ISSN: 0973-7510
E-ISSN: 2581-690X
The aim of the present study is the detection of red beef meat (local and imported) contaminated by E. coli. The microbial load in local and imported beef meat was detected by counting the E. coli total number according to the universal standards. Sixty samples of local and imported beef meat from Indian origin were collected from city meat retailers and butchers that divided into 6 localities (quarters) including AL-Hadbaa, AL-Baladiat, AL-Faisalia, AL-Majmoaa, AL-Jazair and AL-Wahdaa in Mosul city. Ten samples were collected from each locality (five local samples, five imported ones). All samples were subjected for morphological, biochemical (including API20E) and confirmed by molecular identification (PCR ). Results were revealed that imported beef samples were more contaminated with E. coli bacteria than local samples since they ranged from 61 x 102 – 22 x 103 CFU /g and 13 x 102 to 65 x 102 CFU / g respectively.
PCR, E. coli, Bacterial count, Microbial contamination, Beef meat.
World is growing up fast, therefore, the demands for finding other sources were observed; to insurance the protein requirements for next decades1. The red meat consider as necessary food of human all over the world which be as one of the main sources of protein as well as lipid, vitamin, fats, mineral, and salts, also meat is one of the most valuable nutrients in human nutrition, the meat is the best media for growing deferent types of microorganisms causing fast spoilage of meat (food poisoning result from secretion of bacterial poison) leading to economic losses, the more important microorganisms which cause these loses are Escherichia coli (E. coli) and Salmonella2-4. E. coli in food is a pointer of bad sanitary situations5. Meat can become contaminated during slaughter or during processing. Infection can occur from eating contaminated undercooked meat, especially ground beef.
The Escherichia coli return to genus Escherichia which belong to family of Enterobacteriacae, these bacteria are Gram-negative bacteria, fermenting the sugar and producing acid and gas without H2S production. These bacteria have a big rate as a normal inhabitant in the gastrointestinal tract, also causes a disease to many species of animals, for example causes septicemia in lambs, horse, calf and dogs, so these bacteria consider as opportunistic bacteria6. Most important pathogens that play a role in food borne diseases are of animal origin, 2.2 million peoples each year were dead from food and water born diarrheal diseases (especially children) in developing countries7.
The aim of this study to evaluate the bacterial load on a local and imported beef meet and compare the total bacterial count of beef meat with the permissible limit of central organization and quality control.
A-Sampling
In the current study, from June to November 2018, a total of 60 samples of beef meat collected from markets of Mosul city/ Iraq, included 30 samples from imported meat (Indian origin) and 30 samples from local meat were randomly collected. Samples were collected from local markets and booths, transported by sterilized container to microbiological laboratory / College of Veterinary Medicine, University of Mosul, for examination
B-Microbiological Analysis
Culturing and Staining
All samples were cultured on the following media.
- Brain heart infusion broth, (LAB/UK)
- MacConkey agar (LAB/UK).
- Violet Red bile Glucose Agar {VRBGA (LAB/UK) (Media for enumeration of Enterobacteriacae in food}
- Eosin methylene blue (EMB, LAB/UK) for a metallic sheen phenomenon8.
- The bacterial colonies were stained by Gram’s stain9.
Biochemical Tests
Carried out according to a procedure10, 11. The biochemical tests included:
Indole production test, Methyl red, Voges proskcauer, citrate Utilization, Urease test, Hydrogen sulphide production, oxidase test, and catalase test.
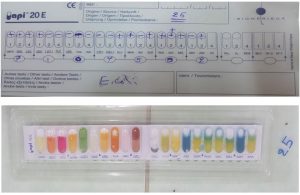
Detection of E. coli by API20 (Biomerix-France)
This method was used for the identification of the members of Enterobacteriaceae. The system has a capacity of 23 biochemical tests12,9. In this method, 20 samples (10 samples from local and 10 samples from imported beef meat) were tested.
Bacterial Counting
The total bacterial count was calculated using an aerobic plate count according to a procedure13, 25 g of sample was taken and added to 225 ml of buffer peptone water (BPW), mixed well for (3-5) min. Then decimal dilution was carried out up to the tenth dilution. 0.1 ml of each dilution was taken and placed on petri dish contained selective culture media (VRBGA, EMB) spreading well by glass spreader and incubated at 37°C for 24 hours { the medium of three Petri dishes from each diluted tube was performed (30-300 colonies)}, results of each dilution were read and fixed14. The results of bacterial counting accepted within the standard mean; when 3 of 5 samples were under the limited permissible account (5 x 102 – 5 x 103 CFU/g) 15.
DNA Extraction
Ten E. coli isolates were taken randomly from isolated bacteria, 5 isolates from local beef meat and 5 isolates from imported beef meat. Extraction was done by using a specific kit (Jena Bioscience / Germany). isolates were cultured and incubated at overnight, then extraction was done according to the kit instructions as below:
Transfer 1ml of cultured cells into a 1.5 ml microtube, centrifuged at 15, 000 g for 1 min and discard the supernatant. Then re-suspend the pellet in 300 µl of Cell Lysis Solution. Added 1.5µl of RNase A Solution and mixed by inverting. Incubated at 37°C for 15-30 min and cooled on ice for 1 min. Then added 100µl of Protein Precipitation Solution and vortex vigorously for 20- 30sec. Again, centrifuged at 15, 000 g for 5min. After that transferred the supernatant to a clean 1.5 ml microtube containing 300 µl Isopropanol (99%), mix the sample by inverting gently for 1min. Centrifuged at 15, 000 g for 1min (DNA should be visible as a small white pellet). The supernatant was discarded and tube drained briefly on clean absorbent paper. Then added 500µl Washing Buffer and inverted the tube several times to wash the DNA pellet. Final centrifugation was done at 15, 000g for 1min. After that, the ethanol was discarded carefully and let to air dry at room temperature for 10-15 min. Then added 50-100 µl of DNA Hydration Solution to the dried DNA pellet. Hydrated the DNA by incubating at 65°C for 60min. After that cooled to 4°C, and stored the sample at -80°C.
Polymerase Chain Reaction Procedure (PCR)
Polymerase chain reaction for detection of E. coli was performed according to16. Specific primers were used in this study ECO223F (5’ATCAACCGAGATTCCCCCAGT 3’) and ECO223R (5’TCACTATCGGTCAGTCAGGAG3’) (produced by IDT- USA). Amplification of DNA was performed in a total volume of 25µl containing: 12.5µl HS Prime Taq Premix (2X) (GeNetBio, Korea), 2µl of each forward and reverse primers, 6.5µl of PCR grade water and finally 2µl of DNA template. Positive and negative controls were also included in separate tubes. The mixture was subjected to amplification using thermocycler (QLS, UK) consisting of 1 cycle of initial denaturation at 95°C for 10 min, then 30 cycles of denaturation at 95°C for 30 sec, annealing at 58°C for 30sec and extension at 72°C for 1 min. The final extension was done at 72°C for 1min followed by cooling at 4°C forever. After amplification, the DNA products were analyzed by 1.5% agarose gel electrophoresis (Major Science, Taiwan) using 7 µl DNA ladder as a molecular weight marker (Gena Bioscience, Germany). The gel was visualized by UV transilluminator then photographed by Gel documentation system (Biometra, Germany).
Bacterial Isolation
Fifty-six samples were appeared positive for E. coli isolation (93.3%) according to selective media and results of biochemical tests and API20E (Fig. 1, 2), included 29 samples of local meat and 27 samples of imported beef meat.
Detection for E. coli, group of tests by (API20)
The isolates from local and imported beef meat were subject to farther more confirmative API reaction. Ten isolates from each of them gave the presumptive E. coli bacterial reaction, they are expressed in Fig. 3, 4.
Bacterial Counting
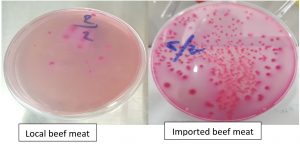
The results of bacterial count showed that 4 localities out of 6 of local beef meat (30 samples) have an acceptable limits of E. coli number (5 x 102 – 5 x 103 CFU\g), while all imported beef meat (30 samples) (India) were out of ICSMF and EEC limits since they were 61 x 102 – 22 x 103 CFU/g 15. Fig. 5, 6, Table 1.
Table (1):
Bacterial count cfu/1g in local and imported beef meat.
| Bacterial count in imported beef meat | Bacterial count in local beef meat | |||||
|---|---|---|---|---|---|---|
| Number of samples | origin | Bacterial count (cfu) | Places | Number of samples | Bacterial count (cfu) | Places |
| 1 | (India) | 120 x 102 | Jazair quarter | 1 | 81 × 102 | Jazair quarter |
| 2 | (India) | 72 x 103 | Jazair quarter | 2 | 78 × 102 | Jazair quarter |
| 3 | (India) | 67 x 102 | Jazair quarter | 3 | No growth | Jazair quarter |
| 4 | (India) | 113 x 102 | Jazair quarter | 4 | 44 × 102 | Jazair quarter |
| 5 | (India) | 80 x 102 | Jazair quarter | 5 | 122 × 102 | Jazair quarter |
| Mean | 22 x 103 | Mean | 65 X 102 | |||
| 6 | (India) | No growth | Hadbaa quarter | 6 | 84×102 | Wahdaa quarter |
| 7 | (India) | 79 x 102 | Hadbaa quarter | 7 | 80 ×102 | Wahdaa quarter |
| 8 | (India) | 47 x 102 | Hadbaa quarter | 8 | 90 × 101 | Wahdaa quarter |
| 9 | (India) | 37 x 103 | Hadbaa quarter | 9 | 62×102 | Wahdaa quarter |
| 10 | (India) | 104x 102 | Hadbaa quarter | 10 | 45× 102 | Wahdaa quarter |
| Mean | 12 x 103 | Mean | 56 X 102 | |||
| 11 | (India) | 109 x 102 | Wahdaa quarter | 11 | 13×102 | Faisalia quarter |
| 12 | (India) | 74 x 102 | Wahdaa quarter | 12 | 49 ×102 | Faisalia quarter |
| 13 | (India) | 66 x 102 | Wahdaa quarter | 13 | 57 × 102 | Faisalia quarter |
| 14 | (India) | 61 x 102 | Wahdaa quarter | 14 | 37 ×102 | Faisalia quarter |
| 15 | (India) | 95 x102 | Wahdaa quarter | 15 | 104 × 102 | Faisalia quarter |
| Mean | 81 x 102 | Mean | 52 X 102 | |||
| 16 | (India) | 44 x 102 | Faisalia quarter | 16 | 12 × 102 | Baladiat quarter |
| 17 | (India) | 62 x 102 | Faisalia quarter | 17 | 60 × 101 | Baladiat quarter |
| 18 | (India) | 63 x 102 | Faisalia quarter | 18 | 78 × 102 | Baladiat quarter |
| 19 | (India) | 80 x 102 | Faisalia quarter | 19 | 50 × 102 | Baladiat quarter |
| 20 | (India) | 91 x 102 | Faisalia quarter | 20 | 89 ×102 | Baladiat quarter |
| Mean | 68 x 102 | Mean | 47X 102 | |||
| 21 | (India) | No growth | Baladiat quarter | 21 | 27 × 102 | Hadbaa quarter |
| 22 | (India) | 93 x 102 | Baladiat quarter | 22 | 55 × 102 | Hadbaa quarter |
| 23 | (India) | 87 x102 | Baladiat quarter | 23 | 73 × 101 | Hadbaa quarter |
| 24 | (India) | 66 x 102 | Baladiat quarter | 24 | 17 × 102 | Hadbaa quarter |
| 25 | (India) | 79 x 102 | Baladiat quarter | 25 | 38 × 102 | Hadbaa quarter |
| Mean | 65 x 102 | Mean | 42 X 102 | |||
| 26 | (India) | 103 x 102 | Majmoaa quarter | 26 | 50 ×101 | Majmoaa quarter |
| 27 | (India) | 41 x 102 | Majmoaa quarter | 27 | 107 × 102 | Majmoaa quarter |
| 28 | (India) | 86 x 102 | Majmoaa quarter | 28 | 88 × 102 | Majmoaa quarter |
| 29 | (India) | 75 x 102 | Majmoaa quarter | 29 | 31 × 102 | Majmoaa quarter |
| 30 | (India) | No growth | Majmoaa quarter | 30 | 140 × 101 | Majmoaa quarter |
| Mean | 61 x 102 | Mean | 13 X 102 | |||
Polymerase Chain Reaction
The results of isolates confirmed by a polymerase chain reaction of ECO.223 gene of E.coli lane M.100 bp DNA ladder were used.Ten isolates were taken randomly (5 isolates from each local and imported beef meat), Lanes 1-5 are positive samples ( 2, 9, 13, 18, 25 ) from local beef meat. Lanes from 6-10 are positive samples (4, 10, 15, 23, 27) from imported beef meat. Lanes 11 is negative control and lanes 12 is positive control (Fig. 7).
Meat is a perfect source of protein in the human diet and it is greatly vulnerable to microbial contaminations, which can cause food poisoning and foodborne infections in human17.
Red meat production is an essential and important livestock production since the livestock production contribute about 22% agricultural production (except fish) in Iraq from 2009 to 20134.
The founding of bacteria in meat has been reported universally, most of the pathogens which play a major role in foodborne diseases have a zoonotic origin1,17. Globally, a number of studies have been conducted in this field, show special importance to bacterial contamination to red meat. The results of the current study were revealed that the imported meat was highly contaminated and unsatisfied for human consumption (61 x 102 – 22 x 103 CFU/g), while the local samples appeared more acceptable when compared with universal standards (13 x 102 to 65×102 CFU/ g). These results confirm that local meat is more fitness for consumption. The results of a local study2, were closed to the present study in term of fact that imported red meat is more polluted than local red meat and attributed to the contamination of E. coli, which may be due to a poor health condition in slaughterhouses and carcasses or during processing. Also, the present study was agreed with the results of the study that carried out by3, which revealed that imported meat was more contaminated than local meat in term of coliform bacteria and deferred from the study carried by18 in case of elevated total number of E.coli bacteria.
The current study was disagreement with the study done by19, which say that no differences in microbial load between the local and imported meat. Meat preservation is necessary during transporting for varied distances to reduce spoiling of texture, color, and nutritional value after the development and rapid growth of supermarkets1.
Also, the present findings have some agreement in general with the study done by18 in case of that the highest number of coliform bacteria.
So we were stressed that the local beef meat is more preferable to be on our tables than the imported beef meat
To control the food-borne illnesses and to keep the microbial load of raw meat in check, the food safety requirements should be followed hardly Hazard analysis critical control point17, 19.
In conclusion the current study indicated that microbial load of E. coli contamination was higher in imported meat (India) than local meat and these may be due to transportation, long period of storage with intermittent electrical cut (bad sanitary condition) and don’t follow the health condition.
ACKNOWLEDGMENTS
The author thanks college of the Veterinary Medicine, University of Mosul, Iraq for their moral support.
FUNDING
None.
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
AVAILABILITY OF DATA
All datasets created or investigated during this study are involved in the manuscript.
- Sharma KP, Chattopadhyay UK. Assessment of Microbial load of raw meat Samples sold in the Open Markets of the city of Kolkata. IOSR Journal of Agriculture and Veterinary Science (IOSR-JAVS), 2015; 8(3): 24-27.
- Abid Ali SAA, Jawd AA, Mahdi S, Mahd S. The Bacteria Contamination of Red Local and Imported Meat. Journal of Iraqi Science, 2013; 54(2): 249-254.
- Mussa AK. Study of Bacteriological Quality of Ground Beef from Different in Basrah Markets. Basrah J. Agric. Sci., 2013; 26(2): 80-88.
Crossref - Ahmad DE, Al-Hani AS. Estimating function supply for red meat in Iraq for the period 1995-2013, Iraqi Journal of Agricultural Sciences, 2016; 94(2): 9911-9059.
- Shekh CS, Deshmukh VV, Waghamare RN, Markandeya NM, Vaidya MS. Isolation of pathogenic E coli from buffalo meat sold in Parbhani city, Maharashtra, India, Vet. World, 2013; 6(5): 277-279.
Crossref - Al-Chalaby AY, pepoyan AZ. The regulatory influence of probiotics on sheep intestinal microflora. (thesis).Armenian national agrarian university, Armenia, Yerevan, 2015; 36-39.
- FAO/WHO. Food safety risk analysis. A guide for national food safety authorities, Food & Nutrition paper 87, FAO, Roma, Italy /WHO, Geneva, Switzerland; 2006; PP: 1-2.
- Brooks GF, Butel JS, Morse SA Jawetz, Melnik and Adelberges. Medical Microbiology 22nd Ed. Lange medical books Mc Graw-Hill New York-Chicago Sanfrancisco; 2001; 225-228.
- Brown AE. Gram staining (Exercise 15) in Benson, s microbiological applications-9th edition- Boston, NewYork, USA. Mc Graw-Hill. I. Companies, 2005; 103-106. Scott Roben Publisher;1488.
- Koneman EW, Allen SD, Janda WM, Schreckenbergo PC, Winn WC. Color Atlas and Textbook of Diagnostic Microbiology, 5th ed., Philadelphia, New York: J.B. Lippin, 1997.
- Mac Fadden JF. Biochemical Tests for Identification of Medical Bacteria, Philadelphia, London, New York: The Williams and Wilkins Company, Waverly, press Inc., 1979; 527.
- Quinn PJ, Carter ME, Markey B, Carter GP. Methods for identification of Bacteria in: Clinical Veterinary Microbiology Edinburg, London, Newyork: Mospy, 2004; 59-63.
- Ranjan KD. Textbook of Diagnostic Microbiology, Medical college and hospital medical publishers (p) Lid Newdulhi, 2007; 124.
- Ali RK. Detection of microbial contaminants of some locally produced fruit juices, ice creams, and yogurts, Al-Mustansiriyah Journal of Pharmaceutical Sciences (AJPS), 2013; 14(2): 124-128.
- ICMSF and EEC. Microbial standards for meat, 1986. microbial in food, v. 2. last modified, 4/2012.
- Riffon R, Sayasith K, Khalil H, Dubreuil P, Drolet M. Development of a Rapid and Sensitive Test for Identification of Major Pathogens in Bovine Mastitis by PCR. Journal of Clinical Microbiology, 2001; 39(7): 2584.
Crossref - Ahmad MUD, Sarwar A, Najeeb MI, Nawaz M, Anjum AA, Ali MA, Mansur N. Assessment of microbial load of raw meat at abattoirs and retail outlets, J. Anim. Plant Science, 2013; 23(3): 745-748.
- Al- Timimi SS, Mohammed SJ, Ali IF. The Effect of Marketing for Indian Frozen Meat in the Local Market and Its Reflection on the Microbial Load, Journal of Iraqi Science, 2014; 55(4): 1517-1527.
- Zulfikar S, Burhandin N, Abu Baker NF. Bacterial Contamination on Beef Sold at Selected Wet Markets in Selangor and Kuala Lumpur city. Journal of Agricultural Science, 2017; 9(13): 89-95. Published by Canadian Centre of Science and Education.
Crossref
© The Author(s) 2020. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.