ISSN: 0973-7510
E-ISSN: 2581-690X
Acinetobacter infection with multidrug resistant strains is an emerging infection of global concern as it leads to serious disease. They are also important pathogens causing hospital acquired infections. Information on the prevalence, spectrum of illness and antibiotic sensitivity pattern of Acinetobacter is important for appropriate management of patients. We aimed to determine the prevalence of Acinetobacter species and evaluate the clinical profile and antibiotic sensitivity pattern of Acinetobacter species from various clinical samples. From October 2018 to September 2019, various clinical samples received in the microbiology laboratory were studied from the electronic records and the data on the isolation of Acinetobacter from these samples and its antibiotic sensitivity pattern was collected and analysed. The clinical data was also collected to determine the clinical spectrum. The prevalence of Acinetobacter species from various clinical samples was found to be 8.9%. Isolates were more common in general wards than in ICUs. The Acinetobacter infections occurred significantly in male patients (65.7%) than in female patients (34.3%), with male: female ratio of 1.9:1. The most common infection caused by Acinetobacter species was Wound infection (54.36%) followed by Respiratory tract infection (34.27%). Multidrug resistance was seen in 75 % of the isolates. Significant prevalence of multidrug resistant Acinetobacter infections was noted in our study. The findings emphasize the need for strict hospital infection control practices and the restricted use of antibiotics to prevent the occurrence of these infections.
Acinetobacter Species, Multidrug Resistant Organisms, Antibiotic Sensitivity Pattern
Acinetobacter species are widespread in environment and are usually present in soil, sewage, water, animal, and human samples. They are non-fermenting gram negative coccobacilli and are strictly aerobic.1 Among the non fermenters, Acinetobacter species is the second most common microorganism isolated from patients. They are known to cause nosocomial infections such as ventilator associated pneumonia (VAP), septicemia, urinary tract infection and wound infection.2
Acinetobacter species have a unique property of survival in hospital environment for longer time, during which they may acquire resistance to broad spectrum antibiotics.3 Currently, they have developed resistance against most of the known antibiotics, and thus becomes difficult to treat infections caused by such resistant strains. Emergence of multidrug resistant (MDR) strains not only narrow downs the choice of antibiotics for treatment but also increases economic burden to the patients as well as hospitals.4,5 With increasing trend of infections from this organism worldwide, we planned to do this study with special reference to detection of MDR Acinetobacter strains. This helps us to know the spectrum of illness and antibiotic sensitivity pattern of Acinetobacter species in our region which aids in formulation of antibiotic policy and guide the clinicians for administering appropriate empirical antibiotic treatment.
Objectives
1. To determine the prevalence of Acinetobacter species from various clinical samples
2. To evaluate the clinical profile and antibiotic sensitivity pattern of Acinetobacter species from various clinical samples
This cross sectional, retrospective study was done at the department of Microbiology, Sri Devaraj Urs Medical College, Kolar, Karnataka and was approved by Institutional Ethics committee (Approval no.: No. DMC/KLR/IEC/318/2021-22). Various samples collected from patients like endotracheal aspirate, sputum, blood, pus, and urine received for aerobic bacterial culture and sensitivity at the microbiology laboratory from October 2018 to September 2019 were included in the study. Data of the samples received with regard to their clinical history was collected. Isolates were identified as Acinetobacter species based on standard microbiological techniques which included Gram stain of direct smears, colony morphology on Blood agar and MacConkey agar and standard biochemical reactions.6,7 Antibiotic sensitivity pattern was determined by Kirby Bauer Disc Diffusion method as per CLSI guidelines.7,8 Following antibiotics were tested for antibiotic sensitivity; Piperacillin (100µg), Piperacillin-Tazobactam (100/10µg), Ceftazidime (30µg), Cotrimoxazole (1.25/23.75µg), Imipenem (10µg), Meropenem (10µg), Amikacin (30µg), Gentamicin (10µg), Ciprofloxacin (5µg), Levofloxacin (5µg), Ampicillin-Sulbactam (10/10µg) and Cefepime (30µg).
Statistical analysis
The data was tabulated in Microsoft Excel and analysed using SPSS Software version 22. Descriptive statistics was used to summarize the results of the study.
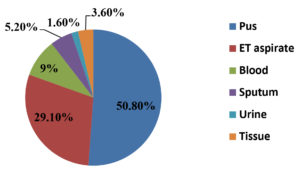
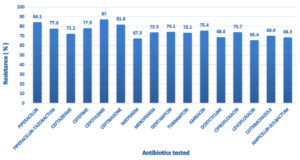
During the study period, a total of 3473 bacteria were isolated from various clinical samples of which, 309 were identified as Acinetobacter species accounting for a prevalence of 8.9%. Out of these 309 isolates, 197 (64%) were from patients located in general wards and 112 (36%) isolates were from patients admitted in Intensive Care Units (ICU). Majority of the isolates were from male patients (65.7%) than in female patients (34.3%), with Male: Female ratio of 1.9:1. With regard to clinical samples, Acinetobacter species was most commonly isolated from pus samples (50.8%), followed by endotracheal aspirate (29.1%), blood (9.06%), sputum (5.17%), urine (1.6%) and tissue samples (3.6%), as shown in Figure 1. The most common infection caused by Acinetobacter species in our study was wound infection (54.36%) followed by Respiratory tract infection (34.27%). Sample wise distribution and clinical profile of Acinetobacter species is shown in Table. Antibiotic sensitivity pattern of the isolates showed that most of the Acinetobacter species were resistant to Cefotaxime (87%), Piperacillin (84.1%), Ceftriaxone (81.8%), Ceftazidime (72%), Cefepime (77.9%), Piperacillin-Tazobactam (77.6%) and Amikacin (75.4%) which is depicted in Figure 2.
Table:
Clinical profile of Acinetobacter infections.
| Clinical profile | Sample type | Number | Percentage |
|---|---|---|---|
| Wound infection | Pus | 168 | 54.36 |
| Ulcer | 46 | 27.38 | |
| Surgical site infections | 38 | 12.3 | |
| Cellulitis | 28 | 16.66 | |
| Diabetic foot | 21 | 6.8 | |
| Abscess | 20 | 6.5 | |
| Wet gangrene | 9 | 2.9 | |
| Amputation | 5 | 1.6 | |
| SSG* infections | 1 | 0.3 | |
| Respiratory tract infection | Sputum & ET aspirate | 106 | 34.3 |
| Pneumonia | 82 | 26.53 | |
| Secondary infection in Bronchial asthma | 12 | 3.8 | |
| ARDS** | 9 | 2.9 | |
| Acute bronchitis | 3 | 0.9 | |
| Septicaemia | Blood | 28 | 9.06 |
| Urinary tract infection | Urine | 05 | 1.6 |
| CAUTI*** Pyelonephritis |
4 1 |
1.3 0.3 |
*SSG-Split skin graft; **ARDS- Acute Respiratory Distress syndrome
***CAUTI-Catheter associated urinary tract infection
Among the non-fermenters, Acinetobacter species is the second most common organism isolated from human specimens following Pseudomonas aeruginosa.9 In our study, prevalence of Acinetobacter species was 8.9% which was comparable with a study done by Chitrabanu et al who reported as 6.2%.10 Majority of Acinetobacter species in our study were isolated from ICUs (36%) followed by various General wards (64%). Similar finding was also observed in a study done at Kenya where in 48% of Acinetobacter species were isolated from critical care units.11 Infections due to Acinetobacter in our study were more common among male patients (65.7%), similar findings were observed in other studies.1,12 Most common clinical specimen, yielding growth of Acinetobacter species in our study was pus sample (50.8%) followed by endotracheal aspirate (29.1%), which is in concordance with the findings in a study done at Nepal where majority of the Acinetobacter species were isolated from pus samples (40%) followed by respiratory samples (21.8%).13 However in few other studies Acinetobacter was commonly isolated from respiratory specimen.1,2,3 The most common clinical manifestation due to Acinetobacter in our study was wound infection (50.8%) followed by Respiratory tract infection (34.2%), which is similar to the findings of other studies.1,14 The variation in the clinical spectrum could be due to difference in sources, usage of antibiotics, hospital environment and infection control practices in different hospitals.15,16,17 Other factors could also contribute to the occurrence of Acinetobacter infections such as patients with immunocompromised conditions, severe underlying disease, prolonged hospital stay and more frequent use of antibiotics.1
Acinetobacter species has caused a major public health problem as antibiotic resistance among them has increased substantially in the past decade. They are considered notorious due to their ability to acquire antibiotic resistance.1 More than 75% of the isolates in our study were resistant to Cefotaxime, Piperacillin, Ceftriaxone, Ceftazidime, Cefepime, Piperacillin-Tazobactam and Amikacin. Significant number of isolates were resistant to Fluoroquinolones and Aminoglycosides [Ciprofloxacin (73.7%), Levofloxacin (65.4%), Gentamicin (74.1%), Amikacin (75.4%), and Tobramycin (73.1%)]. Similar resistance pattern was observed in study done in Nepal.13 In our study, 67.3% and 73.5% of the isolates were resistant to Imipenem and Meropenem respectively, which is similar to the findings in a study done by Mathai A S et al.3 Such Carbapenem resistant strains can be treated by reserved drugs such as Colistin, Polymyxin B and Tigecycline.18 Antibiotic susceptibility testing against these reserved drugs was not done in our study. However, Raut S et al in their study showed 100% sensitivity of Acinetobacter isolates to Colistin, Polymyxin B, and Tigecycline.13 Recent emergence of few Acinetobacter strains resistant to all the commercially available antibiotics is a worrisome development.19,20
In developing countries like India, health care settings especially the public health settings are overcrowded and have limited resources regarding nursing care, hospital infection prevention and control practices. Considering this, clinicians should reduce the usage of broad spectrum antibiotics for empiric management and also follow antibiotic sensitivity testing report for appropriate treatment. In addition, regular analysis of antibiotic sensitivity pattern is essential to form antibiotic policy and change the antibiotic prescription practices accordingly, for which strengthening of diagnostic capabilities of microbiology laboratory is essential.
Our study showed a significant prevalence of Acinetobacter infections in our region. Multidrug resistance was seen in 75% of the Acinetobacter isolates which is of high concern as it leads to treatment failure and adverse outcomes among patients. The findings emphasize the importance of improving microbiological tests for accurate and early identification of such organisms. The study calls for strengthening of antibiotic stewardship program and hospital infection prevention and control practices in the health care settings which may help in preventing the occurrence of infections with multidrug resistant Acinetobacter.
ACKNOWLEDGMENTS
The authors would like to thank Laboratory Technicians for data support.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors have made a direct, substantial, and intellectual contribution to the work, and approved it for publication
FUNDING
None.
DATA AVAILABILITY
All the data sets generated and analysed during this study are included in the manuscript
ETHICS STATEMENT
This study was approved by the Institutional Ethics Committee, Sri Devraj Urs Medical College, Tamaka, Kolar, India. (DMC/KLR/ICE/318/2021/22)
- Tripathi PC, Gajbhiye SR, Agrawal GN. Clinical and antimicrobial profile of Acinetobacter spp.: An emerging nosocomial superbug. Adv Biomed Res. 2014;3:13.
Crossref - Kumari M, Batra P, Malhotra R, Mathur P. A 5-year surveillance on antimicrobial resistance of Acinetobacter isolates at a level-I trauma centre of India. J Lab Physicians. 2019;11(1):34-38.
Crossref - Mathai AS, Oberoi A, Madhavan S, Kaur P. Acinetobacter infections in a tertiary level intensive care unit in northern India: Epidemiology, clinical profiles and outcomes. J Infect Public Health. 2012;5(2):145-152.
Crossref - Liu WL, Liang HW, Lee MF, et al. The impact of inadequate terminal disinfection on an outbreak of imipenem-resistant Acinetobacter baumannii in an intensive care unit. PLoS One. 2014;9(9):e107975.
Crossref - Souli M, Galani I, Giamarellou H. Emergence of extensively drug-resistant and pandrug-resistant Gram-negative bacilli in Europe. Euro Surveill. 2008;13(47):19045.
Crossref - Procop GW, Church DL, Hall GS, et al. The Non fermentative Gram-negative bacilli. Koneman’s Colour Atlas and textbook of Diagnostic Microbiology. 7th Edition. China: Wolters Kluwer 2017;325-327.
- Collee JG, Duguid JP, Fraser AG, Marmion BP, Simmons A. Neisseria, Moraxella, Acinetobacter. Mackie and Mc Cartney Practical Medical Microbiology. 14th edition, Churchill Livingstone Elsevier, New Delhi. 2007;294-296.
- Clinical and Laboratory Standards Institute. CLSI performance standards for antimicrobial susceptibility testing. 13th ed. CLSI standard M02. Wayne, PA: Clinical and Laboratory Standards Institute, 2018.
- Hsu LY, Apisarnthanarak A, Khan E, Suwantarat N, Ghafur A, Tambyah PA. Carbapenem-Resistant Acinetobacter baumannii and Enterobacteriaceae in South and Southeast Asia. Clin Microbiol Rev. 2017;30(1):1-22.
Crossref - Chitrabanu NA, Mallya S. Identification, Speciation and Antibiogram along with Detection of Metallo Beta-lactamase Production in Acinetobacter Isolated from Clinical Samples in a Tertiary Care Hospital. J Pure Appl Microbiol. 2021;15(2):839-844.
Crossref - Musyoki VM, Masika MM, Mutai W, Wilfred G, Kuria A, Muthini F. Antimicrobial susceptibility pattern of Acinetobacter isolates from patients in Kenyatta National Hospital, Nairobi, Kenya. Pan Afr Med J. 2019;33:146.
Crossref - Prashanth K, Badrinath S. Nosocomial infections due to Acinetobacter species: clinical findings, risk and prognostic factors. Indian J Med Microbiol. 2006;24(1):39-44.
Crossref - Raut S, Rijal KR, Khatiwada S, Karna S, Khanal R, Adhikari J, Adhikari B. Trend and Characteristics of Acinetobacter baumannii Infections in Patients Attending Universal College of Medical Sciences, Bhairahawa, Western Nepal: A Longitudinal Study of 2018. Infect Drug Resist. 2020;13:1631-1641.
Crossref - Bennani B, Selmani R, Mahmoud M, Nejjari C, Kanjaa N. Nosocomial pneumonia in mechanically ventilated patients: Prospective study in intensive care unit of Fez University Hospital. Saudi J Anaesth. 2008;2(2):46-51. https://www.saudija.org/text.asp?2008/2/2/46/51855
- Amatya R, Acharya D. Prevalence of tigecycline resistant multidrug resistant Acinetobacter Calcoaceticus- Acinetobacter baumanii complex from a tertiary care hospital in Nepal. Nepal Med Coll J. 2015;17(1-2):83-86.
- Van TD, Dinh QD, Vu PD, et al. Antibiotic susceptibility and molecular epidemiology of Acinetobacter calcoaceticus-baumannii complex strains isolated from a referral hospital in northern Vietnam. J Glob Antimicrob Resist. 2014;2(4):318-321.
Crossref - Shrestha M, Khanal B. Acinetobacter Species: phenotypic characterization and antimicrobial resistance. J Nobel Med Coll. 2013;2(1):43-48.
Crossref - Doi Y, Murray GL, Peleg AY. Acinetobacter baumannii: evolution of antimicrobial resistance-treatment options. Semin Respir Crit Care Med. 2015;36(1):85-98.
Crossref - Lolans K, Rice TW, Munoz-Price LS, Quinn JP. Multicity outbreak of carbapenem-resistant Acinetobacter baumannii isolates producing the carbapenemase OXA-40. Antimicrob Agents Chemother. 2006;50(9):2941-2945.
Crossref - Talbot GH, Bradley J, Edwards JE Jr, Gilbert D, Scheld M, Bartlett JG; Antimicrobial Availability Task Force of the Infectious Diseases Society of America. Bad bugs need drugs: an update on the development pipeline from the Antimicrobial Availability Task Force of the Infectious Diseases Society of America. Clin Infect Dis. 2006;42(5):657-68.
Crossref
© The Author(s) 2022. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.