ISSN: 0973-7510
E-ISSN: 2581-690X
Recently, a novel coronavirus (SARS-CoV-2) appeared which is conscientious for the current outbreak in China and rapidly spread worldwide. Unluckily, there is no approved vaccine found against SARS-CoV-2. Therefore, there is an urgent need for designing a suitable peptide vaccine constituent against the SARS-CoV-2. In this study, we characterized the spike glycoprotein of SARS-CoV-2 to obtain immunogenic epitopes. In addition, we used 58 SARS-CoV-2 isolates were retrieved from the Global Initiative on Sharing All Influenza Data (GISAID) and National Center for Biotechnology Information (NCBI), then aligned to obtain the conserved region of SARS-CoV-2 spike glycoprotein. The interaction between the conserved region with ACE2 receptor, a SARS-CoV-2 receptor on the host cell, has been evaluated through molecular docking approach. The B-cell epitope was identified using the immune epitope database (IEDB) web server. Interestingly, we recommend Pep_4 ADHQPQTFVNTELH as a epitope-based peptide vaccine candidate to deal with the SARS-CoV-2 outbreak. Pep_4 has a high level of immunogenicity and does not trigger autoimmune mechanisms. Pep_4 is capable of forming BCR/Fab molecular complexes with the lowest binding energy for activation of transduction signal the direct B-cell immune response. However, further study is suggested for confirmation (in vitro and in vivo).
Coronavirus, Epitope-based peptide vaccine, Immunoinformatics, SARS-CoV-2
On 30 December 2019, Chinese government reported an outbreak of pneumonia disease in Wuhan (Lam et al., 2020). Furthermore, the causative agent identified by International Committee on Taxonomy of Viruses (ICTV) and named as SARS-CoV-2 (Gorbalenya et al., 2020). The new virus has rapidly spread across China and other countries (Wang et al., 2020). Then, WHO announced a new name for the disease caused by SARS-CoV-2 as coronavirus disease 2019 (COVID-19) (Lai et al., 2020).
Up until today, there are more than three millions people are infected by SARS-CoV-2 worldwide. This data refferring to online interactive dashboard hosted by the Center for Systems Science and Engineering (CSSE) at Johns Hopkins University, Baltimore, USA. This interactive web-based dashboard is tracking the reported cases of COVID-19 in real-time (Dong et al., 2020). Presently, there are three coronaviruses to cause disease in the human, such as severe acute respiratory syndrome coronavirus (SARS-CoV), Middle-East respiratory syndrome coronavirus (MERS-CoV), and SARS-CoV-2 (Walls et al., 2020).
Coronaviruses belong to the Coronaviridae family in the Nidovirales order. In addition, the structural proteins are encoded by the four structural genes, including membrane (M), spike (S), nucleocapsid (N), and envelope (E) (Shereen et al., 2020). Interestingly, the spike glycoprotein plays an essential role in binding receptors on the host cell and the major target of neutralizing antibodies (Phan, 2020; Walls et al., 2020). Thus, the spike glycoprotein is recently considered as as the greatly expectant antigen formulation to the SARS-CoV-2 vaccine construction.
Vaccines have been proven to decrease morbidity and mortality levels of many infectious diseases (Ansori and Kharisma, 2020). In addition, Callaway (2020) stated that approximately 90 vaccines are being developed against SARS-CoV-2 by research groups in universities and companies worldwide. Furthermore, scientists are trialing diverse technologies and some of which haven’t been used in a licensed vaccine before. On the other hand, Shang et al. (2020) demonstrated that inactivated virus vaccines and live-attenuated virus vaccines have their own side effects and disadvantages. New vaccine designs based on the protective antigens derived from SARS-CoV-2 should be considered.
The development of an effective vaccine against the SARS-CoV-2 infection is urgently required. The progress in molecular biology and biotechnology is driving the construction of the novel concepts in vaccinology. Synthetic recombinant proteins containing epitopes could be produced efficiently by using modern biotechnology methods (Oyarzún and Kobe, 2016). Therefore, the epitope-based peptide vaccine is suggested as new potential candidates for effective vaccine of SARS-CoV-2. In this study, we characterized the spike glycoprotein of SARS-CoV-2 to obtain immunogenic epitopes.
Sample Retrieval from Database
The SARS-CoV-2 spike glycoprotein sequences were obtained from the Global Initiative on Sharing All Influenza Data (GSAID), Germany and National Center for Biotechnology Information (NCBI), USA.
The Screening of Conserved Domain
The conserved domain in the SARS-CoV-2 viral spike glycoprotein sequences were identified through protein alignment with Molecular Evolutionary Genetics Analysis (MEGA) X, then the 3D structure was modeled using the SWISS-MODEL (https://swissmodel.expasy.org) with homology modeling approach (Schwede et al., 2003; Adianingsih and Kharisma, 2019). Thus, the results are 3D models displayed in PyMol software (Himmah et al., 2016; Putra et al., 2020).
Immunoinformatics Prediction
This study predictions based on linear B-cell epitopes, including BepiPred, Emini Surface Accessibility, Parker Hydrophilicity, and Karplus & Schulz Flexibility on the Immune Epitope Database Analysis Resource (IEDB-AR) web server (http://tools.iedb.org/main/bcell) (Hilyatuz et al., 2016; Adianingsih and Kharisma, 2019; Ansori and Kharisma, 2020). Then we predicted the characteristics of candidate epitopes that can act as protective antigens using VaxiJen v2.0 web server (http://www.ddgpharmfac.net/vaxijen/VaxiJen/VaxiJen.html) (Hilyatuz et al., 2016; Ansori and Kharisma, 2020). Prediction of similarity with proteins in the cells of Homo sapiens is performed on vaccine candidate peptide via the Basic Local Alignment Protein Search Tool (BLASTp) (https://blast.ncbi.nlm.nih.gov/Blast.cgi). Peptides must have a low score below 20% to allow for the prevention of autoimmune (Himmah et al., 2016; Adianingsih and Kharisma, 2019).
Protein Docking
Molecular interaction simulation between the host cell receptor, angiotensin-converting enzyme 2 (ACE2), and SARS-CoV-2 spike glycoprotein during viral attachment and mechanism for binding peptide vaccine candidate with B-cell receptors (BCR), using the protein docking on the Cluspro 2.0 web server (https://cluspro.org/login.php). Docking is used to determining the binding energy of molecular complexes and biological responses (Kozakov et al., 2017; Adianingsih and Kharisma, 2019).
Conserved Identification from SARS-CoV-2 Spike Glycoprotein
A total of 58 isolates of SARS-CoV-2 spike glycoprotein sequences in the FASTA format were successfully obtained from the NCBI database with the prefixes “MN” and “EPI_ISL_” GISAID (Table 1). Then, protein alignment performed with MEGA X software to identify conserved domains until 229-mer sequences were identified
MFVFLLLSVPRDVVLTGRVLVYIRGFNCGNVYFYLFKNVKHILDFLIRLSAYVLKALTVFLNRVYNTFKSLTASVEAQATCGGLFDIYPCGNLSGPVLSFAGTNVKCTGTGTFGDTDPTCGVPQLDLTYYSYTQAYTNSPISMYECSLTNVTFQLSDFLIDRCALTLLITSLLMITYLISSNVFIVLRLSLILVGKLA DHQPQTFVNTELHLIKLQLQLLICCCGCDVH in SARS-CoV-2 spike glycoprotein as a conserved domain.
Table (1):
SARS-CoV-2 isolates from the biological databases (GISAID and NCBI).
| Sample ID | |
|---|---|
| MN908947.3 | EPI_ISL_403928 |
| MN938384.1 | EPI_ISL_402132 |
| MN975262.1 | EPI_ISL_403963 |
| MN996527.1 | EPI_ISL_403962 |
| MN996528.1 | EPI_ISL_402120 |
| MN996529.1 | EPI_ISL_402121 |
| MN996530.1 | EPI_ISL_402119 |
| MN996531.1 | EPI_ISL_406031 |
| MT007544.1 | EPI_ISL_406531 |
| MN994467.1 | EPI_ISL_406533 |
| MN994468.1 | EPI_ISL_406534 |
| MN997409.1 | EPI_ISL_406535 |
| MN988668.1 | EPI_ISL_406536 |
| MN988669.1 | EPI_ISL_406538 |
| EPI_ISL_404253 | EPI_ISL_406592 |
| EPI_ISL_404895 | EPI_ISL_406593 |
| EPI_ISL_403937 | EPI_ISL_406594 |
| EPI_ISL_403936 | EPI_ISL_406595 |
| EPI_ISL_403935 | EPI_ISL_406596 |
| EPI_ISL_403934 | EPI_ISL_406597 |
| EPI_ISL_403933 | EPI_ISL_406862 |
| EPI_ISL_403932 | EPI_ISL_406798 |
| EPI_ISL_404228 | EPI_ISL_406800 |
| EPI_ISL_404227 | EPI_ISL_406801 |
| EPI_ISL_402123 | EPI_ISL_406970 |
| EPI_ISL_403931 | EPI_ISL_406973 |
| EPI_ISL_403930 | EPI_ISL_407071 |
| EPI_ISL_403929 | EPI_ISL_407073 |
| EPI_ISL_407084 | EPI_ISL_407193 |
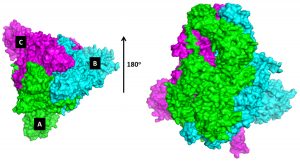
In this study, the identification of the conserved domain in 3D conducted via two stages, consisted of SARS-CoV-2 spike glycoprotein modeling and visualization in PyMol software with a representative structure (Adianingsih and Kharisma, 2019). Homology modeling on the SWISS-MODEL web server has a positive predictive score of approximately 20% (Schwede et al., 2003; Waterhouse et al., 2018). This study obtained the model of glycoprotein from a query protein sequence (MN908947.3) with a template (ID 6ACD) around 76.47% identified as a spike spike glycoprotein, so that the spike glycoprotein model was similar to the template and homologous. The 3D structure of SARS-CoV-2 glycoprotein is displayed in the form of rigid surfaces with a color selection based on its protein chains (Fig. 1).
Fig. 1. 3D model of SARS-CoV-2 spike glycoprotein. Protein structure consists of A (green), B (cyans), and C (magenta) chains.
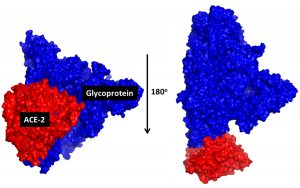
The process of docking proteins in Cluspro is to determine the binding energy that is formed between spike glycoprotein and ACE2 receptor when the SARS-CoV-2 virion attachment occurs in the host cell. The results showed that the molecular complex arrangement was identified to have the lowest binding energy of 1152.6 kcal/mol to achieve a achieve of stability (Table 2), in the glycoprotein_ACE2 binding formation. The more negative binding energy values can trigger the formation of stable molecular complexes (Kozakov et al., 2017; Adianingsih and Kharisma, 2019). This stability can trigger the activation of biological responses to target proteins such as mediated endocytosis during the SARS-CoV-2 viral entry (Cao et al., 2020). The complex of molecular docking displayed in PyMol with representative structures and color selections to identify between receptors (ACE2) and ligands (SARS-CoV-2 spike glycoprotein) (Fig. 2).
Table (2):
Binding energy of ACE-2 with SARS-CoV-2 spike glycoprotein.
Molecular Complex |
Cluster |
Member |
Lowest Energy (kcal/mol) |
|---|---|---|---|
ACE-2_Glycoprotein |
0 |
36 |
-993.2 |
ACE-2_Glycoprotein |
1 |
35 |
-1152.6 |
ACE-2_Glycoprotein |
2 |
31 |
-933.5 |
ACE-2_Glycoprotein |
3 |
29 |
-1042.0 |
ACE-2_Glycoprotein |
4 |
26 |
-912.2 |
ACE-2_Glycoprotein |
5 |
24 |
-925.0 |
Fig. 2. Molecular visualization of the ACE2 (red) molecular complex and SARS-CoV-2 spike glycoprotein (blue). Both are displayed in a rigid surface structure with specific colour selection.
The B-cell Immunogenicity Predictions of Peptide Vaccine Candidate
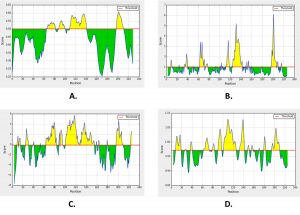
The conserved domain on SARS-CoV-2 spike glycoproteins analyzed for the potential of B-cell immunogenicity to obtain specific peptides as vaccine candidates. Prediction of B-cell immunogenicity consists of BepiPred (≥0.222), Emini Surfaces Accessibility (≥1.000), Parker Hydrophilicity (≥2.068), and Karplus & Schulz Flexibility (≥1.017) parameters, each has a threshold score for predicting B-cell epitopes in linear sequences (Sanchez-Trincado et al., 2017; Adianingsih and Kharisma, 2019; Ansori and Kharisma, 2020). B-cell epitope prediction shows on BepiPred parameters, it has 0.608 threshold points, Emini Surfaces Accessibility 1.000, Parker Hydrophilicity 6.138, and Karplus & Schulz Flexibility 1.078 (Fig. 3). The scores show positive B-cell epitopes prediction because no one has a threshold score below the standard, peptides are obtained Pep_1 FKSLTASVEAQATCG, Pep_2 CGNLSGPVLSFA, Pep_3 TGTFGDTDPT CGVPQLDLTYYSYTQAYTNSPI, and Pep_4 ADHFFQQHHF. We predicted the antigenicity of the peptides in VaxiJen v2.0, and obtained peptides number 1, 3, and 4 have antigenic properties. Furthermore, the peptide was analyzed for its similarity to the cell surface receptors of the body of Homo sapiens, via the BLASTp server. Peptides number 1, 3, and 4 did not have similarities to the cell surface receptors of the body of Homo sapiens with a score of less than 20%, so it is predicted that it will not initiate autoimmune occurrence when produced in vaccine form (Himmah et al., 2016; Adianingsih and Kharisma, 2019).
Fig. 3. The results of B-cell epitope prediction. Green areas showed negative prediction and yellow areas for positive prediction. (A) BepiPred; (B) Emini Surfaces Accessibility; (C) Parker Hydrophilicity; and (D) Karplus & Schulz Flexibility.
Molecular Interaction between Peptide-BCR/Fab
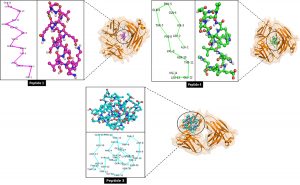
The positive predicted peptide results on B-cell immunogenicity, then the 3D structure is modeled via PEP-FOLD using the fold recognition modeling (Hilyatuz et al., 2016). Then, the structure is obtained and stored in the .pdb format, then the 3D sample of the antigen-binding fragment BCR receptor (Fab) (ID 5IFH) is obtained from the PDB database. Protein-peptide docking simulation using Cluspro to determine the binding energy is needed in the formation of stable complexes and activation of biological responses at BCR receptors (Kozakov et al., 2016; Adianingsih and Kharisma, 2019). The results show that the binding energy produced by the Pep_1-BCR/Fab molecule complex is -572.9 kcal/mol, Pep_3-BCR/Fab -661.3 kcal/mol, and Pep_4-BCR/Fab -686.5 kcal/mol, the results are displayed in PyMol software by the structural and color selection (Figure 4). The analysis showed that Pep_4 ADHQPQTFVNTELH was predicted as a SARS-CoV-2 vaccine candidate with a length of 14-mer. Pep_4 has the potential as a epitope-based peptide vaccine candidate because it has the lowest binding energy score. The lowest binding energy allows the formation of molecular complexes (Fu et al., 2018). Binding energy is influenced by weak bonds such as hydrogen and hydrophobicity (Pourseif et al., 2019). The formation of a stable molecular complex between Pep_4-BCR/Fab allows for the activation of a direct immune response by B cell (Bros et al., 2019). Specific antibodies for neutralization of SARS-CoV-2 and the role of memory cells to keep the body formed from future infections.
In conclusion, we recommend Pep_4 ADHQPQTFVNTELH as epitope-based peptide vaccine candidate to deal with the SARS-CoV-2 outbreak. Pep_4 has a high level of immunogenicity and does not trigger autoimmune mechanisms. Pep_4 is capable of forming BCR/Fab molecular complexes with the lowest binding energy for activation of transduction signal the direct B-cell immune response.
ACKNOWLEDGMENTS
This study supported by the Directorate General of Higher Education, Ministry of Education and Culture of the Republic of Indonesia and Generasi Biologi Indonesia (GENBINESIA) Foundation, Indonesia. We thank EJA – Professional Translation Services for editing the manuscript.
CONFLICT OF INTEREST
The listed authors declare no conflict of interest in any capacity, including competing or financial.
AUTHORS’ CONTRIBUTION
All listed authors have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
PMDSU Scholarship Batch III by the Directorate General of Higher Education, Ministry of Education and Culture of the Republic of Indonesia..
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
AVAILABILITY OF DATA
Not applicable.
- Adianingsih OR, Kharisma VD. Study of B cell epitope conserved region of the Zika virus envelope glycoprotein to develop multi-strain vaccine. J Appl Pharm Sci. 2019;9:98-103.
Crossref - Ansori ANM, Kharisma VD. Characterization of Newcastle disease virus in Southeast Asia and East Asia: Fusion protein gene. Eksakta. 2020;1:20-28.
- Bros M, Haas K, Moll L, Grabbe S. RhoA as a key regulator of innate and adaptive immunity. Cells. 2019;8:733.
Crossref - Callaway E. The race for coronavirus vaccines: a graphical guide. Nature. 2020;580:576-577.
Crossref - Cao Y, Li L, Feng Z, et al. Comparative genetic analysis of the novel coronavirus (2019-nCoV/SARS-CoV-2) receptor ACE2 in different populations. Cell Discov. 2020;6:11.
Crossref - Dong E, Du H, Gardner L. An interactive web-based dashboard to track COVID-19 in real time. Lancet Infect Dis. 2020:S1473-3099(20):30120-30121.
- Fu Y, Zhao J, Chen Z. Insights into the molecular mechanisms of protein-ligand interactions by molecular docking and molecular dynamics simulation: A case of oligopeptide binding protein. Comput Math Methods Med. 2018;2018:3502514.
Crossref - Gorbalenya AE, Baker SC, Baric RS, et al. The species severe acute respiratory syndrome-related coronavirus: Classifying 2019-nCoV and naming it SARS-CoV-2. Nat Microbiol. 2020;5:536-544.
Crossref - Hilyatuz Z, Ma’rup A, Tambunan USF, Parikesit AA. Immunoinformatics approach in designing epitope-based vaccine against meningitis-inducing bacteria (Streptococcus pneumoniae, Neisseria meningitidis, and Haemophilus influenzae Type B). Drug Target Insights. 2016;10:19-29.
Crossref - Himmah K, Fitriyah, Ardiyati T, Deocaris C, Widodo N. Polytope prediction for dengue vaccine candidate based on conserved envelope glycoprotein of four serotypes of Dengue virus and its antigenicity. J Pure App Chem Res. 2016;5(2):101-107.
Crossref - Kozakov D, Hall DR, Xia B, et al. The ClusPro web server for protein-protein docking. Nat Prot. 2017;12:255-278.
Crossref - Lai CC, Shih TP, Ko WC, Tang HJ, Hsueh PR. Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and coronavirus disease-2019 (COVID-19): The epidemic and the challenges. Int J Antimicrob Agents. 2020;55:105924.
Crossref - Lam TT, Shum MH, Zhu HC, et al. Identifying SARS-CoV-2 related coronaviruses in Malayan pangolins. Nature. 2020.
Crossref - Oyarzun P, Kobe B. Recombinant and epitope-based vaccines on the road to the market and implications for vaccine design and production. Hum Vaccin Immunother. 2016;12(3):763-767.
Crossref - Phan T. Genetic diversity and evolution of SARS-CoV-2. Infect Genet Evol. 2020;81:104260.
Crossref - Pourseif MM, Yousefpour M, Aminianfar M, Moghaddam MG, Nematollahi A. A multi-method and structure-based in silico vaccine designing against Echinococcus granulosus through investigating enolase protein. Bioimpacts. 2019;9:131-144.
Crossref - Putra WE, Kharisma VD, Susanto H. Potential of Zingiber officinale bioactive compounds as inhibitory agent against the IKK-B. AIP Conf Proc. 2020;2231:040048.
Crossref - Sanchez-Trincado JL, Gomez-Perosanz M, Reche PA. Fundamentals and methods for T- and B-cell epitope prediction. J Immunol Res. 2017;2017:2680160.
Crossref - Schwede T, Kopp J, Guex N, Peitsch MC. SWISS-MODEL: An automated protein homology-modeling server. Nucleic Acids Res. 2003;31:3381-3385.
Crossref - Shang W, Yang Y, Rao Y, Rao X. The outbreak of SARS-CoV-2 pneumonia calls for viral vaccines. NPJ Vaccines. 2020;5:18.
Crossref - Shereen MA, Khan S, Kazmi A, Bashir N, Siddique R. COVID-19 infection: Origin, transmission, and characteristics of human coronaviruses. J Adv Res. 2020;24:91-98.
Crossref - Walls AC, Park YJ, Tortorici MA, Wall A, McGuire AT, Veesler D. Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell. 2020;181:281-292.e6.
Crossref - Waterhouse A, Bertoni M, Bienert S, et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 2018;46(W1): W296-W303.
Crossref
© The Author(s) 2020. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.