ISSN: 0973-7510
E-ISSN: 2581-690X
The efficiency of anaerobic night soil (human excreta) biodegradation depends on the characteristics of the inoculum used. This anaerobic microbial inoculum (AMI) contains a microbial consortium, including rate-limiting methanogens whose growth depends on various physicochemical and biological parameters necessitating the inoculum characterisation essential for optimum night soil (NS) biodegradation. The inoculum contained 24.567 g/L (±0.07) total solid, 5.022 (±0.57) g/L total dissolved solid, 18.148 (±0.0.058) g/L volatile solid (VS), and 7.0±0.2 pH. The Gas chromatography analysis confirmed the presence of a small amount of volatile fatty acid (VFA), (≤ 4 mM except propionate) showing rapid conversion of VFA to methane, and 55% methane. Further, the presence of methanosaetaceae group of methanogens was detected through Taqman probe-based real-time PCR. The inoculum performance was evaluated for NS biodegradation in semicontinuous mode of feeding in four 2 L anaerobic biodigesters (2D HRT, 4D HRT, 5D HRT, and 10D HRT) with varying hydraulic retention time (HRT) (2, 5, 8, and 10 days, respectively) at mesophilic temperature (35°C). After 5 HRT, VS reduction for 2D HRT and 5D HRT was <40% while for 8D HRT and 10D HRT was >45-50% and >45-60%, respectively. The methane content was <20% in biogas for 2 and 5 days HRT while for 8 and 10 days >45-50% and 50-65% methane, respectively. The % VS reduction and methane content showed the overfeeding of NS in biodigesters 2D HRT and 5D HRT while 8D HRT showed towards optimum and 10D HRT optimum with improved VS reduction and methane content.
Anaerobic Biodigester, HRT, Inoculum, Methane, Methanogen, Night Soil
The microbial biodegradation of human excreta (also known as night soil) in absence of molecular oxygen is an effective and energy-efficient process. The biodegradation of night soil (NS) in anaerobic conditions occurs in a closed vessel known as a biodigester. The biodigester can be operated at mesophilic temperature (35°C) at various places either with low capacity at the family level or with large capacity at a community level, including various public transport of airways, roadways, and railways.1 The biodegradation of NS is a sequential process of hydrolysis, acidogenesis, acetogenesis, and methanogeneis resulting from the microbial interaction catalysing sequential reactions among four major groups of microorganisms.2-4 The formers produce hydrogen and organic acids from the hydrolysis of complex organic materials and later produce methane from its intermediates.5 Thus, the consortium of microorganisms exists and catalyses the long-chain organic waste molecules to simpler carbon dioxide, methane, and water molecules through a series of biochemical reactions. The more reduced metabolites (e.g., propionate, butyrate, lactate, ethanol) accumulated at higher substrate feeding rates, i.e., start-up and periods of overload.6 During this stage, the hydrogen produced after fermentation and acetogenesis fail to consume by hydrogenotrophic class of methanogens.7-13 Feedstock rich in lipid content produces more fatty acids after the hydrolysis of triglycerides. The accumulation of acidic volatile fatty acid (VFA) can shift pH toward an acidic environment, inhibiting methanogen growth and hence methanogenesis.14,15 Thus the methanogenic population is key indicator for organic waste biodegradation.16 This imbalance will allow the growth of another group of hydrogen-removing microorganisms, namely sulfate-reducing bacteria (SRB) and acetogens in the anaerobic systems with inefficient NS biodegradation.11,17 The efficient NS biodegradation depends upon the physicochemical and biological characteristics (specially methanogens) of inoculum and NS strength. The balance between methanogens and NS strength fed to the biodigester plays an important role and If any one of these inclines toward lower (methanogens) or higher (NS) direction will cause an imbalance of ratio and thus lead to failure of the digester. If this imbalance is detected in the early stage, before souring, the digester failure problem can be overcome. Therefore, the presence of methanogens in the inoculum and biodigester is desirable for generating methane (biogas) and energy.18,19 The presence of methanogen becomes more critical during start-up and overload periods. The consortium of microorganisms should contain sufficient levels of methanogens to expedite the start-up and prevention of biodigester failure and maximise energy recovery.7-9 Therefore, for efficient biodegradation of NS and maximum energy recovery in terms of methane, the starting culture, i.e., inoculum, should contain sufficient methanogens along with optimum organic waste fed to the anaerobic digester. The feeding of organic waste to the digester can be done in various ways including biological oxygen demand, chemical oxygen demand, total solid, volatile solid (VS), and hydraulic retention time (HRT) etc.20 The biodegradation of organic waste depends upon the retention time of solid content in the digester. A lower retention time of solid indicates low HRT (overfeeding) while a high retention time indicates higher HRT (underfeeding). The optimum HRT will allow the optimum retention time of NS for optimum NS biodegradation. Therefore the inoculum performance was evaluated for maximum NS biodegradation with fixed VS and varying HRT (2, 5, 8, and 10 days) in a semicontinuous mode of operation. Therefore, the present study focuses on the physicochemical and biological characterisation of inoculum and HRT optimisation of NS biodegradation.
Inoculum and night soil (NS) feed
The cold active anaerobic microbial inoculum (AMI)21 was used as inoculum. Night soil was collected from the local region of Jhansi, India. It was weighed and an equal volume of tap water was added to it (1:1 dilution w/v). It was aliquoted, packed in a polyethene bag, and stored at -80°C in the refrigerator for feeding to the digester. Before feeding, the temperature of aliquoted NS was allowed to bring at room temperature.
Physicochemical characterisation of Inoculum
Total Solid (TS), Volatile Solid (VS), and Total dissolved solid (TDS)
A silica crucible was dried at 105°C in a hot air oven for 1 hour and allowed to cool in an air-tight desiccator. The crucible was weighed (w1) to the constant weight. The inoculum sample of 25 mL was poured into a crucible and was allowed to dry at 105°C for 4 hr and cooled in a desiccator for weighing till the constant weight (w2). The TS (gm/ L) was calculated with the difference in weight (w2-w1) between the dried sample along with the crucible (w2) and the empty crucible (w1).22
The crucible was further put at 550°C in a muffle furnace for 1 hr and was allowed to cool in a desiccator followed by weighing till constant weight (w3). The VS (gm/ L) was calculated with the difference in weight (w3-w1).22 The total dissolved solid was measured through a portable TDS meter. All the experiment of TS and VS was done in triplicate.
pH measurement
Inoculum (100 mL) was taken in a 250 mL beaker along with magnetic bar. It was put on stirrer for mixing and the pH was measured through a pH meter (Metler Toledo, Germany).
Methane estimation from biogas
For the collection of produced biogases, a 10 mL serum vial (filled with water) was sealed with butyl rubber and an aluminium crimp. The biogas was collected in the serum vial through water displacement and kept in an inverted position for methane analysis.
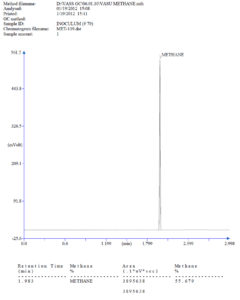
The determination of the methane content within the biogas was performed by GC analysis. For this, 30 µL of collected biogas from a serum vial was injected into a Thermo GC Ultra gas chromatograph with a gas-tight syringe (Hamilton). The GC was equipped with a flame ionisation detector (FID) using Free Fatty Acid Phase-Fused Silica Capillary Column (0.25mm X 0.25 µm X 30 m). The temperature of the column was kept at 50°C in the first 50 sec followed by a linear increase of 4°C sec-1 to 120°C and the final temperature was kept at 200°C with a retention time of 3 min. Nitrogen was used as the carrier gas with a flow rate of 30 mL/ min. The calibration of the GC was performed with standard methane gas (Sigma Gases and Services, New Delhi, India).
Volatile Fatty Acid (VFA) analysis
Volatile fatty acids (VFAs) analysis was performed with a Thermo GC Ultra Gas chromatograph equipped with a Free Fatty Acid Phase-Fused Silica Capillary Column (30 m × 0.25 mm, film thickness of 0.25 µm) and a flame ionisation detector (FID) at 200°C with helium (30 mL/ min) as mobile phase.
1 mL of inoculum sample was centrifuged at 10000 rpm for 10 min. The supernatant was acidified with 3N O-phosphoric acid. 1.0 µL of it was injected at a 1:200 split ratio with the inlet port operated at 150°C and the carrier gas helium at 30 mL/ min. The column oven program started at 60°C and was held for 4 min, then increased to 150°C with a ramp of 10°C/min. Afterwards, the temperature was raised to 180°C with a ramp of 40°C/min and held for 15 min. The comparative standards were a volatile fatty acid (Acetate, Propionate, Isobutyrate, Butyrate, Isovalerate, Valerate, Caproate, and Hexanoic acid) standard mix (Sigma Aldrich) from high-purity single reagents. Data was acquired with Trace GC Ultra software.
Biological characterization of Inoculum
Reference strain
The 16S rDNA of various members of the methanogenic group, namely methanobacteriale (MBT), methanococcale (MCC), methanosarcinaceae (MSc), and methanosaetaceae (MSt), were retrieved from the National Centre for Biotechnology Information (NCBI) gene data bank and were allowed to be aligned. The strains of all four methanogenic groups, namely MBT, MCC, MSc, and MSt were selected (Table 1). The corresponding 16S rDNA gene of each methanogenic group was oligosynthesized (as culturing of anaerobic methanogen is tedious and cumbersome) from Biotek Desk Pvt. Ltd., India and inserted into the pUC57 vector. This oligo-synthesised 16S rDNA gene was used as a methanogens reference DNA for real-time PCR analysis of inoculum.
Real-time PCR analysis
A serial dilution of the oligo-synthesised reference DNA was made from 100 to 105 copy numbers of the gene in duplicate. Quantitative real-time PCR (Cepheid Smart Cycler, U.S.A.) was executed with the serially diluted reference DNA as a positive control (and milliQ sterile water as negative control). It was performed in a 25 µL reaction mixture containing 1000 nM each of forward and reverse primers, 200 nM probe (Table 1), and 25 ng/µL template with Takara Master mix. Amplification was performed in a two-step thermal cycling procedure: predenaturation of 10 min at 94°C followed by 40 cycles of 15 sec at 94°C and 30 sec at 56°C.
The fluorescence and threshold cycle (Ct) values were observed and a standard curve was plotted between the log concentration of gene copy number and Ct value. The minimum limit of detection (LOD) of the gene was also observed for each group. This graph was further used to determine the number of 16S rDNA gene copy numbers in inoculum and night soil genomic DNA isolated by using the commercially available Zymo Research DNA kit along with positive (106 gene copy number of 16S rDNA of reference strain) and negative (sterile Milli-Q water) controls.
Methanogenic Microbial Count of Inoculum
The methanogenic counting of inoculum was done through most probable number (MPN) method. The estimation of culturing of methanogen was done as per the laboratory manual-Microbiological Aspects of Anaerobic Digestion 1988,23 and Hungate.24 Estimation of the number of acetate utilizing methanogens was done by using Medium 25 (in g/L) KH2PO4, 0.3; NH4Cl, 1.0; NaCl, 0.6; MgCl2.6H2O, 0.1; CaCl2 anhydrous, 0.08; Yeast extract, 0.5; Trypticase, 0.5; Trace element, 1.0 ml (Nitrilotriacetic acid, 4.5; FeCl2.4H2O, 0.4; MnCl2.4H2O, 0.1; CaCl2, 0.02; CoCl2.6H2O, 0.17; ZnCl2, 0.1; H3BO3, 0.019; Sodium molybdate 0.01); Trace vitamin, 1.0 mL (Biotin, 2.0 mg; Folic acid, 2.0 mg; Vitamin B12, 0.1 mg, Pyridoxine HCl, 10.0 mg; Thiamine, 5.0 mg; Riboflavin, 5.0 mg; Nicotinic acid, 5.0 mg; DL- calcium pantothenate, 5.0 mg; p-aminobenzoic acid, 5.0 mg; Lipoic acid, 5.0 mg); Sodium acetate-6.8; Resazurin (0.2%) solution (redox indicator)-1.0 mL; Reducing Agent 12.5 mL (Cystein hydrochloride, 2.0 adjst pH 9; Sodium sulphide. 9H2O, 2.0) with N2: CO2 (80-:20) as the gas phase.
Tenfold serial dilution of the well mixed inoculum was made in the sterile dilution medium ((g/L) NaH2PO4.2H2O, 2.964; Na2HPO4, 11.49; reducing agent, 12.5 mL (Cysteine-hydrochloride, 2.0 adjust pH 9; and sodium sulphide.9H2O, 2.0); Resazurin (0.2% solution), 1.0 mL. Dilutions ranging between 10-1 and 10-9 were prepared. One mL of each dilution was inoculated in a replicate of five serum vial bottles (capped with butyl rubber and sealed with aluminum crimp) containing 9 mL sterile liquid medium.25 At least three consecutive dilutions were inoculated. All the inoculated bottles, along with uninoculated control, were incubated at 35°C temperature for 15 days. Prior to inoculation the medium was reduced with sterile anaerobic solution containing: cysteine-HCl and Na2S.9H20. Cysteine was first dissolved in 500 ml distilled water and pH adjusted to 9 with NaOH solution.
At the end of incubation period, the gas from the headspace was analysed for the presence of methane on a gas chromatograph. The methanogenic MPN is computed on the basis of bottles showing positive test.
Hydrolase activity
The hydrolase activity was done through zymography.26-27 For the hydrolase activity, 15 mL of well mixed inoculum was taken in a 15 mL centrifuge tube and it was centrifuged at 10,000 g for 10 min at 4°C. The supernatant was used for the hydrolase (amylase and cellulase) activity after filtering with 0.45 µm filter membrane. The supernatant (20 µL) was loaded in the well of native gel of 10% polyacrylamide gel i.e. zymogram. This was allowed to electrophoresed (Bio-Rad Tetra cells electrophoresis apparatus) and run till the end of native gel. After completion of electrophoresis the gel was submerge for 30 min at 4°C in the substrate (both gel strips were submerged separately in starch and CMC substrate dissolved in 0.05 M Tris-HCl buffer, pH 8) i.e., 2% starch solution and carboxy methyl cellulose (CMC) with gentle shaking. The extracellular hydrolase activity of the enzyme, i.e. amylase and cellulase, was observed with incubation at 35°C with gentle shaking for 1 h. Before staining the zymogram, it was rinsed gently with sterile distilled water. For observation of the hydrolase activity, zymogram was stained with respective iodine solution (5.7% KI, 0.6% I2) for 5 min 26 and 0.1% (w/v) congo red solution for 10 min.27 The excess solution was washed-off gently with distilled water. The hydrolytic zone was observed under Gel Doc Imaging System (Bio-Rad) in visible light.
Biodigester assembly
Two 2 litre flask (biodigester and biogas flask) was taken and fitted with cork and glass rod (L shaped; 15×8 cm each) and interconnected with lazem tube (50 cm). The Biogas flask was fitted with another L-shaped glass rod (30×10 cm) touching the bottom of the flask for water displacement through biogas. Cork and glass rod was made airtight and sealed with Araldite epoxy adhesive.
Biodigester operation
The biodigester operating volume was 2 L. The four biodigesters (2D HRT, 5D HRT, 8D HRT, and 10D HRT) were set up in replicate and filled with 1 L AMI inoculum while the biogas flask was filled with tap water. The four biodigesters (2D HRT, 5D HRT, 8D HRT, and 10D HRT) were operated at varying hydraulic retention times (HRT) of 2, 5, 8 and 10 days with NS feeding, at fixed TS (6.140 g/100mL or 61.40 g/L) and VS (5.333 g/100mL or 53.338 g/L), volumes of 1000 mL, 400 mL, 250, and 200 mL, respectively in fed-batch mode. Initially, biodigester NS feeding was done up to 2 L capacity and later on in semicontinuous mode i.e. an equal volume of the digested slurry was taken out and the same amount was fed with NS. Before and after every feeding, biodigester was mixed for 30 sec. The digestion experiment was allowed to operate at least for 5 HRT and the sample (For TS, VS, and biogas) was collected at each HRT. All the digestion experiment was executed in a controlled environment of a BOD incubator at 35±2°C.
The inoculum was characterized for physicochemically and biologically (for methanogens) and its efficiency was checked after setting up a biodigester assembly at a mesophilic temperature in a BOD incubator. After 5 days, produced biogas was checked and burned with a blue flame.28 This gave the preliminary indication for the presence of enough amount of methane in the biogas and methane-producing methanogen in the biodigester. Further, the methane content in biogas was analysed to 55% through Thermo GC Ultra gas chromatography (Figure 1) showing the presence of a sufficient number of methanogens in the biodigester/inoculum.28 The estimation of acetate utilizing methanogens was done through anaerobic culturing in air tight serum vials. The vials were incubated for 15 days at 35°C for methanogenic activity. The headspace gas of three consecutive dilutions, 10-1, 10-2, and 10-3, (in five replicates) were analysed through GC Ultra gas chromatography. This showed 5,5,4 positive results in subsequent inoculated dilutions. The MPN index table showed the presence of 1600 methanogen (acetate utilizing)/ 100 mL of inoculum at 95% confidence limit.22 The inoculum was also observed for hydrolase (amylase and cellulase) activity through zymography. The 2% starch solution was hydrolysed by amylase and a zone was shown after Iodine solution. A clear hydrolytic zone was showing the presence of amylase activity. Likewise, CMC substrate was hydrolysed by CMCase (cellulase) and a clear zone was visualized after staining with 0.1% Congo red. The hydrolytic zone produced by corresponding enzyme zymogram showed the presence of amylase and CMCase (cellulase) hydrolase activity in the inoculum. The inoculum was characterised physicochemically and showed 24.65 (±0.09) g/L TS, 5.022 (±0.57) g/L TDS, 18.12 (±0.072) g/L VS, and 7.0 (±0.2) pH (Table 2). The VFA mixture (acetate, propionate, isobutyrate, butyrate, isovalerate, valerate, caproate, and hexanoic acid) analysis through Gas Chromatography showed a total 50 mM concentration of VFA mixture. Among these, zero presence of butyrate, valerate, caproate and hexanoic acid and ≤ 4 mM presence of acetate, isobutyrate and isovalerate.29 The absence of volatile fatty acid indicates the complete and rapid conversion of it to methane-containing biogas while the other volatile fatty acid mixture (except propionate) was converted at average speed. On the other hand, Table 2 shows the propionate concentration was more than 4 mM in inoculum indicating a slow consumption rate.6,11-13 The inoculum was characterised biologically for the presence of methanogen at the level of order and family through real-time PCR with corresponding primer and probe set. Table 2 showed the presence of MSt group of methanogen in the range of LOD (Figure 2) while the absence (or <LOD) of MBT, MCC, and MSc group (Figure 2 and Figure 3) of methanogen in inoculum.5,8,9,31-33 In the knowledge of the author, the presented study is the first effort to characterise the night soil adapted inoculum at a biological level.
Table (1):
Primer and probe set used in real time-PCR for quantification of various groups of methanogens
Set name/target group |
Sequence (5’→3’) a |
Amplicon (bp) |
Reference strain |
|
|---|---|---|---|---|
MBT-set/ Methanobacteriales |
F-CGWAGGGAAGCTGTTAAGT T-FAM-AGCACCACAACGCGTGGA-BHQ1 R-TACCGTCGTCCACTCCTT |
343 |
Methanothermobacter thermautotrophicus str. Delta H |
|
MCC-set/ Methanococcales |
F-TAAGGGCTGGGCAAGT T-FAM-TAGCGGTGRAATGYGTTGATCC-BHQ1 R-CACCTAGTYCGCARAGTTTA |
337 |
Methanocaldococcus jannaschii (DSM 2661) |
|
Msc-set/ Methanosaetaceae |
F-GAAACCGYGATAAGGGGA T-FAM-TTAGCAAGGGCCGGGCAA-BHQ1 R-TAGCGARCATCGTTTACG |
408 |
Methanosarcina barkeri strain (DSM 800) |
|
Mst-set/ Methanosarcinaceae |
F-TAATCCTYGARGGACCACCA T-FAM-ACGGCAAGGGACGAAAGCTAGG-BHQ1 R-CCTACGGCACCRACMAC |
164 |
Methanothrix soehngenii, Opfikon (DSM 2139) |
aF, T, and R indicate forward primer, Taqman probe, and reverse primer, respectively.
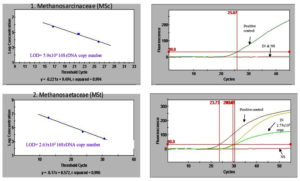
Figure 2. Quantitative real-time PCR of inoculum and night soil, 1) Standard curve of Methanosarcinaceae (MSc) group (left); Fluorescence of Methanosaetaceae with the positive control, inoculum and night soil (right); 2) Standard curve of Methanosaetaceae (MSt) group; Fluorescence of Methanosaetaceae with the positive control, inoculum and night soil. LOD (Limit of detection), NS (Night Soil), IN (Inoculum)
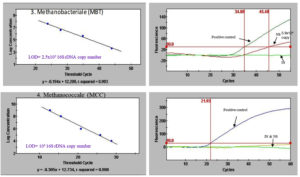
Figure 3. Quantitative real-time PCR of inoculum and night soil, 3) Standard curve of Methanobacteriale (MBT) group (left); Fluorescence of Methanobacteriale with the positive control, inoculum and night soil (right); 4) Standard curve of Methanococcale (MCC) group; Fluorescence of Methanococcale with the positive control, inoculum and night soil. LOD (Limit of detection), NS (Night Soil), IN (Inoculum)
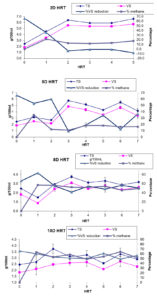
The AMI inoculum21 was used for the optimization of NS biodegradation at mesophilic temperature (35°C) at fixed VS (5.333 g/100 mL or 53.338 g/L) NS feeding in four biodigesters namely, 2D HRT, 5D HRT, 8D HRT and 10D HRT with varying HRT of 2, 5, 8, and 10 days, respectively. The biodegradation of NS was operated at least for 5D HRT. The performance of inoculum for NS biodegradation was evaluated in terms of percent VS reduction and methane content in biogas. Initially, during the first HRT % VS reduction increased rapidly due to the low VS content of inoculum (18.815 g/L) and the high VS content of NS feed (53.338 g/L) while no spent out of slurry was done from the biodigester till the volume of biodigester reach 2 L (biodigester content only 1 L inoculum) with NS feeding. Figure 4 shows that after the first HRT, the biodigester start getting stabilized. In 2D HRT biodigester, after the 2nd HRT the % VS reduction was nil and the methane content of biogas get decreased to < 20%. This showed a failure of 2D HRT biodigester due to overfeeding and the complete removal of methanogens from the biodigester. In 5D HRT biodigester, after 3rd HRT the % VS reduction was < 40% and methane content in biogas <20%. This again shows a failure of biodigester due to overfeeding. In 8D HRT biodigester, after 2nd HRT the % VS reduction was 45-60% and methane content in biogas was 45-55%. This showed that 8D HRT biodigester works optimally and has sufficient numbers of methanogens to produce 60% methane. In 10D HRT biodigester, after 2nd HRT the % VS reduction was 45-60% and methane content in biogas was 50-65%. This shows that 10D HRT biodigester works more optimally and has more numbers of methanogens than either of the biodigester and hence produce more methane (up to 65%). Thus, the inoculum is efficient for night soil biodegradation. Further, 2D HRT and 5D HRT biodigester can be recovered by the inclusion of more methanogens by the addition of more inoculum to the biodigester.16
Table (2):
Physicochemical and biological Characteristics of anaerobic inoculum and night soil
| Parameters | Inoculum | Night Soil (Feed) |
|---|---|---|
| Physical Characteristics | ||
| Total Solid (g/ L) | 24.567 (±0.69) | 61.408 (±3.396) |
| Volatile Solid (g/ L) | 18.815 (±0.58) | 53.338 (±5.132) |
| Total dissolved solid (g /L) | 5.022 (±0.57) | 5.490 (±0.240) |
| Chemical Characteristics | ||
| pH | 7.0 (±0.2) | 6.4 (±0.2) |
| Methane (%) | 55.68 | – |
| Acetate (mM) | 3.620 | 21.239 (±2.20) |
| Propionate (mM) | 42.366 | 24.750 (±3.75) |
| Isobutyrate (mM) | 2.012 | 1.605 (±0.32) |
| Butyrate (mM) | 0.000 | 18.178 (±3.74) |
| Isovalerate (mM) | 4.091 | 1.911 (±0.42) |
| Valerate (mM) | 0.000 | 2.953 (±0.64) |
| Caproate (mM) | 0.000 | 0.000 (±0.00) |
| Hexanoic acid (mM) | 0.000 | 0.000 (±0.00) |
| Biological Characteristics (methanogens) | ||
| Methanobacteriale* | 0.000 | 5.9 x104 |
| Methanococcale* | 0.000 | 0.000 |
| Methanosarcinacae* | 0.000 | 0.000 |
| Methanosaetacae * | 2.75 x 103 | 0.000 |
Figure 4. Biodegradation of four night soil biodigester (2D HRT, 5D HRT, 8D HRT, and 10D HRT) operating at fixed volatile solid with varying hydraulic retention time of 2, 5, 8 and 10 days, respectively at mesophilic temperature (35°C). (TS- toatal solid (g/100 mL); VS- volatile solid (g/100 mL); % VS reduction- per cent volatile solid reduction; % methane- per cent methane; g/L=10x(g/100 mL))
This study shows that 10D HRT biodigester works more optimally and has more numbers of methanogens than either of the biodigester and hence produce more methane (up to 65%). Thus, the inoculum is efficient for night soil biodegradation. Further, 2D HRT and 5D HRT biodigester can be recovered by the inclusion of more methanogens by the addition of more inoculum to the biodigester. The optimized condition along with inoculum can be used for onsite (either small capacity at family or at large capacity of community including roadways, airways, and railways) treatment of night soil.
ACKNOWLEDGMENTS
None.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
Both authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
None.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
Not applicable.
- Han G, Shin SG, Lee J, Lee C, Jo M, Hwang S. Mesophilic Acidogenesis of Food Waste-Recycling Wastewater: Effects of Hydraulic Retention Time, pH, and Temperature. Appl Biochem Biotechnol. 2016;180(5):980-999.
Crossref - Kim J, Shin SG, Han G, O’Flaherty V, Lee C, Hwang S. Common key acidogen populations in anaerobic reactors treating different wastewaters: Molecular identification and quantitative monitoring. Water Res. 2011;45(8):2539-2549.
Crossref - Kawa NC, Ding Y, Kingsbury J, et al. Night Soil: Origins, Discontinuities, and Opportunities for Bridging the Metabolic Rift. Ethnobiol Lett. 2019;10(1):40-49.
Crossref - Park JA, Hur JM, Son BS, Lee JH. Effective treatment of night soil using anaerobic sequencing batch reactor (ASBR). Korean J Chem Eng. 2001;18(4):486-492.
Crossref - Lee C, Kim J, Hwang K, O’Flaherty V, Hwang S. Quantitative analysis of methanogenic community dynamics in three anaerobic batch digesters treating different wastewaters. Water Res. 2009;43(1):157-165.
Crossref - Khan MA, Ngo HH, Guo W, et al. Optimization of hydraulic retention time and organic loading rate for volatile fatty acid production from low strength wastewater in an anaerobic membrane bioreactor. Bioresour Technol. 2019;271:100-108.
Crossref - McMahon KD, Zheng D, Stams AJM, Mackie RI, Raskin L. Microbial population dynamics during start-up and overloading of anaerobic digesters. Biotechnol Bioeng. 2004;87(7):823-834.
Crossref - Griffin ME, Mcmahon KD, Mackie RI, Raskin L. Methanogenic Population Dynamics during Start-Up of Anaerobic Digesters Treating Municipal Solid Waste and Biosolids, Biotechnol Bioengg. 1998;57(3):342-355.
https://doi.org/10.1002/(SICI)1097-0290(19980205)57:3<342::AID-BIT11>3.0.CO;2-I - Angenent LT, Sung S, Raskin L. Methanogenic Population Dynamics during Startup of a Full-Scale Anaerobic Sequencing Batch Reactor Treating Swine Waste. Water Res. 2002;36(18):4648-4654.
Crossref - Chachkhiani M, Dabert P, Abzianidze T, et al. 16S rDNA characterisation of bacterial and archaeal communities during start-up of anaerobic thermophilic digestion of cattle manure. Bioresour Technol. 2004;93(3):227-232.
Crossref - Harmsen HJM, Akkermans ADL, Stams AJM, De Vos WM. Population Dynamics of Propionate-Oxidizing Bacteria under Methanogenic and Sulfidogenic Conditions in Anaerobic Granular Sludge. Applied Environ Microbiol. 1996;62(6):2163-2168.
Crossref - Boone DR, Bryant MP. Propionate-Degrading Bacterium, Syntrophobacter wolinii sp. nov. gen. nov., from Methanogenic Ecosystems. Appl Environ Microbiol. 1980;40(3):626-632.
Crossref - Dhaked RK, Waghmare CK, Alam SI, Kamboj DV, Singh L. Effect of propionate toxicity on methanogenesis of night soil at phychrophilic temperature. Bioresour Technol. 2003;87(3):299-303.
Crossref - Shyan LL, Mat Nanyan NS, Ismail N, et al. Effort to Mitigate Volatile Fatty Acid Inhibition by Using Mixed Inoculum and Compost for the Degradation of Food Waste and the Production of Biogas. Sustainability. 2023;15(2):1185.
Crossref - Sato H, Hirose T, Kimura T, Moriyama Y, Nakashima Y. Analysis of Malodorous Volatile Substances of Human Waste: Feces and Urine. J Health Sci. 2001;47(5):483-490.
Crossref - Negro V, Mainero D, Caredda C, Anedda E, Franchitti E, Traversi D. Analyses of Methanogenic Population as Key Indicator for Anaerobic Digestion: A Case-Study in the Organic Fraction of Municipal Solid Waste Treatment. Waste Biomass Valorization. 2022;14:2461-2468.
Crossref - Briones AM, Daugherty BJ, Angenent LT, Rausch KD, Tumbleson ME, Raskin L. Microbial diversity and dynamics in multi- and single-compartment anaerobic bioreactors processing sulfate-rich waste streams. Environ Microbiol. 2007;9(1):93-106.
Crossref - Singh L, Maurya MS, Ram MS, Alam SI. Biogas production from night soil – Effects of loading and temperature. Bioresour Technol. 1993;45(1):59-61.
Crossref - Zhang X, Jiao P, Wang Y, Wu P, Li Y, Ma L. Enhancing methane production in anaerobic co-digestion of sewage sludge and food waste by regulating organic loading rate. Bioresour Technol. 2022;363:127988.
Crossref - Zhang X, Jiao P, Zhang M, et al. Impacts of organic loading rate and hydraulic retention time on organics degradation, interspecies interactions and functional traits in thermophilic anaerobic co-digestion of food waste and sewage sludge. Bioresour Technol. 2023;370:128578.
Crossref - Verma PV, Vasudevan V, Kashyap BK, et al. Direct lysis glass milk method of genomic dna extraction reveals greater archaeal diversity in anaerobic biodigester slurry as assessed through denaturing gradient gel electrophoresis. Journal of Experimental Biology and Agricultural Sciences. 2018;6(2):315-323.
Crossref - Rice EW, Baird RB, Eaton AD. Standard Methods for the Examination of Water and Wastewater. New York (N.Y.) : American water works association; 2012. http://lib.ugent.be/catalog/rug01:002066269
- A laboratory manual – Microbiological Aspects of Anaerobic Digestion. Edited by Ranade DR, Gadre RV, Yeole TY, Nagarwala NN. Maharashtra Association for the cultivation of science, Research Institute, Pune, India. 1988.
- Hungate Re. The anaerobic mesophilic cellulolytic bacteria. Bacteriol Rev. 1950;14(1):1-49.
Crossref - Touzel JP, Albagnac G. Isolation and characterization of Methanococcus mazei strain MC3. FEMS Microbiol Lett. 1983;16(2-3):241-245.
Crossref - Phetlum S, Champasri C. Purification and characterization of amylases from three freshwater fish species providing new insight application as enzyme molecular markers for zymography. Fish Physiol Biochem. 2023;49(6):1257-1276.
Crossref - Grigorevski-Lima AL, Da Vinha, FNM, Souza DT et al. Aspergillus fumigatus Thermophilic and Acidophilic Endoglucanases. Appl Biochem Biotechnol. 2009:155(1-3):18-26.
Crossref - Hinton N, Stone R. Laminar burning velocity measurements of methane and carbon dioxide mixtures (biogas) over wide ranging temperatures and pressures. Fuel. 2014;116:743-750.
Crossref - Alam SI, Singh L, Maurya MS. Fatty Acids Profile during Anaerobic Digestion of Night Soil-I Effect of Temperature, Calcium Carbonate and Selectively-enriched Inoculum. 1996;46(1):21-26.
Crossref - Raskin L, Stromley JM, Rittmann BE, Stahll’ DA. Group-Specific 16S RRNA Hybridization Probes To Describe Natural Communities of Methanogens. Appl Environ Micrbiol. 1994. 60(4):1232-1240.
Crossref - Hatamoto M, Imachi H, Yashiro Y, Ohashi A, Harada H. Diversity of anaerobic microorganisms involved in long-chain fatty acid degradation in methanogenic sludges as revealed by RNA-based stable isotope probing. Appl Environ Microbiol. 2007;73(13):4119-4127.
Crossref - Stroot PG, Mcmahon KD, Mackie RI, Raskin L. Anaerobic Codigestion of Municipal Solid Waste and Biosolids under Various Mixing Conditions-I. Digester Performance. Water Res. 2001;35(7):1804-1816.
Crossref - Mcmahon KD, Stroot PG, Mackie RI, Raskin L. Anaerobic Codigestion of Municipal Solid Waste and Biosolids Under Various Mixing Conditions- II Microbial Population Dynamics. Water Res. 2001;35(7):1817-1827.
Crossref
© The Author(s) 2024. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.