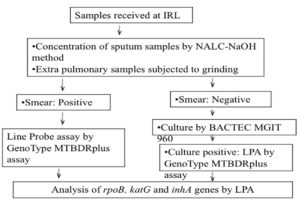
Homogenization, digestion and decontamination of sputum samples were carried out by N-acetyl-L-cysteine-sodium citrate-Sodium hydroxide procedure.14 Digested-decontaminated sputum samples were subjected to smear microscopy. The extra-pulmonary samples were minced and grinded in a class II B biosafety cabinet in the TB containment lab and then subjected to smear microscopy. As per PMDT guidelines sputum and extra-pulmonary smear positive samples were subjected to DNA extraction and LPA.9 AFB smear negative specimens were inoculated into the BACTEC MGIT 960 system.15 MGIT tube Cultures giving positive alert beep were confirmed by serpentine cording (Fig. 2) on smear microscopy and rapid immunochromatographic card test for MPT64 TB Ag (Fig. 3).Culture confirmation was followed by DNA extraction and LPA. The Mtb H37Rv (ATCC 27294) and sterile nuclease free water served as positive and negative quality controls for each batch of DNA isolates tested. The study workflow is depicted in Fig. 4.
DNA extraction
Mycobacterial pellets obtained by the centrifugation (10,000g for 15 minutes) of 500 µl of smear positive samples /1000 µl liquid broth from flagged MGIT tubes were treated with 100 µl Genolyse lysis buffer. Thermal squelching at 95°C for 5 minutes was performed. Subsequently neutralization buffer was added and samples were centrifuged for 5 min at 13,000g.9 Supernatant containing mycobacterial DNA was collected in sterile screw capped tubes for LPA based hybridization and storage at -20°C.
Mastermix preparation
GenoType MTBDRplus kit version 2 constituent reagents were condensed and reconstituted as per manufacturer’s instructions. 5 µl extracted DNA isolate was cued into the polypropylene tubes containing mastermix and subjected to amplification.
Amplified DNA was denatured and nucleotides were reverse-hybridized to oligonucleotide probes on the LPA (nitrocellulose) strip. Conjugate reaction and substrate addition using GenoType MTBDRplus version 2 kit for PCR product band detection was performed.
Results were interpreted by the binding pattern of the isolate to wild type probes and specific mutant probes by using the reporting card. Absence of specific wild-type-probebands (WT) /presence of mutant-probe-bands (MUT) indicate resistance to the particular antibiotic. Mycobacterial isolates having presence of mutant bands alongside with wild type bands were termed to be heteroresistant.
Data thus obtained was scrutinized for statistical significance by P value estimation using ANOVA calculator software.
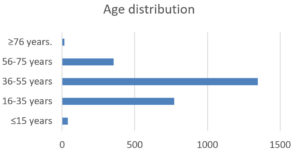
5413 diagnostic samples acknowledged at the Intermediate Reference Laboratory from January 2019 to March 2020 were retrospectively analyzed. 2528 DNA isolates were subjected to Line Probe assay by GenoType MTBDRplus kit version 2. Male: Female ratio was found to be 3.4:1. 53.28% of the DNA isolates analyzed were obtained from the age group 36-55 years. Age-wise distribution has been demonstrated in Fig. 5.
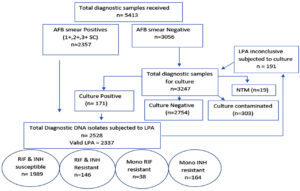
Among the 5413 diagnostic samples received 4661 (86.1%) were sputum samples and 752 (13.89%) were extrapulmonary samples. 43.54% of samples were found to have acid alcohol resistant bacillus (BAAR) on AFB smear microscopy and were subjected directly to LPA. The remaining 56.46% of samples were subjected to MGIT culture along with 191 samples which were deemed to be inconclusive on LPA, as described in Fig. 6 (process of workflow). Among the 191 LPA inconclusive samples subjected to culture 18 (9.42%) showed MGIT contamination. 152 samples (79.5%) showed no growth and 21 of the LPA inconclusive samples flagged positive on MGIT.
2357 smear positive isolates and 171 MGIT flagged samples positive for acid fast bacilli were consigned to LPA. 2337 (92.44%) showed valid LPA results of which 1989 (85.1%) isolates were susceptible to both Rifampicin & Isoniazid. 146 (5.8%) isolates were resistant to both the above mentioned drugs. 38 (1.5%) isolates were mono Rifampicin resistant. 164 (6.5%) isolates were mono Isoniazid resistant.
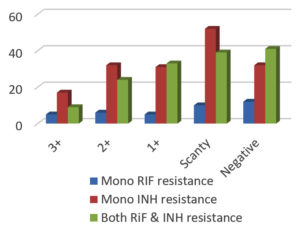
MDR isolates/Rifampicin resistance was 2.7 times more common among males as compared to females. Mono rifampicin resistance was significantly lower (p= 0.01864) among the study isolates in comparison to mono isoniazid & MDR isolates. However there was not significant difference between the existence of mono-isoniazid resistance in comparison with MDR isolates (p= 0.85492) The presence of drug resistance was predominant in samples having smear positivity grades 1+ and below as depicted in Fig. 7 (p= 0.005541).
The study found the absence of rpoB WT8 with the presence of rpoB MUT3 associated with S531L mutation in 111/184 (60.32%) rifampicin resistant isolates. Co-existence of rpoB MUT3 gene with documentation of all (WT) wildtypes was found in 2/184 (1.08%) isolates. (Table 1) Absent wildtype bands with no documented (MUT) mutant bands was seen in 59/184 (32%) RIF resistant isolates. Missing WT8 without known mutation was seen in 15/184 (8.15%) RIF resistant isolates. Missing WT7 and WT2/3 accounted for 11/184 (5.9%) each followed by WT2 accounted for 8/184 (4.3%), WT3/4 for 5/184 (2.7%), WT5/6 for 4/184 (2.17%) WT4/5 for 3/184 (1.6%) of the RIF resistant isolates.
Table (1):
rpoB gene patterns detected by GenoType MTBDRplus assay.
| No | codons analysed | Failing rpoB WT band(s) | Mutant (MUT) band developed | Associated Mutations | Study isolates |
|---|---|---|---|---|---|
| 1 | 505-509 | 1 | F505L, T508A, S509T | 2 | |
| 2 | 510-513 | 2 | L511P* | 8 | |
| 3 | 510-517 | 2/3 | Q513L*, Q513P, DEL514-516 | 11 | |
| 4 | 513-519 | 3/4 | 1 | D516V | 0 |
| D516Y, del515 | 5 | ||||
| 5 | 516-522 | 4/5 | del 518*, N518I | 3 | |
| 6 | 518-525 | 5/6 | S522L, S522Q | 4 | |
| 7 | 526-529 | 7 | 2A | H526Y | 8 |
| 2B | H526D | 4 | |||
| H526R,H526P*, H526Q, H526N, H526L, H526S, H526C | 11 | ||||
| 8 | 530-533 | 8 | 3 | S531L | 111 |
| S531Q*, S531W, L533P | 15 | ||||
| 9 | nil | 3 | Heteroresistance | 2 |
*RARE THEORITICAL MUTATION (IN SILICO)
High level Isoniazid16 resistance (was found in 223/2528 DNA isolates (8.8%) of MTB isolates. 98.2% of the katG resistant isolates were due to S315T1 mutation as demonstrated in table 2. Hetero-resistance to katG was demonstrated in two isolates showing presence of all wildtypes along with presence of katGMUT1.
Table (2):
katG Mutation patterns by GenoType MTBDRplus assay.
| No | Absent WT | codons analyzed | Mutant (MUT) developed | Mutations | Study isolates |
|---|---|---|---|---|---|
| 1 | katGWT | 315 | katGMUT1 | S315T1 | 219 |
| katGMUT2 | S315T2 | 4 | |||
| 2 | Nil | 315 | katGMUT1 | Heteroresistance | 2 |
Low level isoniazid16 resistance and inferred ethionamide/prothionamide resistance was found in 94/2528 (3.71%) of the isolates, most of which could be accounted due to C15T mutations as described in table 3. 9/2528 (0.35%) isolates showed presence of katG as well as inhA mutant bands. All 9 isolates accounted for S315T1 mutation associated with either C15T (7/9) or T8C (2/9) mutations. The 4 isolates with S315T2 mutation did not demonstrate any inhA mutant bands.
Table (3):
Mutations detected by GenoType MTBDRplus assay in inhA promoter region consistent with low level isoniazid resistant Mycobacterium tuberculosis.
| No | Absent WT band(s) | nucleic acid position | Mutant band developed | Mutations | Study isolates |
|---|---|---|---|---|---|
| 1 | inhAWT1 | -15 | inhAMUT1 | C15T | 89 |
| -16 | inhAMUT2 | A16G | 0 | ||
| 2 | inhAWT2 | -8 | inhA MUT3A | T8C | 3 |
| inhAMUT3B | T8A | 1 | |||
| 3 | Nil | -8 | inhA MUT3A | Heteroresistance | 1 |
Mutants showing S531L mutation in rpoB gene were more significantly (p value <0.0001) associated with co-existing isoniazid resistance. 94/111 (84.68%) isolates exhibiting S531L mutation were multidrug-resistant isolates showing the co-presence of S315T1 mutation in 82.9% cases and C15T in 17% cases. 121/146 (82.87%) MDR isolates exhibited katG mutations with absent inhA mutations. 21/38 (55.26%) Rifampicin mono resistant isolates showed the presence of S531L mutation.
TB elimination activities are facing a daunting challenge due to complex diagnostic & treatment challenges associated with drug resistance. The possibility of institutional/household transmission and rapid mortality associated with such infections highlight the requirement for application of rapid strain-susceptibility testing. Molecular line probe assay (LPA) is an advanced nucleic acid amplification technology endorsed by WHO6 which detects resistance associated mutations. LPA has revolutionized DR-TB diagnostics by bringing in a steep reduction in the laboratory Turn Around Time (TAT).9,10 This reduction of TAT is favorable for quick treatment initiation thereby reducing the time from clinical suspicion to treatment initiation.
Report on Global tuberculosis 2019 described the highest burden in men (aged ≥ 15 years) accounting for 57% of total TB cases in 2018. Present study documented a 3.4 times higher chance of TB positivity among males in comparison to females. Individuals in the economically productive age group of 36-55 years accounted for 53.28% of the total cases thereby translating to significant socio-economic implications. Studies by R. Singhal et al,9 Sharma et al.17 also documented a higher positivity rate among males in comparison to females. However while maximum isolates were from the age group of 16-35 in the study by Singhal et al.9, our study documents the highest sample size in the age group of 36-55 years.
Globally as per 2019 data1 61% people with bacteriologically confirmed TB have been offered diagnostics for drug resistance determination. Present study describes the RIF, INH resistance rates by the application of LPA among samples received at the Reference laboratory in Kerala under the NTEP network. In our study 5.8% isolates were found to be MDR. Mono RIF resistance was seen in 1.5% isolates & Mono INH resistance was seen in 6.5% isolates. Studies by Desikan P et al.18 on MTB isolates from central India demonstrated 7.6% Mono-isoniazid resistant, 18% MDR isolates & 11.6% mono-rifampicin resistant isolates. Studies by Singhal et al.9 on MTB isolates from Punjab demonstrated 6.6% mono INH resistance 15.5% MDR isolates & 7.1% mono RIF resistance. The variations in drug resistance rates among the different study settings may be attributed to strain variations and selection pressure in different geographical study settings. The findings of Singh et al.19 demonstrating 102 distinct MTB spoligo-patterns from different study settings all over India backs this possibility. Attitude of patients, long-term treatment follow-up and co-morbidities are the other factors influencing the varied rates of drug resistance.
The MDR-TB/ RIF resistance rates was predominant in samples having smear positivity grades 1+ and below. Studies from Punjab9 showed high incidence of MDR-TB/ RIF resistance corresponding to cases with higher bacterial counts. This may be attributed to the geographical setting and the early health seeking behaviour of the inhabitants in our region. Reciprocity of bacterial load with MDR-TB/ RIF resistance rates needs further evidence based substantiation in different geographical settings.
S531L mutation associated rifampicin resistance was predominantly evidenced among 60.32% study isolates resistant to rifampicin. Studies from Vietnam20 showed 50% association of S531L mutation with rifampicin resistance. Studies from Punjab9 and Central India18 showed 58.4% and 65.88% association of S531L mutation with rifampicin resistance respectively. Isoniazid resistance due to katG gene mutation was documented in 72.58% INH resistant study isolates in agreement with studies from Punjab 93.2%,9 France 62.5%.21 S315T1 mutation was the most common (98.2%) mutation responsible for high level isoniazid resistance. Low level INH resistance with inferred ethionamide/prothionamide resistance was found in 30.3% of isoniazid resistant isolates.
LPA apart from its listed advantages comes with a set of limitations. Since LPA is a DNA based assay only nucleic acid screening is possible by this assay. Silent/ synonymous mutations with no amino acid sequence change may also produce absence of wild type probes thereby rendering inferred resistance result for sensitive isolates. One such example is the L533P mutation which does not render RIF resistance despite failing rpoB WT8 wild type band. Such cases will require phenotypic resistance determination. In our study 32% cases of absent mutant probes alongside with absent wild probes were documented, in sync with studies from Punjab (27%),9 Delhi (26.1%),10 France (29%).21 However, studies from Central India documented 5.9%18 of such rpoB non hot spot region mutations. Geographical incidence information of mutations outside the rpoB 81 base pair region could help formulate diagnostic algorithms for TB-DST laboratories.
Hetero-resistance was documented in 1.8% of rifampicin resistant isolates and 0.9% of isoniazid resistant isolates which was marginally lower (insignificant p value = 0.65) than those documented in other Indian studies by Singhal et al.9,10 in Punjab and central India respectively. Early initiation of MDR regimen among patients harboring heteroresistant isolates in comparison to MDR TB isolates has documented better treatment outcome rates.22 Further studies to elucidate these mutations would enhance our understanding of the epidemiology and transmission dynamics of MDR-strains.
In conclusion, high prevalence of mono isoniazid resistance is documented among our study isolates. Information about common resistance imparting mutations against first line antimycobacterial agents in our geographical area has been collated to envisage the formulation of effective drug resistance control & prevention strategies in this geographical region.
ACKNOWLEDGMENTS
The authors appreciate the efforts of the staff members in IRL, Trivandrum, and the field workers of NTEP, Kerala state.
CONFLICT OF INTEREST
The authors declares that there is no conflicts of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
None.
ETHICS STATEMENT
Not applicable.
AVAILABILITY OF DATA
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
- World Health Organization. Global Tuberculosis Report 2019. Geneva, Switzerland. 2019. https://www.who.int/tb/publications/global_report/en/. Accessed on: 26 February 2020.
- Bifani PJ, Mathema B, Kurepina NE, Kreiswirth BN. Global dissemination of the Mycobacterium tuberculosis W-Beijing family strains. Trends Microbiol. 2002;10(1):45-52.
Crossref - Zaw MT, Emran NA, Zaw Lin. Mutations inside rifampicin-resistance determining region of rpoB gene associated with rifampicin-resistance in Mycobacterium tuberculosis. Journal of Infection and Public Health. 2018;11(5):605-610.
Crossref - Laurenzo D, Mousa SA. Mechanisms of drug resistance in Mycobacterium tuberculosis and current status of rapid molecular diagnostic testing. Acta Trop. 2011;119(1):5-10.
Crossref - Cardoso RF, Cooksey RC, Morlock GP, et al.Screening and characterization of mutations in isoniazid-resistant Mycobacterium tuberculosis isolates obtained in Brazil. Antimicrob Agents Chemother. 2004;48(9):3373-3381.
Crossref - World Health Organization. Policy statement; Molecular line probe assays for rapid screening of patients at risk of multi drug resistant tuberculosis (MDRTB): WHO Report 2008 WHO/ HTM/ TB Geneva, Switzerland; 2008. https://www.who.int/tb/features_archive/policy_statement.pdf
- Morgan M, Kalantri S, Flores L, Pai M. A commercial line probe assay for the rapid detection of rifampicin resistance in Mycobacterium tuberculosis: A systematic review and meta-analysis. BMC Infect Dis. 2005;5:62.
Crossref - Ling D, Zwerling AA, Pai M. GenoType MTBDR assays for diagnosis of multidrug-resistant tuberculosis: A meta-analysis. Eur Respir J. 2008;32:1165-1174.
Crossref - Singhal R, Arora J, Sah GC, et al. Frequency of multi-drug resistance and mutations in Mycobacterium tuberculosis isolates from Punjab state of India. Journal of Epidemiology and Global Health. 2017;7(3):175-180.
Crossref - Singhal R, Myneedu VP, Arora J, et al. Early detection of multi-drug resistance and common mutations in Mycobacterium tuberculosis isolates from Delhi using GenoType MTBDRplus assay. Indian J Med Microbiol. 2015;33(Suppl. 1):S46-52.
Crossref - Singhal R, Myneedu VP, Arora J, Singh N, Sah GC, Sarin R. Detection of multi-drug resistance & characterization of mutations in Mycobacterium tuberculosis isolates from North- Eastern States of India using GenoType MTBDRplus assay. Indian J Med Res. 2014;140(4):501-506.
- Tuberculosis laboratory biosafety manual. World Health Organization. 2012.
- Guidance on regulations for the Transport of Infectious Substances 2017-2018. © World Health Organization. 2017.
- Kent PT, Kubica GP. Public health mycobacteriology. A guide for the level III laboratory. Atlanta, GA: Centers for Disease Control and Prevention. 1985. https://ntrl.ntis.gov/NTRL/dashboard/searchResults/titleDetail/PB86216546.xhtml
- Revised National Tuberculosis Control Programme (RNTCP), Central Tuberculosis Division, Directorate General of Health Services, Ministry of Health and Family Welfare. Guidelines on Programmatic Management of Drug Resistant TB (PMDT) in India. New Delhi; 2017. https://tbcindia.gov.in/WriteReadData/NSP%20Draft%2020.02.2017%201.pdf
- WHO treatment guidelines for isoniazid-resistant tuberculosis. Version: 24 April 2018 Geneva, World Health Organization. 2018. https://www.who.int/tb/publications/2018/FAQ_TB_policy_recommendations_guidelines.pdf
- Sharma SK, Kaushik G, Jha B, et al. Prevalence of multidrug-resistant tuberculosis among newly diagnosed cases of sputum positive pulmonary tuberculosis. Indian J Med Res. 2011;133:308-311.
- Desikan P, Kharate A, Panwalkar N, et al. Frequency of mutations in rifampicin and isoniazid resistant isolates of M. tuberculosis: an analysis from Central India. Germs. 2016;6(4):125-131.
Crossref - Singh J, Sankar MM, Kumar P, Couvin D, Rastogi N, Singh S. Genetic diversity and drug susceptibility profile of Mycobacterium tuberculosis isolated from different regions of India. J Infect. 2015;71(2):207-219.
Crossref - Huyen MNT, Tiemersma EW, Nguyen TNL, et al. Validation of the GenoType MTBDRplus assay for diagnosis of multi-drug resistant tuberculosis in South Vietnam. BMC Infect Dis. 2010;10:149.
Crossref - Brossier F, Veziris N, Pernot CT, Jarlier V, Sougakoff W. Performance of the GenoType MTBDR line probe assay for detection of resistance to rifampicin and isoniazid in strains of Mycobacterium tuberculosis with low and high level resistance. J Clin Microbiol. 2006;44(10):3659-3664.
Crossref - Tolani MP, D’souza DTB, Mistry NF. Drug resistance mutations and heteroresistance detected using the GenoType MTBDRplus assay and their implication for treatment outcomes inpatients from Mumbai, India. BMC Infect Dis. 2012;12:9.
Crossref
© The Author(s) 2021. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.