ISSN: 0973-7510
E-ISSN: 2581-690X
SARS-CoV-2 is continually evolving with the emergence of new variants with increased viral pathogenicity. The emergence of heavily mutated Omicron (B.1.1.529) with spike protein mutations are known to mediate its higher transmissibility and immune escape that has brought newer challenges for global public health to contain SARS-CoV-2 infection. One has to come up with a therapeutic strategy against the virus so as to effectively contain the infection and spread. Natural phytochemicals are being considered a significant source of bioactive compounds possessing an antiviral therapeutic potential. Being a promising anticancer and chemo-preventive agent, Silybin holds a significant potential to be used as a therapeutic. In the present study, molecular docking of Silybin with Omicron spike protein (7QNW) was carried out. Molecular docking results showed greater stability of Silybin in the active site of the Omicron spike protein with suitable binding mode of interactions. The study reveals that Silybin has the potential to block the host ACE2 receptor-viral spike protein binding; thereby inhibiting the viral entry to human cells. Therefore, Silybin may be further developed as a medication with the ability to effectively combat SARS-CoV-2 Omicron.
SARS CoV-2 Omicron, Spike Protein, Phytochemicals, Computational Approach, Antiviral Drugs
The COVID-19 pandemic is a serious global menace that has been affecting the human population. Acute respiratory distress and infection caused by COVID-19, which is thought to have begun in Wuhan, China, in December 2019 and has since spread around the world, is the current leading cause of morbidity and mortality globally.1,2 Infectious disease known as COVID-19 is caused by a newly described virus in the β-coronavirus genus of the coronaviridae family. This virus had phylogenic similarity with severe acute respiratory syndrome coronavirus (SARS-CoV), but later renamed as SARS-CoV-2.3 Some mutations in SARS-CoV-2 have led to the emergence of new viral subtypes with unique properties. Vaccines and other pharmaceutical formulations often allow “variants of concern” (VOCs) or “variants of interest” (VOIs) to escape.4 Moreover, they are more transmissible, and may lead to further severity of the disease.5 In addition to being classified a new VOC by the World Health Organization on November 26, 2021, Omicron was also identified as a variant of SARS-CoV-2 called B.1.1.529.6,7
Omicron SARS CoV-2 known to be originated from South Africa, is a highly divergent variant of the virus with more than 60 mutations.8 Amongst all the SARS-CoV-2 variants, Omicron has been seen to have many mutation prone sites with more than 50% mutations were detected in the spike protein regions.9,10 There are 32 mutations in spike protein which have been confirmed to be involved in transmissibility, immune evasion and disease severity. Moreover, 15 out of 32 mutations have been identified at the spike protein receptor binding region, responsible for making the virus sensitive to neutralizing antibodies.11,12 These mutations may allow the immune response to be bypassed, which could lead to multiple infections leading to several breakthrough infections or re-occurrence with mutant virus strains.8,13 Because of this, the scientific community and pharmaceutical industry have been tirelessly searching for new medications that can effectively combat the infection.14 One effective method of preventing viral infection is to block the process by which virus particles connect to host cells. As the virus spike glycoprotein is a major virulent factor for facilitating viral entry inside the human cells, it is considered as one of the primary drug targets to neutralise the variant of the virus.3,15
Natural products containing phytochemicals have been consistently being considered as primary source of drug development against the virus.16,17 Many natural plant-based phytochemicals including alkaloids, flavonoids, and other compounds, have been reported to be active against SARS-CoV-2.18,19 In vitro and even in vivo studies have revealed that flavonoids have the anti-viral potential as well.20 Silybin, a component of flavonolignan extract isolated from Silybum marianum, has been shown to have hepato-protective effects. Moreover, it has been reported to be a potent anticancer and chemo-preventive agent.21 Silybin was also reported to inhibit Hepatitis C and Influenza A virus infection displaying its antiviral potential.22 Molecular dynamics (MD) simulations are an effective approach in molecular docking for virtual screening of target-ligand binding ability prediction.23,24 Silybin was molecularly docked with SARS-CoV-2 Omicron spike glycoprotein in the current investigation. The objective of the study was to discover Silybin as a potent natural compound that targets omicron spike protein to hinder its interaction with the cellular receptor of the host.
Receptor preparation
The protein responsible for the virus pathogenicity especially the Omicron variant spike glycoprotein which facilitates the viral entry into the host cell, was selected as the target. The 3-D crystal structure of the target spike glycoprotein was retrieved from the Research Collaboratory Structural Bioinformatics-Protein Data Bank (RCSB-PDB) with PDB ID: 7QNW.25,26 The water molecules and heteroatoms were excluded in order to clean the model.
Docking via Auto dock
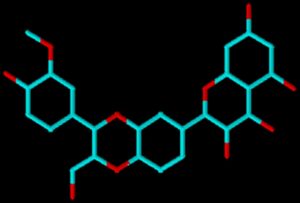
We have selected Silybin as the lead molecule (Figure 1). The docking studies of the ligand design and its fitting in the binding pocket were performed using Autodock tools. The same is designed to check out the binding of drug candidates to a receptor of a known 3D structure.27,28 Visualization studies at every level have been performed using Autodock, PYMOL and SPDBV tools.29,30 It was found that the designed probable best ligand/compound has shown the best binding affinity of the ligand, i.e. ΔG-8.2Kcal/mole around the binding cleft, which proves that the more negative energy corresponds to the better ligand features (Table). Based on our experimental calculations on ligand binding free energy, favoured interactions with protein atoms, the results of the study are consistent with those found in the existing literature.31
Table:
Ligands run with Autodock, showing Binding Energies with RMSD values after docking.
Ligand Mode |
Binding Affinity |
Rmsd/ub |
Rmsd/lb |
|---|---|---|---|
1 |
-7.6 |
12.785 |
9.204 |
2 |
-7.4 |
14.706 |
7.302 |
3 |
-7.3 |
10.423 |
5.437 |
4 |
-7.3 |
11.211 |
7.323 |
5 |
-7.2 |
8.156 |
6.144 |
6 |
-7.1 |
9.9 |
1.935 |
7 |
-7.1 |
11.562 |
5.918 |
8 |
-7.1 |
12.276 |
6.164 |
9 |
-8.2 |
16.988 |
14.213 |
MD simulation Studies
For MD simulations, GROMACS 2020.2 was used on the 7QNW-Silybin complex with the highest docking score, which demonstrates an exceptionally strong binding affinity. The purpose of this study was to learn more about the structural variations that occur in a fast-paced environment on a time scale of 10 ns.32 Interaction energy, free energy of solvation (DGsolv), root mean square fluctuation (RMSF), solvent accessible surface area (SASA), radius of gyration (Rg), and root mean square deviation (RMSD) values as a function of time, and average angle and angle distribution of ligand in the active site of the receptor protein were used as the basis for the MD simulations. The RMSF, SASA values, radius of gyration, and RMSD obtained were in agreement with the previous studies.33
SARS-CoV-2 is continuously evolving into newer variants with its Omicron variant carrying an unusual number of mutations, mainly on the spike protein which could lead to altered transmissibility and immune evasion, imposing a newer challenge for the humanity.34 Spike glycoprotein serves as the major virulence factor that facilitates virus entry into human cells (attached to the cellular receptor ACE2) and mediates its infectious cycle.23 Therefore, the spike glycoprotein was looked at as a promising therapeutic target to prevent human viral entry and transmission. We adopted the in-silico approach as computational virtual screening has been very cost-effective and takes relatively lesser time to reach to the market.18,35
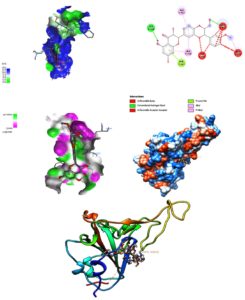
In the quest to produce effective antiviral drugs, natural substances are widely regarded as the most important resource due to their wide availability and low risk of side effects. Phytochemicals like flavonoids abundantly present in our daily diet, have been extensively studied as a possible therapeutic alternative against viruses.20 Silybin (a flavonolignan extract component) has been reported as a potent anticancer and chemo-preventive agent.21 The antiviral activities of Silybin showed its potential to be used as a broad-spectrum antiviral therapeutic.22 Therefore, the molecular docking of Silybin with the target protein was carried out using the Lamarckian Genetic algorithm and Local Search default parameters.24 Developed ligands displayed significant binding as evident from the Gibbs free energy values having lower RMSD values in comparison to the original conformation. Moreover, IC50 values (i.e. activity concentration) of analyzed ligands was also reported to be highly satisfactory consistent with earlier reports in literature (Table) (Figure 2).
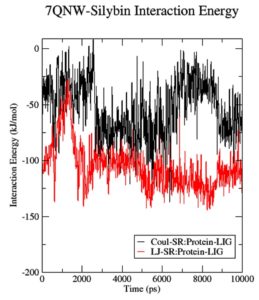
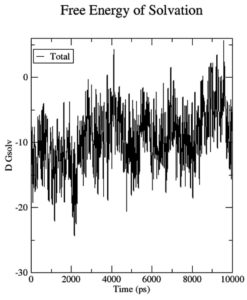
Initial fluctuations in the simulations led to an average Coulomb’s short-range (Coul-SR) value of -53.32 KJ/mol for the complex 7QNW-Silybin. As a whole, the complex has a Lennard-Jones short-range (LJ-SR) value of -105.69 KJ/mol. LJ-SR value was reportedly higher at the start of the simulation and got stable to -100 KJ/mol after two nanosecond simulations, indicating that the 7QNW-Silybin interaction got stable as the simulation proceeded (Figure 3). With an average solvation-free energy of -10 DGsolv, the complex has not changed (Figure 4). To insert a solute molecule into the solvent at constant temperature and pressure, the energy value obtained was reportedly significant and most prominent in keeping the complex static in an environment required.36
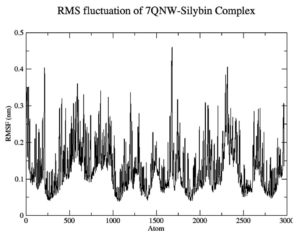
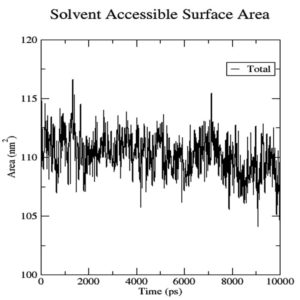
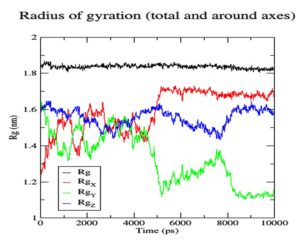
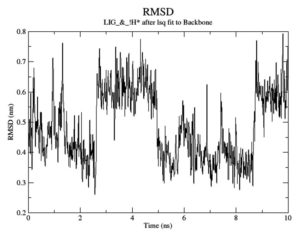
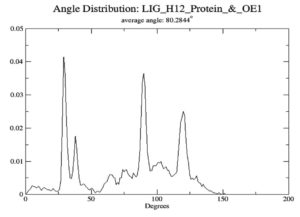
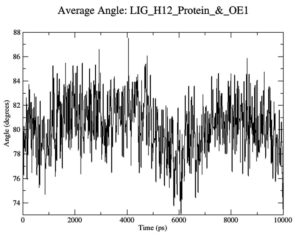
To gauge atomic malleability, we calculated RMSF values for ligand-protein complexes. Under 0.4 nm was maintained for RMSF values in the ligand-protein combination (Figure 5). At least in theory, the SASA can reveal how easily a protein can be dissolved in water. As the simulations progressed, SASA for the complex stayed about 110 nm.2 SASA values for the 7QNW-Silybin complex begin to lower down after 7 ns (Figure 6). Based on the decreasing SASA values, we can infer that the ligand Silybin interacts more with the system as the simulation progresses. The protein’s compactness during MD simulations was visualized by analyzing a plot of Rg during a time interval of 10 ns.37 The 7QNW-Silybin complex has a constant Rg of 1.8 nm, which is an indicator of its stability during simulations (Figure 7). The RMSD of ligands was tracked in order to monitor ligand stability in simulations. Average RMSD for Silybin was 0.5 nm, with a range of 0.3–0.7 nm suggesting the ligand’s stability during the simulation (Figure 8). Literature also supports the range of RMSD values obtained as a generally acceptable range. The average angle of Silybin in the protein’s active site was recorded as 80.28° (Figure 9, 10).38
Studies revealed that Silybin has many health-related beneficial properties. It has also been observed that silybin inhibits the NS5B RNA-dependent RNA polymerase of the Hepatitis C virus as well as the replication of the Influenza A virus.21,22 Computational docking studies of Silybin resulted in binding with the receptor binding domain of Omicron glycoprotein which further supports its use as an adjuvant therapy to inhibit virus spike glycoprotein and angiotensin converting enzyme (ACE)-2 interactions after experimental confirmations.
Recent emergence of the Omicron variant has added new difficulties to the worldwide fight against SARS-CoV-2 infection because this variant is extremely divergent with a significantly higher number of mutations especially in the spike protein region, which are associated with its immune escape potential and higher transmission rate. Selecting or prioritizing possible drug-like compounds for experimental research has been facilitated by computer-aided drug development, which has proved both time- and cost-efficient. In the active site of the Omicron variant spike protein, MD analysis shows that Silybin has lower binding energy, more non-bonded interaction capability, and increased stability. The results suggested that Silybin could be utilized to inhibit the interaction between the spike glycoprotein of SARS-CoV-2 Omicron variant and ACE-2, hence reducing the prevalence and spread of the virus. Silybin shows promise as a potential treatment for treating COVID-19 viral infection, but further research is needed in the form of in-vitro and in-vivo trials.
ACKNOWLEDGMENTS
The authors would like to thank Department of Biotechnology, MMEC, Maharishi Markandeshwar (Deemed to be University), Mullana-Ambala, India; Division of Pathology, ICAR-Indian Veterinary Research Institute, Izatnagar, UP, India; and Sri Guru Gobind Singh College, Chandigarh, India for their help and support.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
AS, SR and DC collected the literature and arranged references. PB and VS performed molecular docking part. AP, RK, DC and AD performed molecular simulation studies. AKS and KD critically analysed the data. PB, VS, AKS wrote the manuscript. KD critically edited the manuscript. All authors read and approved the final manuscript for publication.
FUNDING
None.
DATA AVAILABILITY
The datasets generated and/or analysed during the current study are available from the corresponding author on reasonable request.
ETHICS STATEMENT
This article does not contain any studies on human participants or animals performed by any of the authors.
- Kim J, Zhang J, Cha Y, et al. Advanced bioinformatics rapidly identifies existing therapeutics for patients with coronavirus disease-2019 (COVID-19). J Transl Med. 2020;18(1):257.
Crossref - Sharma AK, Sharma V, Sharma A, Pallikkuth S. Current paradigms in covid-19 research: Proposed treatment strategies, recent trends and future directions. Curr Med Chem. 2021;28(16):3173-3192.
Crossref - Bansal P, Kumar R, Singh J, Dhanda S. In silico molecular docking of SARS-CoV-2 surface proteins with microbial non-ribosomal peptides: Identification of potential drugs. J Proteins Proteom. 2021;12(3):177-184.
Crossref - Deepak J, Sharma V, Varruchi S, Sharma J, Kumar S, Sharma AK. Gender-bias susceptibility of corona virus disease: peeping into the factors responsible and emerging trends from Indian perspective. Biochem Cell Arch. 2020:4451-4456.
- Wolfe M, Hughes B, Duong D, et al. Detection of SARS-CoV-2 Variants Mu, Beta, Gamma, Lambda, Delta, Alpha, and Omicron in Wastewater Settled Solids Using Mutation-Specific Assays Is Associated with Regional Detection of Variants in Clinical Samples. Appl Environ Microbiol. 2022;88(8):e00045-22.
Crossref - Araf Y, Akter F, Tang Yd, et al. Omicron variant of SARS-CoV-2: genomics, transmissibility, and responses to current COVID-19 vaccines. J Med Virol. 2022;94(5):1825-1832.
Crossref - Jha D, Sharma V, Sharma J, et al. Plausible State-Specific Plans and Recommendations to Avert COVID-19 Community Transmission. Bulletin of Pure & Applied Sciences-Zoology. 2020;(2):447-454.
Crossref - Sharma V, Sharma AK, Yadav M, et al. Prediction models based on miRNA-disease relationship: Diagnostic relevance to multiple diseases including COVID-19. Curr Pharm Biotechnol. 2022.
Crossref - Shao W, Zhang W, Fang X, Yu D, Wang X. Challenges of SARS-CoV-2 Omicron Variant and appropriate countermeasures. J Microbiol Immunol Infect. 2022;55(3):387-394.
Crossref - Chakraborty C, Bhattacharya M, Dhama K. Cases of BA.2.75 and recent BA.2.75.2 subvariant of Omicron are increasing in India: Is it alarming at the global level? Ann Med Surg. 2022;84:104963.
Crossref - Hussain S, Tyagi S, Kaur G, Bahl G. The Evolving COVID-19: Omicron. Int J Biomed Res.2022;2(6).
Crossref - Wang P, Nair MS, Liu L, Iketani S, Luo Y, Guo Y, Wang M, Yu J, Zhang B, Kwong PD, Graham BS. Antibody resistance of SARS-CoV-2 variants B. 1.351 and B. 1.1. 7. Nature. 2021;593(7857):130-135.
Crossref - Zhang L, Li Q, Liang Z, et al. The significant immune escape of pseudotyped SARS-CoV-2 variant Omicron. Emerg Microbes Infect. 2022;11(1):1-5.
Crossref - Rabiul Islam M, Nasreen W, Anjum R, et al. Characteristics of the SARS-CoV-2 Omicron (B.1.1.529) Variant and Emerging Impact on Global Public Health. Clin Pathol. 2022;15.
Crossref - Chakraborty C, Bhattacharya M, Sharma AR, Dhama K, Lee SS. The rapid emergence of multiple sublineages of Omicron (B.1.1.529) variant: Dynamic profiling via molecular phylogenetics and mutational landscape studies. J Infect Public Health. 2022;15(11):1234-1258.
Crossref - Owis AI, El-Hawary MS, El Amir D, Aly OM, Abdelmohsen UR, Kamel MS. Molecular docking reveals the potential of Salvadora persica flavonoids to inhibit COVID-19 virus main protease. RSC Advances. 2020;10(33):19570-19575.
Crossref - Mani JS, Johnson JB, Steel JC, et al. Natural product-derived phytochemicals as potential agents against coronaviruses: A review. Virus Res. 2020;284:197989.
Crossref - Bansal P, Tuli HS, Sharma V, et al. Targeting Omicron (B. 1.1. 529) SARS CoV-2 spike protein with selected phytochemicals: an in-silico approach for identification of potential drug. J Exp Biol Agric Sci. 2022:396-404.
Crossref - Tuli H, Shivani S, Abhishek P, et al. Molecular Docking studies of Apigenin, Kaempferol, and Quercetin as potential target against spike receptor protein of SARS COV. J Exp Biol Agric Sci. 2022:144-149.
Crossref - Zakaryan H, Arabyan E, Oo A, Zandi K. Flavonoids: promising natural compounds against viral infections. Arch Virol. 2017;162(9):2539-2551.
Crossref - Bijak M. Silybin, a major bioactive component of milk thistle (Silybum marianum L. Gaernt.)-Chemistry, bioavailability, and metabolism. Molecules. 2017;22(11):1942.
Crossref - Liu C-H, Jassey A, Hsu H-Y, Lin L-T. Antiviral activities of silymarin and derivatives. Molecules. 2019;24(8):1552.
Crossref - Gurung AB, Ali MA, Elshikh MS, Aref I, Amina M, Lee J. An in silico approach unveils the potential of antiviral compounds in preclinical and clinical trials as SARS-CoV-2 omicron inhibitors. Saudi J Biol Sci. 2022;29(6):103297.
Crossref - Sharma V, Panwar A, Gupta GK, Sharma AK. Molecular docking and MD: mimicking the real biological process. Phys Sci Rev. 2022.
Crossref - Dejnirattisai W, Huo J, Zhou D, et al. SARS-CoV-2 Omicron-B. 1.1. 529 leads to widespread escape from neutralizing antibody responses. Cell. 2022;185(3):467-484.
Crossref - Sharma D. Bioinformatics and its applications in environmental science and health and its applications in other disciplines. Sambodhi. 2021;4:88-93.
- Sharma V, Panwar A, Sharma AK. Molecular dynamic simulation study on chromones and flavonoids for the in silico designing of a potential ligand inhibiting mTOR pathway in breast cancer. Curr Pharmacol Rep. 2020;6(6):373-379.
Crossref - Sharma V, Sharma AK. An In-Silico Approach for Designing a Potential Antagonistic Molecule Targeting β 2-adrenoreceptor Having Therapeutic Significance. Lett Appl NanoBioScience. 2021;10:2063-2069.
Crossref - Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS and Olson AJ. AutoDock4 and AutoDockTools4: Automated Docking with Selective Receptor Flexiblity. Journal of Computational Chem. 2009;30:2785-2791.
Crossref - Guex N, Peitsch MC. SWISS-MODEL and the Swiss-Pdb Viewer: an environment for comparative protein modeling. Electrophoresis. 1997;18(15):2714-2723.
Crossref - Raghav M, Sharma V, Gupta S, et al. In-silico designing of a potent ligand molecule against PTEN (Phosphatase and tensin homolog) implicated in Breast Cancer. J Exp Biol Agric Sci. 2022;10(5):1016-1023.
Crossref - Makarewicz T, Kazmierkiewicz R. Molecular dynamics simulation by GROMACS using GUI plugin for PyMOL. ACS Publications. 2013;53(5):1229-1234.
Crossref - Sargsyan K, Grauffel C, Lim C. How molecular size impacts RMSD applications in molecular dynamics simulations. J Chem Theory Comput. 2017;13(4):1518-1524.
Crossref - Xu Z, Liu K, George FG. Omicron variant of SARS-CoV-2 imposes a new challenge for the global public health. Biosafety and Health. 2022;4(3):147-149.
Crossref - Sharma V, Panwar A, Sankhyan A, Ram G, Sharma AK. Exploring the Potential of Chromones as Inhibitors of Novel Coronavirus Infection Based on Molecular Docking and Molecular Dynamics Simulation Studies.BRIAC. 2022;13(2):104-112.
Crossref - Goncalves PF, Stassen H. Calculation of the free energy of solvation from molecular dynamics simulations. Pure and Applied Chemistry. 2004;76(1):231-240.
Crossref - Sharma V, Panwar A, Sharma A, et al. A comparative molecular dynamic simulation study on potent ligands targeting mTOR/FRB domain for breast cancer therapy. Biotechnol Appl Biochem. 2021;69(4):1339-1347.
Crossref - Dixit SB, Ponomarev SY, Beveridge DL. Root mean square deviation probability analysis of molecular dynamics trajectories on DNA. J Chem Inf Model. 2006;46(3):1084-1093.
Crossref
© The Author(s) 2023. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.