ISSN: 0973-7510
E-ISSN: 2581-690X
Candida albicans is a yeast that is an opportunistic fungal pathogen and also identified as ubiquitous polymorphic species that is mainly linked with major fungal infections in humans, particularly in the immunocompromised patients including transplant recipients, chemotherapy patients, HIV-infected patients as well as in low-birth-weight infants. Systemic Candida infections have a high mortality rate of around 29 to 76%. For reducing its infection, limited drugs are existing such as caspofungin, fluconazole, terbinafine, and amphotericin B, etc. which contain unlikable side effects and also toxic. This review intends to utilize advanced bioinformatics technologies such as Molecular docking, Scaffold hopping, Virtual screening, Pharmacophore modeling, Molecular dynamics (MD) simulation for the development of potentially new drug candidates with a drug-repurpose approach against Candida albicans within a limited time frame and also cost reductive.
Benzofurans, Benzothiazoles, Biofilm, Inhibitors, Myristoylation, Simulation
Invasive infections caused by fungal pathogens are life threatening opportunistic infections, having a high rate of mortality and morbidity in patients. It infects billions of individuals and is responsible for 1.5-2 million deaths annually1-4. Fungal infections have risen dramatically in patients over the last decades which are immunocompromised, because of cancer chemotherapy, solid and hematologic organ transplantation, broad use of antibiotics, surgery, and long-term use of corticosteroids5. Invasive fungal infections are particularly exposed to patients receiving cancer treatment, transplant recipients, intensive care unit (ICU) care, and also with acquired immune deficiency syndrome (AIDS). The immunocompromised hosts are having a great risk of these infections with mortality rates from 20% to 40% and it continues to be high, which is relied on what kind of infecting fungal species and the clinical treatment. Several fungal species are present in the world; however, some species, including Candida, Cryptococcus, as well as Aspergillus, lead to life threatening infection in more than 90% of the population. One species of fungus well known as Candida albicans, an ascomycete, and a polymorphic fungus. It is capable of reversibly transforming to various morphologies, include (1) yeast forms, (2) pseudohyphae forms, and (3) true hyphae forms. It’s both commensal as well as opportunistic pathogen among humans and ranks as the fourth most common threat of nosocomial bloodstream infections in modern hospitals with roughly 40% death rates3, 6-17. Pathogenicity of invasive infection caused by Candida albicans is regulated by several factors namely invasive (a) filamentation, (b) biofilm development, and (c) the ability to escape from the immune system18. Studies of metabolic labeling state that Candida albicans synthesize protein N-myristoyl (20-kDa). Myristoyl-CoA: N-myristoyl transferase (NMT), was reported as a target for antifungal as well as antiviral treatment19. Antifungal drugs may be used to handle such infections; however, the mortality rates remain high around 50% and there was also a high prevalence of Invasive fungal infections. Discussing treatment options, Antifungals Azoles, echinocandins, and polyenes are existing for the curing of fungal infections which are limited and clinically available18. Azoles and polyenes target different biological fungal processes relevant to ergosterol metabolism as well as echinocandins targets cell wall β-1,3 glucan production. 5-flurocytosine is usually used as adjunctive therapy. Fazly et al. described filastatin (a small molecule) that prevents filamentation, adhesion, and virulence of Candida albicans20. Garcia et al. reported (N1-(3,5-dichlorophenyl)-5-chloro-2 hydroxybenzamide) halogenated salicylanilide and its analogs Niclosamide, an antifilament molecules that inhibited Candida albicans’ biofilm development and had similar antibiofilm and anti-filamentation activities21. Siwek et al. investigated the antifungal effect of 4-arylthiosemicarbazides and found the isoquinoline-thiosemicarbazide compound to exhibit greater affinity compared to the native ligand22. These antifungal agents have significant clinical failures such as unfavorable pharmacokinetic profiles, restricted antifungal range, significant side effects, minimal clinical effectiveness, drug-drug interactions, as well as increased drug-resistance. Therefore it is an urgent need to use all the advanced Bioinformatics tools and techniques to improve the existing fungal drugs or designing novel drugs against it. Existing drugs and the same structural analogous shows the resistant problem on the antifungal targets. Therefore, searching out the new inhibitor is the most promising approach to tackle the resistant fungal infections23-35
This review is the effort to use advanced bioinformatics techniques such as Molecular docking, Scaffold hopping, Virtual screening, Pharmacophore modeling, Molecular simulation for developing novel drug candidates with drug repurposing approach against Candida albicans within a short period, cost-reducing and solve the resistant problem in fungal infections.
Candida Albicans: Biofilm Formation
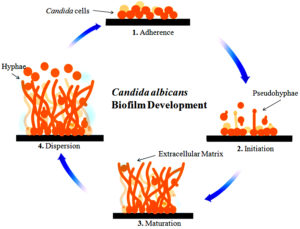
Earlier, microbiologists have studied planktonic cells which are free-floating cells in pure culture. Later they have discovered that there is a link available between sessile cells, microbial pathogenesis, and infections associated with humans and it differs basically from a planktonic cell present in the same species36. A broad variety of fungi alternately connecting planktonic cells (freely suspended cells) and multicellular populations, known as biofilms37. Biofilms are characterized as well-structured microorganism populations that are interconnected with the surface as well as enclosed by an extracellular matrix (ECM) produced by themselves38. The biofilms-associated microorganism is related with several human diseases such as cystic fibrosis, native valve endocarditis and to colonize an extensive range of medical devices which taking into consideration that these structures are very much associated with antimicrobial-resistant and it is very difficult to manage such kind of infections within the clinical setting39. A short time ago it has been understood that fungal species form biofilms and it is associated with the escalating clinical problem38,40,41. So many Candida Species have been identified but the most famous studied species is Candida albicans as a well-developed biofilm activity with the most adaptable opportunistic pathogen42. There have been so many Candida species reported, but as a well-developed biofilm operation with the most adaptable opportunistic pathogen, the most popular species studied is Candida albicans. Biofilm is developed on various medical devices such as dentures, neurosurgical shunts, speech prostheses, breast implants, prosthetic joints, endotracheal tubes, intracardiac prosthetic devices, urinary catheters, dialysis catheters for peritoneal and hemodialysis, peripheral and venous catheters43. It exists in various types, such as yeast, hyphae, and multicellular biofilm44. Candida albicans adherence and colonization to denture acrylic substrates as well as oral mucosa is the first step of pathogenesis45-48. Candida albicans’ initial attachment to the surface is limited by the pH, osmolarity, flow of the nearby medium, such as urine, antimicrobial agents, bacteria, saliva, Mucus, temperature, blood, as well as the host immune factors49-54. Candida albicans biofilm formation having different phases of development. It contains substrate adhesion, colonization, extracellular material production, and maturation53-62. Biofilm development has been shown in Fig. 1.
Fig. 1. Better describe the development of different phases such as early, intermediate and Mature. Adhesion and germination occurred in the early phase. Hyphal development as well as Extra cellular material production in the intermediate phase and the last one is maturation phase, in which dispersal occurred. In this Fig. light blue color represent ECM, circle correspond to Candida albicans cells, hyphae is also able to be seen.
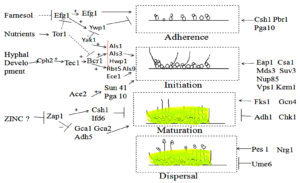
The yeast cell’s ability to shape biofilms on the implanted medical devices or on the surface in the host enhances its virulence. Candida albicans adhere to the surface with the support of Eap1p (cell wall protein) and Als3p (agglutinin like sequence protein)63,64. Als3p and Eap1p are initiations to the formation of microcolonies and further Efg1 regulatory protein is essential for the production of biofilm and its development of pseudo-and true-hyphae to form a complex association of hyphal structures with budding yeast-like cells spread throughout52,65. Further, the growth and the maturation of Biofilms, Candida albicans biofilm cells encompass a beta-glucan rich extracellular matrix that protects from environmental stresses, antimicrobial agents, and host defenses66. The existence of the hypoxic environment is correlated with the maturation of biofilm and this condition induces Tye7p-dependent up-regulation of glycolytic genes required to respond to hypoxia and prevent uncontrolled hyphal formation67. In the final step, planktonic yeast cells dispersed from the mature biofilm and established a new colony on a new surface to grow a new biofilm from Candida68. Candida albicans Biofilm formation has been presented in Fig. 2.
Fig. 2. Different genes are presented here which showed function in Biofilm formation. It has four steps (I) Adherence, (II) initiation, (III) maturation, and (IV) dispersal. In the right-hand side part of the diagram, the genes are connected and involve in pathway but in the left hand side part, the genes may not attach to an established pathway but function in a particular step. Arrows signify positive connection, the Dashed line signify repression by an indirect mechanism. “+” sign indicates that an upstream gene stimulates the expression of the downstream target and “-” sign is opposite of it. “T-shaped” indicated a negative relationship (repression by an indirect process).
The diverse transcription factors such as Efg1p, Ace2p, Zap1p, and Bcr1p are the regulator which controlled the formation of Biofilm63, 69-71. The various genes have been presented in Fig. 2, for controlling and maintaining the development of biofilm. The most important thing is to understand the mechanism of those genes so inhibition such kind of infections in the populations. The key sites of the infections are biomaterials43, wounds72, Urinary tract73,74, Gastrointestinal tract75, lower respiratory tract76, upper respiratory tract77, 78, oral cavity79, etc.
Candida albicans: NMT [N-Myristoyl-transferase] as a drug target
Post-translational modification is a very important step for proteins to function in a specific way through further modification. Post-translational modification occurs at the protein’s C- or N-terminal or on the amino acid side chain80. The modification occurs during the post-translational modification are different according to the different transformation such as C terminal amidation, N-terminal acetylation, phosphorylation of threonine, tyrosine or serine residues mainly in kinases, methylation of arginine and lysine residues mostly in histones, acylation of lysine residues and oxidation mainly in proline residues81. A less common type of post-translational modification is lipidation. Lipidation is the covalent attachment of the lipid moiety to the protein. Lipidation increases stability, membrane interaction, protein hydrophobicity, changes in conformation, trafficking, etc. Different types of lipidation are known, differing according to the group being attached and the position of attachment82. Lipidation attachment has been presented in (Table 1).
Regarding the attachment of longer chain fatty acid acylation, myristoylation (attachment of linear chain c-14), and palmitoylation (c-16)86, 87.
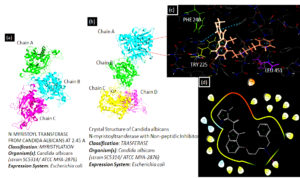
Fig. 3. PDB ID [1NMT] represents NMT from Candida albicans species at 2.45Ao and PDB ID [1IYL] represents Candida albicans NMT with Non-peptidic Inhibitor. The Ligplot interaction diagram has been generated using the Schrodinger software suit. The ligand is showing its major interaction with a certain amino acid such as PHE 240, TRY 225, and LEU 451.
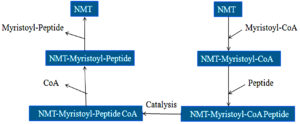
Researchers did extensive research and identified Candida albicans as an antiviral and antifungal therapy target. N-myristoyl transferase is indeed a monomeric cytosolic enzyme that is vital for the function and growth of fungi88,89. NMT is present in eukaryotes such as animals, protozoa, and fungi excluding bacteria. Protein N-myristoyl transferase is associate with the Gcn5-related N-acetyltransferases superfamily90. Candida albicans NMT contains 451 residues of amino acids and 45% of the human enzyme sequence identity. NMT is a compact globular, wedge-formed structure in which a big saddle-shaped beta-sheet present and it occupies the center of the protein structure, also, it is surrounded by several helices means consisting of an N-terminal strand, preceded by two helices, three anti-parallel beta strands, preceded by a signature (central helix) and last beta-strand91. The NMT protein structure has been illustrated in Fig. 3. C-terminal half is crucial for the peptide binding site and N-terminal half is important to form mainly Myristoyl-CoA binding site90. N-Myristoyl-transferase catalysis reaction is catalyzed by N-myristoyl-transferase, the co-translational addition of myristic acid (14-C saturated fatty acid) to the N-terminal Glycine (GLY) residue of the substrate protein via amide bonding. The N-myristoyl transferase catalysis reaction is performed by the ordered Bi-Bi reaction mechanism, the enzyme forming a high binary selectivity complex (Myristol-CoA-NMT). This binary complex is essential to the further interaction of N-Myristoyltransferase with peptide and produces a ternary complex recognized as NMT-Myristoyl-CoA-Peptide, following the catalytic transfer of myristate to the peptide substrate. The first free CoA is released, followed by the N-myristylated protein. In general, Myristoylation is irreversible as well as a significant post-translational modification is defined as N-terminal lipidation of eukaryotic and viral proteins92. Myristoylation mechanism has been shown in Fig. 4.
Myristoylation involved in anchoring and directing proteins to membranes and their effects such as signal transduction, cellular regulation, numerous pathologic processes caused by viruses, apoptosis, and translocation93, 94. The binding of myristoyl residues enables hydrophobicity to affect protein partitioning to the cell membrane and promote the interaction of protein-proteins. It is important for the overall biological expression of viral and cellular protein activity95-97. In Fungi, the myristoyl is associated with the cellular membrane and myristoyl-protein interactions. This protein takes part in protein and vesicular trafficking and signals transduction cascade. In Candida albicans with defective NMT unable to infect mice98,99. Genetic studies showed that the enzyme is important for pathogenic Candida albicans to grow vegetatively100. NMT is a good antifungal agent target because it is responsible for systemic fungal infections and lacking its expression is related to significantly decrease cell growth and increased cell death.
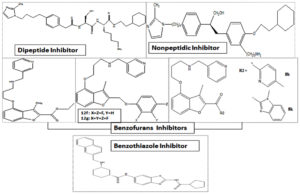
Candida albicans: NMT inhibitors
Several potent and selective inhibitors have been identified against Candida albicans NMT which showed low inhibitory activity against hNMT. All NMT polypeptides have similar folding but different inhibitor binding sites because of their particular amino acid differences90,101-104. As we studied earlier that NMT is responsible for the survival and growth of diverse fungal species, therefore so many different inhibitors have also been identified for reducing its fungal activity such as Benzofurans inhibitors101-103, Benzothiazole inhibitors104, Myristic acid analogs105,106, Peptidomimetic inhibitors88,107, p-toluene sulphonamide inhibitors108, etc. Devadas et al. reported a peptidomimetic inhibitor against Candida albicans NMT. This inhibitor was structured dependent on octapeptide substrate GLYASKLS-NH2 that was obtained from the N-terminal fragment of ARF2 (ADP ribosylation factor 2) and its analogous ALYASKLS-NH2107. Due to the lower antifungal activity of peptidomimetic inhibitors, Devadas et al. explored new forms of non-peptide inhibitors which represent simply one chiral core and demonstrate fungicidal activity109. Parang et al. have tested myristic acid analogs as putative inhibitors of NMT. Quite a lot of (+)-2-halotetradecanoic acids including (+)-2-bromotetradecanoic acid presented strong activity against Candida albicans (MIC = 39μM). These compounds illustrated antifungal action in vitro but not showed in vivo106. A new class of inhibitor was also reported named as p-toluene sulfonamides110.
In the journey of finding out the novel compounds with high selectivity, Benzofurans and Benzothiazoles are more promising than the previously reported compounds102-104. From the viewpoint of the development of antifungal drug candidates, other inhibitors were also developed such as 4-arylthiosemicarbazides derivatives22, novel benzofuran-semicarbazide hybrids and 1,3-dialkoxybenzene-semicarbazide hybrids, etc.111. Candida albicans NMT inhibitors are presented in Fig. 5.
Computational drug discovery approaches really works for finding out the novel agents as new medications. Day by Day pharmaceutical and biotech companies are growing to help the society but the major problems we have to face today is the cost of the drugs are increasing and the expenses which we have to pay for the medicine are increasing. The drug productivity measures are unable to meet the increasing demands. Thus, advanced bioinformatics tools and techniques were discussed to find out the novel antifungal agents using drug repurposing approach.
Table (1):
Representation of the structures of lipidation attachments.
Attachment to the |
Attachment |
Post translation |
|---|---|---|
N-terminus |
Myristoylation83 |
|
Cysteine |
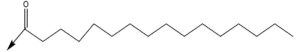 |
Palmitoylation84 |
N-terminus |
Palmitoylation85 |
|
Serine |
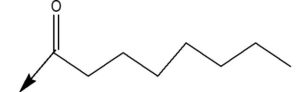 |
Octanoylation85 |
Virtual Screening and molecular docking
Protein-ligand docking is a technique commonly used to determine a drug candidate’s binding orientation to their specific target. In our survey, we are in consideration of Candida albicans NMT as a drug target for drug designing purpose. Typically molecular docking technique is performed either to searched out that how a specific ligand molecule bind to a target protein or illustrate binding interaction with the target-specific amino acid residues either H-bonding, Hydrophobic interaction, disulfide bond formation, salt bridge, pi-pi interaction or to find out the potent compound from the available databases that can bind with the target protein112-119. The docking can be categorized into two key steps, the initial positioning of the ligand structure at the active site of the target protein using the docking algorithm. followed by uses of the scoring function to assess the potency of the binding interaction.
There are a huge number of docking algorithms, tools, techniques are available to highlight the diverse orientation of the interaction between the ligand and the target structure as shown in (Table 2).
Table (2):
List of the protein-ligand docking software.
S. No. |
Docking Programs |
Conformational searching methods |
Scoring Function |
Investigated by |
|---|---|---|---|---|
1. |
AutoDock |
Genetic algorithm |
Empirical |
(Morris et al., 1998)120 |
2. |
Dock |
Incremental construction |
Force field |
(Ewing et al., 2001)121 |
3. |
FlexX |
Incremental construction |
Empirical |
(Rarey et al., 1996)122 |
4. |
Glide |
Incremental construction / Monte Carlo optimization |
Empirical |
(Friesner et al., 2004)112 |
5. |
Gold |
Genetic Algorithm |
Force field |
(Jones et al., 1997)123 |
6. |
Surflex |
Incremental construction, surface-based molecular similarity |
Empirical |
(Jain, 2003)124 |
7. |
ICM |
Monte Carlo simulation |
Force field/ Empirical |
(Abagyan et al., 1994)125 |
8. |
LigandFit |
Monte Carlo Simulation |
Empirical |
(Venkatachalam et al., 2003)126 |
9. |
eHiTS |
Exhaustive systematic |
Knowledge-based/ Empirical |
(Zsoldos et al., 2007)127 |
In the early days, the docking algorithm did not treat the protein and ligand as flexible objects, only the six translations and also the rotational degree of freedom was incorporated. Currently, we are having more consistent docking methods which give the options for flexible docking that the target protein is treated as fixed during the docking process, but the ligand is capable to move around the target.
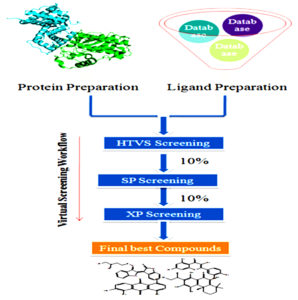
In this situation, the active site of the protein shall not be considered to undergo any significant changes in conformity with the binding of the ligand. The flexible docking is broadly used with parallel computing resources to relatively, accurately, and quickly search databases for potential ligands to a target protein. The more precise algorithms which consider both ligand and receptor flexibility are very time-consuming, therefore have not been developed extensively. The algorithm which treats flexibility of the ligand is partitioned into three categories, for example, stochastic methods, systematic methods, and simulation methods. The docked poses are ranked and assessed using docking scoring functions which estimate a ligand’s binding free energy to a receptor, which is very important to differentiate the right poses from incorrect ones. The scoring function incorporates diverse sorts of terms that express electrostatic interactions, solvation effects, non bonded interactions, and van der Waals interactions128. The structure-based virtual screening framework was presented in Fig. 6.
Pharmacophore modeling
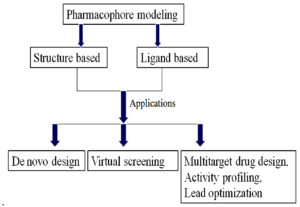
The initial idea of a pharmacophore was developed by Paul Ehrlich during the late 1800s. The theory in the past was that in a molecule there were some chemical groups or functions that were responsible for a biological effect and that certain effect molecules even had similar functions. Fig. 7, revealed the pharmacophore, with its applications.
Later in 1960, Schueler coined the term pharmacophore in his book “Chemobiodynamics and Drug Design”. It elucidated that a molecular structure that expresses the essential characteristics liable for the biological activity of the drug. The Pharmacophore has been illustrated by IUPAC since 1997129.
It projected that a pharmacophore is the collection of features known steric and electronic that are original to ensuring the mainly desirable supramolecular contacts linked with the desired protein target and blocking its biological activity130.
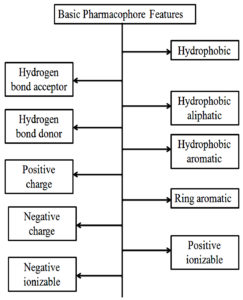
The pharmacophore reveals an abstract idea. it relies on the features shared by a group of active molecules or it is the pattern of the features of a molecule that is responsible for a biological effect. Forms of molecular features patterns are hydrogen-bond acceptors, hydrogen-bond donors, hydrophobic, anionic, cationic, aromatic plus any such type of possible combinations presented in Fig. 8.
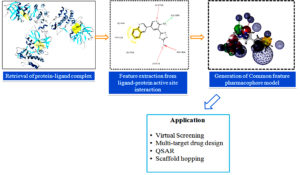
Pharmacophore modeling (Structure-based)
The structure-based approach to pharmacophore modeling describes the relevant presentation of important interactions in a protein-binding pocket. This pharmacophore modeling is appropriate in aspects of a free structure or a complex target-ligand structure. The free structure is classified as apo and the holo known as the target-ligand complex. The structural pharmacophore modeling was performed using free ligand without protein, using only protein-active site details. and when the pharmacophore modeling uses protein-ligand structure complexes utilize the possible interactions involving protein and ligand, shown in Fig. 9. Structure-based pharmacophore modeling is a very effective tool for virtual screening such as multi-target drug design, scaffold hopping, parallel screening, QSAR, and multi-target drug development131, 132.
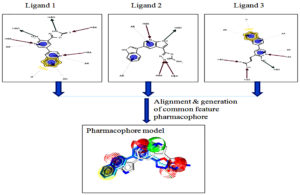
Pharmacophore modeling (Ligand-Based)
Pharmacophore modeling based on ligand structure is a powerful computational tool of great importance for helping to discover a new drug compound. It is done by extracting the important and crucial chemical features, among the set of ligands. The ligands have been divided into training and test for alignment and generating a pharmacophore model, presented in Fig. 10. This model can be utilized further for the virtual screening process for finding a similar feature molecule that behaves like a drug133, 134.
Software available for performing pharmacophore modeling
There are diverse software and tools are available to perform structure and ligand-based pharmacophore modeling, presented in (Table 3–4).
Table (3):
Structure-based pharmacophore modeling software.
S. No. |
Software |
Molecular Alignment |
Commercialization |
References |
|---|---|---|---|---|
1. |
LigandScout |
Complex-based |
Marketed by Inte: Ligand |
135 |
2. |
GBPM |
Complex-based |
Not commercialized |
136 |
3. |
Pocket v.2 |
Complex-based |
Not commercialized |
137 |
Table (4):
Ligand-based pharmacophore modeling software.
S. No. |
Software |
Molecular Alignment (methods) |
Commercialization |
Reference |
|---|---|---|---|---|
1. |
DISCO |
Bron-Kerbosh Clique detection algorithm |
Tripos Inc., Sybyl interface |
138 |
2. |
APOLLO |
Feature-based |
Not commercialized |
139 |
3. |
GALAHAD |
Atom-based |
Tripos Inc., Sybyl interface |
140 |
4. |
HipHOP |
Feature-based |
Discovery Studio (Biovia) |
138 |
5. |
MOE |
Property-based |
Chemical Computing Group |
141 |
6. |
MPHIL |
Atom-based |
Not commercialized |
142 |
7. |
HypoGen |
Feature-based |
Discovery Studio (Biovia) |
143 |
8. |
HypoRefine |
Feature-based |
Discovery Studio (Biovia) |
144 |
9. |
Apex-3D |
Feature-based |
Catalyst (Biovia) |
145 |
10. |
CLEW |
Feature-based |
Not commercialized |
146 |
11. |
GAMMA |
Atom-based |
Not commercialized |
147 |
12. |
GASP |
Atom-based |
Tripos Inc., Sybyl interface. |
138 |
13. |
PHASE |
Feature-based |
Schrodinger Inc. |
148-149 |
14. |
PharmaGist |
Feature-based |
http://bioinfo3d.cs.tau.ac.il/PharmaGist/ |
150 |
15. |
LigandScout |
Matching pattern-based alignment |
Marketed by Inte: Ligand |
151 |
Scaffold Hopping
Schneider et al. (1999) presented Scaffold hopping, in 1999. It is a method for the discovery of isofunctional molecular structures by way of considerably different molecular backbones152. Traditionally, a large fraction of the medicines produced is extracted from natural hormones, other medications, and natural products by scaffolding modification153. Recently published papers and reviewing these relevant examples provide useful guidance for a medicinal chemist for developing a new bioactive molecule. Scaffold hopping, also known as lead hopping154, 155. It is one of those approaches for finding out the new lead candidates156. Scaffold hopping intends to discover a structurally novel substance structure starting from previously identified active compounds by altering the center core structure of the molecule157. Scaffold hopping is commonly used in lead optimization. Since using HTS, so many compounds are unsuccessful compounds with poor PK properties and poor physiochemical properties. To overcome this, side-chain modification is sufficient sometimes, the core structure of the parent molecule or the scaffold may often be changed152,158-160.
Why Scaffold hopping is so important
- Central scaffolds are also specifically involved in target protein interactions. An enhanced binding affinity can result from a change in the scaffold.
- Replacing a lipophilic scaffold by an extra polar one could enhance the solubility of the compounds (lipophilic compound soluble in fats, oils, lipids etc. for increasing solubility, lipophilic scaffold can be change by an extra polar side chain or fragment).
- Replacing a very flexible scaffold known as peptide backbone by an inflexible central scaffold would also considerably advance the binding affinity and on the total DMPK characteristics.
- Changes in the core of the structure may lead to a patentable novel compound.
- Replacing a metabolically labile scaffold via a reduced amount of toxic, and an additional stable one will improve pharmacokinetic properties.
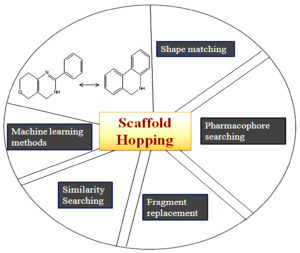
Insilico approaches for scaffold hopping
There are different approaches are available for scaffold hopping but the main idea behind is (1) matching of shape, (2) searching for pharmacophores, (3) replacement of fragments, (4) looking for similarities, and (5) machine learning, etc.
The Shape matching approach describes if the compounds are structurally related means display similar biological activities and if the compounds are more distantly related, the less probable to show the same biological effects. In shape matching, it is possible to find out the compounds which mimic accurately this structure, but it is not possible to get the same features that are significant for binding to the target structure161,162. If the ligands are structurally different however, can adopt similar shape and share common features, It is possible to derive 3D pharmacophores, which is used for shape matching scaffold hopping.
But the drawback is for searching out the 3D pharmacophore using different chemical structure databases is not sufficient to find out the novel scaffold because it searches from the known compound databases163-166. Another approach to scaffold hopping is fragment replacement, in which no need for the replacement of the entire compounds but searching for a replacement of fragment of an active compound167-169. Similarity searching is also used for scaffold hopping. The chemical structures are assembled in these algorithms using fragment joining as well as the novel scaffolds are resolved by their match to the query170-172. Also, machine learning methods are used for scaffolds hopping, methods together with self-organizing maps that allow compound distributions to be visualized173, 174. The computational methods of scaffold hopping are shown in Fig. 11. The software tools is listed out in (Table 5).
Table (5):
List of Software and tools for scaffold hopping.
S. No. |
Software & tools |
Applications |
References |
|---|---|---|---|
1. |
1-Click Scaffold Hop |
It is ready to use drug discovery platform for scaffold hopping |
175 |
2. |
Spark™ |
Spark works in Shape space and electrostatic so it can go with the nature of reference molecules |
176 |
3. |
Core Hopping |
The core-hopping technique is to test several possible scaffolds (also known as protocores) against a template and look for alignments of possible attachment points on the scaffold with the attachment points on the template. |
177 |
4. |
LigCSRre |
LigCSRre is a modern effective and standardized method for 3D matching screening of tiny compounds, the modular plan of which opens the door to lots of improvements. |
178 |
5. |
e-LEA3D |
The approach is perfectly appropriate for scaffold-hopping, this section moreover permits a search for potential substitutes to a selected scaffold. |
179 |
6. |
ChemMapper |
ChemMapper using the user given the chemical structure of the molecules as the query, the highest alike structure in respect of 3D similarity is sent back using related pharmacology annotations. |
180 |
7. |
SHOP |
It is a grid-based technique for Scaffold bouncing. In a database, scaffolds were predictable utilizing 3 types of 3D-descriptors. |
181 |
8. |
LeadGrow+ |
Creating a molecular library for efficient scaffold hopping. |
182 |
9. |
Recore |
Recore is a rapid and flexible scaffold hopping method based on conformations of small molecule crystal structures. |
183 |
10. |
HTSFPs |
(HTSFPs), It is a method that matches patterns of actions in investigational screens. |
184 |
11. |
MORPH |
MORPH is a software tool for scaffold hopping which can scientifically alter aromatic rings in molecular 3-dimensional models exclusive change of the non-hydrogen atom co-ordinates in the rings. |
185 |
Molecular Simulations And Advancement
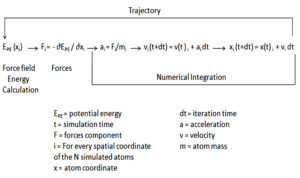
Since the first protein (folded globular protein) MD simulation is discussed in 1977186. In December 1999, IBM declared a five-year intend to build up a massively parallel computer for studying biomolecular phenomena, in which they have discussed for increasing longer simulation time as well as developing computing software and hardware for bimolecular MD simulation187. Certain software packages were also developed for simultaneously scales well-organized MD simulation very well on machines188. A massively parallel machine such as Anton was introduced which was able to reach millisecond time-scale simulation for biomolecular systems189. In recent times graphical processing unit which is known as GPU achieved remarkable progress with high-performance computing capability for MD simulations190,191. currently, MD simulation is very important for studying protein, DNA, and RNA systems. In the MD simulation so many terms are used in which force field is very significant where a protein force field included bonded (bond angle, dihedral angles, bond length) and also non-bonded interactions (electrostatic, van der Waals). The development of improved sampling methods and escalating computational performance came with more inaccuracies in the protein force field192. In these aspects, the classical protein field has been improved with Gromacs188, AMBER193, CHARMM194, and NAMD195. A basic algorithm for MD simulation has been represented Fig. 12 and a list of software has been shown in (Table 6). apart from this MD simulation software, there are other software also available such as Desmond196, TINKER, DL_POLY197, ESPResSo198, etc. so that it is understood that MD is already an important tool in serving to understand biology.
Table (6):
List of major Software and tools for MD Simulation.
About |
GROMACS |
AMBER |
CHARMM |
NAMD |
|---|---|---|---|---|
Developer (s) |
Martin Karplus, Accelrys |
Peter Kollman’s, research group (at first Developed) at the University of California. |
Martin Karplus, Accelrys |
The University of Illinois at Urbana – Champaign |
Initial release |
1991; 29 years ago |
2002; 18 years ago |
1983; 37 years ago |
1995; 25 years prior |
Stable release |
2018.4/12 November 2018; 16 months prior |
Amber 18, Amber Tools 19/ April 26, 2019; 10 months ago |
c40b1, c40b2 / 2015; 5 years ago |
2.12 / December 22, 2016; 3 years ago |
Written in |
C++, C |
C, C++, Fortran 95 |
FORTRAN 77-95, CUDA |
C++ |
Operating system |
Linux, Windows macOS, some other Unix variety |
Windows, OS X, Linux, Unix, CNK |
Unix-like: Linux, macOS, AIX, iOS |
Cross-platform: Windows, Linux, macOS, Unix |
Platform |
Many |
x86, Nvidia GPUs, Blue Gene |
x86, ARM, Nvidia GPU; Cray XT4, XT5 |
x86, x86-64 |
Available in |
English |
English |
English |
English |
License |
LGPL variants >= 4.6, GPL variants
| Amber: Proprietary Amber Tools: public domain , GPL, other open-source |
Proprietary |
Proprietary, freeware for noncommercial use |
Website Sources |
www.gromacs.org |
ambermd.org |
www.charmm.org |
www.ks.uiuc.edu/Research/namd |
The anticipated outcome could be development and searching out the novel, specific inhibitors for Candida albicans MyristoylCoA: Protein N-Myristoyltransferase as anti-fungal agents using advanced computational approaches. A humble beginning made towards this end needs patronage for further development. All the more interesting on this aspect is, still as on date, no successful attempt has been made towards development of a best, specific inhibitor for Candida albicans MyristoylCoA: Protein N-Myristoyltransferase which again augments support.
N-myristoyl transferase is a monomeric cytosolic protein that is vital for the function and growth of fungi. There are so many inhibitors that have been designed against Candida albicans NMT for reducing fungal infections in humans but at present antifungal drugs are not perfect in the expressions of the antifungal spectrum, efficacy, and protection. Drug repurposing is one of the most significant, more affordable, and increasingly proficient techniques in drug discovery. So right now, we have examined in silico drug repurposing approach which joins Molecular docking, Virtual Screening, Pharmacophore demonstrating, Scaffold hopping, and Molecular dynamics (MD) simulation for the advancement of a novel Candida albicans NMT inhibitors.
ACKNOWLEDGMENTS
The Authors would like to offer their sincere thanks to ICMR (Indian Council of Medical Research), New Delhi, Government of India, for financial support related help For the present research and Department of Biotechnology (DBT) New Delhi for offering help as Bioinformatics infrastructure facility. The authors also would like to acknowledge the Schrödinger group for giving software support.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors have made substantial, direct, and intellectual contribution to the work and approve it for publication.
FUNDING
This work was supported by ICMR (Indian Council of Medical Research) Grant number ISRM/11(46)/2019, New Delhi, and Government of India.
ETHICS STATEMENT
Not applicable.
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in this manuscript.
- Brown GD, Denning DW, Gow NAR, Levitz SM, Netea MG, White TC. Hidden Killers: Human Fungal Infections. Sci Transl Med. 2012;4(165).
Crossref - Brown GD, Denning DW, Levitz SM. Tackling Human Fungal Infections. Science. 2012;336(6082):647.
Crossref - Pfaller M, Diekema D. Epidemiology of invasive candidiasis: a persistent public health problem. Clin Microbiol Rev. 2007;20(1):133-163.
Crossref - Negri M, Martins M, Henriques M, Svidzinski TI, Azeredo J, Oliveira R. Examination of potential virulence factors of Candida tropicalis clinical isolates from hospitalized patients. Mycopathologia. 2010;169(3):175-182.
Crossref - Roemer T, Krysan DJ. Antifungal drug development: challenges, unmet clinical needs, and new approaches. Cold Spring Harb Perspect Med. 2014;4(5).
Crossref - Perlroth J, Choi B, Spellberg B. Nosocomial fungal infections: epidemiology, diagnosis, and treatment. Med Mycol. 2007;45(4):321-346.
Crossref - Rueping MJ, Vehreschild JJ, Cornely OA. Invasive candidiasis and candidemia: from current opinions to future perspectives. Expert Opin Investig Drugs. 2009;18(6):735-748.
Crossref - Jacobsen ID, Wilson D, Wachtler B, Brunke S, Naglik JR, Hube B. Candida albicans dimorphism as a therapeutic target. Expert Rev Anti Infect Ther. 2012;10(1):85-93.
Crossref - Borg-von Zepelin M, Kunz L, Ruchel R, Reichard U, Weig M, Gross U. Epidemiology and antifungal susceptibilities of Candida spp. to six antifungal agents: results from a surveillance study on fungaemia in Germany from July 2004 to August 2005. J Antimicrob Chemother. 2007;60(2):424-428.
Crossref - Panizo MM, Reviakina V, Dolande M, Selgrad S. Candida spp. in vitro susceptibility profile to four antifungal agents. Resistance surveillance study in Venezuelan strains. Med Mycol. 2009;47(2):137-143.
Crossref - Richardson M, Lass-Florl C. Changing epidemiology of systemic fungal infections. Clin Microbiol Infect. 2008;14 Suppl 4:5-24.
Crossref - Pappas PG, Kauffman CA, Andes D, et al. Clinical practice guidelines for the management of candidiasis: 2009 update by the Infectious Diseases Society of America. Clin Infect Dis. 2009;48(5):503-535.
Crossref - Pfaller MA, Diekema DJ, Jones RN, et al. International surveillance of bloodstream infections due to Candida species: frequency of occurrence and in vitro susceptibilities to fluconazole, ravuconazole, and voriconazole of isolates collected from 1997 through 1999 in the SENTRY antimicrobial surveillance program. J Clin Microbiol. 2001;39(9):3254-3259.
Crossref - Wisplinghoff H, Bischoff T, Tallent SM, Seifert H, Wenzel RP, Edmond MB. Nosocomial bloodstream infections in US hospitals: analysis of 24,179 cases from a prospective nationwide surveillance study. Clin Infect Dis.2004;39(3):309-317.
Crossref - Mean M, Marchetti O, Calandra T. Bench-to-bedside review: Candida infections in the intensive care unit. Crit Care. 2008;12(1):204.
Crossref - Sheehan DJ, Hitchcock CA, Sibley CM. Current and emerging azole antifungal agents. Clin Microbiol Rev. 1999;12(1):40-79.
Crossref - Schiaffella F, Macchiarulo A, Milanese L, et al. Design, synthesis, and microbiological evaluation of new Candida albicans CYP51 inhibitors. J Med Chem. 2005;48(24):7658-7666.
Crossref - Shapiro RS, Robbins N, Cowen LE. Regulatory circuitry governing fungal development, drug resistance, and disease. Microbiol Mol Biol Rev. 2011;75(2):213-267.
Crossref - Wiegand RC, Carr C, Minnerly JC, et al. The Candida albicans myristoyl-CoA:protein N-myristoyltransferase gene. Isolation and expression in Saccharomyces cerevisiae and Escherichia coli. J Biol Chem. 1992;267(12):8591-8598.
Crossref - Fazly A, Jain C, Dehner AC, et al. Chemical screening identifies filastatin, a small molecule inhibitor of Candida albicans adhesion, morphogenesis, and pathogenesis. Proc Natl Acad Sci USA. 2013;110(33):13594-13599.
Crossref - Garcia C, Burgain A, Chaillot J, Pic E, Khemiri I, Sellam A. A phenotypic small-molecule screen identifies halogenated salicylanilides as inhibitors of fungal morphogenesis, biofilm formation and host cell invasion. Sci Rep. 2018;8(1):11559.
Crossref - Siwek A, Stefanska J, Dzitko K, Ruszczak A. Antifungal effect of 4-arylthiosemicarbazides against Candida species. Search for molecular basis of antifungal activity of thiosemicarbazide derivatives. J Mol Model. 2012;18(9):4159-4170.
Crossref - Paiva JA, Pereira JM. New antifungal antibiotics. Curr Opin Infect Dis. 2013;26(2):168-174.
Crossref - Perfect JR. The antifungal pipeline: a reality check. Nat Rev Drug Discov. 2017;16(9):603-616.
Crossref - Wurtele H, Tsao S, Lepine G, et al. Modulation of histone H3 lysine 56 acetylation as an antifungal therapeutic strategy. Nat Med. 2010;16(7):774-780.
Crossref - Raj S, Krishnan K, Askew DS, et al. The Toxicity of a Novel Antifungal Compound Is Modulated by Endoplasmic Reticulum-Associated Protein Degradation Components. Antimicrob Agents Chemother. 2015;60(3):1438-1449.
Crossref - Alksne LE, Projan SJ. Bacterial virulence as a target for antimicrobial chemotherapy. Curr Opin Biotechnol. 2000;11(6):625-636.
Crossref - Gauwerky K, Borelli C, Korting HC. Targeting virulence: a new paradigm for antifungals. Drug Discov Today. 2009;14(3-4):214-222.
Crossref - Kim K, Zilbermintz L, Martchenko M. Repurposing FDA approved drugs against the human fungal pathogen, Candida albicans. Ann Clin Microbiol Antimicrob. 2015;14:32.
Crossref - Enoch DA, Ludlam HA, Brown NM. Invasive fungal infections: a review of epidemiology and management options. J Med Microbiol. 2006;55(Pt 7):809-818.
Crossref - Blot S, Vandewoude K. Management of invasive candidiasis in critically ill patients. Drugs. 2004;64(19):2159-2175.
Crossref - Arendrup MC, Fuursted K, Gahrn-Hansen B, et al. Seminational surveillance of fungemia in Denmark: notably high rates of fungemia and numbers of isolates with reduced azole susceptibility. J Clin Microbiol. 2005;43(9):4434-4440.
Crossref - Hobson R. The global epidemiology of invasive Candida infections-is the tide turning? J Hosp Infect. 2003;55(3):159-168.
Crossref - Verweij PE, Snelders E, Kema GH, Mellado E, Melchers WJ. Azole resistance in Aspergillus fumigatus: a side-effect of environmental fungicide use? Lancet Infect Dis. 2009;9(12):789-795.
Crossref - Groll AH, De Lucca AJ, Walsh TJ. Emerging targets for the development of novel antifungal therapeutics. Trends Microbiol. 1998;6(3):117-124.
Crossref - Costerton JW, Lewandowski Z, Caldwell DE, Korber DR, Lappin-Scott HM. Microbial biofilms. Annu Rev Microbiol. 1995;49:711-745.
Crossref - Donlan RM. Biofilms: microbial life on surfaces. Emerg Infect Dis. 2002;8(9):881-890.
Crossref - Ramage G, Mowat E, Jones B, Williams C, Lopez- Ribot J. Our current understanding of fungal biofilms. Crit Rev Microbiol. 2009;35(4):340-355.
Crossref - Donlan RM, Costerton JW. Biofilms: survival mechanisms of clinically relevant microorganisms. Clin Microbiol Rev. 2002;15(2):167-193.
Crossref - Sayed SI, Datta S, Deore N, Kazi RA, Jagade MV. Prevention of voice prosthesis biofilms: current scenario and future trends in prolonging prosthesis lifetime. J Indian Med Assoc. 2012;110(3):175-180.
- Fanning S, Mitchell AP. Fungal biofilms. PLoS Pathog. 2012;8(4):e1002585.
Crossref - Kabir MA, Hussain MA, Ahmad Z. Candida albicans: a model organism for studying fungal pathogens. ISRN Microbiol. 2012;2012:538694.
Crossref - Ramage G, Martinez JP, Lopez-Ribot JL. Candida biofilms on implanted biomaterials: a clinically significant problem. FEMS Yeast Res. 2006;6(7):979- 986.
Crossref - Noble SM, Gianetti BA, Witchley JN. Candida albicans cell-type switching and functional plasticity in the mammalian host. Nat Rev Microbiol. 2017;15(2):96-108.
Crossref - Ellepola AN, Samaranayake LP. Adhesion of oral Candida albicans isolates to denture acrylic following limited exposure to antifungal agents. Arch Oral Biol. 1998;43(12):999-1007.
Crossref - Holmes AR, Bandara BM, Cannon RD. Saliva promotes Candida albicans adherence to human epithelial cells. J Dent Res. 2002;81(1):28-32.
Crossref - Radford DR, Challacombe SJ, Walter JD. Denture plaque and adherence of Candida albicans to denture- base materials in vivo and in vitro. Crit Rev Oral Biol Med. 1999;10(1):99-116.
Crossref - Williams DW, Kuriyama T, Silva S, Malic S, Lewis MA. Candida biofilms and oral candidosis: treatment and prevention. Periodontol 2000. 2011;55(1):250-265.
Crossref - Hawser S, Baillie G, Douglas LJ. Production of extracellular matrix by Candida albicans biofilms. J Med Microbiol. 1998;47(3):253-256.
Crossref - Baillie GS, Douglas LJ. Matrix polymers of Candida biofilms and their possible role in biofilm resistance to antifungal agents. J Antimicrob Chemother. 2000;46(3):397-403.
Crossref - Richard ML, Nobile CJ, Bruno VM, Mitchell AP. Candida albicans biofilm-defective mutants. Eukaryot Cell. 2005;4(8):1493-1502.
Crossref - Ramage G, VandeWalle K, Lopez-Ribot JL, Wickes BL. The filamentation pathway controlled by the Efg1 regulator protein is required for normal biofilm formation and development in Candida albicans. FEMS Microbiol Lett. 2002;214(1):95-100.
Crossref - Ramage G, Wickes BL, Lopez-Ribot JL. A seed and feed model for the formation of Candida albicans biofilms under flow conditions using an improved modified Robbins device. Rev Iberoam Micol. 2008;25(1):37-40.
Crossref - Chandra J, Mukherjee PK, Leidich SD, et al. Antifungal resistance of candidal biofilms formed on denture acrylic in vitro. J Dent Res. 2001;80(3):903-908.
Crossref - Chandra J, Kuhn DM, Mukherjee PK, Hoyer LL, McCormick T, Ghannoum MA. Biofilm formation by the fungal pathogen Candida albicans: development, architecture, and drug resistance. J Bacteriol. 2001;183(18):5385-5394.
Crossref - Al-Fattani MA, Douglas LJ. Penetration of Candida biofilms by antifungal agents. Antimicrob Agents Chemother. 2004;48(9):3291-3297.
Crossref - Baillie GS, Douglas LJ. Effect of growth rate on resistance of Candida albicans biofilms to antifungal agents. Antimicrob Agents Chemother. 1998;42(8):1900-1905.
Crossref - Garcia-Sanchez S, Aubert S, Iraqui I, Janbon G, Ghigo JM, d’Enfert C. Candida albicans biofilms: a developmental state associated with specific and stable gene expression patterns. Eukaryot Cell. 2004;3(2):536-545.
Crossref - Nikawa H, Jin C, Makihira S, Egusa H, Hamada T, Kumagai H. Biofilm formation of Candida albicans on the surfaces of deteriorated soft denture lining materials caused by denture cleansers in vitro. J Oral Rehabil. 2003;30(3):243-250.
Crossref - Ramage G, Vande Walle K, Wickes BL, Lopez-Ribot JL. Standardized method for in vitro antifungal susceptibility testing of Candida albicans biofilms. Antimicrob Agents Chemother. 2001;45(9):2475-2479.
Crossref - Blankenship JR, Mitchell AP. How to build a biofilm: a fungal perspective. Curr Opin Microbiol. 2006;9(6):588-594.
Crossref - Ramage G, Vandewalle K, Wickes BL, Lopez-Ribot JL. Characteristics of biofilm formation by Candida albicans. Rev Iberoam Micol. 2001;18(4):163-170.
- Zhao X, Daniels KJ, Oh SH, et al. Candida albicans Als3p is required for wild-type biofilm formation on silicone elastomer surfaces. Microbiology (Reading). 2006;152(Pt 8):2287-99.
Crossref - Li F, Svarovsky MJ, Karlsson AJ, et al. Eap1p, an adhesin that mediates Candida albicans biofilm formation in vitro and in vivo. Eukaryot Cell. 2007;6(6):931-939.
Crossref - Mukaremera L, Lee KK, Mora-Montes HM, Gow NAR. Candida albicans Yeast, Pseudohyphal, and Hyphal Morphogenesis Differentially Affects Immune Recognition. Front Immunol. 2017;8:629.
Crossref - Nett JE, Sanchez H, Cain MT, Andes DR. Genetic basis of Candida biofilm resistance due to drug-sequestering matrix glucan. J Infect Dis. 2010;202(1):171-175.
Crossref - Bonhomme J, Chauvel M, Goyard S, Roux P, Rossignol T, d’Enfert C. Contribution of the glycolytic flux and hypoxia adaptation to efficient biofilm formation by Candida albicans. Mol Microbiol. 2011;80(4):995-1013.
Crossref - Uppuluri P, Chaturvedi AK, Srinivasan A, et al. Dispersion as an important step in the Candida albicans biofilm developmental cycle. PLoS Pathog. 2010;6(3):e1000828.
Crossref - Finkel JS, Mitchell AP. Genetic control of Candida albicans biofilm development. Nat Rev Microbiol. 2011;9(2):109-118.
Crossref - Nobile CJ, Mitchell AP. Genetics and genomics of Candida albicans biofilm formation. Cell Microbiol. 2006;8(9):1382-1391.
Crossref - Fanning S, Xu W, Solis N, Woolford CA, Filler SG, Mitchell AP. Divergent targets of Candida albicans biofilm regulator Bcr1 in vitro and in vivo. Eukaryot Cell. 2012;11(7):896-904.
Crossref - Seth AK, Geringer MR, Hong SJ, Leung KP, Mustoe TA, Galiano RD. In vivo modeling of biofilm-infected wounds: a review. J Surg Res. 2012;178(1):330-338.
Crossref - Kauffman CA, Vazquez JA, Sobel JD, et al. Prospective multicenter surveillance study of funguria in hospitalized patients. Clin Infect Dis. 2000;30(1):
14-18.
Crossref - Wenastein RA, Lundstrom T, Sobel J. Nosocomial candiduria: a review. Clin Infect Dis. 2001;32(11):1602-1607.
Crossref - Damman CJ, Miller SI, Surawicz CM, Zisman TL. The microbiome and inflammatory bowel disease: is there a therapeutic role for fecal microbiota transplantation? Am J Gastroenterol. 2012;107(10):1452-1459.
Crossref - Cugini C, Calfee MW, Farrow JM III, Morales DK, Pesci EC, Hogan DA. Farnesol, a common sesquiterpene, inhibits PQS production in Pseudomonas aeruginosa. Mol Microbiol. 2007;65(4):896-906.
Crossref - Buijssen KJ, van der Laan BF, van der Mei HC, et al. Composition and architecture of biofilms on used voice prostheses. Head Neck. 2012;34(6):863-871.
Crossref - Ell SR. Candida’the cancer of silastic’. J Laryngol Otol. 1996;110(3):240-242.
Crossref - Jakubovics NS. Talk of the town: interspecies communication in oral biofilms. Mol Oral Microbiol. 2010;25(1):4-14.
Crossref - Uy R, Wold F. Posttranslational covalent modification of proteins. Science. 1977;198(4320):890-896.
Crossref - Green KD, Garneau-Tsodikova S. Posttranslational Modification of Proteins. 2010;433-468.
Crossref - Chen B, Sun Y, Niu J, Jarugumilli GK, Wu X. Protein Lipidation in Cell Signaling and Diseases: Function, Regulation, and Therapeutic Opportunities. Cell Chem Biol. 2018;25(7):817-831.
Crossref - Wright MH, Heal WP, Mann DJ, Tate EW. Protein myristoylation in health and disease. J Chem Biol. 2010;3(1):19-35.
Crossref - Aicart-Ramos C, Valero RA, Rodriguez-Crespo I. Protein palmitoylation and subcellular trafficking. Biochim Biophys Acta. 2011;1808(12):2981-2994.
Crossref - Chang S-C, Magee AI. Acyltransferases for secreted signalling proteins. Mol Membr Biol. 2009;26(1- 2):104-113.
Crossref - Magee AI, Courtneidge SA. Two classes of fatty acid acylated proteins exist in eukaryotic cells. EMBO J. 1985;4(5):1137-1144.
- McIlhinney RA, Pelly SJ, Chadwick JK, Cowley GP. Studies on the attachment of myristic and palmitic acid to cell proteins in human squamous carcinoma cell lines: evidence for two pathways. EMBO J. 1985;4(5):1145-1152.
Crossref - Devadas B, Freeman SK, Zupec ME, et al. Design and synthesis of novel imidazole-substituted dipeptide amides as potent and selective inhibitors of Candida albicans myristoylCoA:protein N-myristoyltransferase and identification of related tripeptide inhibitors with mechanism-based antifungal activity. J Med Chem. 1997;40(16):2609-2625.
Crossref - Rocque WJ, McWherter CA, Wood DC, Gordon JI. A comparative analysis of the kinetic mechanism and peptide substrate specificity of human and Saccharomyces cerevisiae myristoyl-CoA:protein N- myristoyltransferase. J Biol Chem. 1993;268(14):9964-9971.
Crossref - Vetting MW, LP SdC, Yu M, et al. Structure and functions of the GNAT superfamily of acetyltransferases. Arch Biochem Biophys. 2005;433(1):212-226.
Crossref - Sogabe S, Masubuchi M, Sakata K, et al. Crystal structures of Candida albicans N-myristoyltransferase with two distinct inhibitors. Chem Biol. 2002;9(10):1119-1128.
Crossref - Rudnick DA, McWherter CA, Rocque WJ, Lennon PJ, Getman DP, Gordon JI. Kinetic and structural evidence for a sequential ordered Bi Bi mechanism of catalysis by Saccharomyces cerevisiae myristoyl-CoA:protein N- myristoyltransferase. J Biol Chem. 1991;266(15):9732-9739.
Crossref - Ducker CE, Upson JJ, French KJ, Smith CD. Two N- myristoyltransferase isozymes play unique roles in protein myristoylation, proliferation, and apoptosis. Mol Cancer Res. 2005;3(8):463-476.
Crossref - Zha J, Weiler S, Oh KJ, Wei MC, Korsmeyer SJ. Posttranslational N-myristoylation of BID as a molecular switch for targeting mitochondria and apoptosis. Science. 2000;290(5497):1761-1765.
Crossref - Peitzsch RM, McLaughlin S. Binding of acylated peptides and fatty acids to phospholipid vesicles: pertinence to myristoylated proteins. Biochemistry. 1993;32(39):10436-10443.
Crossref - Gordon JI, Duronio RJ, Rudnick DA, Adams SP, Gokel GW. Protein N-myristoylation. J Biol Chem. 1991;266(14):8647-8650.
Crossref - Farazi TA, Waksman G, Gordon JI. The biology and enzymology of protein N-myristoylation. J Biol Chem. 2001;276(43):39501-39504.
Crossref - Lodge JK, Johnson RL, Weinberg RA, Gordon JI. Comparison of myristoyl-CoA:protein N- myristoyltransferases from three pathogenic fungi: Cryptococcus neoformans, Histoplasma capsulatum, and Candida albicans. J Biol Chem. 1994;269(4):2996-3009.
Crossref - Nakayama H, Mio T, Nagahashi S, Kokado M, Arisawa M, Aoki Y. Tetracycline-regulatable system to tightly control gene expression in the pathogenic fungus Candida albicans. Infect Immun. 2000;68(12):6712-6719.
Crossref - Weinberg RA, McWherter CA, Freeman SK, Wood DC, Gordon JI, Lee SC. Genetic studies reveal that myristoylCoA:protein N-myristoyltransferase is an essential enzyme in Candida albicans. Mol Microbiol. 1995;16(2):241-250.
Crossref - Masubuchi M, Kawasaki K, Ebiike H, et al. Design and synthesis of novel benzofurans as a new class of antifungal agents targeting fungal N- myristoyltransferase. Part 1. Bioorg Med Chem Lett. 2001;11(14):1833-1837.
Crossref - Ebiike H, Masubuchi M, Liu P, et al. Design and synthesis of novel benzofurans as a new class of antifungal agents targeting fungal N- myristoyltransferase. Part 2. Bioorg Med Chem Lett. 2002;12(4):607-610.
Crossref - Kawasaki K, Masubuchi M, Morikami K, et al. Design and synthesis of novel benzofurans as a new class of antifungal agents targeting fungal N- myristoyltransferase. Part 3. Bioorg Med Chem Lett. 2003;13(1):87-91.
Crossref - Ebara S, Naito H, Nakazawa K, Ishii F, Nakamura M. FTR1335 is a novel synthetic inhibitor of Candida albicans N-myristoyltransferase with fungicidal activity. Biol Pharm Bull. 2005;28(4):591-595.
Crossref - Paige LA, Zheng GQ, DeFrees SA, Cassady JM, Geahlen RL. Metabolic activation of 2-substituted derivatives of myristic acid to form potent inhibitors of myristoyl CoA:protein N-myristoyltransferase. Biochemistry. 1990;29(46):10566-10573.
Crossref - Parang K, Knaus EE, Wiebe LI, Sardari S, Daneshtalab M, Csizmadia F. Synthesis and antifungal activities of myristic acid analogs. Archiv der Pharmazie. 1996;329(11):475-482.
Crossref - Devadas B, Zupec ME, Freeman SK, et al. Design and syntheses of potent and selective dipeptide inhibitors of Candida albicans myristoyl-CoA:protein N- myristoyltransferase. J Med Chem. 1995;38(11):1837-1840.
Crossref - Pathare SS, Bhansali SG, Mahadik KR, Kulkarni VM. Pharmacophore modeling and atom-based 3d-qsar studies of antifungal benzofurans. Int J Pharm Pharm Sci. 2015;7(3):453-458.
- Devadas B, Freeman SK, McWherter CA, et al. Novel biologically active nonpeptidic inhibitors of myristoyl CoA: protein N-myristoyltransferase. J Med Chem. 1998;41(6):996-1000.
Crossref - Karki RG, Kulkarni VM. A feature based pharmacophore for Candida albicans MyristoylCoA: protein N-myristoyltransferase inhibitors. Eur J Med Chem. 2001;36(2):147-163.
Crossref - Xu H, Hou Z, Liang Z, Guo MB, Su X, Guo C. Design, Synthesis and Antifungal Activity of Benzofuran and Its Analogues. Chin J Chem. 2019;37(12):1245-1250.
Crossref - Friesner RA, Banks JL, Murphy RB, et al. Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J Med Chem. 2004;47(7):1739-1749.
Crossref - Halgren TA, Murphy RB, Friesner RA, et al. Glide: a new approach for rapid, accurate docking and scoring. 2. Enrichment factors in database screening. J Med Chem. 2004;47(7):1750-1759.
Crossref - Friesner RA, Murphy RB, Repasky MP, et al. Extra precision glide: Docking and scoring incorporating a model of hydrophobic enclosure for protein− ligand complexes. J Med Chem. 2006;49(21):6177-6196.
Crossref - Kellenberger E, Rodrigo J, Muller P, Rognan D. Comparative evaluation of eight docking tools for docking and virtual screening accuracy. Proteins. 2004;57(2):225-242.
Crossref - Rashid M, Husain A, Shaharyar M, Mishra R, Hussain A, Afzal O. Design and synthesis of pyrimidine molecules endowed with thiazolidin-4-one as new anticancer agents. Eur J Med Chem. 2014;83:630-645.
Crossref - Hussain A, Verma CK. Molecular docking and in silico ADMET study reveals 3-(5-{[4-(aminomethyl) piperidin-1-yl] methyl}-1h-indol-2-yl)-1h-indazole-6-carbonitrile as a potential inhibitor of cancer Osaka thyroid kinase. Biomed. Res. 2017;28(13):5805-5815.
- Hussain A, Verma CK, Chouhan U. Identification of novel inhibitors against Cyclin Dependent Kinase 9/Cyclin T1 complex as: Anti cancer agent. Saudi J Biol Sci. 2017;24(6):1229-1242.
Crossref - Hussain A, Verma CK. Recognition of New Inhibitor of CDK9/Cyclin T1 Complex as Persuasive Anticancer Agent. IJABR. 2016;7(2):654-668.
- Morris GM, Goodsell DS, Halliday RS, et al. Automated docking using a Lamarckian genetic algorithm and an empirical binding free energy function. J Comput Chem. 1998;19(14):1639-1662.
Crossref - Ewing TJ, Makino S, Skillman AG, Kuntz ID. DOCK 4.0: search strategies for automated molecular docking of flexible molecule databases. J Comput Aided Mol Des.2001;15(5):411-428.
Crossref - Rarey M, Kramer B, Lengauer T, Klebe G. A fast flexible docking method using an incremental construction algorithm. J Mol Biol. 1996;261(3):470-489.
Crossref - Jones G, Willett P, Glen RC, Leach AR, Taylor R. Development and validation of a genetic algorithm for flexible docking. J Mol Biol. 1997;267(3):727-748.
Crossref - Jain AN. Surflex: fully automatic flexible molecular docking using a molecular similarity-based search engine. J Med Chem. 2003;46(4):499-511.
Crossref - Abagyan R, Totrov M, Kuznetsov D. ICM-a new method for protein modeling and design: applications to docking and structure prediction from the distorted native conformation. J Comput Chem. 1994;15(5):488-506.
Crossref - Venkatachalam CM, Jiang X, Oldfield T, Waldman M. LigandFit: a novel method for the shape-directed rapid docking of ligands to protein active sites. J Mol Graph Model. 2003;21(4):289-307.
Crossref - Zsoldos Z, Reid D, Simon A, Sadjad SB, Johnson AP. eHiTS: a new fast, exhaustive flexible ligand docking system. J Mol Graph Model. 2007;26(1):198-212.
Crossref - Kitchen DB, Decornez H, Furr JR, Bajorath J. Docking and scoring in virtual screening for drug discovery: methods and applications. Nat Rev Drug Discov. 2004;3(11):935-949.
Crossref - Wermuth C, Ganellin C, Lindberg P, Mitscher L. Glossary of terms used in medicinal chemistry (IUPAC Recommendations 1998). Pure Appl Chem. 1998;70(5):1129-1143.
Crossref - Qing X, Lee XY, De Raeymaecker J, et al. Pharmacophore modeling: advances, limitations, and current utility in drug discovery. J Receptor Ligand Channel Res. 2014;7:81-92.
Crossref - Pirhadi S, Shiri F, Ghasemi JB. Methods and applications of structure based pharmacophores in drug discovery. Curr Top Med Chem. 2013;13(9):1036-1047.
Crossref - Hussain A, Verma CK. Ligand-and structure-based pharmacophore modeling, docking study reveals 2-[[4- [6-(isopropylamino) pyrimidin-4-yl]-1H-pyrrolo [2, 3- b] pyridin-6-yl] amino] ethanol as a potential anticancer agent of CDK9/cyclin T1 kinase. J Cancer Res Ther. 2019;15(5):1131-1140.
Crossref - Vuorinen A, Engeli R, Meyer A, et al. Ligand-based pharmacophore modeling and virtual screening for the discovery of novel 17β-hydroxysteroid dehydrogenase 2 inhibitors. J Med Chem. 2014;57(14):5995-6007.
Crossref - Hussain A, Verma CK. A Combination of Pharmacophore Modeling, Molecular Docking and Virtual Screening Study Reveals 3, 5, 7-Trihydroxy-2- (3, 4, 5-trihydroxyphenyl)-4H-Chromen-4-One as a Potential Anti-Cancer Agent of COT Kinase. Indian J Pharm Educ Res. 2018;52(4):699-706.
Crossref - Wolber G, Langer T. LigandScout: 3-D pharmacophores derived from protein-bound ligands and their use as virtual screening filters. J Chem Inf Model. 2005;45(1):160-169.
Crossref - Ortuso F, Langer T, Alcaro S. GBPM: GRID-based pharmacophore model: concept and application studies to protein-protein recognition. Bioinformatics. 2006;22(12):1449-1455.
Crossref - Chen J, Lai L. Pocket v. 2: further developments on receptor-based pharmacophore modeling. J Chem Inf Model. 2006;46(6):2684-2691.
Crossref - Patel Y, Gillet VJ, Bravi G, Leach AR. A comparison of the pharmacophore identification programs: Catalyst, DISCO and GASP. J Comput Aided Mol Des. 2002;16(8-9):653-681.
Crossref - Jansen JM, Copinga S, Gruppen G, Molinari EJ, Dubocovich ML, Grol CJ. The high affinity melatonin binding site probed with conformationally restricted ligands-I. Pharmacophore and minireceptor models. Bioorg Med Chem. 1996;4(8):1321-1332.
Crossref - Richmond NJ, Abrams CA, Wolohan PR, Abrahamian E, Willett P, Clark RD. GALAHAD: 1. Pharmacophore identification by hypermolecular alignment of ligands in 3D. J Comput Aided Mol Des. 2006;20(9):567-587.
Crossref - Tian Y-S, Kawashita N, Arai Y, Okamoto K, Takagi T. Pharmacophore Modeling and Molecular Docking Studies of potential inhibitors to E6 PBM-PDZ from Human Papilloma Virus (HPV). Bioinformation. 2015;11(8):401-6.
Crossref - Holliday JD, Willett P. Using a genetic algorithm to identify common structural features in sets of ligands. J Mol Graph Model. 1997;15(4):221-232.
Crossref - Guner O, Clement O, Kurogi Y. Pharmacophore modeling and three dimensional database searching for drug design using catalyst: recent advances. Curr Med Chem. 2004;11(22):2991-3005.
Crossref - Schuster D, Laggner C, Steindl TM, Palusczak A, Hartmann RW, Langer T. Pharmacophore modeling and in silico screening for new P450 19 (aromatase) inhibitors. J Chem Inf Model. 2006;46(3):1301-1311.
Crossref - Hariprasad V, Kulkarni VM. A proposed common spatial pharmacophore and the corresponding active conformations of some peptide leukotriene receptor antagonists. J Comput Aided Mol Des. 1996;10(4):284- 292.
Crossref - Dolata D, Parrill A, Walters W. CLEW: the generation of pharmacophore hypotheses through machine learning. SAR QSAR Environ Res. 1998;9(1-2):53-81.
Crossref - Handschuh S, Wagener M, Gasteiger J. Superposition of three-dimensional chemical structures allowing for conformational flexibility by a hybrid method. J Chem Inf Comput Sci. 1998;38(2):220-232.
Crossref - Dixon SL, Smondyrev AM, Rao SN. PHASE: a novel approach to pharmacophore modeling and 3D database searching. Chem Biol Drug Des. 2006;67(5):370-372.
Crossref - Dixon SL, Smondyrev AM, Knoll EH, Rao SN, Shaw DE, Friesner RA. PHASE: a new engine for pharmacophore perception, 3D QSAR model development, and 3D database screening: 1. Methodology and preliminary results. J Comput Aided Mol Des. 2006;20(10-11):647-671.
Crossref - Schneidman-Duhovny D, Dror O, Inbar Y, Nussinov R, Wolfson HJ. PharmaGist: a webserver for ligand-based pharmacophore detection. Nucleic Acids Res. 2008;36(suppl_2):W223-W228.
Crossref - Prasad NK, Kanakaveti V, Eadlapalli S, Vadde R, Meetei AP, Vindal V. Ligand-based pharmacophore modeling and virtual screening of RAD9 inhibitors. J Chem. 2013;2013:679459.
Crossref - Schneider G, Neidhart W, Giller T, Schmid G. Scaffold-Hopping” by Topological Pharmacophore Search: A Contribution to Virtual Screening. Angew Chem Int Ed Engl. 1999;38(19):2894- 2896.
Crossref - Sneader W. Drug prototypes and their exploitation. Eur J Med Chem. 1997;32(1):91.
Crossref - Cramer RD, Jilek RJ, Guessregen S, Clark SJ, Wendt B, Clark RD. “Lead hopping”. Validation of topomer similarity as a superior predictor of similar biological activities. J Med Chem. 2004;47(27):6777-6791.
Crossref - Martin YC, Muchmore S. Beyond QSAR: lead hopping to different structures. QSAR Comb Sci. 2009;28(8):797-801.
Crossref - Schneider G, Schneider P, Renner S. Scaffold-hopping: how far can you jump? QSAR Comb Sci. 2006;25(12):1162-1171.
Crossref - Bohm HJ, Flohr A, Stahl M. Scaffold hopping. Drug Discov Today Technol. 2004;1(3):217-224.
Crossref - Brown N, Jacoby E. On scaffolds and hopping in medicinal chemistry. Mini Rev Med Chem. 2006;6(11):1217-29.
Crossref - Mauser H, Guba W. Recent developments in de novo design and scaffold hopping. Curr Opin Drug Discov Devel. 2008;11(3):365-374.
- Cramer RD, Poss MA, Hermsmeier MA, Caulfield TJ, Kowala MC, Valentine MT. Prospective identification of biologically active structures by topomer shape similarity searching. J Med Chem. 1999;42(19):3919-3933.
Crossref - Johnson MA, Maggiora GM. Concepts and applications of molecular similarity. J Comput Chem. 1990;13(4):539-540.
Crossref - Bohm HJ, Klebe G. What can we learn from molecular recognition in protein-ligand complexes for the design of new drugs? Angew Chem Int Ed Engl. 1996;35(22):2588-2614.
Crossref - Good AC, Mason JS. Three-dimensional structure database searches. Rev Comput Chem. 1996:67-117.
Crossref - Van Drie JH. Strategies for the determination of pharmacophoric 3D database queries. J Comput Aided Mol Des. 1997;11(1):39-52.
Crossref - Kaminski JJ, Rane D, Snow ME, et al. Identification of novel farnesyl protein transferase inhibitors using three- dimensional database searching methods. J Med Chem. 1997;40(25):4103-4112.
Crossref - Barreca ML, Gitto R, Quartarone S, De Luca L, De Sarro G, Chimirri A. Pharmacophore modeling as an efficient tool in the discovery of novel noncompetitive AMPA receptor antagonists. J Chem Inf Comput Sci. 2003;43(2):651-655.
Crossref - Lauri G, Bartlett PA. CAVEAT: a program to facilitate the design of organic molecules. J Comput Aided Mol Des. 1994;8(1):51-66.
Crossref - Pierce AC, Rao G, Bemis GW. BREED: Generating novel inhibitors through hybridization of known ligands. Application to CDK2, p38, and HIV protease. J Med Chem. 2004;47(11):2768- 2775.
Crossref - Lewell XQ, Jones AC, Bruce CL, et al. Drug rings database with web interface. A tool for identifying alternative chemical rings in lead discovery programs. J Med Chem. 2003;46(15):3257- 3274.
Crossref - Rarey M, Dixon JS. Feature trees: a new molecular similarity measure based on tree matching. J Comput Aided Mol Des. 1998;12(5):471-490.
Crossref - Sheridan RP, Kearsley SK. Why do we need so many chemical similarity search methods? Drug Discov Today. 2002;7(17):903-911.
Crossref - Lengauer T, Lemmen C, Rarey M, Zimmermann M. Novel technologies for virtual screening. Drug Discov Today. 2004;9(1):27-34.
Crossref - Geppert H, Vogt M, Bajorath J. Current trends in ligand-based virtual screening: molecular representations, data mining methods, new application areas, and performance evaluation. J Chem Inf Model. 2010;50(2):205-216.
Crossref - Schneider P, Tanrikulu Y, Schneider G. Self-organizing maps in drug discovery: compound library design, scaffold-hopping, repurposing. Curr Med Chem. 2009;16(3):258-266.
Crossref - Yuan S, Chan HS, Hu Z. Implementing WebGL and HTML5 in macromolecular visualization and modern computer-aided drug design. Trends Biotechnol. 2017;35(6):559-571.
Crossref - Floresta G, Rescifina A, Marrazzo A, et al. Hyphenated 3D-QSAR statistical model-scaffold hopping analysis for the identification of potentially potent and selective sigma-2 receptor ligands. Eur J Med Chem. 2017;139:884-891.
Crossref - Wang X-J, Zhang J, Wang S-Q, Xu W-R, Cheng X-C, Wang R-L. Identification of novel multitargeted PPARα/γ/δ pan agonists by core hopping of rosiglitazone. Drug Des Devel Ther. 2014;8:2255-2262.
Crossref - Quintus F, Sperandio O, Grynberg J, Petitjean M, Tuffery P. Ligand scaffold hopping combining 3D maximal substructure search and molecular similarity. BMC Bioinformatics. 2009;10(1):245.
Crossref - Douguet D. e-LEA3D: a computational-aided drug design web server. Nucleic Acids Res. 2010;38(Web Server issue):W615-21.
Crossref - Gong J, Cai C, Liu X, et al. ChemMapper: a versatile web server for exploring pharmacology and chemical structure association based on molecular 3D similarity method. Bioinformatics. 2013;29(14):1827-1829.
Crossref - Ahlstrom MM, Ridderstrom M, Luthman K, Zamora I. Virtual screening and scaffold hopping based on GRID molecular interaction fields. J Chem Inf Model. 2005;45(5):1313-1323.
Crossref - Gurusamy M, Abdul JF. Lead Optimization Studies Towards Finding NS2B/NS3 Protease Target-specific Inhibitors as Potential Anti-dengue Drug-like Compounds. Curr Drug Discov Technol. 2019;16(3):307-314.
Crossref - Maass P, Schulz-Gasch T, Stahl M, Rarey M. Recore: a fast and versatile method for scaffold hopping based on small molecule crystal structure conformations. J Chem Inf Model. 2007;47(2):390-399.
Crossref - Wassermann AM, Lounkine E, Urban L, et al. A screening pattern recognition method finds new and divergent targets for drugs and natural products. ACS Chem Biol. 2014;9(7):1622-1631.
Crossref - Beno BR, Langley DR. MORPH: a new tool for ligand design. J Chem Inf Model. 2010;50(6):1159-1164.
Crossref - McCammon JA, Gelin BR, Karplus M. Dynamics of folded proteins. Nature. 1977;267(5612):585-590.
Crossref - Allen F, Almasi G, Andreoni W, et al. Blue Gene: A vision for protein science using a petaflop supercomputer. IBM Syst J. 2001;40(2):310-327.
Crossref - Hess B, Kutzner C, Van Der Spoel D, Lindahl E. GROMACS 4: algorithms for highly efficient, load- balanced, and scalable molecular simulation. J Chem Theory Comput. 2008;4(3):435-447.
Crossref - Shaw DE, Deneroff MM, Dror RO, et al. Anton, a special-purpose machine for molecular dynamics simulation. Commun ACM. 2008;51(7):91-97.
Crossref - Liu W, Schmidt B, Voss G, Muller-Wittig W. Accelerating molecular dynamics simulations using Graphics Processing Units with CUDA. Comput Phys Commun. 2008;179(9):634-641.
Crossref - Harvey M, De Fabritiis G. A survey of computational molecular science using graphics processing units. Wiley Interdiscip Rev Comput Mol Sci. 2012;2(5):734-742.
Crossref - Monticelli L, Tieleman DP. Force fields for classical molecular dynamics. Biomolecular simulations. Springer. 2013;924:197-213.
Crossref - Case DA, Cheatham III TE, Darden T, et al. The Amber biomolecular simulation programs. J Comput Chem. 2005;26(16):1668-1688.
Crossref - Brooks BR, Brooks III CL, Mackerell Jr AD, et al. CHARMM: the biomolecular simulation program. J Comput Chem. 2009;30(10):1545-1614.
Crossref - Nelson MT, Humphrey W, Gursoy A, et al. NAMD: a parallel, object-oriented molecular dynamics program. The International Journal of Supercomputer Applications and High Performance Computing. 1996;10(4):251-268.
Crossref - Chow E, Rendleman CA, Bowers KJ, et al. Desmond performance on a cluster of multicore processors. DE Shaw Research Technical Report DESRES/TR–2008- 01. 2008.
- Yan T, Burnham CJ, Del Popolo MG, Voth GA. Molecular dynamics simulation of ionic liquids: The effect of electronic polarizability. J Phys Chem B. 2004;108(32):11877-11881.
Crossref - Limbach H-J, Arnold A, Mann BA, Holm C. ESPResSo-an extensible simulation package for research on soft matter systems. Comput Phys Commun. 2006;174(9):704-727.
Crossref
© The Author(s) 2021. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.