The contamination of environmental sully with antibiotics is regarded as a major problem today and predictable to attain more recognition in near future. However, human intervention resulting in antibiotic consumption is being enhancing all around the world. Our review of literature revealed the role of microbiome in sully and how antibiotic resistant genes raised. The structure of antibiotics basically influenced by natural components such as biotic and abiotic push which shifts based on different soils. Therefore, management of microbiome in soil and their expression studies were distinctively revealed. The assessment of antibiotic resistance genes with help of next generation sequencing provided a clear comprehension on genome and transcriptome of the bacterial genes. Thus, interaction of microbiome with soil can also be well understood. The current findings in our study will guide every researcher to follow logical protocol in analyzing microbiota composition is covered as well and also to understand its metagenomic and sequenced with next-generation sequencer which helps to comprehend the diverse micro-flora present in soil and its operation. Finally, later progresses in bioinformatics computer program, flow of work, and applications for analyzing metagenomic information are put in a nutshell.
Microbiome, Antibiotics, Metagenomic, Bioinformatics, Next Genome Sequencing
The soil possesses several micro-flora which are intensely imperative for existence on the Earth, involving diverse functions such as nutrients circulation, carbon dioxide, & nitrogen dioxide.1 The soil contains organic matter along with several mineral and nutrients. Nevertheless, microorganisms residing in the soil environment have a key role as agents of biotransformation of organic matter present in soil. The microbes exert their actions based on the physicochemical properties as well as interactions with ecosystem.2 Although, many microbes main habitant is soil which acts as substrate for its growth on the planet. Though, from earlier times traditional methods such as isolation & growth of microbes on agar plates techniques was frequently used and also examination of the total DNA in the soil displayed a massive multiplicity in the composition of microflora.3 In accordance to Dunbar 1999, culture-based methods provides a comprehension on the existence of microflora in soil, i.e if a gram of soil sample has been taken for investigation was found to possess approximately one hundred species. Controversy existed indicating that less than 1 percent of the species however can be cultivated from the multiple cell lines. Several evidences support to these studies.4 However, deoxyribonucleic based approaches displayed the existence of several fauna in soils usually comprising of 110-1050 microorganisms per gram, which corresponds to bacterial species in thousands number.5 Therefore, current knowledge on taxonomy of microbes helps to understand its metagenomic and sequenced with next-generation sequencer which helps to comprehend the diverse micro-flora present in soil and its operation.6,7,8
In this section, we would investigate the literature reports on the study of antibiotics in soil microbiota along with examination of the profusion diversity of bacterial communities in soil ecosystem and their phenotypic and genotypic resistance to antibiotics.
Role of Soil biota in Ecosystem
Our environmental sully is the basis for growth of living organisms as well as provides advancement in the fields of agriculture and forestry. Both agriculture & forestry which supports human beings, mammals to derive nourishment from our ecosystem. However, the role of ecosystem needs to increase in its production.9 Nevertheless, the soil microbiome must be stable, since augmentation of microflora causes damage to soil properties.10 Nonetheless, Antimicrobial susceptibility examination is necessary for dealing with various sorts of bacterial diseases, specifically focusing on present augmented antibiotic resistance species. However, usually utilized techniques are phenotypic resistance assessment by estimating the intensification of bacteria in troupe with antibiotics.11 Although, main purpose of antibacterial susceptibility testing is to foresee the treatment given with antibacterial/antimicrobial agents on a patient’s malady. Therefore, experimental remedial measures persist to be effectual against few pathogenic bacteria but distinct comprehension on the mechanism of action yet not clear.12
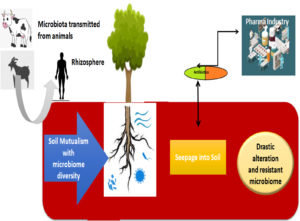
The significant component of earth system is soil which plays a key role in maintaining human health. Since, agricultural crops which provide staple food are grown in soil ecosystem. The plants grown in soil supply essential nutrients, minerals along with trace elements which in turn consumed by individuals.13,14 Microbiota, in soil is based on the physical and chemical & biological perspectives which have revealed to have unswerving influences on individual condition and safety. Although, it’s well acknowledged fact the soil microflora normally exists but agriculturist add certain antibiotics to enhance the growth of their crops. However, certain antibiotics can disturb or cause changes in the microbiome there by affecting the total community. The antibiotics can alter their phylogenetic nature and disturb the ecological functions of micro-flora. Various literature reports have depicted the modifications of microbial ecosystem on addition of antibiotics to the soil ecosystem. (Fig. 1).
Isolation, screening of multi drug resistant microbiome
Consequences that occurred after addition of antibiotics to ecological system along with alteration in their functional activities have also been revealed, involving transformation of nitrogen, biomethanation and reduction of sulphate.15 Several researchers have studied through conventional approaches such as isolation & screening of antimicrobial-producing actinomycetes. However, these traditional methods of isolation and screening helped in discovery of novel antibiotics on soil microflora.16 Nevertheless, due to mutation of microbiota various multidrug-resistant pathogens started to emerge which is insidious to the human population health. Therefore, it has been regarded as a perilous problem for therapy of infectious malady. Elaborate research on ecosystem area is required in order to find unique antibiotics that assist in management of quandary.17
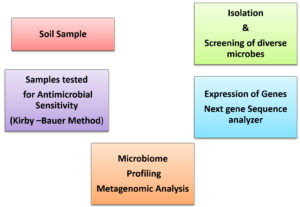
Environmentalists have extensively been fascinated by the enormous modest multiplicity of organisms that exist on planet. Indubitably, comprehension the multifaceted patterns of this diversity, and the prevalent influence to manage them, is a critical focal point in bio-ecology. However, part of ecology always has been in attention and exclusively reviewed intensely with focal point.18 Moreover, essential in comprehending ecosystem through examining the mutual association relating with the microflora and bio-ecology. The environmental science is investigation of life forms in their instinctive existence or locale. Nevertheless, subsistence of microbiome in the soil is considered as asignificant constituent of an ecosystem. Micro-ecological system investigates the bacterial gene expression and mechanism of actions in their native ecosystem.19 Currently, soil microbiome was hindered by defective approaches to examine the vast variety of microbial communities in soil. (Fig. 2).
Antibiotics Resistance
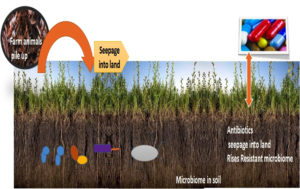
Antibacterial defiance is regarded as principal difficulty in living beings & creature’s currently Moreover, greater antibiotics proportions are emancipated from animal creatures or human body with no discomposure, which relatively meant diverse locale from the man kind to water course or soil environmental which are being contaminated with pharmaceutical substances.20 However, release of medicines after farm handled creatures is piled up in the farm muck and this can in turn used as compost on the plough land.21 The comprehension of antibiotics is very less among the agriculturist as well as poultry man. Therefore, restricted acquaintance on germicidal applications exerted preference for defiance bacterial species in ecosystem.22 Although, administration of medicines in ecosystem were normally minimal but even reduced doses might instigate certain bacterial reactions upon examinations of such reactivity is a focal point of attention. (Fig. 3).
Assessment of Physicochemical Properties
Bonetta23 depicted the survival of microbiome in contaminated sites which specifically relies on intrinsic features such as biochemical, physiological along with its structural properties. Moreover, a genetic modification involves change in morphological trait of cells along with environmental alterations. For instance, elevated quantity of pollution load can influence the measurable calculation of the constituent microbial populace. Antibacterial defiance is most well-known exemplar of rapid modification of bacteria to a latest ecosystem. Nevertheless, the non-pathogenic natures of bacteria are regarded as greatly indispensable as they have extensive function in the waste management area and a broad scope of industrial application. Nonetheless, to handle new variant of organisms which are nonpathogenic in nature with possessions of broad range of exclusive enzymes as well as molecules for industrial application plus degradation of waste. Although, there has been a stable attempt in isolating new bacteria from varied environment. Therefore, this work specifically tends to analyze the physical properties of bacteria which were isolated from the soil ecosystem. In addition, bacterial screening conducted on the basis of enzymatic mechanism and finally checked the antibiotic properties to assure their pathogenic behavior. (Table 1).
Table (1):
Approaches for detection of physicochemical properties.
No. |
Soil Parameters |
Methods for extraction |
References |
|---|---|---|---|
1. |
Oven dry soil |
Gravimetric Method |
Byju, G31 |
2. |
Bulk density, Water holding capacity, Porosity |
Keen Raczkowskii box method |
Wright32 |
3. |
Texture |
International Pipette Method |
Day33 |
4. |
Turbidity ratio |
Turbidimetric Method |
Williams et al34 |
5. |
pH |
1:2.5 Soil: Water Suspension. pH meter |
Page et al35 |
6. |
Organic Carbon |
Chromic acid Digestion Method |
Walkley and Black36 |
7. |
Available nitrogen |
Microdiffusion Method |
Page et al35 |
8. |
Available Phosphorous |
Bray and Kurtz Method, Spectrophotometer |
Bray and Kurtz37 |
9. |
Exchangeable Potassium |
Neutral 1N ammonium acetate, flame photometer |
Page et al35 |
10. |
Labile carbon |
Permanganate Method |
Weil et al38 |
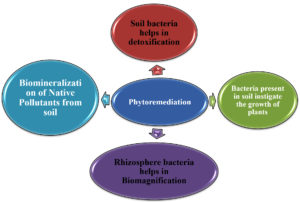
Phytoremediation
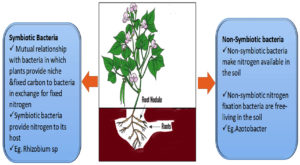
Bacterium present in soil has been utilized for production of crops for many centuries. The major role played by these bacterium according to Davison24 are (1) They provide nutrients to crop plants; (2) They instigate growth of plants, for instance through hormone production; (3) Management and restrain reactions of infectious flora infections; (4) Enhance the configuration of soil (5) biomagnifications/ biomining of inorganics.25,26 However, currently many genetically modified bacilli have been exploited for the bio-mineralization of certain native pollutants, i.e. process is regarded as phytoremediation of contaminated soils.27-30 Modern age for long standing agricultural crops production, associations of microbes with plants in rhizobium participated a fundamental function in mobilization, transfiguration, dissolution of nutrients from a restricted nutritive consortium, and consequently uptake of vital nourishments by flora to comprehend their complete inherited ability. Therefore, utilization of biologic procedures on upsurge for stabilizing and maintaining the physicochemical properties and thereby enhance the yield of crop. In this regard, the role of different microbes has found its potentiality in enhancing the soil conditions for better yield in production of crops.39,40 Nonetheless, broad varieties of symbiotic (Rhizobium sp.) as well as free living bacterium (Azotobacter, Klebsiella, Bacillus, and Azospirillum sp., etc.) are being utilized universally with the purpose of augmenting fecundity and benefit of agricultural crops.28,41 (Fig. 4).
Non-symbiotic and symbiotic bacteria association
The non-symbiotic bacterium in soil provides valuable support for growth of flora are commonly named Plant-Growth Promoting Rhizobacteria (PGPR), which has potential to boost the intensification of rooted plants by colony formation.42,43,44 Although, the PGPR can also be termed as Nodule Promoting Rhizobacteria (NPR) which mainly linked with roots of rhizobium display significant interactions of plant-microbiome in soil ecosystem. However, plant growth is based on rhizobacteria present in soil environment and the association between both plant-microbiome helps to classify these group into two: One Symbiotic bacteria and second is non-symbiotic or also termed as free-living rhizobacteria.45 Nonetheless, symbiotic bacteria reside inside the root nodules of plant cells. These nodules are specialized structures which help in promotion of plants growth. Moreover, the non-symbiotic/free living rhizobacteria do not reside in nodules of plant cells but still induces development of plants.46 Although, supreme microbe for enhancing the development of plants in root nodules are Rhizobium, which is especially available in nodules of leguminous plants. Though, wide variety of microbiome exists in soil environment which intends to enhance the supplements & nourishments to agricultural crops. Bacterium variants like Azorhizobium Rhizobium, Bradyrhizobium, Mesorhizobium, All orhizobium and Sinorhizobium have been effectively & broadly utilized since it allows profitable establishment with soil ecosystem and thereby helps in fixing of nitrogen effectively with support of leguminous plant crop flora.47,48 In contrary, free living nitrogen fixing bacteria like, Azospirillum, Klebsiella & Bacillus species are injected into productive soil specifically to improve productivity in plants.49,50 In addendum, bacterium sp. solubilizing phosphate like of Bacillus & Paenibacillus (earlier called Bacillus) remain concern of study as they particularly improve the phosphorus status of plants.51 Thus, bacteria enhancing its favorable growth in soil on interaction with plant species results in better agricultural productivity. (Fig. 5).
Combined Antibiotics in various maladies
Fish et al52 stated that, on combination of antibiotics the bacterial strains was examined to be resistant to one or both agents. Moreover, synergistic actions were exhibited In-vitro for all antibiotics such as, ciprofloxacin, fluoroquinolones, and levofloxacin which were found to be the most apt drugs for experimental as well as clinical usage, particularly in contradiction to Pseudomonas aeruginosa. The concentration of drug compounds on experimentation reveals the usual dosage. However, Pseudomonas aeruginosa revealed very greater-intensity of resistance to several antibacterial agents and end result is its capability to progress into resistance variants during handling, practical treatment for severe systemic maladies generally included two or more-drug combination strategies. For instance, b-lactam in combination with fluoroquinolone as well as Ciprofloxacin and levofloxacin combinations has received greatest attention as they have apparently displayed higher activity against Pseudomonas aeruginosa by lowering MICs. However, single agent was not successful but the synergic activity revealed better results against various maladies.
The main constrain for antibiotic resistance is due to constant usage of antibiotic agents. It is anticipated that when antibiotics released into ecosystem, soil microbiome undergoes changes in their genetic makeup and acquire resistance to antibiotics. The pharmaceutical company’s manufacture a large number of drugs and release their effluent into ecosystem and even agriculturist use antibiotics to prevent infection during growth of plants. Since, broader existence of antibiotics, and their in appropriate utilization and discarding modifies the soil characteristics as well as the bacterial characteristics. Around the 20th century globally India was the huge antibiotics consumer for human health at approximately (10.7 Units /person). Moreover, next biggest customers were China estimated (7.5 Units/person) pursued by United States at (22.0 Units/person). However, 75% augmentation in world wide antibiotic consumption was involved in the years of 2000 – 2010, due to BRICS nations. Consequently, among these nations, twenty three percent of the upsurge in retail sales of antibiotics in India, & further 57% of augmentation displayed in the medical sector in China (Laxminarayan et al).53 Therefore, antibacterial/antimicrobial medicines used for inhibitory or therapy reasons in agricultural, human, veterinary rationale and also concentrates on survival as well as spread of resistant varieties (Gebreyohannes et al).54 Mostly antibiotics were utilized for agricultural purpose which accounts at least half in United States of America (Lipsitch et al).55 Mainly, the dumping of animal waste and further excessive use of fertilizers results in growth of resistant bacterial species in soil ecosystem. This happens probably through modification in their genetic makeup during evolution where there is lateral gene transformation which results in obdurate antibiotic resistance among microbiome in soil. On the other hand, cattle, poultry, pig hoist with antibiotics protect them from vital populations microbiome, which are communicated through direct contact from animals to humans through their meat, meat products, eggs, milk etc. (Marshall et al and Santajit et al).56,57
Release of Pharmaceutical Products disturbs soil Ecosystem
The pharmaceutical products dosages in runoff, manure, muck, biological-solids, and soil display huge variations which in turn relies on the kind of drug, their metabolism in animals. However, extent of therapy time is associated with the administration time. Although, tetracyclines medicines have greater concentrations and are most commonly exhibited pharmaceutical substance remains in compost.58,59,60 The next set medicinal substances which are present in moderate degrees in compost are fluoroquinolones61,62 & sulfonamides.63 Amongst, macrolide pharmaceuticals, greater degree in compost were estimated for tylosin (Dolliver).64 Nevertheless, in comparison to compost, biological solids possess very lesser quantity of pharmaceuticals (Jones-Lepp & Stevens).65
Multiplicity of microbiome along with profusion of antibiotic resistance genes (ARGs) were greatly manipulated by horticulture.66,67 It has been acknowledged that dispersion of compost from animals administered with various medicines on paddocks influences their quantity as well as multiplicity of antibiotic resistant genes.68,69 For existence of antibiotic resistance genes, management of soil conditions is essential such as the organic matter content, pH, temperature, these are significant factors influencing the growth antibiotic resistant genes in the sully ecosystem.70,71 Genes which revealed defiance to chloramphenicol, streptomycin, minocycline, gentamycin, amikacin, kanamycin, rifampicin, and tetracycline in sully subjugated with compost which shared for approximately seventy percent of the total antibiotic resistant genes. Moreover, greater than half of the antibiotic resistant genes revealed lesser identity, <sixty percent, when sequenced with closest proteins in GeneBank.68 Amongst, antibiotic resistant genes examined (tetO, tetW, ampC, and ermB), standard profusion of ampC & tetO in the compost-administered sully was 421 & 3.3% higher in contrast with an untreated controlsully, correspondingly.69 Broad range of antibiotic resistant genes outlined & instituted among five rice paddy sully around South China.72 Nevertheless, database of antibiotic defiancegenes were performed with next-generation sequencing, total of 16 varieties of antibiotic resistant genes were identified from 110 antibiotic resistant gene sub groups respectively. However, synergistic resistance of genes exhibited in all soils up to 47.5%, specifically resistant to genes for bacitracin, acriflavine, macrolide-lincosamide-streptogramin. (Fig. 6).
Examination of Bacterial Genome
The study reports on providence of antibiotic resistant genes in sully ecosystem concerned with not only examining the multiplicity along with profusion genes of our interest, yet also to find latest antibiotic resistant genes, and explained latest resistance methods and unique enzymes accountable for defiance of bacteria to pharmaceutical products.73,74 However, examination of bacterial genome and comprehend the mechanism of action, functional metagenomics provides support in investigation of antibiotic resistance genes which helps to identify various tetracycline defiance genes from diverse grades in sully. Moreover, twenty-one efflux pump genes facilitator super family predominant in the sequence libraries which were obtained from examined soils. Additionally, two more novel genes engaged in enzymatic deactivation of tetracycline were recognized.74
Torres-Cortes,75 explained unique kind of dihydrofolate reductase protein which was approximately 27k Dabestowing defiance to trimethoprim in soil microbiota. However, with functional meta transcriptomics analysis recognized three latest antibiotic resistance genes providing defiance to ampicillin, gentamicin, chloramphenicol, & trimethoprim from gene libraries produced from 3diverse sully samples. Nonetheless, 9-carbapenem-hydrolizing metallo-beta-lactamases (MBLs) enzymes along with 7 more new enzymes displaying 33 to 59% identity to earlier explained MBLs. However, from six proteobacteria the beginning of resistant communities originated. Two were from Bacteroidetes, & Gemmatimonadetes one. Mostly MBLs were observed more commonly and established in greater diversity among grassland in contrast to tilled soil.76 Examination on Canadian soils by functional metagenomics the resistome of bacterial communities were greater, thirty four novel ARGs with greater similarity to determinants bestowing defiance to sulfonamindes and aminoglycosides with wide spectrum of beta-lactams was recognized.73 Greater-resolvation proteomics associated with functional metagenomics led in identification of unique proline-rich peptide PPPAZI which promote defiance for several macrolides, but not to other ribosome-directed medicines.73 However, Donato77 recognized 2 new dual functional proteins which were accountable for bacterial defiance against ceftazidime & kanamycin, correspondingly.
Zhou et al,78 examined heavy metals performing as a significant feature subscribing to carry on antibiotic resistant genes in the soil ecosystem. Cattle farms were investigated for antibiotic resistance genes exhibited imperative relationship between the plethora of antibiotic resistant genes & metal resistance genes. In dairy farm specifically the quantity of heavy metals such as Cu & Zn in cattle feces were investigated. The heavy metals functions could be determined by immunologic-defiance for bearance of bacillimore than one antibacterial compound.79,80 Nonetheless, expression of cooperative-resistance, positioning for antibiotics defiance genes along with heavy metals defiance encoding genes and other antimicrobial vehicles on selfish genetic elements.79 However, combined-drug outflow drives intervenes speedy ejection of antibiotics along with heavy metals from cell dwindle the vulnerability toward these agents, are instances for immunogenic-defiance.81 However, cross-defiance happens during assortment of DNA nearby on chromosome or else extra chromosomal constituent, which enhances probability of genes to instigate for synergistic transmission via Horizontal Gene Transfer. On the other hand, heavy metals triggers cooperative-selection procedure, but also augment the for bearance levels to pharmaceuticals due to defiance genes management.80 Nevertheless, mechanistic activity might influence the proliferation of antibiotic resistant genes from environmental populace of bacteria with clinical significance even in non-existence of direct pharmaceutical selection.82
Ecological habitats of soil in diverse locations
Dezfully et al,83 mainly examined kochi variety crops and isolated antibiotic generating microorganisms from the rhizosphere zones in Kochi. Soil possessing nitrogen fixing bacteria has chosen for sampling because; microbial communal is relatively abundant in sully in contrast to other ecosystem. However, rhizobial bacterium manifests greater degree of adversary actions. Various scientists have opt for sully which possess new antibiotics and can act as a source of several pharmaceutical manufacturing bacteria involving Actinomycetes84-88 also displayed heterogeneity of soil is outcomes of broader variety of ecological habitats and existence of greater soil microbial variants. However, these results were correlated with our collected sample procedures and their implementation techniques for sample collections in diverse locations. The Random sampling method was described by Williams and Vickers,89 was a traditional methodology for collection of sample; this procedure supports the current study. In order to identify the morphological attributes Grams staining procedure was followed, which is a traditional method conferred by several scientists.90-92 Therefore, further comprehension of its morphological characteristics appropriate staining of soil bacterial isolates with Gram’s staining method showed that mostly bacterial isolates ascertained gram positive. However, outcomes regarded as confirmatory for Wadetwar and Patil,87 stated that most of the soil microbiome after testing found to be gram positive.
Bull et al,93 claimed that most of the indigenous products have unique configuration, they continue to be main assured resource for secondary metabolites and further function as antimicrobial agent in contradictory to pathogenic bacteria.94 However, indigenous products retrieved from microorganisms even now considered as most promising resource for future antibiotics.95 Amongst the diverse uncharted niche, sully is regarded as one of the most appropriate domain for intensification of microbes. Cavalcanti96 isolated the microorganisms from the soil ecosystem which is the foremost source for antibiotic detection. Soil along with plant-connected ecological system embraces many bacteria that release metabolic antibiotic with precise or wide-range actions contradictory to combined microbiota depending on the nutritive abundance, pH, & content of humus. However, actions & multiplicity of life forms existing in soil are specifically regulated by two factors like abiotic & biotic factors. Nevertheless, major abiotic factors are temperature, climate, moisture, structure of soil, texture of soil, pH, and salinity. The atmospheric states further manipulates the physiology of soil microbiome as it diverge transversely in entire globe or, in the same niches, in-between climatic periods. Although, it is obvious that the ecological soil system is abundant with diverse microbiome which has the potential to synthesize antibiotics. However, the regularity of their existence and capability to synthesize happens at significantly different ecologically levels and much clarity is missing. Nonetheless, conventional steps taken of randomly done screening was carried over for past fifty years to generate novel antibiotics preferably for human kinds. Therefore, current study aimed in isolation, characterization of soil microbiome from diverse regions from India.
Broad Spectrum antibiotics
Soil microbiota produces a broad spectrum of specialized metabolites, comprising antibiotics. These bioactive compounds might be chiefly utilized in interaction and antagonism activity with other organisms, along with adaption to ecological alterations.97 Microbiota linked specialized metabolites involves 2 main relative products: peptide secondary metabolites and oligopeptides. These two peptide families explain that most antibiotics, antifungals, antibacterial, along with other antimicrobial agents that were distinguished from bacterial cultures.98,99 However earlier in era of 1940s to 60s, several pharmaceuticals were primarily isolated by examination of streptomycetes from sully for antibacterial actions, involving streptomycin, which was found in 1952 by Winner Selman Waksman, who got Nobel Prize along with his colleagues. Currently, the introduction of genomic and molecular approaches has created renaissance of curiosity in isolating and screening of pharmaceuticals from the environmental sully microbiome. Today, with help of high-through put genomic tools which are now being utilized for investigation of bioactive agents.100-101
The current study has displayed that B. amyloliquefaciens has ascertained to effective resource of pharmaceuticals in contradictory to flora pathogens102-107 and is chiefly connected with plant rhizosphere108 and hence offer sturdy support for our examination. Together with Bacillaene formed by B. amyloliquefaciens has exhibited to be effectual against pathogens like as Serratiamarcescens, S.aureus and K. pneumonia.109 In our findings, B. amyloliquefaciens have resulted in maximum amount of inhibition zone in contrast to S. aureus. Likewise, B. amyloliquefaciens also generated turin A which is effectual in contradictory to fungal pathogens like Rhizoctonia solani.110 Moreover, Bacilysin and Difficidin has displayed antibacterial actions in contrast to Xanthomonas oryzae rice pathogens.111 Nonetheless, B. amyloliquefaciens has been utilized as an antibacterial compound against dairy & veterinary bacterial pathogens in animals.112 Therefore, our study reports as certain and provide sturdy support that B. amyloliquefaciens could function as a potential source of antibiotics.
Molecular level Investigation
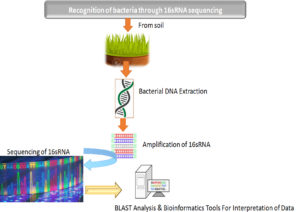
The molecular level of recognition of bacteria was conducted through sequencing & amplification of 16SrRNA. An imperative gain of this procedure is that isolation, characterization, as well as recognition of bacteria in samples takes place within two to three days in contrast to traditional biochemical test, which commonly takes many weeks. Various reports from literature suggested that 16S gene sequence of rRNA investigation was considered as advanced approach for recognition of bacteria in comparison to phenotypic methods.113-114 The consequent sequencing and further using BLAST tool for examining on collected sample as well as stored data of Bacillus amyloliquefaciens. These out comes depicted sturdys caffoldas in earlier investigations which already have confirmed Bacillus as the most pre-dominant bacteria existing in soil.115-116 Thus B.amyloliquefaciensim part as turdy sustenance to the earlier literature reports which have as certained B. amyloliquefaciens as a potential source for antibiotics. Although, Boottanun et al117 has confirmed that B. amyloliquefaciens has the potential to create many peptides of antimicrobial nature along with other secondary metabolites, which bestow broad insights to our results. However, Vijayalakshmi et al102 also confirmed and reported that B. amyloliquefaciens results in utmost proteins (antimicrobial) which displayed greater effect against gram positive S.aureus. It is well acknowledged that bacteria which are gram-positive are more prone to antibiotics since it has an outer layer of peptidoglycan that doesn’t act as an effective blockade. (Fig. 7).
The soils possess their own flora and fauna which generate their own antibacterial metabolites. For instance, diverse antibacterial peptides have also been extracted from tissue of earthworm. 118-119 In some cases, for development of antibiotics genes might be transferred transmitted from bacteria to their hosts. Nonetheless, beta lactam synthesis alleyways have been examined in soil dwelling insects. Moreover, this pathway is not present in densely associated insects. These astounding investigation has been as certained by genomic, phylogenetic and transcriptomic analyses which sturdily recommends horizontal gene transfer from a bacterial contributor in a innate ancestor.120-122
Use of metagenomics to monitor the environment and biobanks for antibiotic resistance
The survival of antibiotic-tolerant pathogens is an alarming situation globally. For example,a gene blaNDM1 confers tolerance against carbapenems and other β-lactam antib Escherichia coli, iotics.123 blaNDM1 was first discovered in India in 2008. (Johnson.) and Woodford 2013. Eventually, various strains containing blaNDM1, such as Klebsiella and Escherichia coli, have been related to a vast geographic area that includes all continents except Antarctica and South America. Related to the outbreak in the hospital.123
In addition, blaNDM1 was found in drinking water samples from the Indian subcontinent. Walsh et al.124 Although the clinical relevance of blaNDM1 is not unique, it is a worrying instance for the rapid spread of ARG in the environment and affects the clinical outcome of human diseases. Metagenomics revealed that blaNDM1 in hospital effluent is a hot spot for ARG. 43 A significant feature of indulging and minimizing the outbreak of antibiotic tolerance is the investigation and monitoring of ecological pools of contagious agents and the accumulation of ARGs. Human microbiota is often exposed to and affected by ecological microbes.125 These microbes are a potent reservoir for ARG, which can be conveyed to pathogens. To expect the occurrence and frequency of ARO and ARG, researchers need to illustrate and calculate the prevalence of ARO and ARG in the environment and their effects on pathogens. Though many locations can serve as pools for contagious sources, it has recently been emphasized that host-related symbioses and reservoirs are particularly important for monitoring.126-127
Metagenomic parameters
Allow long-term and monitoring of rapid deposits that can be focused and confirmed using culture-based approaches. Pets represent a potentially hazardous organic tank for rims and topics due to high threat and the comparative affluence of zoonotic spread to humans.128-129 Antibiotics to both non-therapeutic and therapeutic concentrations are widely used in area of livestock (for example, poultry, cattle, pigs), with an overall annual applications that exceed the use of human antibiotics. The remaining antibiotics compounds and related metabolites are excreted out in manure, waste and sewage.130-131 The metagenomic monitoring of animal dumping products shows an enrichment of bacteria allied with these pet tanks. In 2015, an E. coli surveillance project in pigs in China identified the origin of the resistance colistin gene, MCR1. Colistin is a ballast antibiotic and, prior to the finding of Mcr1, resistance to colistin occurs due to mutations in chromosomes. Since locating MCR1, it is a part of food and animal sources, as well as in competitive infections.132 Monitoring enrichment in animal tanks can help in searching out growing pathogens. Although, it is unclear to what extent the rings or grounds in animal tanks contribute to pathogens and human infections, and further studies are necessary to quantitatively assess the involved risks. Water supplying plants harbor a great variety of microbial organisms and may be contaminated with pathogens that will be classified as urgent or serious antimicrobial threats.126
Because many of the ARGs obtained are placed on conjugative plasmids that can move on their own between different types of bacteria, the aquatic environment provides a reservoir in which ARGs can be exchanged between benign bacteria and human pathogens.126 Wastewater receives various discharges.133-134 These releases include antibiotic residues used by humans and animals, which will increase the selective pressure on the ARG native to the microbial community in the water. Although wastewater is treated to eliminate coliforms, the treatment may not be effective in destroying ARG, resulting in ARG accumulating in living sewage microbial samples downstream of the sewage treatment plant, indicating that wastewater treatment and its disposal are the main reservoir of resistance process in the environment.135 A new report utilized metagenomics to all the more likely comprehend the recurrence of ARO and ARG in the Chaobia River (the primary wellspring of drinking water).136 Metagenomic examination shows that the Chaobia Stream has expanded ARG transmission contrasted and different regions in the district and regions without human collaboration (like remote ocean and Antarctic silt). The primary ARGs found are identified with anti-infection agents generally utilized in human medication, veterinary treatment, and animals.136 The increase of ARGs in foremost water resources may be accompanied by AROs and AROs. The ARG in human drinkable products directly affects millions of people. Metagenomic investigation of savoring water at 25 urban communities on three mainlands uncovered an assortment of ARG types in human drinking water supplies. Most examples have higher ARG concentration than natural silt and soil samples.137 Broadly, these studies demonstrate the pervasiveness of ARG, the incidence of AROs and ARGs in water supplies, as well as the significance of understanding the source and distribution of AROs and ARGs in drinking water. ARO and ARG environmental libraries will become a vicious circle if they are not monitored and spreading ARO and ARG will increase over time and long distances.
Sequencing Technology of Novel Antibiotics and their Resistance Mechanisms
As sequencing technology has advanced, the detection of different lead antibiotic combinations and drug tolerance mechanisms has augmented.138-140 With these advances, databases have expanded with assembled genomes, annotated ORF, and metagenomics, making sequencing something of an emerging. makes pathogens and ARG monitoring methods.141 Monitoring of these harmful bacteria or ARGs, however, is only component of the clinically significant advantage that microbial sequencing could provide. This section describes multiple techniques for discovering antibiotics and the mode of acquired resistance, as well as providing novel clues into previously characterized mechanisms.
Study of different isolates
WGS analysis of bacterial isolates is currently mainly done through vastly multiplexed short-read sequencing technology (such as Illumina, San Diego, CA). You shall formerly use tools such as Bowtie 2.69 to map the reads with reference genome database. This process is valuable since it is quicker and computationally cost-effective than de novo genome assembly.142 If the bacterium is underrepresented in database, or the ARG is a new or deleted homolog, the hit will be unrecognized.143 Furthermore, due to the short read lengths used in certain WGS analyses (500 bp), a few samples may incorrectly coincide with different genes due to local sequence homology and repetitive DNA144. A further effect of low read lengths is the lack of the genetic context around a specific ARG. This is especially concerning for multidrug resistant studies since it decreases the capacity to identify ARG transmission through co-linearity with transposons and plasmids.
Novo set refers to the assembly of short reading genomes with continuous segments (contigs) without the assistance of a reference genome. A new revision examines the well-known genome assemblies. Many of the reading task’s difficulties are outweighed by the capacity to examine huge contigs. The arguments may be detected in the perspective of their neighboring genomic material using the mounted information, and the genomes of species that share the similarity of the low sequence with reference sequences can be examined. This approach, however, has draw backs. Readings assembly is computationally expensive, necessitates high sequence coverage in terms of reading allocation, and its authenticity and accuracy might be undermined by sequencing mistakes and repetitive DNA components.143 An exciting progress in this sector is State the development of LONEAD sequencing technologies such as Singla molecule in real time sequencing and the sequence of nanopore145,146 as an alternative of readings of 50-150 BP characteristically created by technologies as illuminated, these approaches yield readings of 10 KB to more than 100 KB.145-146 One significance of having lengthier readings is the ability to investigate antimicrobial resistance. Bacterial isolates considerably simplify the Novo Assembly question and can even construct a whole genome of bacteria.146 By far the most interesting advancement of these tools is that they generate data significantly faster than the previous technologies.147 Nanopore sequencing, specifically Excel, produces fast data even without sophisticated and specialized equipment, making it a perfect foundation. As a proof of the concept, Bradley et al148 showed that Staphylococcus aureus antibiotic resistance profiles could accurately predict the use of nanopore technology in just 7 hrs
Analysis of metagenomes
The WMS study of microbiota is more complex than the WGS investigation on its alone. Because the WMS data includes information from an unspecified number of creatures, the sequence coverage and depth of numerous species vary, making de novo synthesis a difficult process.149 In a recent review, common methods of metagenomic assembly were compared.150 Once reassembled from WMS, the genome may be reported on and studied in the very same way. The use of NGS to test organisms and metagenomics has ramped up drug discovery process by identifying novel antibiotics and giving data for the design and synthesis of medicines that overcome multidrug resistance. Automated monitoring methods can use genome data to find the genesis and distribution of antibiotic resistance genes (ARG) and develop insights into the evolutionary history of the emergence of particular tolerance mechanisms. It can be accomplished through the use of sequence-based calculating techniques (as mentioned above) or function-based approaches like functional metagenomics. Antibiotic and Secondary Metabolite Analysis Dish (antiSMASH) has been created, which incorporates an HMM-based approach to detect possible BGC.151 In addition, anti-SMASH numerous additional techniques, including as active site prediction, domain analysis, and chemical structure prediction, are also implemented. Anti-SMASH can swiftly detect and describe a wide range of BGCs, including non-ribosomal peptide synthetase and polyketide synthase families. Using this method, investigators discovered over 3,000 small molecule BGCs by using the NIH Human Microbiome Project’s 752 metagenomic groups.152 As prototype, the investigators isolated and characterised an anticipated tone (an antibiotic family which has been found to be extensively dispersed in the microbiota of human diners) and confirmed its antibacterial efficacy against a variety of human harmful bacteria, including S. Aureus. Enterococcus fecili, Corynebacterium aurimucosum and Gardnerella vaginalis.152 When compared to culture-based approaches, sequence-based methods have a higher content of placement and allow for the study of resistant to cultivate agencies. This enables a quick and relatively accurate evaluation of metagenomic data sets for the functionalities of one’s interest. But, bigger-scale analyses continue to need access to expensive sequencing equipment, massive computer resources, and bioinformatics expertise, making clinic dissemination challenging. This is a fast growing field, and the advancement of sequencing technology marks a significant step toward the practical application of Point care. Furthermore, software tools are constantly developed such that they are accessible to a large user base while yet being small enough to fit on a midsize laptop.147 Several software tools described thus far are open with expanding availability outside of big academic institutions.
Investigation of Gut Microbiome
Nonetheless, mankind actions were modified by profusion and diversity of genes of antibiotic resistance in the soil ecosystem.32,153 Nonetheless, local fauna administration of manure to soil environment resulted in exposure of both antibiotics resistant genes as well as antibiotics. Zhu et al154 in his investigation on collembolan gut microbiome after application of manure along with antibiotics exposure for fourteen days resulted in ARGs significantly inside the gut microbiome. Therefore, these collembolans are considerably imperative in accrual and dispersion of antibiotic resistant genes in sully environment. Thus, augmentation of antibiotic resistant genes in the collembolan gut microbiome which is a particular soil fauna and significant in transfer of antibiotic resistant genes through the standard transfer in food chain.
Hacioglu et al155 investigated the antimicrobial compounds which displayed a significant role in therapy of bacterial maladies and maximum attempts have been performed by several scientists in order to screen for new microbes which is capable of producing antibiotics.86 Despite of the fact that many attempts have been made to isolate and examine various antibiotics from soil fauna that have been found to be very effective in fighting pathogenic microbes that aid in the treatment of various maladies. Nonetheless, the appearance of new infections and the reappearance of antibiotic resistant pathogens would diminish the effectivity of currently used clinical antibiotics, necessitating the search for novel antibiotics.156 Antibiotics were classified based on their spectrum as well as the characteristics of bacteria, such as whether they are gramme +ve or -ve. The broad spectrum of antibiotics that can affect gram-positive bacteria, whereas only a narrow range of antibiotics can affect gram-negative bacteria. Nevertheless, several scientists have found the mode of actions of these antibiotics. The mechanism of action includes cell wall formation is inhibited as well as DNA and protein synthesis.157-159 For discovery of antibiotics from diverse natural resource, they can be used as medicinal herbs acting as potential agent and displaying antibacterial, antifungal, anti-inflammatory, anticancer and antioxidant agents160-162 and soil. Nejad et al. described the characteristic features of soil environment as indigenous loose mixture of organic particles and mineral which remains the most significant target for many scientists to find new medicines which have bio-pharmaceutical values. The symbiotic and non-symbiotic bacteria residing in the soil has the capability to produce secondary metabolites which helps in growth of flora. These bacteria exists in the root nodules of legumes that not only helps in fixing nitrogen but in turns supports in growth of plant species and also prevents the invasion of other pathogenic organisms. Linares163, stated the functions of antibiotics, other than protection also helps in communication between cells. Through cell–cell signaling. Subsequently, numerous multi-drug resistance pathogenic organisms has transformed and evoluted over time and there are as it were confined helpful drugs that are appropriate to battle against these pathogens.164 Therefore, continuous research for discovery of novel antibiotics from diverse sources for antibiotics to remain effectual.165 Thus our current study endeavors to find out new antimicrobial compounds from soil ecosystem samples from different countries at diverse localities.
Future Perspectives for control of soil biota from human intervention
Sully microbiota contaminated after release of pharmaceutical products from hospitals, pharma companies led to diverse resistant microorganisms. The microbiome as such is rich source of bioactive agents and due to human actions enhanced resistance at global level. Thus, enhanced the antibiotic resistant genes in the sully ecosystem which has direct repercussion on planetary health. Therefore, management of soil biota is crucially significant in maintaining human health. The augmenting profusion of antibiotic resistant genes in sully has distinct and direct influence on human health with unswerving association between soils and production of food. The resistance determinants increase is because of transfer of antibiotics, pesticides, heavy metal directly into soil and those crops cultivated utilized for human consumption, which in turn triggers the commensals to pathogenic bacteria. Still examination on this issue is being carried out, our article provides broad insights to researchers to assess the soil microbes and study their genome expression through next gene sequencer.
The contamination of environmental sully with antibiotics is regarded as a major problem today and predictable to attain more recognition in near future. However, human intervention resulting in antibiotic consumption is being enhancing all around the world. Our review of literature revealed the role of microbiome in sully and how antibiotic resistant genes raised. The structure of antibiotics mainly gets altered by environmental factors like a biotic and biotic stress which varies based on diverse soils. Therefore, management of microbiome in soil and their expression studies was distinctively revealed. The assessment of ARGs with help of next generation sequencing provided a clear comprehension on genome and transcriptome of the bacterial genes. Thus, interaction of microbiome with soil can also be well understood. The current findings in our study will guide every researcher to follow logical protocol in analyzing microbiota composition.
ACKNOWLEDGMENTS
The authors would like to thank Dr. Sabu Thomas, RGCB, Kerala, India. Dr. Mukesh Doble, IIT, Madras, India and Dr. M.C. Madhusudhan, Mysore, India, for their comments and their insightful suggestions and careful reading of the manuscript.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
None.
ETHICS STATEMENT
Not applicable.
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the manuscript.
- Jansson JK, Hofmockel KS. The soil microbiome-from metagenomics to metaphenomics. Curr Opin Microbiol. 2018;43:162-168.
Crossref - Powlson DS, Hirsch PR, Brookes PC. The role of soil microorganisms in soil organic matter conservation in the tropics. Nutr Cycl Agroecosystems. 2001;61(1):41-51.
Crossref - Torsvik V, Goksoyr J, Daae FL. High diversity in DNA of soil bacteria. Appl Environ Microbiol. 1990;56(3):782-787.
Crossref - Amann RI, Ludwig W, Schleifer KH. Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol Rev. 1995;59(1):143-169.
Crossref - Gans J, Wolinsky M, Dunbar J. Computational improvements reveal great bacterial diversity and high metal toxicity in soil. Science. 2005;309(5739):1387-1390.
Crossref - Baveye PC. To sequence or not to sequence the whole-soil metagenome? Nat Rev Microbiol. 2009;7(10):756.
Crossref - Raynaud X, Nunan N. Spatial ecology of bacteria at the microscale in soil. PLoS One 2014;9(1):e87217.
Crossref - Mandal S, Van Treuren W, White RA, Eggesbo M, Knight R, Peddada SD. Analysis of composition of microbiomes: a novel method for studying microbial composition. Microb Ecol Health Dis. 2015;26:27663.
Crossref - Fischer G, Heilig GK. Population momentum and the demand on land and water resources. Philos Trans R Soc Lond B Biol Sci. 1997;352(1356):869-889.
Crossref - Wagg C, Bender SF, Widmer F, van der Heijden MGA. Soil biodiversity and soil community composition determine ecosystem multifunctionality. Proc Natl Acad Sci U S A. 2014;111(14):5266-5270.
Crossref - Marina R, Garcia-Quintanilla M, Martin-Pena R, Cisneros JM, McConnell MJ. Progress on the development of rapid methods for antimicrobial susceptibility testing. J Antimicrob Chemother. 2013;68(12):2710-2717.
Crossref - Farraro JH. Antimicrobial susceptibility testing: General principles and contemprary practice. Oxford university. 1998.
- Beavington F. Foundation work on soil and human health. Eur J Soil Sci. 2000;51(2):365-366.
Crossref - Steffan JJ, Brevik EC, Burgess LC, Cerda A. The effect of soil on human health: an overview. Eur J Soil Sci. 2018;69:159-171.
Crossref - Ding C, He J. Effect of antibiotics in the environment on microbial populations. Appl Microbiol Biotechnol. 2010;87(3):925-941.
Crossref - Baniya A, Singh S, Singh M, et al. Isolation and screening of antibiotics producing Streptomyces spp. from the soil collected around the root of Alnus nepalensis from Godawari. Nepal Journal of Biotechnology. 2019;6(1):46-56.
Crossref - Ganesan P, David HA, Reegan D, et al. Isolation and molecular characterization of actinomycetes with antimicrobial and mosquito larvicidal properties. Beni-Suef University Journal of Basic and Applied Sciences. 2017;6(2):209-217.
Crossref - Mittelbach GG, Steiner CF, Scheiner SM, et al. What is the observed relationship between species richness and productivity? Ecology. 2001;82(9):2381-2396.
Crossref - Brock TD. Origin and status of microbial ecology. In: Trends In Microbial Ecology. (Guerrero, R. and Pedros-Alio, C. Eds.). Proc VIthInternl Symp Microbial Ecol. Spanish Society for Microbiol. 1993:201-206.
- Martinez JL. Effect of antibiotics on bacterial populations: a multihierachical selection process. F1000Research. 2017;6:51.
Crossref - Larsson DGJ. Antibiotics in the environment. Ups J Med Sci. 2014;119(2):108-112.
Crossref - Bengtsson-Palme J, Larsson DGJ. Concentrations of antibiotics predicted to select for resistant bacteria: proposed limits for environmental regulation. Environ Int. 2016;86:140-149.
Crossref - Bonetta S, Bonetta S, Mosso S, Sampo S, Carraro E. Assessment of microbiological indoor air quality in an Italian office building equipped with an HVAC system. Environ Monit Assess. 2010;161(1-4):473-483.
Crossref - Davison J. Plant beneficial bacteria. Biotechnology. 1988;6:282-286.
Crossref - Brierley JA. Use of microorganisms for mining metals. In: Halvorson HO, Pramer D, Rogul M (eds) Engineered organisims in the environment: scientifc issues. ASM Press, Washington. 1985: 141-146.
- Ehrlich HL. Geomicrobiology, 2nd edn. Dekker, New York, 1990:646.
- Middledrop PJM, Briglia M, Salkinoja-Salonen M. Biodegradation of pentachlorophenol in natural polluted soil by inoculated Rhodococcus chlorophenolicus. Microb Ecol. 1990;20(1):123-139.
Crossref - Burd G, Dixon DG, Glick BR. Plant growth promoting bacteria that decrease heavy metal toxicity in plants. Can J Microbiol. 2000;46(3):237-245.
Crossref - Zhuang X, Chen J, Shim H, Bai Z. New advances in plant growth-promoting rhizobacteria for bioremediation. Environ Int. 2007;33(3):406-413.
Crossref - Zaidi S, Usmani S, Singh BR, Musarrat J. Significance of Bacillus subtilis strains SJ-101 as a bioinoculant for concurrent plant growth promotion and nickel accumulation in Brassica Juncea. Chemosphere. 2008;64(6):991-997.
Crossref - Byju G. Soil Analysis-A Laboratory Manual. Central Tuber Crops Research Institute, Thiruvananthapuram, Kerala. 2001.
- Wright GD. Antibiotic resistance in the environment: a link to the clinic? Curr Opin Microbiol. 2010;13(5):589-594.
Crossref - Day PR. Particle fractionation and particle-size analysis. In: Methods of Soil Analysis. Part 1. Physical and Mineralogical Properties, including Statistics of Measurement and Sampling. Black CA, Evans DD, Ensminger LE, White JL, Clark FE, Dinauer RC. (Eds.). ASA, Madison, Wisconsin, USA. 1965:552-562.
- Williams BG, Greenland DJ, Lindstrom GR, Qurik JP. Techniques for the determination of the stability of soil aggregates. Soil Sci. 1966;101(3):157-163.
Crossref - Page AL, Miller RH, Keeney DR. Methods of Soil Analysis Part 2. Chemical and Microbiological Properties. ASA, SSSA, Madison, Wisconsin, USA. 1982.
- Walkley A, Black IA. An estimation of the degtjareff method for determining soil organic matter and a proposed modification of the chromic acid titration method. Soil Sci. 1934;37:29-38.
- Bray RH, Kurtz LT. Determination of total, organic and available forms of phosphorus in soils. Soil Science. 1945;59(1):39-45.
Crossref - Weil RR, Islam KR, Stine MA, Gruver JB, Liebig SES. Estimating active carbon for soil quality assessment: a simplified method for labortaory and field use. Am J Altern Agric. 2003;18(1):3-17.
Crossref - Sturz AV, Christie BR, Novak J. Bacterial endophytes: potential role in developing sustainable system of crop production. Crit Rev Plant Sci.2000;19:1-30.
Crossref - Shoebitz M, Ribaudo CM, Pardo MA, Cantore ML, Ciampi L, Cura JA. Plant growth promoting properties of a strain of Enterobacter ludwigii isolated from Lolium perenne rhizosphere. Soil Biol Biochem. 2009;41(9):1768-1774.
Crossref - Cocking EC. Endophytic colonization of plant roots by nitrogen-fixing bacteria. Plant Soil. 2003;252(1):169-175.
Crossref - Kloepper JW, Schroth MN. Plant growth promoting rhizobacteria on radishes. Proceedings of the Fourth International Conference on Plant Pathogen Bacteria. INRA. 1978;2:879-882.
- Kloepper JW, Lifshitz R, Zablotowicz RM. Free-living bacterial inocula for enhancing crop productivity. Trends Biotechnol. 1989;7(2):39-43.
Crossref - Cleyet-Marcel JC, Larcher M, Bertrand H, Rapior S, Pinochet X. Plant growth enhancement by rhizobacteria. In: MorotGaudry JF (ed) Nitrogen assimilation by plants: physiological, biochemical and molecular aspects. Science Publishers, Plymouth. 2001:185-197.
- Khan AG. Role of soil microbes in the rhizosphere of plants growing on trace metal contaminated soils in phytoremediation. J Trace Elem Med Biol. 2005;18(4):355-364.
Crossref - Gray EJ, Smith DL. Intracellular and extracellular PGPR: commonalities and distinctions in the plant-bacterium signaling processes. Soil Biol Biochem. 2005;37(3):395-412.
Crossref - Bottomley PJ, Maggard SP. Determination of viability within serotypes of a soil population of Rhizobium leguminosarum biovar trifolii. Appl Environ Microbiol. 1990;56(2):533-540.
Crossref - Bottomley PJ, Dughri MH. Population size and distribution of Rhizobium leguminosarum biovar trifolii in relation to total soil bacteria and soil depth. Appl Environ Microbiol. 1989;55(4):959-964.
Crossref - Lynch JM. Soil biotechnology. Blackwell, Oxford. 1983.
- Lynch JM. Beneficial interactions between miroorganisms and roots. Biotechnol Adv. 1990;8(2):335-346.
Crossref - Brown ME. Seed and root bacterization. Annu Rev Phytopathol. 1974;12:181-197.
Crossref - Fish DN, Choi MK, Jung R. Synergic activity of cephalosporins plus fluoroquinolones against Pseudomonas aeruginosa with resistance to one or both drugs. J Antimicrob Chemother. 2002;50(6):1045-1049.
Crossref - Laxminarayan R, Chaudhury RR. Antibiotic resistance in India: drivers and opportunities for action. PLoS Med. 2016;13(3):e1001974.
Crossref - Gebreyohannes G, Moges F, Sahile S, Raja N. Isolation and characterization of potential antibiotic producing actinomycetes from water and sediments of Lake Tana, Ethiopia. Asian Pac J Trop Biomed. 2013;3(6):426-435.
Crossref - Lipsitch M, Singer RS, Levin BR. Antibiotics in agriculture: when is it time to close the barn door? Proc Natl Acad Sci USA. 2002;99:5752-5754.
Crossref - Marshall BM, Levy SB. Food animals and antimicrobials: impacts on human health. Clin Microbiol Rev. 2011;24(4):718-733.
Crossref - Santajit S, Indrawattana N. Mechanisms of antimicrobial resistance in ESKAPE pathogens. Biomed Res Int. 2016;2016:2475067.
Crossref - Pan X, Qiang Z, Ben W, Chen M. Residual veterinary antibiotics in swine manure from concentrated animal feeding operations in Shandong Province, China. Chemosphere. 2011;84(5):695-700.
Crossref - Chen W, Liu W, Pan N, Jiao W, Wang M. Oxytetracycline on functions and structure of soil microbial community. J Soil Sci Plant Nutr. 2013;13(4):967-975.
Crossref - Masse DI, Saady NMC, Gilbert Y. Potential of biological processes to eliminate antibiotics in livestock manure: an overview. Animals. 2014;4(2):146-163.
Crossref - Zhao L, Dong YH, Wang H. Residues of veterinary antibiotics in manures from feedlot livestock in eight provinces of China. Sci Total Environ. 2010;408(5):1069-1075.
Crossref - Van Doorslaer X, Dewulf J, Van Langenhove H, Demeestere K. Fluoroquinolone antibiotics: an emerging class of environmental micropollutants. Sci Total Environ. 2014;500-501:250-269.
Crossref - Martinez-Carballo E, Gonzalez-Barreiro C, Scharf S, Gans O. Environmental monitoring study of selected veterinary antibiotics in animal manure and soils in Austria. Environ Pollut. 2007;148(2):570-579.
Crossref - Dolliver H, Gupta S, Noll S. Antibiotic degradation during manure composting. J Environ Qual. 2008;37(3):1245-1253.
Crossref - Jones-Lepp TL, Stevens R. Pharmaceuticals and personal care products in biosolids/sewage sludge: the interface between analytical chemistry and regulation. Anal Bioanal Chem. 2007;387(4):1173-1183.
Crossref - Wepking C, Avera B, Badgley B, et al. Exposure to dairy manure leads to greater antibiotic resistance and increased mass-specific respiration in soil microbial communities. Proc R Soc B Biol Sci. 2017;284(1851):20162233.
Crossref - Xiong W, Wang M, Dai J, Sun Y, Zeng Z. Application of manure containing tetracyclines slowed down the dissipation of tet resistance genes and caused changes in the composition of soil bacteria. Ecotoxicol Environ Saf. 2018;147:455-460.
Crossref - Su JQ, Wei B, Xu CY, Qiao M, Zhu YG Functional metagenomic characterization of antibiotic resistance genes in agricultural soils from China. Environ Int. 2014;65:9-15
Crossref - Kyselkova M, Kotrbova L, Bhumibhamon G, et al. Tetracycline resistance genes persist in soil amended with cattle feces independently from chlortetracycline selection pressure. Soil Biol Biochem. 2015;81:259-265.
Crossref - Cermak L, Kopecky J, Novotna J, et al. Bacterial communities of two contrasting soils reacted differently to lincomycin treatment. Appl Soil Ecol. 2008;40(2):348-358.
Crossref - Popowska M, Rzeczycka M, Miernik A, Krawczyk-Balska A, Walsh F, Duffy B. Influence of soil use on prevalence of tetracycline streptomycin, and erythromycin resistance and associated resistance genes. Antimicrob Agents Chemother. 2012;56(3):1434-1443.
Crossref - Xiao K-Q, Li B, Ma L, T, et al. Metagenomic profiles of antibiotic resistance genes in paddy soils from South China. FEMS Microbiol. Ecol. 2016;92(3):fiw023.
Crossref - Lau CH-F, van Engelen K, Gordon S, Renaud J, Topp E. Novel antibiotic resistance determinants from agricultural soil exposed to antibiotics widely used in human medicine and animal farming. Appl Environ Microbiol. 2017;83(16):e00989-e00917.
Crossref - Wang S, Gao X, Gao Y, et al. Tetracycline resistance genes identified from distinct soil environments in China by functional metagenomics. Front Microbiol. 2017;8:1406.
Crossref - Torres-Cortes G, Millan V, Ramirez-Saad HC, Nisa-Martinez R, Toro N, Martinez-Abarca F. Characterization of novel antibiotic resistance genes identified by functional metagenomics on soil samples. Environ Microbiol. 2011;13(4):1101-1114.
Crossref - Gudeta DD, Bortolaia V, Pollini S, et al. Expanding the repertoire of carbapenem-hydrolyzing metallo- b-lactamases by functional metagenomic analysis of soil microbiota. Front Microbiol. 2016;7:1985.
Crossref - Donato JJ, Moe LA, Converse BJ, et al. Metagenomic analysis of apple orchard soil reveals antibiotic resistance genes encoding predicted bifunctional proteins. Appl Environ Microbiol. 2010;76(13):4396-4401.
Crossref - Zhou B, Wang C, Zhao Q, et al. Prevalence and dissemination of antibiotic resistance genes and coselection of heavy metals in Chinese dairy farms. J Hazard Mater. 2016;320:10-17.
Crossref - Chapman JS. Disinfectant resistance mechanisms, cross-resistance, and co-resistance. Int Biodeterior Biodegr. 2003;51(4):271-276.
Crossref - Baker-Austin C, Wright MS, Stepanauskas R, McArthur JV. Co-selection of antibiotic and metal resistance. Trends Microbiol. 2006;14(4):176-182.
Crossref - Martinez JL, Sanchez MB, Martinez-Solano L, et al. Functional role of bacterial multidrug efflux pumps in microbial natural ecosystems. FEMS Microbiol Rev. 2009;33(2):430-449.
Crossref - Herrick JB, Haynes R, Heringa S, Brooks JM, Sobota LT. Coselection for resistance to multiple late-generation human therapeutic antibiotics encoded on tetracycline resistance plasmids captured from uncultivated stream and soil bacteria. J Appl Microbiol. 2014;117(2):380-389.
Crossref - Dezfully NK, Ramanayaka JG. Isolation, identification and evaluation of antimicrobial activity of Streptomyces flavogriseus, strain ACTK2 from soil sample of Kodagu, Karnataka State (India). Jundishapur J Microbiol. 2015;8(2):e15107.
Crossref - Yunus FN, Khalid ZZ, Rashid F, Ashraf A, Iqbal MN, Hussain F. Isolation and screening of antibiotic-producing bacteria from soil in Lahore city. PSM Microbiology. 2016;1:1-4.
- Nike AR, Hassan SA, Ajijolakewu, Bosede AF. Soil screening for antibiotic-producing microorganisms. Adv Environ Biol. 2013;7(1):7-11.
- Oskay M, Tamer AU, Azeri C. Antibacterial activity of some actinomycetes isolated from farming soil of Turkey. African J Bitechnol. 2004;3(9):441-446.
Crossref - Wadetwar RN, Patil AT. Isolation and characterization of bioactive actinomycetes from soil in and around Nagpur. Int J Pharm Sci Res. 2013;4:1428-1433.
Crossref - Constancias F, Terrat S, Saby NPA, et al. Mapping and determinism of soil microbial community distribution across an agricultural landscape. Microbiology Open. 2015;4(3):505-517.
- Williams ST, Vickers JC. The ecology of antibiotic production. Microb Ecol. 1986;12(1):43-52.
Crossref - Kaur S, Kaur J, Pankaj PP. Isolation and characterization of antibiotic producing microorganisms from soil samples of certain area of Punjab region of India. Int J Pharm Clin Res. 2014;6(4):312-315.
- Diddi K, Chaudhry R, Sharma N, Dhawan B. Strategy for identification & characterization of Bartonellahenselae with conventional & molecular methods. Indian J Med Res. 2013;137(2):380-387.
- Kumar S, Chaurasia P, Kumar A. Isolation and characterization of microbial strains from textile industry effluents of bhilwara, india: analysis with bioremediation. J Chem Pharm Res. 2016;8(4):143-150.
- Bull AT, Stach JE. Marine actinobacteria: new opportunities for natural product search and discovery. Trends Microbiology. 2007;15(11):491-499.
Crossref - Pavunraj M, Ramasubbu G, Baskar K. Leucas aspera (willd.) linn.: antibacterial, antifungal and mosquitocidal activities. Trends Phytochem Res. 2017;1(3):133-140.
- Mohammadhosseini M. The ethnobotanical, phytochemical and pharmacological properties and medicinal applications of essential oils and extracts of different Ziziphora species. Ind Crops Prod. 2017;105(1):164-192.
Crossref - de Queiroz Cavalcanti MA, de Oliveira LG, Fernandes MJ, Lima DM. Filamentous fungi isolated from soil in districts of the Xingo region, Brazil. Acta Bot Bras. 2006;20(4):831-837.
Crossref - Audrain B, Farag MA, Ryu C-M, Ghigo J-M. Role of bacterial volatile compounds in bacterial biology. FEMS Microbiol Ecol. 2015;39(2):222-233.
Crossref - Lorena B, Corcione S, Pacini G, Perri GD, D’Avolio A, Rosa FGD. A 30-years review on pharmacokinetics of antibiotics: is the right time for pharmacogenetics? Curr Drug Metab. 2014;15(6):581-598.
Crossref - Guzman JD, Gupta A, Bucar F, Gibbons S, Bhakta S. Antimycobacterials from natural sources: ancient times, antibiotic era and novel scaffolds. Front Biosci. 2012;17(5):1861-1881.
Crossref - Tong Z, Ivask A, Guo K, et al. Crossed flow microfluidics for high throughput screening of bioactive chemical-cell interactions. Lab Chip. 2017;17(3):501-510.
Crossref - Hautbergue T, Jamin EL, Debrauwer L, Puel O, Oswald IP. From genomics to metabolomics, moving toward an integrated strategy for the discovery of fungal secondary metabolites. Nat Prod Rep. 2018;35(2):147-173.
Crossref - Vijayalakshmi K, Premalatha A, Rajakumar GS. Antimicrobial protein production by Bacillus amyloliquefaciens MBL27: optimization of culture conditions using Taguchi’s experimental design. Indian J Sci Technol. 2011;4(8):931-937.
Crossref - Arguelles-Arias A, Ongena M, Halimi B, et al. Bacillus amyloliquefaciens GA1 as a source of potent antibiotics and other secondary metabolites for biocontrol of plant pathogens. Microb Cell Fact. 2009;8:63.
Crossref - Alvarez F, Castro M, Principe A, et al. The plant associated Bacillus amyloliquefaciens strains MEP218 and ARP23 capable of producing the cyclic lipopeptides iturin or surfactin and fengycin are effective in biocontrol of sclerotinia stem rot disease. J Appl Microbiol. 2012;112(1):159-174.
Crossref - Meng QX, Jiang HH, Hanson LE, Hao JJ. Characterizing a novel strain of Bacillus amyloliquefaciens BAC03 for potential biological control application. J Appl Microbiol. 2012;113(5):1165-1175.
Crossref - Shafi J, Tian H, Ji M. Bacillus species as versatile weapons for plantpathogens: a review. Biotechnol Equip. 2017;31(3):446-459.
Crossref - Siahmoshteh F, Siciliano I, Banani H, et al. Efficacy of Bacillus subtilis and Bacillus amyloliquefaciens in the control of Aspergillus parasiticus growth and aflatoxins production on pistachio. Int J Food Microbiol. 2017;254:47-53.
Crossref - Karimi E, Safaie N, Shams-Baksh M, Mahmoudi B. Bacillus amyloliquefaciens SB14 from rhizosphere alleviates Rhizoctonia damping-off disease on sugar beet. Microbiol Res. 2016;192:221-230.
Crossref - Arias CA, Murray BE. Antibiotic-resistant bugs in the 21st century-a clinical super-challenge. N Engl J Med. 2009;360:439-443.
Crossref - Yu GY, Sinclair JB, Hartman GL, Bertagnolli BL. Production of iturin A by Bacillus amyloliquefaciens supressing Rhizoctonia solani. Soil Biol Biochem. 2002;34(7):955-963.
Crossref - Wu L, Wu H, Chen L, Yu X, Borriss R, Gao X. Difficidin and bacilysin from Bacillus amyloliquefaciens FZB42 have antibacterial activity against Xanthomonas oryzae rice pathogens. Scientifc Reports. 2015;5:129755.
Crossref - Vazquez-Mendoza P, Elghandour MMM, Alaba PA, et al. Antimicrobial and bactericidal impacts of Bacillus amyloliquefaciens CECT 5940 on fecal shedding of pathogenic bacteria in dairy calves and adult dogs. Microb Pathog. 2018;114:458-463.
Crossref - Janda JM, Abbott SL. 16S rRNA gene sequencing for bacterial identification in the diagnostic laboratory: pluses, perils, and pitfalls. J Clin Microbiol. 2007;45(9):2761-2764.
Crossref - Lau SKP, Teng JLL, Ho CC, Woo PCY. Gene amplification and sequencing for bacterial identification. Methods Microbiology. 2015;42:433-464.
- Singh AP, Singh RB, Mishra S. Studies on isolation and characterization of antibiotic producing microorganisms from industrial waste soil sample. Open Nutraceuticals J. 2012;5(1):169-173.
Crossref - Abbas S, Senthilkumar R, Arjunan S. Isolation and molecular characterization of microorganisms producing novel antibiotics from soil sample. Eur J Experiment Biol. 2014;4:149-155.
- Boottanun P, Potisap C, Hurdle JG, Sermswan RW. Secondary metabolites from Bacillus amyloliquefaciens isolated from soil can kill Burkholderia pseudomallei. AMB Express. 2017;7(1):16.
Crossref - Li W, Li S, Zhong J, Zhu Z, Liu J, Wang W. A novel antimicrobial peptide from skin secretions of the earthworm Pheretima guillelmi (Michaelsen). Peptides. 2011;32(6):1146-1150.
Crossref - Liu YQ, Sun Z-J, Wang C, Li S-J, Liu Y-Z. Purification of a novel antibacterial short peptide in earthworm Eisenia foetida. Acta Biochim Biophys Sin. 2004;36(4):297-302.
Crossref - Roelofs D, Timmermans MJTN, Hensbergen P, et al. A functional isopenicillin N synthase in an animal genome. Mol Biol Evol. 2013;30(3):541-548.
Crossref - Suring W, Marien J, Broekman R, van Straalen NM, Roelofs D. Biochemical pathways supporting beta-lactam biosynthesis in the spring tail Folsomia candida. Biol Open. 2016;5(12):1784-1789.
Crossref - Suring W, Meusemann K, Blanke A, et al. Evolutionary ecology of beta-lactam gene clusters in animals. Mol Ecol. 2017;26(12):3217-3229..
Crossref - Johnson AP, Woodford N. Global spread of antibiotic resistance: the example of New Delhi metallo-beta-lactamase (NDM)-mediated carbapenem resistance. J Med Microbiol. 2013;62(4):499-513.
Crossref - Walsh TR, Weeks J, Livermore DM, Toleman MA. Dissemination of NDM-1 positive bacteria in the New Delhi environment and its implications for human health: an environmental point prevalence study. Lancet Infect Dis. 2011;11:355-362.
Crossref - Lax S, Gilbert JA. Hospital-associated microbiota and implications for nosocomial infections. Trends Mol Med. 2015;21(7):427-432.
Crossref - Vaz-Moreira I, Nunes OC, Manaia CM. Diversity and antibiotic resistance in Pseudomonas spp. from drinking water. Sci Total Environ. 2012;426:366-374.
Crossref - Wen X, Mi J, Wang Y, et al. Occurrence and contamination profiles of antibiotic resistance genes from swine manure to receiving environments in Guangdong Province southern China. Ecotoxicol Environ Saf. 2019;173:96-102.
Crossref - Ruiz-Fons F. A review of the current status of relevant zoonotic pathogens in wild swine (Sus scrofa) populations: changes modulating the risk of transmission to humans. Transbound Emerg Dis. 2017;64(1):68-88.
Crossref - Richard M, Knauf S, Lawrence P, et al. Factors determining human-to-human transmissibility of zoonotic pathogens via contact. Curr Opin Virol. 2017;22:7-12.
Crossref - Zhu YG, Johnson TA, Su J-Q, et al. Diverse and abundant antibiotic resistance genes in Chinese swine farms. Proc Natl Acad Sci U S A. 2013;110(9):3435-3440.
Crossref - Xie WY, Yang XP, Li Q, Wu LH, Shen QR, Zhao FJ. Changes in antibiotic concentrations and antibiotic resistome during commercial composting of animal manures. Environ Pollut. 2016;219:182-190.
Crossref - Ruan Z, Sun Q, Jia H, et al. Emergence of a ST2570 Klebsiella pneumoniae isolate carrying mcr-1 and blaCTX-M-14 recovered from a bloodstream infection in China. Clin Microbiol Infect. 2019;25(7):916-918.
Crossref - Reddy B, Dubey SK. River Ganges water as reservoir of microbes with antibiotic and metal ion resistance genes: high throughput metagenomic approach. Environ Pollut. 2019;246:443-451.
Crossref - Fernandes T, Vaz-Moreira I, Manaia CM. Neighbor urban wastewater treatment plants display distinct profiles of bacterial community and antibiotic resistance genes. Environ Sci Pollut Res Int. 2019;26(11):11269-11278.
Crossref - Amos GC, Zhang L, Hawkey PM, Gaze WH, Wellington EM. Functional metagenomic analysis reveals rivers are a reservoir for diverse antibiotic resistance genes. Vet Microbiol. 2014;171(3-4):441-447.
Crossref - Chen H, Bai X, Jing L, Chen R, Teng Y. Characterization of antibiotic resistance genes in the sediments of an urban river revealed by comparative metagenomics analysis. Sci Total Environ. 2019;653:1513-1521.
Crossref - Ma L, Jian X-T, Want Y-L, et al. Catalogue of antibiotic resistome and host-tracking in drinking water deciphered by a large scale survey. Microbiome. 2017;5(1):154.
Crossref - Hover BM, Kim S-H, Katz M, et al. Culture-independent discovery of the malacidins as calcium-dependent antibiotics with activity against multidrug-resistant gram-positive pathogens. Nat Microbiol. 2018;3(4):415-422.
Crossref - Pehrsson EC, Tsukayama P, Patel S, et al. Interconnected microbiomes and resistomes in low-income human habitats. Nature. 2016;533(7602):212-216.
Crossref - Gibson MK, Wang B, Ahmadi S, et al. Developmental dynamics of the preterm infant gut microbiota and antibiotic resistome. Nat Microbiol. 2016;1:16024.
Crossref - Liu B, Pop M. ARDB-antibiotic resistance genes database. Nucleic Acids Res. 2009;37:D443-D447.
Crossref - Lin Y, Li J, Shen H, Zhang L, Papasian CJ, Deng HW. Comparative studies of de novo assembly tools for next-generation sequencing technologies. Bioinformatics. 2011;27(15):2031-2037.
Crossref - Gibson MK, Forsberg KJ, Dantas G. Improved annotation of antibiotic resistance determinants reveals microbial resistomes cluster by ecology. ISME J. 2015;9(1):207-216.
Crossref - Bocanegra-Ibarias P, Garza-Gonzalez E, Morfin-Otero R, et al. Molecular and microbiological report of a hospital outbreak of NDM-1-carrying Enterobacteriaceae in Mexico. PLoS ONE. 2017;12(6):e0179651.
Crossref - Eid J, Fehr A, Gray J, et al. Real-time DNA sequencing from single polymerase molecules. Science. 2009;323(5910):133-138.
Crossref - Clarke J, Wu HC, Jayasinghe L, Patel A, Reid S, Bayley H. Continuous base identification for single-molecule nanopore DNA sequencing. Nat Nanotechnol. 2009;4(4):265-270.
Crossref - Brinda K, Callentrello A, Cowley L, et al. Lineage calling can identify antibiotic resistant clones within minutes. bioRxiv. 2018.
Crossref - Bradley P, Gordon NC, Walker TM, et al. Rapid antibiotic-resistance predictions from genome sequence data for Staphylococcus aureus and Mycobacterium tuberculosis. Nat Commun. 2015;6:10063.
Crossref - Ghurye JS, Cepeda-Espinoza V, Pop M. Metagenomic assembly: overview, challenges and applications. Yale J Biol Med. 2016;89(3):353-362.
- van der Walt AJ, van Goethem MW, Ramond JB, Makhalanyane TP, Reva O, Cowan DA. Assembling metagenomes, one community at a time. BMC Genom. 2017;18(1):521.
Crossref - Blin K, Wolf T, Chevrette MG, et al. antiSMASH 4.0-improvements in chemistry prediction and gene cluster boundary identification. Nucleic Acids Res. 2017;45(W1):W36-W41.
Crossref - Donia MS, Cimermancic P, Schulze CJ, et al. A systematic analysis of biosynthetic gene clusters in the human microbiome reveals a common family of antibiotics. Cell. 2014;158(6):1402-1414.
Crossref - Chen QL, Fan X-T, Zhu D, An X-L, Su J-Q, Cuia L. Effect of biochar amendment on the alleviation of antibiotic resistance in soil and phyllosphere of Brassica chinensis L. Soil Biol. Biochem. 2018;119:74-82.
Crossref - Zhu D, An X-L, Chen QL, et al. Antibiotics disturb the microbiome and increase the incidence of resistance genes in the gut of a common soil collembolan. Environ Sci Technol. 2018;52(5):3081-3090.
Crossref - Hacioglu N, Dulger B. Occurence and antibiotic susceptibility of some bacteria in Saricay stream (Canakkale, Turkey). European Journal of Experimental Biology. 2011;1(4):158-163.
- Roberts MC. Antibiotic resistance mechanisms in bacteria of oral and upper respiratory origin. Int J Antimicrob Agents. 1998;9:255-267.
Crossref - Lambert PA. In: Hugo WB, Russell AD, (Eds.), Pharmaceutical microbiology 5th ed. (Oxford, Blackwell Scientific Publications, 1977) 1977:189.
- Brooks GF, Butel JS, Morse SA, Jawetz, Melnick, Adelberg’s medical microbiology 22nd ed. (San Francisco, The McGraw-Hill Companies, Inc., 2001).
- Tortora GJ, Funke BR, Case CL. Microbiology: An introduction 9th ed. San Francisco, Pearson Education, Inc, 2007. 192.
- Sharma M, Kumar A, Sharma B, Akshita, Dwivedi N. Evaluation of phytochemical compounds and antimicrobial activity of leaves and fruits Tribulus terrestris. Eur J Exp Biol. 2013;3(5):432-436.
- Rajaperumal S, Nimmi M, Kumari BDR. In vitro studies on antimicrobial and antioxidant effect of methanolic extract of Indigofera aspalathoides (Vahl ex DC) and its cytotoxic property against human lung cancer cell line NCI H460. European Journal of Experimental Biology. 2013;3(3):18-29.
- Dua A, Garg G, Mahajan, R. Polyphenols, flavonoids and antimicrobial properties of methanolic extract of fennel (Foeniculum vulgare Miller). Eur J Exp Biol. 2013;3(4):203-208.
- Linares JF, Gustafsson I, Baquero F, Martinez JL. Antibiotics as intermicrobial signaling agents instead of weapons. Proc Natl Acad Sci U S A. 2006;103(51);19484-19489.
Crossref - Demain AL, Sanchez S. Microbial drug discovery: 80 years of progress. J Antibiot. 2009;62(1):5-16.
Crossref - Van der Waaij D, Nord CE. Development and persistence of multi-resistance to antibiotics in bacteria; an analysis and a new approach to this urgent problem. Int J Antimicrob Agents. 2000;16(3):191-197.
Crossref
© The Author(s) 2022. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.