ISSN: 0973-7510
E-ISSN: 2581-690X
Antimicrobial resistance gene profile characterization and dissemination offer useful detail on the possible challenge in treating bacteria. The development of aminoglycoside modifying enzymes (AMEs) is considered as the primary mechanism of resistance to aminoglycosides, in addition to the 16S rRNA methylases. This study aimed at isolation and characterization of aminoglycosides resistant clinical isolates of enterobacteriaceae family from different clinical samples. Over a period of 24 months, thirty samples were collected and 49 clinical isolates of E. coli [n=25], Klebsiella [n=13], Enterobacter species (n=7) and Proteus species (n=4) were isolated from Egyptian clinical laboratories. The identities of the cultures were confirmed following standard microbiological procedures. Resistance of the isolates to aminoglycosides was determined by the disc diffusion method and isolates with highest resistance (n=9) were selected and investigated for 16S rRNA methylase and AMES encoding genes by polymerase chain reaction (PCR) and sequencing. In general, aminoglycoside resistance was found in 95% of the isolates; the isolates displayed the highest rate of resistance to netilmicin (75%) and kanamycin (55%), while resistance to gentamycin (18%) and tobramycin (16%) was low. A total of 9 isolates have the highest aminoglycoside resistant rate, showed the highest appearance for aac(6′)-Ib as well as ant (3″)-Ia resistant genes, with aac (3)-II (44%) and ant (4′)-IIb (34%) following closely. The high prevalence of AMEs observed among resistant isolates in this study suggests the urgent need for more efficient treatment designs to mitigate the selection burden as well as improved care of patients who have been infected with these drug-resistant organisms.
Aminoglycoside modifying enzymes, antibiotic resistance, E. coli, Klebsiella pneumoniae, 16S RNA
Gram-negative bacteria (particularly E.coli and Klebsiella) have long been suspected major causes of nosocomial infections, including infections of the urinary tract (UT), respiratory system and bacteremia.1,2 Resistance of antibiotics, particularly to Gram-negative bacteria, has arisen as worldwide epidemic in the Twenty-First Century. Their resistance to diverse antibiotic classes limits available therapeutic options for their control. In particular, antibiotic-resistant pathogens isolated from hospitals are becoming increasingly common, which is a most health concern around the world.3 A report published in 2014 revealed that, antimicrobial resistance causes more than 700,000 deaths worldwide per year, with that Fig. expected to increase to 10 million by 2050.4 There have been a recent development in the management of severe infections by older antibiotics, especially aminoglycosides.5 This is because the relatively low use of older antibiotics may have helped maintain their effectiveness against certain bacterial isolates that became more resistant to newer antibacterial agents.6 Aminoglycoside molecules bind irreversibly to the ribosome 30S subunit, leading to complete protein synthesis inhibition and final bacterial death. Moreover, the mRNA translation interference by these antibiotics contributes to misreading of the codons of mRNA.7 The study of molecular and genetic resistance determinants has a key role in the interpretation, controlling and distribution of resistance pathogens. Antimicrobial resistance mechanisms involving aminoglycosides include the following: addition of aminoglycoside-modifying enzymes leads to the antibiotic being inactive, Changes in ribosomal high affinity sites, the down-regulation of porin genes has decreased antibiotic uptake and efflux pumps.8,9 Resistance to aminoglycosides may be caused by different mechanisms, particularly enzymatic modification, which is the most significant pathway and is classified into three categories; aminoglycoside nucleotidyl transferase (ant), aminoglycoside acetyltransferase (aac) and aminoglycoside phosphoryl transferase (aph); production of these enzymes is encoded by genes located on bacterial chromosomes or plasmids.10 Aminoglycoside modifying enzymes catalyze the modification at –OH or –NH2 groups of the 2-deoxystreptamine nucleus or the sugar moieties and can be acetyltransferases (AACs), nucleotidyl- tranferases (ANTs), or phosphotransferases (APHs).11 Overall, aac (3)-Ia, aac (6′)-II, aac (6′)-Ib and aac (3)-II are among the common aminoglycoside in-activating enzymes in several Gram-negative clinical isolates, such as Klebsiella pneumoniae, whereas, aph (3′), ant(3)I or ant(4)Пb are less common.12,13 Generation of 16S rRNA methyltransferase (16S RMTase) is another pathway of aminoglycoside resistance, which methylates the drug’s binding site, making bacteria more resistant to clinically significant members of this group of antibiotics like gentamicin, tobramycin and amikacin.14 The 16S rRNA methylases (16S-RMTase) have been shown to be responsible for high-level resistance against a range of aminoglycosides in Gram-negative bacilli. 16S rRNA methylases have emerged as a novel resistance mechanism to aminoglycosides.15 Eight 16S-RMTase genes have been discovered in numerous enterobacteriaceae species and identified as npmA, rmtF, rmtE, rmtD, rmtC, rmtB, rmtA and armA. Particularly, rmtB and armA genes have been found to be the most predominant and extensively widespread throughout Asia.16 However, in Egypt, little research has been reported on the existence of AMEs and the genes that encode enzymes for aminoglycoside resistance. This study is therefore intended to determine levels of and genes encoding aminoglycoside resistance among E. coli and K. pneumoniae isolates from patients with various infectious diseases.
Collection, isolation and identification of bacteria
Thirty clinical samples were obtained from three laboratories located in the Beni-Swef Governorate, Egypt. The ethics committee at the faculty of medicine, Beni-suef University, NU. Beni-suef, Egypt, approved the study. The study protocol conformed to the ethical guidelines of the 1975 Declaration of Helsinki. All the samples were collected from October 2016 to November 2018. These clinical samples were first grown aerobically on tryptone soya broth (TSB) then streaked on Mac-Conkey agar (Oxoid, UK) and incubated for 24-hours at 37°C. Basic microbiological techniques, such as Gram staining and colony morphology, and some biochemical tests, were used to confirm the identity of the isolates.
Molecular identification and Phylogenetic analysis
Extraction, purification, and analysis of DNA of the isolates was performed as previously described.17 The universal primers 1525R (5′-AGAAAGGAGGTGATCCAGCC-3′)18 and 357F (5′-TACGGGAGGCAGCAG-3′)19 were used for the 16S rRNA detection. Using the BLAST program, the sequence of purified PCR product was aligned to closely related strains.20 The Neighbor-joining procedure was used to establish a phylogenetic tree. The Kimura-2-parameter model was used to quantify sequence divergences between strains. To deal with gaps, the ‘‘Complete Deletion” option was used. For testing the neighbor-joining method data, Bootstrap analysis (1000 replicates) was selected.21-23
Antibiotic Susceptibility Testing
Antibiotic susceptibility testing was achieved using disc diffusion method (Kirby-Bauer test) with Mueller-Hinton agar (MHA) (Oxoid, UK) as described before.24 The following antibiotic discs (Oxoid, UK) were used: amikacin 30µg, gentamicin 10µg, spectinomycin 100µg, netlimicin 30µg, tobramycin 10 µg, kanamycin 30µg, streptomycin 10µg, neomycin 30µg, rifampicin 5µg/ml, ampicillin 10µg/ml, amoxicillin/clavulanicacid 30µg/ml, clarithromycin 15µg/ml, nalidixic acid 30 µg/ml, doxycycline 30µg/ml, chloramphenicol 30µg/ml, cefotaxime 30µg/ml, cefoperazone 15µg/ml, ceftriaxone 30µg/ml, levofloxacin 5µg/ml and ciprofloxacin 5µg/ml. Each test was done in triplicates, the mean zone of inhibition interpreted as sensitive, intermediate resistant and resistant in accordance with clinical and Laboratory Standards Institute (CLSI)25 The isolates with the highest resistance rate were selected for further investigation.
PCR Detection and Sequencing of AMEs and 16S-RMTases Encoding Genes
Polymerase chain reaction was employed to detect the genes encoding AMEs [ant(4′)-IIb, ant(2″)-Ia, ant(3”)-Ia, aac(3)-Ia, aac(6′)-Ib, aac(6′)-II, aac(3)-II, aac(6′)-aph(2″), aph(3′)-IIIa and aph(3′)VI] and 16S rRNA methylases (rmtB and armA) in aminoglycosides resistant isolates. Primers employed are presented in Table 1. Extraction of DNA was performed in accordance to the instructions of the manufacturer of the extraction kit (Bioneer Company, Korea). PCR cycling was done in VeritiTM thermocycler (Applied Biosystems, USA) with initial denaturation at 96°C for 7 minutes followed by 35 cycles of the following sequence; a denaturation step at 96°C for one minute, annealing at 52°C for one minute, chain elongation at 72°C for one minutes, with final extension at 72°C for 10 minutes. An amplification reaction at a complete volume of 25 μl was formulated using 12.5 μl 2X Master Mix (Thermo Fisher Scientific, USA), 2.5 μl of template DNA (50 pg concentration), 0.5 μM of both primers (reverse and forward) and 9 μl of nuclease-free water. Electrophoresis of PCR products was performed at 100 volts for 40 minutes on a 1.5 % agarose gel buffer, then stained by ethidium bromide dye and viewed using an iBrightTM system of gel documentation (UVtec, UK).
Table (1):
Sequences of primers used for detection of AMEs and 16S-RMTases genes.
| Target genes | Primer sequence (5′-3′) | Product (bp) |
Annealing temp. (°C ) |
Ref. |
|---|---|---|---|---|
| aac(6′)-Ib | F: TTG CGA TGC TCT ATG AGT GGC TA | 482 | 55 | 37 |
| R: CTC GAA TGC CTG GCG TGT TT | ||||
| aac(6′)-II | F: CGACCATTTCATGTCC | 542 | 55 | 38 |
| R: GAAGGCTTGTCGTGTTT | ||||
| aac(3)-II | F: TGA AAC GCT GAC GGA GCC TC | 370 | 57 | 39 |
| R: GTC GAA CAG GTA GCA CTG AG | ||||
| aac(3)-Ia | F: ATG GGC ATC ATT CGC ACA TGTAGG | 465 | 59 | 40 |
| R: TTA GGT GGC GGT ACT TGG GTC | ||||
| aac(6′)-aph(2″) | F: GAAGTACGCAGAAGAGA | 491 | 57 | 41 |
| R: ACATGGCAAGCTCTAGGA | ||||
| aph(3′)VI | F: ATGGAATTGCCCAATATTATT | 780 | 55 | 42 |
| R: TCAATTCAATTCATCAAGTTT | ||||
| aph(3′)-IIIa | F: AAATACCGCTGCGTA | 242 | 55 | 43 |
| R: CATACTCTTCCGAGCAA | ||||
| ant(2″)-Ia | F: ATG GAC ACA ACG CAG GTC GC | 535 | 56 | 43 |
| R: TTA GGC CGC ATA TCG CGA CC | ||||
| ant(4′)-IIb | F: TAT CTC GGC GGC GGT CGA GT | 364 | 59 | 40 |
| R: CAC GCG GGG AAA CGC GAG AA | ||||
| ant(3”)-Ia | F: CATCATGAGGGAAGCGGTG | 787 | 55 | 42 |
| R: GACTACCTTGGTGATCTCG | ||||
| armA | F: CCGAAATGACAGTTCCTATC | 846 | 55 | 42 |
| R: GAAAATGAGTGCCTTGGAGG | ||||
| rmtB | F: ATGAACATCAACGATGCCCTC | 769 | 60 | 42 |
| R: CCTTCTGATTGGCTTATCCA |
A direct sequencing of both strands of pure PCR products was performed for confirmation of the detected genes by the Macrogen Company (Seoul, South Korea). Sequence analysis and alignment were carried out by the BLAST program online (http://blast.ncbi.nlm.nih.gov/Blast), (NCBI).
Isolation rates of bacteria from the clinical samples
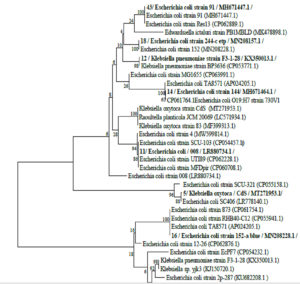
Thirty clinical samples from separate laboratories were used to obtain 49 enterobacterial isolates throughout the study period. The bacterial agents isolated are E.coli [51% (25/49)], Klebsiella [27% (13/49)], Enterobacter species [14% (7/49)] and Proteus species [8% (4/49)]. The high rate of bacterial isolation was from urine specimens at 69% (35/49), then sputum 18% (9/49) and pus 10% (5/49). Among the studied 49 isolates, 9 (18%) had the highest resistance rate. On the basis of 16S rRNA gene sequence analysis, seven of these nine isolates were identified as E. coli and assigned accession numbers as follows: E. coli strain 008 (Accession number LR880734.1), Escherichia coli strain 144 (Accession number MH671464.1), Escherichia coli strain EcPF7 (Accession number CP054232.1), Escherichia coli strain 91 (Accession number MH671447.1), Escherichia coli strain54 (Accession number MH671431.1), Escherichia coli strain152-a blue (Accession number MN208228.1), Escherichia coli strain SCU-103 (Accession number CP054457.1). The remaining two isolates were Klebsiella pneumoniae strain F3-1-28 (Accession number MK824895.1) and Klebsiella oxytoca strain CdS (Accession number MT271953.1). Fig. 2 shows the phylogenetic tree.
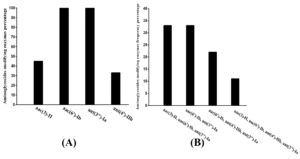
Fig. 1. Distribution of resistance genes in the highest resistant isolates:
A- Percentage of single gene in isolates
B- Percentage of co-existence genes isolates
Fig. 2. The phylogenetic tree of the highest resistance rate of isolates based on 16S rRNA gene sequence analysis. Bootstrap values, shown at the nodes, were calculated from 1000 replicates. The scale bar indicates substitutions per nucleotide. The GenBank accession numbers for the 16S rRNA sequences are given in parentheses after the strain.
Resistance profile of the isolates
Table 2 shows that, forty-nine isolates (100%) were resistant to rifampicin, 44 isolates (90%) are resistant to ampicillin, 33 isolates (67%) are resistant to amoxicillin/clavulanic acid, clarithromycin, 26 isolates (53%) are resistant to nalidixic acid and doxycycline and 14 isolates (29%) are resistant to ciprofloxacin . As shown in Table 3, 36 isolates (73.5%) are resistant to streptomycin, 27 isolates (55.1%) are resistant to Kanamycin, 18 isolates (36.7%) are resistant to gentamycin, 18 isolates (36.7%) are resistant to Neomycin, 16 isolates (32.6%) are resistant to tobramycin, 15 isolates (30.6%) are resistant to Spectinomycin, 4 isolates (8.2%) are resistant to Netlimicin and 2 isolates (4.1%) are resistant to Amikacin.
Table (2):
Antimicrobial resistance profile of Enterobacterial isolates.
No. |
Antimicrobial Agents & Disc potency (µg) |
Susceptible No. |
Intermediate No. |
Resistance No. |
|---|---|---|---|---|
1 |
Nalidixic acid (30) |
20 |
3 |
26 |
2 |
Ciprofloxacin (5) |
31 |
4 |
14 |
3 |
Chloramphenicol (30) |
25 |
1 |
23 |
4 |
Ampicillin (10) |
5 |
0 |
44 |
5 |
Amoxycilin / clavulanic acid (30) |
9 |
7 |
33 |
6 |
Levofloxacin (5) |
33 |
1 |
15 |
7 |
Doxycycline (30) |
18 |
5 |
26 |
8 |
Rifampicin (5) |
0 |
0 |
49 |
9 |
Ceftriaxone (30) |
29 |
1 |
19 |
10 |
Clarithromycin (15) |
4 |
12 |
33 |
11 |
Cefotaxime (30) |
27 |
2 |
20 |
12 |
Cefoperazone (75) |
19 |
10 |
20 |
Table (3):
Resistance profile of Enterobacterial isolates to different aminoglycosides.
No. |
Antibiotic Agents & Disc potency (µg) |
No (%) Susceptible |
No (%) Intermediate |
No (%) Resistance |
|---|---|---|---|---|
1 |
Amikacin (30) |
41 (83.7) |
6 (12.2) |
2 (4.1) |
2 |
Gentamicin (10) |
30 (61.2) |
1 (2.1) |
18 (36.7) |
3 |
Spectinomycin (100) |
29 (59.2) |
5 (10.2) |
15 (30.6) |
4 |
Netlimicin (30) |
37 (75.5) |
8 (16.3) |
4 (8.2) |
5 |
Tobramycin (10) |
29 (59.2) |
4 (8.2) |
16 (32.6) |
6 |
Kanamycin (30) |
12 (24.5) |
10 (20.4) |
27 (55.1) |
7 |
Streptomycin (10) |
5 (10.2) |
8 (16.3) |
36 (73.5) |
8 |
Neomycin (30) |
7 (14.3) |
24 (49) |
18 (36.7) |
Detection of AMEs 16S rRNA methylase encoding genes
PCR analysis and sequencing for the nine isolates that had the highest resistance rate to aminoglycoside, demonstrated the existence of the genes of resistance in each of these isolates (100%). Among those, nine (100%) carried ant (3”)-Ia and aac (6′)-Ib genes, four (44%) isolates carried aac (3)-II gene and three (33%) carried ant (4′)-IIb gene. ant(3”)-Ia, aac(3)-II, and aac(6′)-Ib genes were detected in three isolates (33%). Moreover, three isolates (33 %) were found to contain both ant (3″)-Ia and aac (6′)-Ib genes. Furthermore, ant (4′)-IIb, aac (6′)-Ib and ant (3”)-Ia genes were detected in two (22%) isolates. In addition, aac (3)-II gene was found to co-exist with, ant (3”)-Ia, aac (6′)-Ib, and ant (4′)-IIb genes in one (11%) isolate. The isolates used in the study were negative for armA, rmtB, ant(2″)-Ia, aph(3′)VI, aac(6′)-IIa, aac(3)-Ia, aph (3′)-IIIa and aac(6′)-aph(2″) genes as shown in (Table 4 with Fig. s 1 and 3).
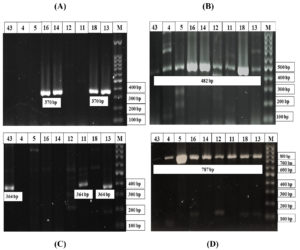
Fig. 3. Representative Image of gel electrophoresis of Polymerase Chain Reaction (PCR) for detecting aminoglycoside modifying enzyme (AME) genes:
(A) Lane M: molecular marker (ladder: 100bp), lane 13, 18, 14 and 16: positive; showing band at 370 bp aac (3)-II gene.
(B) Lane M: molecular marker (ladder: 100bp), lane 13, 18, 11, 12, 14, 16, 5, 4, and 43: positive; showing band at 482 bp aac (6′)-Ib gene.
(C) Lane M: molecular marker (ladder: 100bp), lane 13, 11, and 43: positive; showing band at 364 bp ant (4′)-IIb gene.
(D) Lane M: molecular marker (ladder: 100bp), lane 13, 18, 11, 12, 14, 16, 4, 5, and 43: positive; showing band at 787 bp ant (3′′)-Ia gene.
Table (4):
Distribution of AMEs and 16S-RMTases genes among non-susceptible isolates.
Gene name / Isolate & accession No. |
Strain number |
aac(3)-Ia |
aac(3)-II |
aac(6′)-Ib |
ant(2″)-Ia |
ant(4′)-IIb |
aph(3‘)VI |
aph(3′)-IIIa |
aac(6‘)-aph(2″) |
armA |
rmtB |
aac(6′)-II |
ant (3”)-Ia |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
E. coli (MW599814) |
4 |
– |
– |
+ |
– |
– |
– |
– |
– |
– |
– |
– |
+ |
Kleb. Oxytoca (MW599815) |
5 |
– |
– |
+ |
– |
– |
– |
– |
– |
– |
– |
– |
+ |
E. coli (MW599816) |
11 |
– |
– |
+ |
– |
+ |
– |
– |
– |
– |
– |
– |
+ |
Kleb. Pneumonae (MW599817) |
12 |
– |
– |
+ |
– |
– |
– |
– |
– |
– |
– |
– |
+ |
E. coli (MW599818) |
13 |
– |
+ |
+ |
– |
+ |
– |
– |
– |
– |
– |
– |
+ |
E. coli (MW599819) |
14 |
– |
+ |
+ |
– |
– |
– |
– |
– |
– |
– |
– |
+ |
E. coli (MW599820) |
16 |
– |
+ |
+ |
– |
– |
– |
– |
– |
– |
– |
– |
+ |
E. coli (MW599821) |
18 |
– |
+ |
+ |
– |
– |
– |
– |
– |
– |
– |
– |
+ |
E. coli (MW599822) |
43 |
– |
– |
+ |
– |
+ |
– |
– |
– |
– |
– |
– |
+ |
Total |
0 |
4 |
9 |
0 |
3 |
0 |
0 |
0 |
0 |
0 |
0 |
9 |
Despite the growing resistance rates and several side effects of aminoglycosides, they are still effective antibacterial agents, particularly to treat bacterial infections. Aminoglycosides (gentamicin, amikacin, and tobramycin) resistance percentage was 78% for K. pneumoniae and 33% for E. coli among cerebrospinal fluid and blood clinical isolates in Montenegro, according to the CAESAR Annual Report 2018.26 Phylogenetic tree showed that, isolates of this study were closely related to K. pneumoniae and E. coli. Previous studies have reported antibiotic resistance of E. coli and Klebsiella pneumoniae.27-29 This study was done on Gram-negative isolates (particularly E. coli and Klebsiella spp.), which showed resistance to aminoglycosides as amikacin, gentamicin, netilmicin and tobramycin. In addition, the prevalence of resistance genes was assessed. When the susceptibility of the tested isolates to several aminoglycosides was tested, the high rates of resistance were found for kanamycin (72%) and tobramycin (42%), followed by gentamycin (38%) and netilmicin (24%). These results were in the same line with Eftekhar et al., who discovered that 78.5% of clinical isolates were resistant to kanamycin.30 Another study done by Estabraghi et al. found that clinical isolates exhibited an elevated resistance rate to gentamicin (24%) and amikacin (93%).31 One of the most commonly reported aminoglycoside resistance mechanisms in clinical isolates is aminoglycoside-modifying enzymes (AME).32 In diverse members of resistant bacteria as Acinetobacter baumannii, Enterococcus species,33 Pseudomonas aeruginosa,34 and methicillin-resistant Staphylococcus aureus (MRSA),33 the occurrence of 16S rRNA methylases and AME is becoming increasingly established. Overall, the high rates of aminoglycoside resistance indicate that overuse of these antibiotics in hospitals has resulted in the emergence and spread of resistant isolates. Additionally, the data of the present study emphasize the need for establishing a local and national antimicrobial resistance surveillance system for monitoring the administration of antimicrobials and emergence of antibiotic resistance within the bacterial isolates present in our hospital settings. The knowledge yielded by hospital infection and antibiotic monitoring teams on bacterial resistance can assist doctors in improving antimicrobial management, resulting in more efficient usage of antibiotics. In addition, the current investigation indicated a notable elevation in AMEs prevalence (100%) in clinical isolates that were aminoglycosides resistant. Totally, ant (4′)-IIb (33%), aac (3)-II (45%) and aac (6′)-Ib with ant (3”)Ia (100%) resistant genes were shown to be the most frequently expressed genes among non-susceptible clinical isolates, either in combination or alone. Conversely, an Iranian study found that aac (3) – II as well as aac (6′)-Ib genes were highly expressed in 35.1 % and 42.5% of clinical isolates, respectively.35 Moreover, Liang et al. found that the ant (3″)-I, aac (6′)-Ib, aac (3)-II and ant (2″)-I genes were detected in 13.6 %, 30.2 %, 4.3 % and 19.8 % of clinical isolates in China, respectively.36 Furthermore, Lindemann et al. showed that the aac (3)-IIa gene was detected in 79.3 % while aac (6′)-Ib gene was expressed only in 37.9 % of clinical isolates from their studies, in Western Norway.12 Collectively, these results suggest that aac(3)-II and aac (6′)-Ib, are of great importance among aminoglycoside resistance-inducing genes and are globally significant in aminoglycoside resistant isolates located in various geographic regions.
The current study showed that Klebsiella and Escherichia coli isolated from clinical samples exhibited a high rate of resistance to aminoglycosides. In this study, it was shown that AMEs as well as 16S rRNA methylase encoding genes were positively expressed in E. coli as well as Klebsiella isolates that may be responsible for the aminoglycosides resistance. Moreover, our findings showed that resistance to aminoglycoside was essentially due to AMEs in clinical isolates; with ant (3″)-Ia and aac (6′)-Ib being the most prevalent resistance encoding genes. 16S rRNA methyl transferases do not appear to play any role in aminoglycoside resistance among the studied bacteria. Routine aminoglycoside resistance monitoring and antimicrobial management actions can contribute to the reduction of the distribution of resistant bacteria, and the improvement of treatments.
ACKNOWLEDGMENTS
Authors are thankful to Dr. Ali Mohammed Ahmady, Professor of Microbiology and Immunology, Faculty of Pharmacy, Cairo University, for his assistance in sample collection and Dr. Mohamed Salem Soltan, Assistant Professor of Infectious Diseases, Faculty of Veterinary Medicine, Suez-canal University, for his sincere assistance in genes sequencing.
CONFLICT OF INTEREST
All authors declares that here is no conflict of interest.
AUTHORS’ CONTRIBUTION
MF collected samples, analyzed, performed the microbiological identification, interpreted the results and drafted the manuscript. WB was responsible for the idea and concept of the research, revised the whole manuscript and participated in the supervision of the practical work. EE participated in the design of the study, supervision of practical work, manuscript editing and overall manuscript revision. TD participated in the supervision of the practical work, performed overall revision on the study and all authors read and approved the final version of the manuscript to be published.
FUNDING
None.
ETHICS STATEMENT
The study was approved by the ethics committee at the faculty of medicine, Beni-suef University, NU. Beni-suef, Egypt. The study protocol conformed to the ethical guidelines of the 1975 Declaration of Helsinki.
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the manuscript.
- Diab M, Salem D, El-Shenawy A, et al. Detection of high level aminoglycoside resistance genes among clinical isolates of Enterococcus species. Egypt J Med Hum Genet. 2019;20(1):28.
Crossref - Baldır G, Ozturk Engin D, Kucukercan M, et al. High-level resistance to aminoglycoside, vancomycin, and linezolid in enterococci strains. J Microbiol Infect Dis. 2013;3(3):100-103.
Crossref - Nasiri G, Peymani A, Farivar TN, Hosseini P. Molecular epidemiology of aminoglycoside resistance in clinical isolates of Klebsiella pneumoniae collected from Qazvin and Tehran provinces, Iran. Infect Genet Evol. 2018;64:219-224.
Crossref - O’neill J. Antimicrobial resistance. Tackling a Cris Heal wealth nations. 2014. https://amr-review.org/sites/default/files/AMR%20Review%20Paper%20-%20Tackling%20a%20crisis%20for%20the%20health%20and%20wealth%20of%20nations_1.pdf
- Breijyeh Z, Jubeh B, Karaman R. Resistance of Gram-negative bacteria to current antibacterial agents and approaches to resolve it. Molecules. 2020;25(6):1340.
Crossref - Gad GF, Mohamed HA, Ashour HM. Aminoglycoside resistance rates, phenotypes, and mechanisms of gram-negative bacteria from infected patients in upper Egypt. PLoS One. 2011;6(2):e17224.
Crossref - Shakil S, Khan R, Zarrilli R, Khan AU. Aminoglycosides versus bacteria-a description of the action, resistance mechanism, and nosocomial battleground. J Biomed Sci. 2008;15(1):5-14.
Crossref - Garneau-Tsodikova S, Labby KJ. Mechanisms of resistance to aminoglycoside antibiotics: overview and perspectives. Medchemcomm. 2016;7(1):11-27.
Crossref - Samadi N, Pakzad I, Sefidan AM, Hosainzadegan H, Tanomand A. Study of aminoglycoside resistance genes in enterococcus and salmonella strains isolated from ilam and milad hospitals, Iran. Jundishapur J Microbiol. 2015;8(4):e18102.
Crossref - Ramirez MS, Tolmasky ME. Aminoglycoside modifying enzymes. Drug Resist Updat. 2010;13(6):151-171.
Crossref - Yeo J, Zhou L, Ratnasabapathy Y. Stroke care delivery at North Shore Hospital, Waitemata District Health Board 2014. N Z Med J. 2016;129(1431):67-79.
- Lindemann PC, Risberg K, Wiker HG, Mylvaganam H. Aminoglycoside resistance in clinical Escherichia coli and Klebsiella pneumoniae isolates from Western Norway. Apmis. 2012;120(6):495-502.
Crossref - Haldorsen BC, Simonsen GS, Sundsfjord A, Samuelsen O. Increased prevalence of aminoglycoside resistance in clinical isolates of Escherichia coli and Klebsiella spp. in Norway is associated with the acquisition of AAC (3)-II and AAC (6′)-Ib. Diagn Microbiol Infect Dis. 2014;78(1):66-69.
Crossref - Galimand M, Courvalin P, Lambert T. Plasmid-mediated high-level resistance to aminoglycosides in Enterobacteriaceae due to 16S rRNA methylation. Antimicrob Agents Chemother. 2003;47(8):2565-2571.
Crossref - Xia J, Sun J, Cheng K, et al. Persistent spread of the rmtB 16S rRNA methyltransferase gene among Escherichia coli isolates from diseased food-producing animals in China. Vet Microbiol. 2016;188:41-46.
Crossref - Tada T, Miyoshi-Akiyama T, Kato Y, et al. Emergence of 16S rRNA methylase-producing Acinetobacter baumannii and Pseudomonas aeruginosa isolates in hospitals in Vietnam. BMC Infect Dis. 2013;13(1):251.
Crossref - Birri DJ, Brede DA, Nes IF. Salivaricin D, a novel intrinsically trypsin-resistant lantibiotic from Streptococcus salivarius 5M6c isolated from a healthy infant. Appl Environ Microbiol. 2012;78(2):402-410.
Crossref - Liesack W, Weyland H, Stackebrandt E. Potential risks of gene amplification by PCR as determined by 16S rDNA analysis of a mixed-culture of strict barophilic bacteria. Microb Ecol. 1991;21(1):191-198.
Crossref - Muyzer G, De Waal EC, Uitterlinden AG. Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol. 1993;59(3):695-700.
Crossref - Morgulis A, Coulouris G, Raytselis Y, Madden TL, Agarwala R, Schäffer AA. Database indexing for production MegaBLAST searches. Bioinformatics. 2008;24(16):1757-1764.
Crossref - A Rzhetsky, M Nei, A Simple Method for Estimating and Testing Minimum-Evolution Trees. Molecular Biology and Evolution. 1992;9(5): 945.
Crossref - Kuster E, Williams ST. Selection of media for isolation of streptomycetes. Nature. 1964;202(4935):928-929.
Crossref - Felsenstein J. Confidence limits on phylogenies: A justification. Evolution (N Y). 1985;39:783-791.
Crossref - Patel JB, Cockerill FR, Bradford PA. Performance standards for antimicrobial susceptibility testing: twenty-fifth informational supplement. 2015. http://file.qums.ac.ir/repository/mmrc/CLSI2015.pdf
- Wayne A. Clinical and Laboratory Standards Institute; CLSI. 2011. Performance standards for antimicrobial susceptibility testing. 20th Informational Supplement. CLSI Doc. 2010. https://www.researchgate.net/file.PostFileLoader.html?id=50f0395de39d5e8368000022& assetKey=AS%3A272179910905873% 401441904155623
- Organization WH. Central Asian and Eastern European Surveillance of Antimicrobial Resistance: Annual report 2018. https://www.euro.who.int/en/health-topics/disease-prevention/antimicrobial-resistance/publications/2018/central-asian-and-eastern-european-surveillance-of-antimicrobial-resistance-annual-report-2018-2018
- El-Badawy MF, Tawakol WM, El-Far SW, et al. Molecular Identification of Aminoglycoside-Modifying Enzymes and Plasmid-Mediated Quinolone Resistance Genes among Klebsiella pneumoniae Clinical Isolates Recovered from Egyptian Patients. Int J Microbiol. 2017;2017:8050432.
Crossref - Yoon E-J, Gwon B, Liu C, et al. Beneficial Chromosomal Integration of the Genes for CTX-M Extended-Spectrum β-Lactamase in Klebsiella pneumoniae for Stable Propagation. Msystems. 2020;5(5):e00459-20.
Crossref - Stephens C, Arismendi T, Wright M, et al. F plasmids are the major carriers of antibiotic resistance genes in human-associated commensal Escherichia coli. Msphere. 2020;5(4):e00709-20.
Crossref - Eftekhar F, Seyedpour SM. Prevalence of qnr and aac (6′)-Ib-cr Genes in clinical isolates of Klebsiella Pneumoniae from Imam Hussein Hospital in Tehran. Iran J Med Sci. 2015;40(6):515-521.
- Estabraghi E, Salehi TZ, Aminiq K, Jamshidian M. Molecular identification of extended-spectrum β-lactamase and integron genes in Klebsiella pneumonia. J Nepal Med Assoc. 2016;54(202):72-78.
Crossref - Zembower TR, Noskin GA, Postelnick MJ, Nguyen C, Peterson LR. The utility of aminoglycosides in an era of emerging drug resistance. Int J Antimicrob Agents. 1998;10(2):95-105.
Crossref - Behnood A, Farajnia S, Moaddab SR, Ahdi-Khosroshahi S, Katayounzadeh A. Prevalence of aac (6′)-Ie-aph (2 ″)-Ia resistance gene and its linkage to Tn5281 in Enterococcus faecalis and Enterococcus faecium isolates from Tabriz hospitals. Iran J Microbiol. 2013;5(3):203-208.
- Vaziri F, Peerayeh SN, Nejad QB, Farhadian A. The prevalence of aminoglycoside-modifying enzyme genes (aac (6′)-I, aac (6′)-II, ant (2″)-I, aph (3′)-VI) in Pseudomonas aeruginosa. Clinics. 2011;66(9):1519-1522.
Crossref - Peerayeh SN, Rostami E, Siadat SD, Derakhshan S. High rate of aminoglycoside resistance in CTX-M-15 producing Klebsiella pneumoniae isolates in Tehran, Iran. Lab Med. 2014;45(3):231-237.
Crossref - Liang C, Xing B, Yang X, Fu Y, Feng Y, Zhang Y. Molecular epidemiology of aminoglycosides resistance on Klebsiella pneumonia in a hospital in China. Int J Clin Exp Med. 2015;8(1):1381-1385.
- Park CH, Robicsek A, Jacoby GA, Sahm D, Hooper DC. Prevalence in the United States of aac (6′)-Ib-cr encoding a ciprofloxacin-modifying enzyme. Antimicrob Agents Chemother. 2006;50(11):3953-3955.
Crossref - Gow SP, Waldner CL, Harel J, Boerlin P. Associations between antimicrobial resistance genes in fecal generic Escherichia coli isolates from cow-calf herds in western Canada. Appl Environ Microbiol. 2008;74(12):3658-3666.
Crossref - Arpin C, Dubois V, Coulange L, et al. Extended-spectrum β-lactamase-producing Enterobacteriaceae in community and private health care centers. Antimicrob Agents Chemother. 2003;47(11):3506-3514.
Crossref - Hujer KM, Hujer AM, Hulten EA, et al. Analysis of antibiotic resistance genes in multidrug-resistant Acinetobacter sp. isolates from military and civilian patients treated at the Walter Reed Army Medical Center. Antimicrob Agents Chemother. 2006;50(12):4114-4123.
Crossref - Padmasini E, Padmaraj R, Ramesh SS. High level aminoglycoside resistance and distribution of aminoglycoside resistant genes among clinical isolates of Enterococcus species in Chennai, India. Sci world J. 2014;2014:329157.
Crossref - Hu X, Xu B, Yang Y, et al. A high throughput multiplex PCR assay for simultaneous detection of seven aminoglycoside-resistance genes in Enterobacteriaceae. BMC Microbiol. 2013;13(1):58.
Crossref - Emaneini M, Taherikalani M, Eslampour M-A, et al. Phenotypic and genotypic evaluation of aminoglycoside resistance in clinical isolates of staphylococci in Tehran, Iran. Microb Drug Resist. 2009;15(2):129-132.
Crossref
© The Author(s) 2021. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.