ISSN: 0973-7510
E-ISSN: 2581-690X
Bovine abortion is economically one of the most devastating problems faced by dairy farmers. Apart from non-infectious causes, several infectious pathogens are responsible for abortions, which sometimes manifests as abortion storms. Vaccine against several pathogens is available, in spite of that, abortions cause huge economic losses for the dairy sector. Timely and accurate identification of the etiological agent helps in adopting the mitigation steps to control the damage caused. In addition to the common abortion-causing pathogens such as Brucella abortus, Bovine herpesvirus-1 (BHV-1), bovine viral diarrhea virus (BVDV), several emerging viral causes are being investigated for their possible role in abortion, either exclusively or as co-infection. Molecular methods are widely accepted for the identification of the involved pathogens. However, these assays require individual screening against each pathogen which is time-consuming and uneconomical, hence the multiplex format of PCR assays has been adopted by several laboratories. Multiplexing in real-time PCR is a sensitive and reliable technique, but it requires trained manpower and sophisticated equipment which is largely unavailable in regional disease diagnostic laboratories in India. Hence, in this study, a user-friendly, ready-to-use, gel-based RT-PCR multiplex assay was developed for simultaneous detection of three common pathogens (B. abortus, BHV-1, and BVDV) and two emerging pathogens; bluetongue virus (BTV) as a cause of abortions in bovine and Schmallenberg virus (SBV). After the standardization of the assay, a panel of 211 samples was screened. A high degree of concordance was observed which indicates the developed multiplex PCR assay is reliable and has the potential for screening at regional diagnostic laboratories.
Multiplex PCR, Bovine abortion, Schmallenberg virus, BHV-1, BTV, Brucella abortus
Livestock plays an important role in supporting the Indian economy. India has the largest livestock population of more than 500 million1. The contribution of the livestock sector in the country’s gross domestic product (GDP) is nearly 4.11%1. However, there are serious threats like abortions to animal health and production, which cause huge economic losses. The majority of animals are reared under an un-organised sector where abortions due to infectious pathogens are a significant problem. The burden of maintaining unproductive females, for longer periods, causes huge financial losses to the poor farmers3.
Various viral, bacterial and protozoan pathogens are responsible for abortions in the cattle population in India4. Such pathogens include viruses like Bovine herpesvirus-1 (BHV-1), and bovine viral diarrhea virus (BVDV) as the most common viral causes of abortions5-7. Bacterial agents such as Brucella abortus are considered as the most important bacterial zoonotic pathogen, causing abortions8. Apart from these, emerging viruses such as Schmallenberg virus (SBV) and the role of bluetongue virus (BTV) in causing abortion are now being extensively investigated9-11.
Transmission of bovine brucellosis is estimated to occur at an endemic equilibrium in India, with disease prevalence estimated to be 13.5%12. The seroprevalence of BHV-1 and BVDV in India has been estimated to about 38.0%5 and 15.29%13, respectively. Although BTV is endemic in the Indian population of sheep and goats with an estimated seroprevalence of 39-43% with frequent reporting of clinical outbreaks from various parts of India, clinical outbreaks in cattle have never been reported. However, seroprevalence in cattle has been reported up to 34%14. Recently, involvement of BTV in bovine abortion is being highlighted and is a matter of investigation11.
A recently identified viral agent, Schmallenberg virus (SBV; an Orthobunyavirus) causes abortions in cattle, sheep, and goat population15. Although the presence of SBV in India is yet to be established, serological studies have indicated circulation of antibodies against SBV in the Indian livestock population16.
The infectious diseases causing abortions are preventable, provided the prompt and accurate diagnosis is available and mitigation efforts are adopted during the early course of the disease. Veterinary diagnostic laboratories at the regional and national level plays an important role in obtaining an accurate and timely diagnosis.
Molecular methods are becoming a handy tool for providing a specific and rapid diagnosis. During the last few decades, sensitivity and specificity of nucleic acid-based diagnostic methods have improved tremendously. However, the involvement of different abortion- causing pathogens requires individual detection, which is time-consuming and uneconomical. Simultaneous detection of up to six viral pathogens, namely, foot-and-mouth disease virus (FMDV), BTV, Vesicular stomatitis virus (VSV), BVDV, bovine rotaviruses (BRVs), and BHV-1 have been demonstrated recently, by multiplex real-time PCR assays17. However, probe-based assays require a specific set of equipment and expertise which is largely unavailable at the regional laboratories in India. Hence, conventional gel-based multiplex PCR has been utilized principally for molecular diagnosis.
Considering the challenges faced for rapid identification of bovine abortion causative agents in the regional laboratories in India, a multiplex one-step PCR assay was developed to provide simultaneous detection of B. abortus, BHV-1, BVDV, BTV, and SBV.
In order to obtain distinct amplification products for easy differentiation in agarose gel electrophoresis for each pathogen, primer sets were designed for BHV-1, BVDV, and BTV whereas, recommended primers were used for specific detection of SBV and B. abortus. The details of primers used in this study are described in table 1. Representative gene sequences of BTV, BHV, and BVDV were downloaded from the NCBI database and aligned with the Mega7 multiple alignment tool. Primer pairs were designed using the Hyden tool18. NCBI primer blast tool was used for evaluating the cross-specificity (https://www.ncbi.nlm.nih.gov/tools/primer-blast/). The primer pairs were synthesized from Eurofins (Bangalore, India).
Table (1):
Primers used in the study.
| S No | Primer name | Sequence | Target organism | Gene | PCR amplicon size (bp) | Ref |
|---|---|---|---|---|---|---|
| 1 | BVDV-1F | GGGATCCGATTTAGCCATGC | BVDV | 5’UTR | 204 | Present study |
| BVDV-1R | TTTTCACCTGAACGACCCCC | |||||
| 2 | BT_G01F | GCTGCATTCGCATCGTACGCAGAAGC | BTV | Segment 10 | 459 | Present study |
| BT_G01R | AAACCTCGGGGCGCCACTCTACCTAC | |||||
| 3 | SBV-S-FW | TCAGATTGTCATGCCCCTTGC | SBV | S fragment | 88 | 31 |
| SBV-S-RW | TTCGGCCCCAGGTGCAAATC | |||||
| 4 | BHV-gNF | CAGCAGCTTTCCAACCGCCA | BHV-1 | UL49.5 gene | 362 | Present study |
| BHV-gNR | CGCAGCGTTGCGGTCAATCA | |||||
| 5 | IS711- for | CGTGTCTGCATTYAACGTAACC | Brucella | IS711 gene | 498 | 32 |
| IS711-rev | TCCTCATTGWCAGCACCATATC |
The designed primers were optimised for the detection of the known positive controls. Virus isolates of BHV-1 and BTV10 were revived in MDBK and BHK-21 cells, respectively. With the appearance of obvious cytopathic effects (CPE), viruses were harvested and subjected to the end-point dilution method of titration as described by Spearman and Karber19,20. Total nucleic acid (TNA) was extracted from virus isolates with a pre-determined titer, using magnetic beads-based MagMax TNA extraction kit (Thermo Scientific), as per the manufacturer’s instructions.
The 5’UTR of BVDV and partial S-segment of SBV of size 282 bp and 702 base pairs, respectively, were chemically synthesized commercially (Biotech Desk India) as virus isolates were unavailable in the laboratory. The synthesized genes were cloned into the pGEMT easy vector system (Promega, USA) as per the manufacturer’s recommendation followed by transformation into competent E. coli cells. Plasmids were extracted and quantified using a Qubit spectrophotometer and were used as positive controls. B. abortus strain available in the laboratory was revived on selective Brucella agar media and confirmed by PCR assay.
Various conditions such as annealing temperature, magnesium ion concentration, and primer concentrations were optimized for each pathogen with the positive controls to obtain a single amplicon. Then, a pool was prepared consisting of BHV-1 and BTV10 viruses in concentrations 2.1×103.5 TCID50 each and B. abortus 1010 CFU/ml. Synthetic gene vector constructs of BVDV and SBV (100ng each) were also spiked in the pool. The pool was serially 10-fold diluted in 2% FBS EMEM media. Total nucleic acid extracted was tested with the multiplex format of primers with optimized conditions. Briefly, the reaction mixture was assembled in 0.2ml PCR tubes on ice, using One-Step RT-PCR kit (Qiagen, Germany) as per the manufacturer’s instructions with minor modifications. 10 X reaction buffer was mixed with 05 pmol of forward and reverse primers (Table 1), each and enzyme mixture. Magnesium chloride concentration was adjusted to 3mM final concentration followed by the addition of nucleic acid template. The amplification cycles were as follows; reverse transcription step at 50°C for 30 min followed by initial denaturation at 95°C for 15 min, 35 cycles of denaturation at 95°C for 15 sec, annealing at 60°C for 30 sec and extension at 72°C for 30 sec. A final extension for 10 min at 72°C was provided.
Panel (n =211) of tissue samples/swabs consisting of BHV-1 (n =41), B. abortus (n =82), and BVDV (n =19) were processed. Positive and negative samples for above-mentioned pathogens were obtained from a sample repository maintained at the Center for Animal Disease Research and Diagnosis (CADRAD; IVRI, Bareilly, India). Additionally, nasal swab samples (n =35), positive for BTV from sheep were also included in the study. Details of these samples are presented in Table 2. The total nucleic acid was extracted from these samples and tested with the multiplex format of RT-PCR using optimized concentrations and conditions. The amplified products were resolved in 1.5% agarose gel.
Table (2):
Details of samples used for validation in this study.
S No |
Pathogen |
Stomach content of aborted foetus |
Vaginal Swab |
Placenta |
Semen |
Nasal swabs |
Total |
|---|---|---|---|---|---|---|---|
1 |
BHV-1 |
10 |
6 |
12 |
8 |
5 |
41 |
2 |
B. abortus |
46 |
13 |
18 |
5 |
0 |
82 |
3 |
BVDV |
1 |
3 |
15 |
0 |
0 |
19 |
4 |
BTV |
0 |
0 |
0 |
0 |
35 |
35 |
Further, to develop a user-friendly and rapid method for performing one-step RT-PCR assay, the reagents were freeze-dried with supplementation of 100mM trehalose (Merck Millipore) as stabilizer, as described by Sharma and co-workers21. The lyophilized mixture containing the constituents for the detection of all five pathogens; primer sets, reverse transcriptase and polymerase enzymes, and other PCR components were prepared. These components required reconstitution with 100µl nuclease-free water in each vial, before setting up the reactions. For a 25µl reaction, 23µl of reconstituted reaction mix was mixed with 2µl of extracted TNA from the samples. The freeze-dried master mix was used for testing TNA extracted from samples of the above-mentioned panel and the results were analyzed.
The lowest detection limit was estimated by spiking the known concentrations of BTV, BHV, and B. abortus in previously known negative placental tissues. For BVDV and SBV, known concentrations of plasmid were spiked in negative placental tissues. TNA was extracted as described above and subjected to amplification using lyophilized reaction mixture.
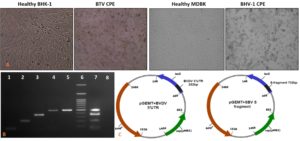
The virus isolates of BTV-10 and BHV-1, available at the laboratory were revived in the cell culture system (Fig. 1(A)). BTV-10 was found to contain 3 × 104.5 TCID50/ml whereas the titer for BHV-1 was 4.4×107.5 TCID50/ml. B. abortus strain isolated from abortion consisted of 1010 CFU/ml. Since the confirmed isolates of BVDV and SBV were not available, a synthetic gene construct of size 282 bp of 5´UTR of BVDV and 702bp of S fragment of SBV were chemically synthesized and cloned into pGEMT easy vector.
Clearly, distinct bands of PCR amplicons were obtained in multiplexing format with the designed primers pairs at a common annealing temperature of 60°C and extension at 72°C for 30 sec (Fig. 1(B)).
Fig. 1. (A) Propagation of BTV10, and BHV-1 in BHK-21 cells and MDBK cells, respectively. BHV-1 and BTV-10 viruses showed characteristic CPE at 48-72 hpi. (B) Agarose gel (2%) for singleplexed and multiplexed amplicons of abortion causing pathogens. 1- 88bp SBV, 2- 204bp BVDV, 3- 362bp BHV-1, 4- 459bp BTV, 5- 498bp B. abortus, 6- 100bp DNA marker, 7- 88bp SBV; 204bp BVDV; 362bp BHV-1; 459bp BTV; 498bp B. abortus, multiplexed. 8- No template control for multiplexed PCR. (C) A representation of insertion of synthetic gene constructs for BVDV 5´UTR and SBV; S segment in pGEMT vector system.
A panel of samples consisting of positive samples, as detected by OIE described method for each pathogen (Table 2) was used for the validation of the multiplex PCR developed in the study. A high degree of concordance was observed when the panel of previously known samples (n =211) was tested with the optimized method. Out of 41 positive samples for BHV-1, 39 (95.1% concordance) resulted in the amplification of 362 bp amplicon with multiplex PCR whereas none other sample showed amplification (no false positive), indicating the high specificity of the assay with little lower sensitivity. Similarly, out of 19 BVDV positive samples, 18 (94.7% concordance) resulted in amplification of 204 bp amplicon. All 82 B. abortus positive samples resulted in amplification of 498 bp (100% concordance) amplicon, however, one additional sample was also found positive which was considered as false positive (1.2% discordance).
Additional swab samples (n= 35) known to be positive for BTV were also tested with the developed multiplex PCR assay. All 35 samples showed clear amplification without any cross-amplification with other pathogens (100% concordance) indicating the high specificity of the assay for BTV. However, no tissue samples obtained from aborted animals resulted in positive amplification. All the samples were found negative for SBV. The lowest detection limit by freeze-dried PCR assay was equivalent to 6 TCID50 for BHV, 5 X 102.0 for BTV, 100 CFU/ml for B. abortus and 60fg each for SBV and BVDV. We also compared the wet reagent and freeze-dried reagent PCR mixtures with our panel of known samples. We did not find any differences while comparing the samples with wet reagents and lyophilized reagents (Table 3)
Table (3):
Analogy between the freeze-dried reagent and wet reagent multiplexed PCR reaction mixtures.
S No |
Pathogen |
Total samples |
Multiplexed PCR results (using wet reagent PCR mix) |
Multiplexed PCR results (using freeze dried reagent PCR mix) |
|---|---|---|---|---|
1 |
BHV-1 |
41 |
39 |
39 |
2 |
B. abortus |
82 |
82 |
82 |
3 |
BVDV |
19 |
18 |
18 |
4 |
BTV |
35 |
35 |
35 |
Apart from non-infectious causes, several pathogens are responsible for abortions in cattle and buffaloes4. Most of these pathogens cause abortion storms in the intensive form of animal rearing. In an un-organized sector, where only 2-3 animals are reared per household, several pathogens are endemic and occasionally lead to abortions. Although careful clinical and post-mortem observations may shortlist the tentative pathogens involved in the abortions, laboratory confirmation is a must to implement suitable mitigation measures.
Accurate and rapid diagnosis of the abortion-causing agents in bovines can prevent the huge economic losses incurred by the dairy industry. Nucleic acid amplification methods are considered as the most reliable and practical means for pathogen detection, especially when multiple pathogens are suspected. We regularly receive the samples for identification of pathogens in the cases of abortions, and the diagnosis is provided utilizing OIE recommended methods. However, we understand that rapid and user-friendly kits should be available to the resource-poor regionally located laboratories so that the time spends on transportation of samples can be utilized for early diagnosis and treatment. However, considering the lack of suitable infrastructure and trained manpower to carry out the real-time PCR assay, it was realized that user-friendly assay then real-time PCR should be developed. Therefore, in this study, we developed a ready-to-use multiplex RT-PCR assay for simultaneous detection of five pathogens. Three, BHV, BVDV, and B. abortus are common pathogens responsible for abortions in most cases, whereas SBV and BTV are emerging pathogens associated with bovine abortions11,15. A set of the multiplex standard PCR-assay was developed for the detection of many bacterial and viral pathogens associated with bovine abortions by Tramuta and co-workers22. Over a period of time, a ready-to-use gel-based multiplex PCR assay for detection and typing of FMD at regional laboratories has significantly contributed in the control of FMD in India21.
The assay was developed such that the size of the PCR amplicon for each pathogen could be distinct, to avoid confusion in the agarose gel electrophoresis analysis. Therefore, new primer pairs were designed for BTV, BHV-1, and BVDV, whereas previously published primers for SBV and B. abortus were used (table 1). After verification of primer specificity, the assay was optimized for use in freeze-dried format.
Reagents such as polyethylene glycol (PEG) and glycerol are widely used for the preservation of PCR components. Also, trehalose has been proved as an excellent agent for the cryo-preservation of PCR reagents in the ready-to-use format21. Therefore, in this study, we used trehalose (100mM) as a stabilizer. Lyophilization of PCR reagents not only reduces the complexity for use but also enhances the stability of reagents which is essential for the transportation of the reagents to distant laboratories. It also minimizes the cold storage cost as the lyophilized reagents can be stored at ambient temperature. Furthermore, lyophilization reduces the chances of contamination during step-wise preparations and as the necessary reagents are present in appropriate concentration in freeze-dried form, the errors due to improper wet reagent handling can also be reduced. Lyophilization did not compromise the analytical sensitivity of the assay. The lowest detection limit in placental tissue spiked samples for the freeze-dried PCR components were 6 TCID50 for BHV and 5 X 102.0 BTV, 100 CFU/ml in B. abortus, and 60 fg of spiked plasmid which were comparable with the earlier studies where fresh master mix was prepared for PCR reactions23,24.
The primers designed in the study resulted in specific amplification of targets without any cross specificity, which indicates their suitability for wider usage. However, further extensive validation (including inter-laboratory and field laboratory validation) is required before they can be accepted as universal primers.
The genomic content of pathogens involved in abortions in bovines hugely varies. The genomic DNA of Brucella abortus can be isolated relatively easily when compared to viral pathogens such as BHV-1 in which capsid is largely glycosylated and genomic DNA is rich in GC content (~72%). Similarly, the genomic RNA extraction from BVDV and SBV is relatively simpler when compared to BTV in which double-stranded RNA genome is packed in multiple layers of capsids. Hence, a robust and reliable system of total nucleic acid extraction was required. We evaluated several kits including manual and column-based nucleic acid extraction methods. However, all these methods were found to suffer from one or other demerits such as time consumption or low yield. Magnetic beads-based kits offer a solution to this problem. The guanidium-based lysis buffer uses a proteinase-K solution which in presence of SDS efficiently lyse multiple complex protein capsids. These kits do not require high-speed refrigerated centrifuge and are much rapid in extraction as compared to manual methods. The reliability of these kits was assured by testing the large number of samples in the laboratory.
A high degree of correlation (0.95-1.0) was observed between the established methods of pathogen identification and the newly developed assay. A maximum level of correlation was observed for detection of B. abortus followed by BTV. Detection of a false positive in the case of B. abortus can be linked to laboratory contamination or handling error. However, one semen sample positive for BHV-1 was found negative in the newly developed assay. The PCR contaminants especially the high amount of zinc salts leads to variable results and the use of desalting columns or charcoal has been advised for semen testing25. All the swab samples positive for BTV resulted in clear amplification of BTV specific amplicon size and no cross-reactivity was observed. We emphasize here that OIE-recommended primers for the detection of BTV23 did not detect all the samples as several mutations have been inserted in the primer binding regions over a period of time. Hence, revision of primers for BTV detection is required as OIE-recommended primers might not be applied universally. We also found that the specificity of multiplexed PCR using the lyophilized reagents was similar to PCR using the wet reagents.
Although all the aborted samples were found negative for SBV and BTV, the sample size was too small to arrive at any conclusion. However, the capability of the multiplex PCR in the detection of multiple pathogens will help in screening further aborted content for the identification of emerging pathogens.
Several reports have been published for rapid detection and characterization of abortion-causing pathogens26,27 and of which many assays have been developed in the multiplex format22,28. In addition, several isothermal and chromatography-based assays are now available for onsite diagnosis29,30. The potential of polymerase chain reaction (PCR) for the diagnostic purpose has been shown over some time. Multiplexing for simultaneous detection and typing of many pathogens in a single reaction is one of the advantages of PCR assay. However, PCR-based assays suffer with some limitations to adopt as an on-site assay. One of the limitations is the preparation of complex PCR reaction mixture and avoiding cross- contamination while setting up the reaction. Potential application of freeze-dried based PCR reaction mixture has been demonstrated earlier for detection and serotyping of FMD virus21. Inclusion of freeze-dried positive controls removes the barrier to keep virus for positive control.
We conclude that the developed multiplex PCR assay for the detection of abortion-causing pathogens (BHV-1, BVDV, BTV, SBV and B. abortus) can be used in the field-based laboratory diagnosis as the ready-to-use, freeze-dried pre-mix for PCR-based detection is rapid, sensitive, specific, and easy to use.
ACKNOWLEDGMENTS
The authors are thankful to the Indian Council of Agricultural Research (ICAR), Government of India, and Department of Animal Husbandry, Dairying, and Fisheries for providing the necessary facility for this work. The authors are thankful to the Indian Council of Medical Research (ICMR).
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
GKS, CP, and SM designed the work. GKS, and PD conducted the experiments and wrote the manuscript. SN, VC and KPS contributed to the reagents, kits, and proofread the manuscript. SK, ARC, and SMG helped in cell culture work. All authors approved the manuscript.
FUNDING
Indian Council of Agricultural Research (ICAR) and DST-SERB (EMR/2016/006782) provided necessary funding under institute funded project. Indian Council of Medical Research (ICMR) provided a fellowship for the Ph.D. candidate.
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
AVAILABILITY OF DATA
Data generated during the study is available from the corresponding author on reasonable request.
- DAHDF. 19th livestock census report. 2012. http://dahd.nic.in/sites/default/filess/Livestock%20%205.pdf
- Shanmathy M, Gopi M, Beulah P. Contribution of Animal Husbandry to Indian Economy, Its Characteristics and Future: A Review. Asian J Agr Econ Ext Sociol. 2018;27(1):1-7.
Crossref - Anderson ML. Infectious causes of bovine abortion during mid-to late-gestation. Theriogenology. 2007;68(3):474-486.
Crossref - Nandi S, Kumar M, Yadav V, Chander V. Serological Evidences of Bovine Herpesvirus-1 Infection in Bovines of Organized Farms in India. Transbound Emerg Dis. 2011;58(2):105-109.
Crossref - Mishra N, Rajukumar K, Tiwari A, et al. Prevalence of Bovine viral diarrhoea virus (BVDV) antibodies among sheep and goats in India. Trop Anim Health Prod. 2009;41(7):1231-1239.
Crossref - Chakraborty AK, Mukherjee P, Karam A, et al. Evidence of BVDV in Pigs from North Eastern Part of India-Genetic Profiling and Characterisation. Op Virol J. 2018;12:110-120.
Crossref - Mittal M, Sharma V, Nehra K, et al. Abortions in an organized dairy farm from North India reveal the possibility of breed susceptibility to Bovine Brucellosis. One Health. 2018;5:1-5.
Crossref - Wernike K, Holsteg M, Schirrmeier H, Hoffmann B, Beer M. Natural infection of pregnant cows with Schmallenberg virus-a follow-up study. PloS One. 2014; 9(5):e98223.
Crossref - Endalew AD, Faburay B, Wilson WC, Richt JA. Schmallenberg Disease-A Newly Emerged Culicoides-Borne Viral Disease of Ruminants. Viruses. 2019;11(11):1065.
Crossref - Maclachlan NJ, Osburn BI. Teratogenic bluetongue and related orbivirus infections in pregnant ruminant livestock: timing and pathogen genetics are critical. Curr Opin Virol. 2017;27:31-35.
Crossref - Kang GJ, Gunaseelan L, Abbas KM. Epidemiological modeling of bovine brucellosis in India. Proc IEEE Int Conf Big Data. 2014;6-10.
Crossref - Sudharshana KJ, Suresh KB, Rajasekhar M. Prevalence of bovine viral diarrhoea virus antibodies in India. Revue Scientifique et Technique (International Office of Epizootics). 1999;18(3):667-671.
Crossref - Rupner RN, VinodhKumar OR, Karthikeyan R, et al. Bluetongue in India: a systematic review and meta-analysis with emphasis on diagnosis and seroprevalence. Vet Quart. 2020;40(1):229-242.
Crossref - Meyers A, Tatu U. Schmallenberg virus. Resonance. 2014;19(9):814-820.
Crossref - NIHSAD. Annual Report, 2015-16. 2016. http://nihsad.nic.in/pdf/NIHSAD%20Annual%20Report%202015-16.pdf
- Fan Q, Xie Z, Xie Z, et al. Development of a GeXP-multiplex PCR assay for the simultaneous detection and differentiation of six cattle viruses. PloS One. 2017;12(2):e0171287.
Crossref - Linhart C, Shamir R. Degenerate Primer Design: theoretical analysis and the HYDEN program. Methods Mol Biol. 2007;402:220-244.
Crossref - Spearman C. The method of right and wrong cases (constant stimuli) without Gauss’s formulae. Brit J Psychol. 1908;2(3):227-242.
Crossref - Kärber G. Beitrag zur kollektiven Behandlung pharmakologischer Reihenversuche. Naunyn-Schmiedebergs Archiv für experimentelle pathologie und pharmakologie.1931. 162(4):480-483.
- Sharma GK, Mahajan S, Das B, et al. Comparison of stabilisers for development of a lyophilised multiplex reverse-transcription PCR mixture for rapid detection of foot and mouth disease virus serotypes. Rev Sci Tech. 2014;33(3):859-867.
Crossref - Tramuta C, Lacerenza D, Zoppi S, et al. Development of a set of multiplex standard polymerase chain reaction assays for the identification of infectious agents from aborted bovine clinical samples. J Vet Diagn Invest. 2011;23(4):657-664.
Crossref - Dangler CA, de Mattos CA, de Mattos CC, Osburn BI. Identifying bluetongue virus ribonucleic acid sequences by the polymerase chain reaction. J Virol Met. 1990;28(3):281-292.
Crossref - Johnson DJ, Wilson WC, Paul PS. Validation of a reverse transcriptase multiplex PCR test for the serotype determination of US isolates of bluetongue virus. Vet Microbiol. 2000;76(2):105-115.
Crossref - Sharma GK, Subramaniam S, De A, et al. Detection of foot-and-mouth disease virus in semen of infected cattle bulls. Indian J Anim Sci. 2012. 82(12):1472-1476.
- Wolf-Jackel GA, Hansen MS, Larsen G, Holm E, Agerholm JS, Jensen TK. Diagnostic studies of abortion in Danish cattle 2015-2017. Acta Veteren Scand. 2020;62:1.
Crossref - Reichel MP, Wahl LC, Hill FI. Review of diagnostic procedures and approaches to infectious causes of reproductive failures of cattle in Australia and New Zealand. Front Vet Sci. 2018;5:222.
Crossref - Richtzenhain LJ, Cortez A, Heinemann MB, et al. A multiplex PCR for the detection of Brucella spp. and Leptospira spp. DNA from aborted bovine fetuses. Vet Microbiol. 2002;87(2):139-147.
Crossref - Moeini-Zanjani A, Pournajaf A, Ferdosi-Shahandashti E, et al. Comparison of loop-mediated isothermal amplification and conventional PCR tests for diagnosis of common Brucella species. BMC Research Notes. 2020;13:533.
Crossref - Pawar SS, Meshram CD, Singh NK, Saini M, Mishra BP, Gupta PK. Loop-Mediated Isothermal Amplification for Rapid Detection and Differentiation of Wild-Type Bovine Herpesvirus-1 and Glycoprotein E-Deleted Marker Vaccine Strain. Anim Biotechnol. 2015;26(4):268-272.
Crossref - Varela M, Pinto RM, Caporale M, et al. Mutations in the Schmallenberg Virus Gc Glycoprotein Facilitate Cellular Protein Synthesis Shutoff and Restore Pathogenicity of NSs Deletion Mutants in Mice. J Virol. 2016;90(11):5440-5450.
Crossref - Das A, Kumar B, Chakravarti S, Singh KP, Abhishek, Shrinet G. Single-tube duplex-PCR for specific detection and differentiation of Brucella abortus S19 vaccine strain from other Brucella spp. Indian J Anim Res. 2019;53(6):821-826.
Crossref
© The Author(s) 2021. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.