ISSN: 0973-7510
E-ISSN: 2581-690X
The purpose of this study was to evaluate the antibiotic susceptibility of clinically significant Staphylococcus aureus and its association with biofilm production. The antibiotic resistance pattern and biofilm production by S. aureus isolated from invasive sites such as deep tissue and bone, deep seated pus, blood and other sterile body fluids were studied. The prevalence of multidrug resistant strains and the associated risk factors and co-morbidities were noted. Samples were subjected to antibiotic susceptibility testing using modified Kirby-Bauer disc diffusion method and biofilm production was detected by using microtiter plate assay. Of the total 80 clinically significant invasive S. aureus strains, resistance to penicillin was observed in 70(88.6%) isolates and 38 (47.5%) isolates were resistant to cephalothin. Resistance to erythromycin was observed in 42(52.5%) isolates and 14(17.5%) isolates were resistant to clindamycin. Resistance to ciprofloxacin was 79.5%(n=63). Resistance to rifampicin was observed in 1 isolate (2.1%) and 1 isolate (1.3%) was resistant to teicoplanin. All the isolates were sensitive to vancomycin, tigecycline and linezolid. Out of the 80 S. aureus strains, 21 (26.3%) strains were biofilm producers and 59(73.75%) strains were non-biofilm producers. Among the biofilm producers, resistance to penicillin (57.14%), cephalothin (57.1%) and ciprofloxacin (57.14%) was higher compared to non-biofilm producers. (penicillin 45.7%, cephalothin 35.6% and ciprofloxacin 38.9%,). The rate of MRSA isolated from invasive infections was high (41.25%). We conclude that MRSA and biofilm-producing strains exhibit higher resistance to antibiotics and hence beta-lactams may not be a good empirical antibiotic of choice, especially in biofilm producers. Clindamycin may be an effective alternative substitute to vancomycin forin MSSA and MRSA treatment. Since the patients improved after appropriate antibiotic treatment, we support the role of an early start of appropriate and adequate antibiotic therapy for better patient outcome. We conclude that S. aureus strains exhibited a high resistance to penicillin, β-lactam, macrolide and fluoroquinolones. The rate of MRSA was found to be 41.25%. MRSA and biofilm producing strains exhibit higher resistance to antibiotics. The high prevalence of MDRSA was high (53.75%), which could potentially pose beas a threat to public health, antibiotic use and patient outcome.
Invasive Staphylococcus aureus, MRSA, Antibiogram, Biofilm production, Resistance
Staphylococcus aureus is a commensal as well as an opportunistic pathogen responsible for infections that range from mild skin and soft tissue infection to severe sepsis, toxic shock syndrome and pneumonia1. It is one of the most frequently isolated pathogens in major causes of nosocomial infections (E.g: surgical site infections, pneumonia) and community-acquired infections (skin and soft tissue infections, blood stream infections)2.
Invasive Staphylococcus aureus are pathogenic isolates from sites such as pleural fluid, synovial fluid, cerebrospinal fluid, deep-seated abscesses or bone and blood3.
Methicillin-Resistant Staphylococcus aureus (MRSA) is a major pathogen in nosocomial infections, causing serious problems in hospitals worldwide4. MRSA is of significance due to its resistance to β-lactam antibiotics and its association with multidrug resistance in Staphylococcus aureus5.
MRSA is classified into two types, Community-Acquired MRSA(CA-MRSA) and Healthcare-Associated MRSA (HA-MRSA). CA-MRSA can be defined as the MRSA infection in a person who has not yet recently been hospitalised or has not undergone a medical procedure. HA-MRSA can be defined as the MRSA infection in which the Staphylococcus aureus strain is resistant to several antibiotics and was recovered from patients who frequently visit healthcare facilities6.
Biofilm is defined as clusters of bacteria, lodged in an extracellular polysaccharide matrix7,9. Biofilm formation can lead to an uncompromising infection and poor outcome7. The clinical importance of biofilm infections induced by biofilm producing pathogens relies on in the fact that bacteria in biofilms aid the resistance the action of pathogens to antimicrobial compounds by forming a protective sheath around them and persist despite sustained host defence and hence (Ref) infections caused by biofilm-producing microorganisms express prolonged hard-to- treat infections7. Biofilm formation facilitates increases the mutation rates and thus helps in the spread of antibiotic resistance8. (Ref)Biofilm infections are initially mild but later may act as reservoirs of infection through sloughing8. Such biofilm producing bacteria may be transported to other sterile body sites of the carrier or transmitted to other patients with inbuilt medical devices or immunocompromised state. This would complicate the treatment options, especially in resource limited settings where assays for the biofilm detection are unavailable. Besides, standard in vitro antibiotic susceptibility tests may not be predictive of the therapeutic outcome of biofilm associated infections8.
Staphylococcus aureus and Staphylococcus epidermidis are common biofilm producers. Biofilm formation helps the attachment of microorganisms to biomaterial and protects them from host immune response9. Biofilm producing strains tend to exhibit resistance to antibiotics, disinfectant and germicides10. Hence it is considered to be a virulence factor.
Hence the ability of a strain to produce biofilm needs to be detected in order to give effective treatment due to increasing trends of resistance to antibiotics that is observed in S. aureus strains.
A total of 80 clinically significant Staphylococcus aureus samples were collected from Kasturba Medical College Hospital (Ambedkar Circle and Attavara), a tertiary care center situated in Mangalore, Karnataka, over a period of 6 months from November 2018-April 2019 from invasive sites such as deep tissue, bone, deep seated pus, blood and other sterile body fluids of patients from the hospitals
Collection and processing of specimen
The received clinical samples were inoculated on 5% sheep blood agar, chocolate agar and MacConkey’s agar and incubated for 24 hrs at 37°C. Staphylococcus aureus strains were identified using standard biochemical tests.
Antibiotic susceptibility test for all clinically significant Staphylococcus aureus isolates was performed by modified Kirby-Bauer disc diffusion method on Mueller Hinton Agar (MHA).Antibiotics that was used are penicillin(1µg), cephalothin(30µg), cotrimoxazole (25µg), erythromycin (15µg), clindamycin (2µg), ciprofloxacin (5µg), tetracycline (30µg), linezolid (30µg), vancomycin(30µg), teicoplanin ( 30µg), tigecycline(15µg), rifampicin (5µg) and gentamicin (10µg)5. The results were interpreted according to the Clinical Laboratory Standards Institute (CLSI) guidelines11. Staphylococcus aureus ATCC 25923 was the control strain used5.
Detection of Methicillin resistance
The S. aureus strains were screened for resistance to methicillin by using a cefoxitin(30µg) disk on Mueller–Hinton agar plates. The zone of inhibition was determined after 16–18 h incubation at 37°C. Zone size was interpreted as per CLSI criteria: susceptible ≥22 mm; resistant ≤21 mm11.
Detection of Vancomycin susceptibility using E-Strips
All the MRSA strains were tested for Vancomycin susceptibility using Vancomycin E-test® strips (BioMerieux) and interpreted according to CLSI guidelines11.
Detection of Inducible clindamycin resistance
D- test was conducted to test the resistance to erythromycin and clindamycin according to CLSI guidelines11.
Detection of biofilm production
Biofilm assay was done by using microtiter wells10.
The bacterial colony was inoculated into Brain-Heart Infusion (BHI) broth and incubated at 37°C 24 hours. The cultures were further diluted using fresh medium and adjusted to 0.5 MacFarlands Standard. 200µl of these diluted suspensions were transferred into 96-well polystyrene microtiter plate in triplicate and incubated at 35°C for 24 hours10. Sterile broth was used as blank. The plates were then tapped gently. The contents of the wells were emptied. 200µl of phosphate buffer saline (PBS) was used to wash the wells. Wells were washed 3-4 times to remove free floating bacteria. 100µl of 2% Sodium acetate was added as a fixative. 0.1% crystal violet was used to stain the wells. Deionised water was used to wash off the excess stain. 33% of Glacial Acetic Acid was added to individual wells and optical density (OD)value of the stained adherent biofilm was read using a micro ELISA autoreader at a wavelength of 570nm.An OD value of more than 0.240 was considered as strong biofilm producers, 0.120-0.240 as moderate biofilm producers and less than 0.120 as non-biofilm producers10.
Data collection
Data, including age, sex, duration of hospital stay, history of chronic disease, history of instrumentation or device implantation(the risk factors and co-morbidities) collected from the medical records department were assessed. Bacteriological culture from other sites will be noted to trace the probable source of sepsis. The treatment, outcome in the patients were followed up.
Statistical analysis
p-value was calculated by ANOVA using SPSS v.25 statistical analysis software (IBM Corporation New York, USA). Biofilm formation, drug resistance was compared by Pearson’s chi-square test.
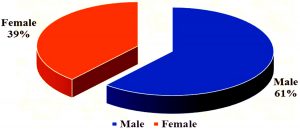
Out of the total 80, clinically significant invasive Staphylococcus aureus isolates majority were skin and soft tissue infections (n=64,80%) followed by bloodstream infections (n=12,15%) and bone infections (n=4, 5%). The study group belonged to the age < 1yr up to 80 yrs. The mean age was 46.8 and as shown in Fig. 1, 31(38.8%) were females and 49(61.3%) were males.
The resistance pattern of 80 Staphylococcus aureus isolates is shown in table 1. All the isolates were sensitive to vancomycin, tigecycline and linezolid. Vancomycin MIC was noted and found to be in the range from 0.5 to 2µg/ml. Of the 33 MRSA isolates,39.4%(n=13) exhibited vancomycin MIC of 0.5µg/ml, 33.3% (n=11) exhibited a MIC of 1µg/ml,12.1%(n=4) exhibited a MIC of 1.5 µg/ml and 15.2%(n=5) MIC of 2 µg/ml.
Table (1):
Antibiotic resistance pattern in Staphylococcus aureus.
Antibiotics tested |
Number of resistant isolates (%) |
|---|---|
Penicillin |
70(88.6%) |
Cephalothin |
38 (47.5%) |
Co-trimoxazole |
12(15%) |
Erythromycin |
42(52.5%) |
Clindamycin |
14(17.5%) |
Cefoxitin |
33(41.2%) |
Ciprofloxacin |
63(79.5%) |
Tetracycline |
4(5.7%) |
Linezolid |
0(0) |
Vancomycin |
0 |
Teicoplanin |
1(1.3%) |
Tigecycline |
0 |
Rifampicin |
1(2.1%) |
Gentamicin |
18(23.4%) |
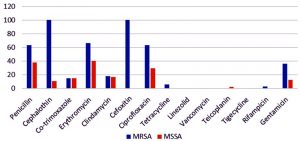
Antibiotic resistance observed in MRSA was higher compared to MSSA and the distribution is as shown in Fig. 2.
Methicillin resistance (MRSA) was observed in 33 (41.3%) isolates and 47(58.8%) isolates were MSSA. Of the 33 MRSA isolates, 5(15.2%) were HA-MRSA infections and 28(84.8%) were CA-MRSA infections.
The distribution of MLSB phenotypes in MRSA and MSSA is shown in Table 2.
Table (2):
Distribution of MLSB phenotypes in MRSA and MSSA.
Organisms |
HA-MRSA |
CA-MRSA |
MSSA |
Total |
|---|---|---|---|---|
iMLSB |
0(0%) |
2(7.14%) |
5(10.63) |
7(8.8%) |
cMLSB |
0(0%) |
3(10.7%) |
3(6.38) |
6(7.5%) |
MSB |
4(80%) |
14(50%) |
17(36.17%) |
35(43.8%) |
CL |
0 |
0 |
0 |
0 |
NR |
1(20%) |
9(32.1%) |
22(46.8%) |
32(40%) |
TOTAL |
5(100%) |
28(100%) |
47(100%) |
80(100%) |
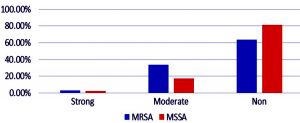
Twenty-one (26.3%) out of the 80 S. aureus strains were biofilm producers and 59(73.75%) strains were non-biofilm producers. Of the 21 biofilm producers, 19(23.8%) strains produced moderate biofilm and 2(2.5%) strains produced strong biofilm. The two strains that exhibited strong biofilm production were isolated from samples that belonged to the extremities of age i.e <1 and 80 years old.
Twelve out of the 21 biofilm producing strains were MRSA and 9 were MSSA. The distribution of biofilm in MRSA and MSSA is as shown in Fig. 2. The p-value is 0.222 and is not significant.
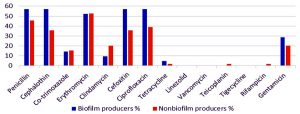
The association of MRSA with biofilm production and antibiotic resistance pattern in biofilm and non-biofilm producers are as shown in the Fig. 3 and 4 respectively. Among the 21 biofilm producers, 12(57.1%) isolates, were resistant to penicillin and 12(57.1%) were resistant to cephalothin . Among the 59 non-biofilm producers, 27 (45.7%) isolates were resistant to penicillin and 21 (35.6%) were resistant to cephalothin. Thus higher percentages of beta lactam resistance were observed in biofilm producers.
In this study, 43 isolates (53.75%) were Multidrug-Resistant Staphylococcus aureus (MDRSA). Extensively Drug Resistant Staphylococcus aureus (XDRSA) was not found in our study.
Table (3):
Risk factors and Co-morbidities observed in patients with Staphylococcus aureus invasive infections.
Risk factors |
Number |
Percentage (%) |
|---|---|---|
Urinary Tract Infection |
13 |
16.3 |
Skin and Soft Tissue Infection (SST) |
10 |
12.5 |
Respiratory Tract Infection |
10 |
12.5 |
Bone Infection |
19 |
23.8 |
CNS |
7 |
8.8 |
Cardiac |
6 |
7.5 |
GIT |
7 |
8.8 |
Dialysis |
2 |
2.5 |
Diabetes mellitus |
38 |
47.5 |
Hypertension |
29 |
36.3 |
Immunocompromised |
11 |
13.7 |
Priorsurgery/ Hospitalisation |
43 |
53.8 |
Neutropenia |
2 |
2.5 |
Hypalbuminaemia |
7 |
8.8 |
Blood transfusion |
7 |
8.8 |
The predominant co-morbidities observed are shown in Table 3. The list of antibiotics used in the empirical therapy are shown in Table 4.
Table (4):
Empirical antibiotics used in the various infections studied.
| Antibiotics given | Infections | Total | |||
|---|---|---|---|---|---|
| Skin & soft tissue | Blood | Bone | |||
| Amoxicillin-clavulanic acid | Count | 14 | 3 | 0 | 17 |
| % | 25.9% | 30.0% | 0.0% | 25.4% | |
| Clindamycin | Count | 19 | 4 | 1 | 24 |
| % | 35.2% | 40.0% | 33.3% | 35.8% | |
| Vancomycin | Count | 5 | 1 | 0 | 6 |
| % | 9.3% | 10.0% | 0.0% | 9.0% | |
| Linezolid | Count | 7 | 0 | 2 | 9 |
| % | 13.0% | 0.0% | 66.7% | 13.4% | |
| Cefuroxime | Count | 3 | 0 | 0 | 3 |
| % | 5.6% | 0.0% | 0.0% | 4.5% | |
| Ciprofloxacin | Count | 2 | 0 | 0 | 2 |
| % | 3.7% | 0.0% | 0.0% | 3.0% | |
| Others | Count | 4 | 2 | 0 | 6 |
| % | 7.4% | 20.0% | 0.0% | 9.0% | |
| Total | Count | 54 | 10 | 3 | 67 |
| % | 100.0% | 100.0% | 100.0% | 100.0% | |
Out of the total 80 patients with Staphylococcus aureus invasive infections, 73(91.3%) patients improved with antibiotic treatment. There was no significant improvement in 5(6.3%) patients. Mortality was observed in 2(2.5%) of the patients.
The obtained results revealed high rates of resistance to penicillin (88.6%), β-lactam (53.2%), macrolide (52.5%) and fluoroquinolone (79.5%). A multicentre study in 2016, by Mendem et al., showed higher resistance to penicillin (97.64%), macrolides (56%), β-lactams (61%) and fluoroquinolones (50%)12.
S.aureus strains were susceptible to linezolid and vancomycin. This is in par with a study by Belbase et al., published in 2017, in which 100% susceptibility was observed towards vancomycin and linezolid thus making it the antibiotic of choice in MRSA infections13.
Results revealed a low rate of resistance to clindamycin (17.5%) in contrast to the study by Al-Zoubi et al., where the isolates showed a clindamycin resistance rate of 81.8%14. Only 1 isolate (2.1%) exhibited resistance to rifampicin which supports the results obtained from a previous study from Kuwait Hospitals where an insignificant increase in rate of resistance to rifampicin from 0.1% to 1.6% , during the period of 2011-2015 was recorded15.
Our study revealed higher percentages of antibiotic resistance in MRSA compared to MSSA. This is in par with a study by Saba et al in the year 2017, where MRSA was found to be more resistant in comparison to MSSA16.
Among the 33 MRSA isolates, the majority (84.8%) were CA-MRSA and the rest as compared to (15.2%) were are HA-MRSA. A study by Huang et al., demonstrated a higher measure of CA-MRSA (45%) among MRSA in their setup17.
This study revealed 26.25% of the Staphylococcus aureus isolates as biofilm producers, classified into 12 MRSA and 9 MSSA. Biofilm production was observed to be higher in MRSA (p=0.222), Signifying no association between biofilm production and MRSA). Higher percentages of resistance to penicillin, beta lactams and ciprofloxacin were observed among the biofilm producers. This was similar to results observed in a study by Khan et al., in the year 2011, wherein biofilm production was higher in MRSA and resistance to all groups of antibiotics were exhibited by biofilm producers18.
The high prevalence of MDRSA (53.75%) observed in our study is in par, which supports the results obtained from a study in 2015 by Bhattacharya et al., where the prevalence of MDRSA was high (57%)5.
Regarding the current study,The common empirical antibiotics used in these 80 patients were, clindamycin (35.8%,n=24), amoxicillin-clavulanic acid (25.4%,n=17), linezolid (13.4%, n=9) and vancomycin (9.0%,n=6). We observed that β-lactams alone were not used as empirical therapy in any of these patients, instead a combination of β-lactam inhibitor antibiotics like amoxicillin-clavulanic acid were used to treat in 25.9%(n=14) of the SSTI.
Clindamycin was used to treat in 35.2%(n=19) of the SST infections, 40%(n=4) of the bloodstream infections and 33.3%(n=1) of bone infections. Macrolides such as erythromycin are inactive against MRSA, due to development of resistance among the strains and hence not recommended for the treatment of SST infections. Clindamycin is a choice, is often administered as a first-line antibiotic to treat SST infections and for MSSA osteomyelitis19. Considering the low rate of clindamycin resistance observed in our study, clindamycin could be used as an effective empirical as well as therapeutic alternative to vancomycin, thus limiting vancomycin use, which would prevent the emergence of VISA and VRSA. Also, clindamycin is effective on MRSA as well as MSSA.
In our study, vancomycin was used to treat SST infections (9.3%) and bloodstream infections (10%). Vancomycin and other gylcopeptides are commonly used in the eradication of S. aureus infections. vancomycin has always been preferred for treatment of MRSA infections. A study by Rayner et al., indicates that vancomycin is effectively used to treat MRSA infection, but not MSSA infections19.
Linezolid was used to treat bone infections (66.7%) and SST infections (13%). According to the study in 2012 by Watkins et al., linezolid was evaluated and considered to be an alternate and effective treatment to MRSA infections due to the appearance of strains that exhibited reduced vancomycin susceptibility20.
From the empirical therapy administered to 67 patients, the antibiotics were changed in 3 patients, in one patient, amoxicillin-clavulanic acid was changed to clindamycin as it was a bone related infection and for the other two patients, clindamycin was changed to linezolid for one as it was an MRSA isolate that was resistant to clindamycin and in the other, vancomycin was changed to clindamycin as it was MSSA infection and significant improvements were observed in the patients after administration .
Mortality observed in our study though low (2.5%, n=2), were secondary to Staphylococcal bacteremia who had not received empirical antibiotic therapy. This supports the role of antibiotic therapy in better patient outcome.
The rate of MRSA isolated from the invasive infections in our study was high (41.25%). Our study supports the view that MRSA exhibited higher resistance to antibiotics. Biofilm producers showed higher percentages of resistance to penicillin, cephalothin and ciprofloxacin compared to non-biofilm producers. Hence, we conclude that MRSA and biofilm producing strains exhibit higher resistance to antibiotics and hence beta-lactams may not be a good empirical antibiotic of choice especially in biofilm producers.
The prevalence of MDRSA in our study was high (53.75%), which could be a threat to the antibiotic use and patient outcome. Owing to the lower rate of resistance to clindamycin, we conclude that clindamycin may be an effective alternative substitute to vancomycin in MRSA treatment.
It was observed that patients improved after appropriate antibiotic treatment. The mortality, though low (2.5%), was observed in those who had not received empirical antibiotic therapy. This supports the role of an early start of appropriate and adequate antibiotic therapy for better patient outcome.
ACKNOWLEDGMENTS
The authors are grateful to Manipal Academy of Higher Education for the support.
CONFLICT OF INTEREST
All authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
GVP: collected the data, performed the experiment and has gathered the literature and drafted the manuscript. ABK has supervised the work, analyzed the data, reviewed the manuscript. SB has supervised the work, analyzed the data and reviewed the manuscript.
FUNDING
None.
ETHICS STATEMENT
The above study has obtained ethical clearance from the Institutional ethical committee.
AVAILABILITY OF DATA
All data sets generated or analysed during the study are included in the manuscript and is saved as Excel sheets .
- RVP, MJ. A Comparative Analysis of Community Acquired and Hospital Acquired Methicillin Resistant Staphylococcus aureus. J Clin Diagnostic Res. 2013;7(7):1339-42.
- Mirzaee M, Peerayeh SN, Ghasemian A-M. Detection of icaABCD Genes and Biofilm Formation in Clinical Isolates of Methicillin Resistant Staphylococcus aureus. Iran J Pathol. 2014;9(4):257-62.
- Jacobsson G, Gustafsson E, Andersson R. Outcome for invasive Staphylococcus aureus infections. Eur J Clin Microbiol Infect Dis. 2008;27:839-48.
Crossref - Palavecino E. Community-acquired methicillin-resistant Staphylococcus aureus infections. Clin Lab Med. 2004;24:403-18.
Crosssref - Bhattacharya S, Bir R, Majumdar T. Evaluation of Multidrug Resistant Staphylococcus aureus and their Association with Biofilm Production in a Tertiary Care Hospital, Tripura, Northeast India. J Clin Diagnostic Res. 2015;9(9):1-5.
Crossref - David MZ, Glikman D, Crawford SE, et al. What Is Community-Associated Methicillin-Resistant Staphylococcus aureus? J Infect Dis. 2008;197:1235-43.
Crossref - Cha J, Il J, Sik J, Chung H, et al. Investigation of Biofilm Formation and its Association with the Molecular and Clinical Characteristics of Methicillin-resistant Staphylococcus aureus. Osong Public Heal Res Perspect. 2013;4(5):225-32.
Crossref - Eyoh AB, Toukam M, Atashili J, et al. Relationship between mutiple drug resistance and biofilm formation in Staphylococcus aureus isolated from medical and non-medical personnel in Yaounde,Cameroon. Pan African Med J. 2014;8688:1-6.
Crossref - Mathur T, Singhal S, Khan S, Upadhyay D, Fatma T, Rattan A. Detection Of Biofilm Formation Among The Clinical Isolates of Staphylococcus: An Evaluation Of Three Different Screening Methods. Indian J Med Microbiol. 2006;24(1):25–9.
Crossref - Bose S, Khodke M, Basak S, Mallick SK. Detection Of Biofilm Producing Staphylococci : Need Of The Hour. J Clin Diagnostic Res. 2009;Dec(3):1915-20.
- CLSI. Performance Standards for Antimicrobial Susceptibility Testing. 29th ed. CLSI supplement M100.Wayne, PA : Clinical and Laboratory Standards Institute; 2019.
- Mendem SK, Gangadhara TA, Shivannavar CT, Gaddad SM. Antibiotic resistance patterns of Staphylococcus aureus : A multi center study from India. Microb Pathog [Internet]. 2016;98:167-70.
Crossref - Belbase A, Pant ND, Neupane B, Baidhya R, Baidya R. Antibiotic resistance and biofilm production among the strains of Staphylococcus aureus isolated from pus / wound swab samples in a tertiary care hospital in Nepal. Ann Clin Microbiol Antimicrob. 2017;1-5.
Crossref - Al-zoubi MS, Al-tayyar IA, Hussein E, Jabali A Al, Khudairat S. Antimicrobial suceptibility pattern of Staphylococcus aureus isolated from clinical specimens in Northern area of Jordan. Iran J Microbiol. 2015;7(5):265-72.
- Udo EE, Boswihi SS. Antibiotic Resistance Trends in Methicillin-Resistant Staphylococcus aureus Isolated in Kuwait Hospitals: 2011-2015. 2017;13110:485–90.
Crossref - Saba CKS, Amenyona JK, Kpordze SW. Prevalence and pattern of antibiotic resistance of Staphylococcus aureus isolated from door handles and other points of contact in public hospitals in Ghana. Antimicrob Resist Infect Control. 2017;6(44):1-6.
Crossref - Huang H, Flynn NM, King JH, Monchaud C, Morita M, Cohen SH. Comparisons of Community-Associated Methicillin-Resistant Staphylococcus aureus (MRSA) and Hospital-Associated MSRA Infections in Sacramento, California. J Clin Microbiol. 2006;44(7):2423-7.
Crossref - Khan F, Shukla I, Rizvi M, Mansoor T, Sharma SC. Detection of Biofilm Formation in Staphylococcus aureus. Does it have a Role in Treatment of MRSA Infections? Trends Med Res. 2011;6(2):116–23.
Crossref - Rayner C, Munckhof WJ. Antibiotics currently used in the treatment of infections caused by Staphylococcus aureus. Intern Med J. 2005;35:S3-16.
Crossref - Watkins RR, Lemonovich TL, Jr TMF. An evidence-based review of linezolid for the treatment of methicillin-resistant Staphylococcus aureus (MRSA): place in therapy. 2012;131-43.
Crossref
© The Author(s) 2020. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.