ISSN: 0973-7510
E-ISSN: 2581-690X
Healthcare-Associated Infections (HAIs) are of global concern in this present era and white coats play an important role in the transmission of HAIs. The most common healthcare-associated bacteria are Enterococcus species, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumanni, Pseudomonas aeruginosa and Enterobacter species (ESKAPE bugs). These healthcare-associated bacteria are capable of surviving on white coats which could act as fomites in the transmission of HAIs. The purpose of this study was to compare the duration of survival of healthcare-associated bacteria on different materials used for making white coats. Materials like pure cotton, artificial cotton, cotton silk, spun and crepe were cut into swatches of 1cm2 size and sterilized by autoclaving. Five different bacteria isolated from clinical specimens were grown on 5 % sheep blood agar and bacterial suspensions were made in sterile physiological saline. The swatches were immersed in bacterial suspension and kept in petri plates at 25°C. The viable counts of bacteria were determined at definite time intervals by surface plate method. The present study shows that among the healthcare-associated bacteria, S.aureus survived the maximum up to 52 days. The duration of survival of S.aureus was significantly longer than P.aeruginosa and A.baumannii (p <0.05). A.baumannii survived only up to 20 days maximum. All the healthcare-associated bacteria significantly survived for the shortest duration of time on crepe. Therefore, crepe could be a better material used for making white coats.
Healthcare-Associated bacteria, Survival, White coat materials, Fomites, Gram-positive cocci, Gram-negative bacilli
Healthcare-Associated Infections (HAIs) are mainly caused by healthcare-associated bacteria belonging to the “ESKAPE bugs” which include Enterococcus species, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumanni, Pseudomonas aeruginosa and Enterobacter species.1 Different modes of transmission of these bacterial pathogens in the health care settings include direct and indirect contact, inhalation, ingestion and inoculation.2 Fomites like white coats, gloves, masks and instruments play a significant role in the transmission of healthcare-associated bacteria in the healthcare settings. The white coat is a sign of purity, but still could function as a fomite in the transmission of microorganisms in the healthcare settings.2,3
The duration of survival of different bacteria on white coat may vary based on the type of bacteria, number of bacteria, material used for making white coat, environmental temperature and humidity. Gram-positive bacteria like S.aureus and Enterococcus spp. has known to survive on white coats.4 Methicillin resistant S.aureus (MRSA) which has the capability of spreading HAIs has also been isolated from the white coats.5 Among the gram negative bacteria E.coli, P.aeruginosa and A.baumannii are the common healthcare-associated bacteria known to cause HAIs.6
Traditionally, the white coats worn by Healthcare workers (HCW’s) are made from cotton, but nowadays many other materials like spun, artificial cotton, cotton-silk and crepe are also used. These healthcare-associated bacteria have survived on materials like cotton, cotton silk, and spun for an extended period of time and can cause HAIs by transferring from gowns to others.7,8 With the increased rate of infection caused by these bacteria, studying the inanimate surfaces including the HCW’s attire is also necessary for reducing the rate of HAIs.9 There is a paucity of studies done on the comparison of survival of healthcare-associated bacteria on white coat materials. Therefore, the purpose of the study was to compare the duration of survival of selected healthcare-associated bacteria on different materials used for making white coat.
Study setting and study design
The descriptive study was conducted in Department of Microbiology, Kasturba Medical College, Mangalore, during January 2021 to May 2021.
Preparation of white coat materials
White coat materials such as pure cotton, artificial cotton, spun, cotton-silk and crepe were used. Swatches of size 1cm2 size were cut from these materials.9 Then the swatches were sterilised by autoclaving for 15 minutes at 121°C. After sterilisation, the swatches were spreaded on sterile petri plates and dried in incubator at 37°C. After drying, the edges of the plates were covered using a cellophane tape to avoid contamination.
Preparation of bacterial inoculum
The test bacteria such as S.aureus, E.faecalis, E.coli, P.aeruginosa and A.baumannii used in were procured from clinical microbiology specimens and were grown in 5% sheep blood agar, at 37°C for 24hours. These isolates were identified by standard bacteriological methods. The fresh bacterial colonies obtained from the culture were picked using a sterile wire and suspended in sterile physiological saline and turbidity was matched with McFarland 0.5 standard (bacterial count 1.5 x 108 CFU/ml). The bacterial suspension was then further diluted 1 in 1000 in sterile physiological saline to get bacterial count 1.5 x 105 CFU/ml.4
Test for survival of bacteria on white coat material
This method was based on a previously published article but few modifications were made.4 Here, the swatches were soaked in different bacterial suspensions and then they were spreaded on sterile petri dishes. The edges of the plates were covered with cellophane tape to prevent contamination and were kept in room temperature (24-26°C). The swatches were then transferred to 1ml nutrient broth and vortexed once in week. Using a calibrated loop of 4mm internal diameter (0.01ml) the inocula were streaked on 5% sheep blood agar and then colony count was determined. As the colony count reduced, sub-culturing was done on the alternate days. The nutrient broth containing the swatch was incubated at 37°C for 24 hours and then was checked for turbidity. Nutrient broth showing no turbidity was subcultured to 5% sheep blood agar to check for bacterial growth. No growth was confirmed as the plate showed no bacterial colony. The mean and standard deviation of the number of days the bacteria survived was calculated and a graph was plotted with number of days on x-axis and log viable count on y-axis.
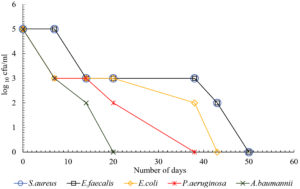
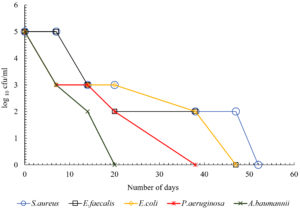
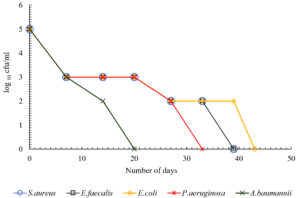
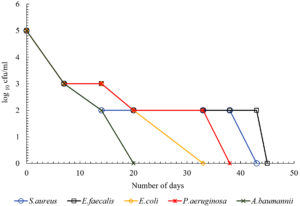
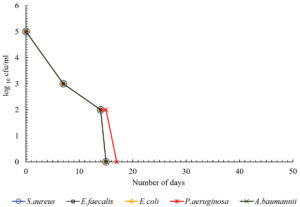
The present study showed that there was difference in the duration of survival of different bacteria on white coat. The data from the study showed that S.aureus survived the maximum duration for up to 52 days. The duration of survival of S.aureus was significantly more when compared with P.aeruginosa and A.baumannii (p<0.05). Among the gram negative bacteria, E.coli survived the maximum up to 47 days. The duration of survival of E.coli was significantly more when compared with P.aeruginosa and A.baumannii (p<0.05). A.baumannii survived the shortest duration for up to 15 days on crepe and 20 days on all other materials (Table 1). All HA bacteria have survived the shortest duration of time on crepe. P.aeruginosa survived for 17 days and all other bacteria for 15 days on crepe. Fig. 1 – 5 depicts the log viable count of different types of bacteria on different white coat materials like pure cotton, artificial cotton, cotton silk, spun and crepe respectively. The graph shows the reduction in the log viable count of the bacteria.
Table (1):
Duration of survival of HAI causing bacteria on different white coat materials.
| Bacteria | Duration of survival of HA bacteria on different white coat materials | ||||
|---|---|---|---|---|---|
| (Days – Mean ± standard deviation) | |||||
| Pure cotton | Artificial cotton | Cotton silk | Spun | Crepe | |
| S.aureus | 50.0 ± 4.3 | 52.0 ± 3.4 | 39.0 ± 2.6 | 43.0 ± 6.2 | 15.0 ± 0 |
| E.faecalis | 50.0 ± 3.3 | 47.0 ± 3.1 | 39.0 ± 1.7 | 45.0 ± 1.7 | 15.0 ± 0 |
| E.coli | 43.0 ± 6.2 | 47.0 ± 5.7 | 43.0 ± 6.2 | 33.0 ± 4.9 | 15.0 ± 0 |
| P.aeruginosa | 38.0 ± 2.4 | 38.0 ± 3.1 | 34.0 ± 0 | 38.0 ± 0 | 17.0 ± 0 |
| A.baumannii | 20.0 ± 0 | 20.0 ± 0 | 20.0 ± 0 | 20.0 ± 0 | 15.0 ± 0 |
The present study was conducted to compare the duration of survival of healthcare-associated HA bacteria on different materials used for preparing white coats. Materials like pure cotton, artificial cotton, cotton silk, spun and crepe were used for the study. The data from the study shows that the healthcare-associated bacteria could survive up to 52 days. The data was comparable to a previous study which also showed that the duration of survival of these bacteria depend upon the materials used for making white coats.10 Among the healthcare-associated bacteria S.aureus survived the maximum up to 52 days. The duration of survival of S.aureus was significantly more when compared with P.aeruginosa and A.baumannii (p<0.05). A previous study showed that white coats were contaminated with S.aureus, E.coli and P.aeruginosa and could act as fomites in the transmission of HAIs.11 The ability of S.aureus to survive could be because of their structural morphology. Moisture and humidity also help in the survival of these bacteria on white coats. Previous studies have showed that S.aureus is a major contaminant of white coats.12-14 Therefore, longer viability of S. aureus on white coats can lead to the spread of this bacteria.
The present study also shows that the healthcare-associated bacteria have survived on pure cotton and artificial cotton comparatively longer than other materials. This is because of the hydrophilic structure of the material that can trap water, oxygen and moisture.15,16 A previously published study showed that S.aureus has high binding capacity to polyester and very low binding capacity on cotton.17 But in this study S.aureus survived the longest on natural fibres like cotton and spun (cotton-polyester blend) is seen to be the third material after pure cotton and artificial cotton where it survived for 43 days. Followed by S.aureus, E.faecalis survived the maximum up to 50 days. This was not consistent with the previous study which suggested that enterococci survived longer than staphylococci.3
Among the gram negative bacteria, E.coli survived the maximum up to 47 days. It might be because of the lack of sunlight and low temperature which helped in the growth of gram negative bacteria on white coat materials.18 A previous study has also showed that coliform bacteria has good adherence on cotton.17 The duration of survival of E.coli was significantly longer than P.aeruginosa and A.baumannii (p<0.05). In the case of P.aeruginosa, it survived the maximum up to 38 days in pure cotton, artificial cotton and spun. The least surviving organism was found to be A.baumannii and survived the maximum up to 20 days, except on crepe where it survived for up to 15 days and this could be because of the room temperature as the optimum temperature for their growth is 37°C.
Of all these materials, all the healthcare-associated bacteria survived the shortest duration of time on the material crepe. P.aeruginosa survived up to 17 days and all other bacteria up to 15 days, suggesting that crepe could be a better material used for making white coat because it sustains the survival of these bacteria for shortest period of time. This could be because of the manufacturing of the material which involves tighter twisting of the yarns or because of the chemical treatment with caustic soda. So overall, the data shows that gram positive bacteria survived longer than gram negative bacteria and it was similar to a previous study.19 This could be because of the rigid body of gram positive bacteria. Fig. 1 – 5 shows the log viable count of different types of bacteria on different white coat materials. The graph shows the reduction of the log viable count. A previous study showed that, even though higher inoculum is used for checking the survival rate of bacteria, the number of bacteria dying in the high inoculum increases to sustain the growth of viable cells so that the viable cells can survive from the nutrients of the dying cells.4 This could explain the decrease in log viable count.
The present study showed that the survival of healthcare-associated bacteria on white coat materials depends on the type of bacteria and type of white coat material. S.aureus survived significantly more than other bacteria. All the bacteria survived shortest duration of time on crepe. Therefore, crepe could be the material used for making white coat because it sustains survival of these bacteria for the shortest period. Selection of white coat material and frequent laundering of white coats are crucial in controlling HAIs transmitted through white coats.
ACKNOWLEDGMENTS
The authors would like to express gratitude to all the teaching and non-teaching staffs of Department of Microbiology, Coastal Karnataka, India for their co-operation throughout the study.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
Both the authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
None.
ETHICS STATEMENT
This study was approved by Institutional Ethics Committee (IEC), Kasturba Medical College, Mangalore, India with Protocol number-IECKMCMLR-12/2020/401.
AVAILABILITY OF DATA
All datasets analysed during the study are included in the manuscript.
- Hanczvikkel A, Toth A. Quantitative study about the role of environmental conditions in the survival capability of multidrug-resistant bacteria. J Infect Public Health. 2018;11(6):801-806.
Crossref - Olvera-Lopez E, McCaffery E, Kasubhai M. White coat wearing and laundering frequency among internal medicine department physicians. Am J Infect Control. 2020;48(7):834-836.
Crossref - Mitchell A, Spencer M, Edmiston C. Role of healthcare apparel and other healthcare textiles in the transmission of pathogens: A review of the literature. J Hosp Infect. 2015;90(4):285-292.
Crossref - Neely AN, Maley MP. Survival of Enterococci and Staphylococci on hospital fabrics and plastic. J Clinic Microbiol. 2000;38(2):724-726.
Crossref - Mwamungule S, Chimana HM, Malama S, Mainda G, Kwenda G, Muma JB. Contamination of health care workers’ coats at the University Teaching Hospital in Lusaka, Zambia: the nosocomial risk. J Occup Med Toxicol. 2015;10(1):34.
Crossref - Peleg AY, Hooper DC. Hospital-Acquired Infections Due to Gram-Negative Bacteria. N Engl J Med. 2010;362(19):1804-1813.
Crossref - Suleyman G, Alangaden G, Bardossy AC. The Role of Environmental Contamination in the Transmission of Nosocomial Pathogens and Healthcare-Associated Infections. Curr Infect Dis Rep. 2018;20(6):12.
Crossref - Gupta P, Bairagi N, Priyadarshini R, Singh A, Chauhan D, Gupta D. Bacterial contamination of nurses’ white coats after first and second shift. Am J Infect Control. 2017;45(1):86-88.
Crossref - Wiener-Well Y, Galuty M, Rudensky B, Schlesinger Y, Attiasm D, Yinnon AM. Nursing and physician attire as possible source of nosocomial infections. Am J Infect Control. 2011;39(7):555-559.
Crossref - Chacko L, Jose S, Isac A, Bhat KG. Survival of nosocomial bacteria on hospital fabrics. Indian J Med Microbiol. 2003;21(4):291.
Crossref - Kumar PA, Chougale RA, Sinduri I. Bacterial Contamination of White Coats among Medical Personnel- A Cross Sectional Study in Kolhapur, India. J Pure Appl Microbiol. 2020;14(2):1405-1411.
Crossref - Treakle AM, Thom KA, Furuno JP, Strauss SM, Harris AD, Perencevich EN. Bacterial contamination of health care workers’ white coats. Am J Infect Control. 2009;37(2):101-105.
Crossref - Banu A, Anand M, Nagi N. White coats as a vehicle for bacterial dissemination. J Clin Diagn Res. 2012;6(8):1381-1384.
Crossref - Boyce JM. Environmental contamination makes an important contribution to hospital infec-tion. J Hosp Infect. 2007;65(2):50-54.
Crossref - Badr AA. Anti-microbial and durability characteristics of socks made of cotton and re-generated cellulosic fibres. Alex Eng J. 2018;57(4):3367-3373.
Crossref - Shin Y, Park J, Park W. Sterilization efficiency of pathogen-contaminated cottons in a laun-dry machine. J Microbiol. 2020;58(1):30-38.
Crossref - Takashima M, Shirai F, Sageshima M, Ikeda N, Okamoto Y, Dohi Y. Distinctive bacteria-binding property of cloth materials. Am J Infect Control. 2004;32(1):27-30.
Crossref - Colclasure VJ, Soderquist TJ, Lynch T, et al. Coliform bacteria, fabrics, and the environ-ment. Am J Infect Control. 2015;43(2):154-158.
Crossref - Malini M, Thomas T, Bhargava D, Girija S. Microbiology of the white coat in a dental operatory. Ind J Dent Res. 2012;23(6):841.
Crossref
© The Author(s) 2022. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.