ISSN: 0973-7510
E-ISSN: 2581-690X
The aim of the present study was to characterize extended-spectrum b-lactamase (ESBLs) genes in multidrug resistant enterobacterial pathogens as well as commensal isolates from the Sudan during the period 2003 to 2007. ESBL production was determined phenotypically by the combined disc method, and was characterized genotypically by the detection of blagenes by PCR and nucleotide sequencing. Transferability was examined by conjugation with nalidixic-acid resistant E. coli K12. The results showed that a total of 106 of the 113 (94%) isolates including E. coli, Klebsiella pneumoniae, Proteus spp., Enterobacter cloacae, Providencia spp. and Morganella morganii, were positive for blagenes including the prototype blaTEM. Eleven isolates (28%) of the 113 were ESBL producers encoding blaSHV genes (SHV5, SHV5a, SHV12, SHV26, SHV28 and SHV38), 90 isolates (80%) were CTX-M positive. All, but only one (CTX-M9) were CTX-M15. Only 3(2.7%) of the isolates were Amp-C producers (CMY-4 and DHA-1). Plasmid transfer of the multiple resistance patterns was achieved among all the isolates. These findings demonstrated that ESBLs were highly produced by multi-resistant enterobacterial isolates from the Sudan; among both clinical pathogens as well as stool commensals. This is the first report of ESBLs genes characterization from the Sudan.
Multidrug, blagenes, enterobacteria, ESBLs, pathogens, commensals, resistance.
Extended-spectrum b-lactamases (ESBLs) are b-lactamase enzymes capable of conferring bacterial resistance to the broad-spectrum oxyiminocephalosporins. They hydrolyse extended-spectrum cephalosporins, as well as aztreonam. They are inhibited by b-lactamase inhibitors such as clavulanic acid1,2. ESBLs are not active against cephamycins and most strains expressing ESBLs are susceptible to cefoxitin and cefotetan. However ESBL producing strains can become resistant to cephamycins due to acquisition of plasmid-mediated Amp-C b-lactamases or to the loss of outer membrane porin proteins3. Carbepenems are less susceptible to hydrolysis by ESBLs than any other b-lactam antimicrobials, although resistance has been reported among mutants with porin loss3.
ESBLs represent a major health problem worldwide4,5. Over 60% of bacterial nosocomial isolates mostly E. coli and Klebsiella spp are claimed to be capable of beta-lactamase production6,7. ESBL-related infections are on the rise and has been observed throughout the globe on both hospital and community settings8,9,10,11. Community-acquired infections or colonization with ESBL-producing E. coli, were reported in several studies12 as an important cause of community onset urinary tract infections13, as well as blood stream infections14. Multi-drug resistant E. coli15,16 and ESBL producing Enterobacteriaceae have been reported from different hospitals in Sudan17,18. Recently, Musa has demonstrated predominance of blaCTX-M gene among ESBL producing E.coli and Klebsiella spp from Sudan19. We have undertaken this study in order to determine the prevalence and mechanism of antimicrobial resistance among Enterobacteriaceae from Sudan.
Bacterial strains
This study was carried out in two major tertiary care referral hospitals in the State of Khartoum (Omdurman and Khartoum teaching hospitals) in order to determine the prevalence and mechanisms of antimicrobial resistance among Enterobacteriaceae from Sudan. Clinical isolates were collected from patients and healthy individuals during the period 2003-2007. Two hundred clinical isolates, were obtained from each hospital Microbiology Laboratory and 121 normal fecal strains from inpatients and 500 fecal strains from healthy individuals in the community were included in the study. All 1021 bacterial strains were identified to the species level by standard microbiology methods20 and confirmed by API-20E and were stored at -20°C in Cryopreservers until tested.
Antibiotic susceptibility testing
All 1021 bacterial isolates were tested for antibiotic susceptibilities by disc diffusion according to BSAC standardised methods using Iso-Sensitest agar for 18 antimicrobials (Ampcillin, Cephalexin, Cefuroxime, Cefotaxime, Ceftriaxone, Ceftazidime, Meropenem, Co-amoxyclav, Nalidixic acid, Ciprofloxacin, Strep-tomycin, Gentamicin, Amikacin, Chlora-mphenicol, Trimethoprim, Co-trime-thoprim, Tetracyclin and Rifampicin) (Oxoid Ltd, Bas-ingstoke, UK)21. Minimum inhibitory conce-ntrations (MICs) were determined to Ampicillin, Cefotaxime, Ceftazidime, Cefuroxime, Co-amoxiclav, Chloramphenicol, Ciprofloxacin, Gentamicin, Tetracycline and Trimethoprim. Following the BSAC agar incorporation standardised technique22.
ESBL-Screening test
Bacterial strains resistant to ceftazidime and/or cefotaxime (113) were further screened for the production of Extended Spectrum Beta-Lactamases (ESBLs), according to BSAC guide lines23. Screening test was carried out using cefpodoxime (10µg) and cefoxitin (30µg) (Oxoid) antibiotic discs. Bacterial strains that were resistant to cefpodoxime were considered as ESBL producers while resistance to Cefoxitin indicates Amp-C production23.
ESBL-Confirmatory test
The Bacterial strains (113) were then subjected to the confirmatory test by the combined disc method. Three sets of antibiotic discs (Mast Laboratories Ltd, Bootle, Merseyside, UK) were used23.
Plasmid analysis and conjugation studies
Conjugal transfer of resistance determinants was performed by broth culture with E. coli K12 as a recipient. After 24 hours incubation, mating mixtures were plated onto agar supplemented with nalidixic acid (30µg/ml) and ampicillin (10 µg /ml). Plasmid DNA was prepared from donors and transconjugants using a commercial kit (Plasmid Mini Kit, Qiagen GmbH, Hilden, Germany).
PCR amplification and sequencing
DNA for PCR was extracted by boiling of suspended bacteria on sterile water for 10 minutes. PCR was carried out for the identification of the bla genes coding for extended spectrum b-lactamases (TEM, SHV, and CTX-M) and for Amp-C Beta-Lactamases. PCR products were detected by electrophoresis on 1.5% (w/v) agarose gels visualized under U.V light. Sequence determination of amplicons was performed using the respective PCR primers for both directions using ABI 3100 automated sequencer (Warrington, UK) and compared on BLAST.
Statistical Analysis
Results were analysed using SPSS software program.
Ethics approval and consent to participate
Not applicable, all study samples (isolates) were obtained from microbiology laboratories.
Isolates included in this study were resistant to ampicillin (MICs>64), first and second generations cephalosporins (MICs>4) and to expanded-spectrum cephalosporins (MICs>4), in addition to three or more antimicrobials such as gentamicin (MICs>4), ciprofloxacin (MICs>4), trimethoprim (MICs>8), tetracycline (MICs>4) and co-amoxyclav (MICs>64). All isolates were sensitive to meropenems.
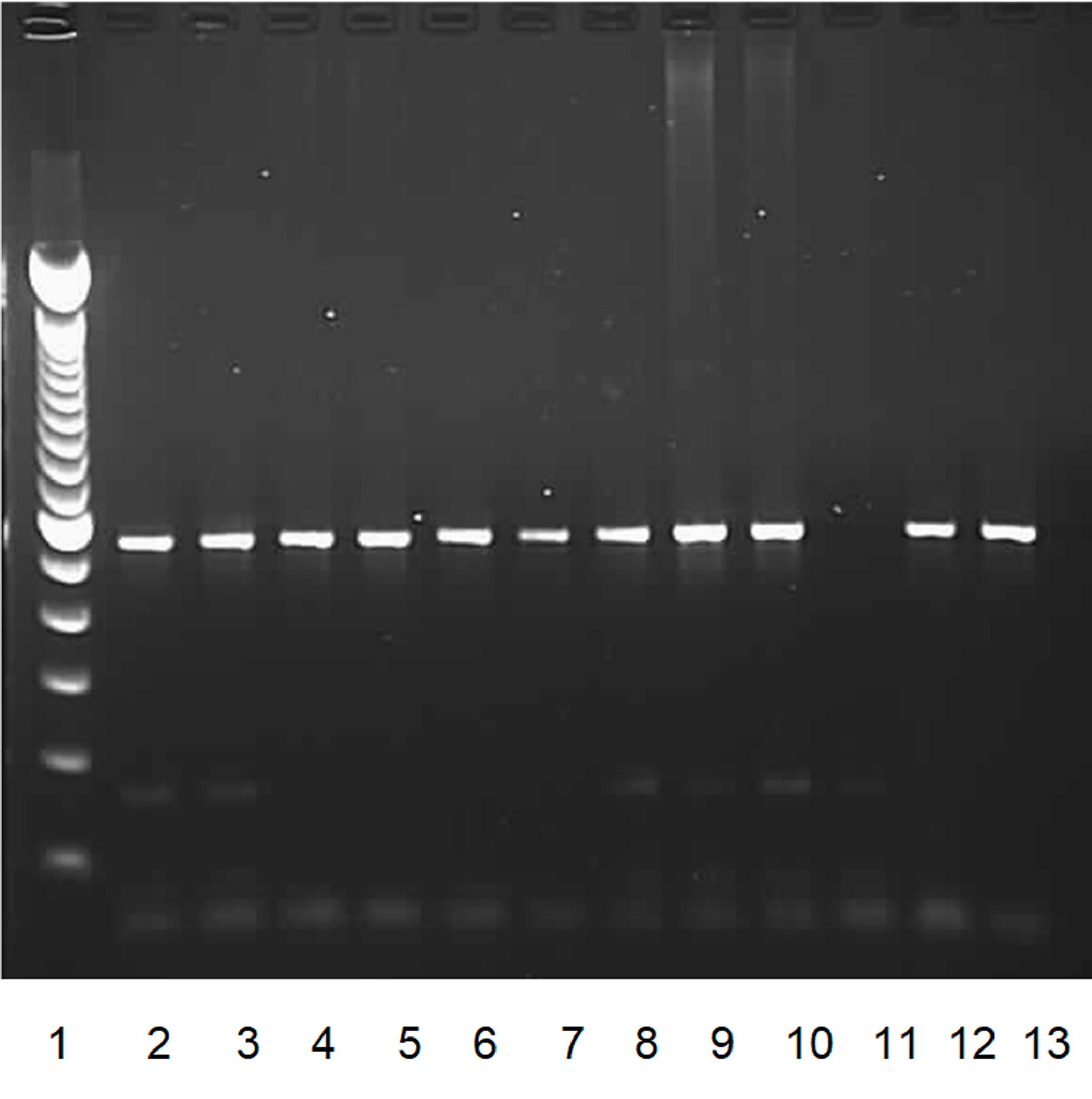 Fig. 1. PCR results of blaCTX-M positive isolates.
Fig. 1. PCR results of blaCTX-M positive isolates.Lane 1: Lambda DNA of a multiple 100bp size marker.
Lanes 2 to 10: blaCTX-M positive isolates.
Lane 11: negative control.
Lane 12, 13: positive controls.
Over all 106(94%) isolates were confirmed by the Combined disc method for ESBL production while 7(6%) isolates were tested negative (Table 1).
ESBLs-prodcing strains
For the characterization of ESBL only multidrug resistant strains were resistant to cefotaxime or/and ceftazidime antimicrobials were included. The total number was One-hundred and Thirteen strains (113). Consisted of 103 (91%) clinical isolates and ten stool commensals (9%), eight of which were from hospitalized patients and two were from healthy individuals (table-2).
bla genes were detected by PCR in 94% of the isolates (106/113), it includes different enterobacterial species comprising E.coli, Klebsiella pneumoniae, Proteus spp, Enterobacter spp, Salmonella spp and few others. Some isolates were found to possess more than one type of ESBL as well as the prototypic TEM-1 b-lactamase enzyme (Table 3).
DNA-sequencing results revealed CTX-M-15 was detected among all CTX-M positive isolates except for an E.coli isolate which was CTX-M9 producer. In all 39(34.5%) of the isolates tested for ESBLs were blaSHV positive, only 11(28%) encode for ESBLs (Table 4). DNA-sequencing results for blaTEM positive isolates revealed blaTEM-1 prototype.
Table (1):
Comparison between cefpodoxime screening test and the combined disc test results.
Combined disc +ve |
Combined disc -ve |
Total |
|
|---|---|---|---|
Cefpodoxime resistant |
106 |
5 |
111 |
Cefpodoxime sensitive |
0 |
2 |
2 |
Total |
106 |
7 |
113 |
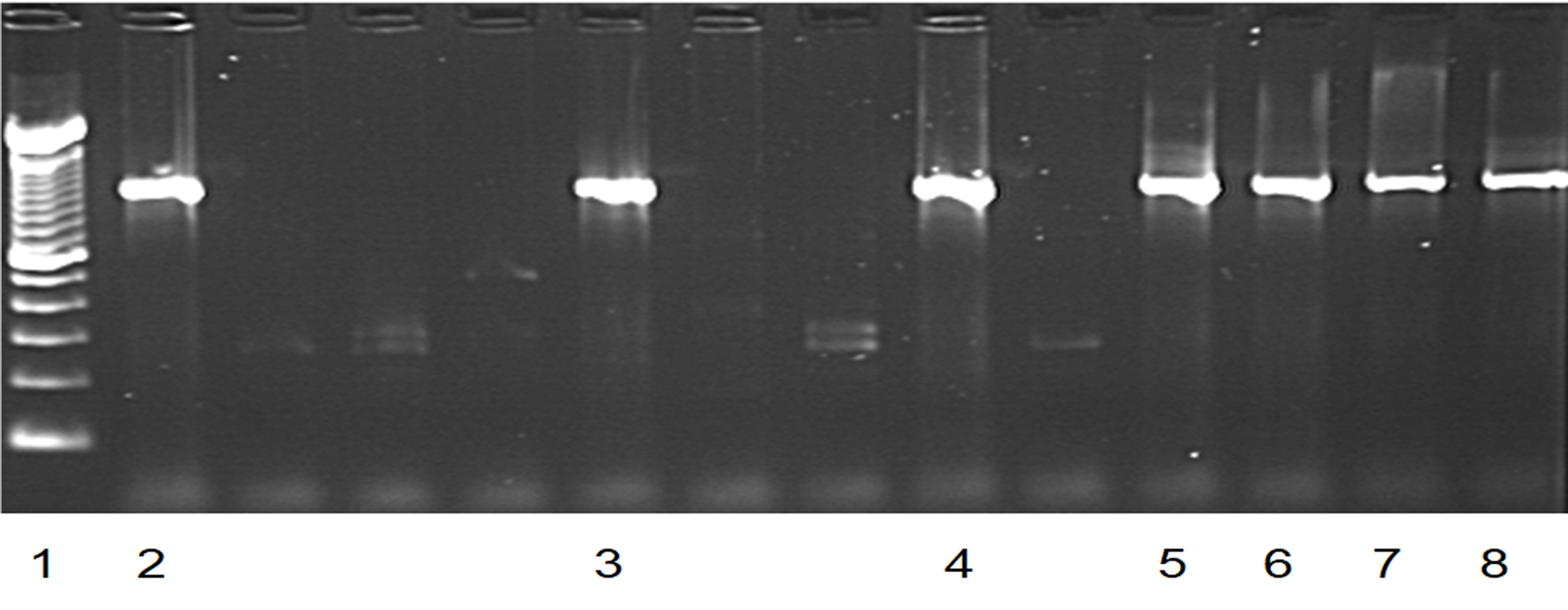 Fig. 2. PCR results of blaSHV positive isolates.
Fig. 2. PCR results of blaSHV positive isolates.Lane 1: Lambda DNA of a multiple 100bp size marker.
Lanes 2, 3, 4, 5, 6 and 7: blaSHV positive isolates.
Lane 8: positive control.
Resistance transfer to the expanded-spectrum cephalosporins was achieved among all the tested isolates, as well as the resistance to the other antimicrobial groups which was also co-transferred to the recipient organisms.
Table (2):
Number of bacterial isolates included in the study
| Clinical pathogens | 103 (91%) | ||
|---|---|---|---|
| Stool commensals | Inpatients | 8 |
10 (9%) |
| Community | 2 | ||
| Total | 113 | ||
Table (3):
Distribution of blagenes in different enterobacterial species detected by PCR.
blagene |
E.coli |
Klebs |
Proteus |
Enterob |
C-freu |
Proved |
Morg |
Sum |
|---|---|---|---|---|---|---|---|---|
CTX-M |
50 |
30 |
3 |
4 |
2 |
0 |
1 |
90 |
SHV |
11 |
21 |
0 |
7 |
0 |
0 |
0 |
39 |
TEM |
46 |
32 |
3 |
5 |
2 |
1 |
1 |
91 |
Table (4):
Sequencing results for blaSHV-positive isolates.
| SHV-TYPE | Total No. isolate | ORGANISMS | ||
|---|---|---|---|---|
| E.coli | Klebsiella | Enterobacter | ||
| SHV-1 | 12 | 6 | 6 | 0 |
| SHV-1a | 4 | 0 | 4 | 0 |
| SHV-5 | 4 | 0 | 3 | 1 |
| SHV-5a | 7 | 1 | 2 | 4 |
| SHV-11 | 3 | 2 | 1 | 0 |
| SHV-12 | 3 | 1 | 1 | 1 |
| SHV-26 | 2 | 0 | 2 | 0 |
| SHV-28 | 2 | 1 | 1 | 0 |
| SHV-38 | 2 | 0 | 2 | 1 |
We have undertaken this study in order to find out the Prevalence of ESBL-producing Enterobacteriaceae among hospitalized patients and healthy subjects in the community and to characterize ESBL genes, as the first study of its kind in Sudan. The prevalence among Klebsiella spp., E.coli, Proteus spp. and Morganella morganii isolates from hospitalized patients was exceptionally high when compared to that from the community. Similar high ESBL prevalence among Enterobacteriaceae in Sudan was reported later by Musa and Yousif18,19. ESBL-producing organisms, mostly Klebsiella spp. and E.coli. have been detected widely worldwide, but a few years later ESBL production by other Enterobacterial species have been reported24,25,26. The most frequently encountered ESBL genes were CTX-M. This is consistent with the findings of Musa and Yousif from Sudan. However, our study was more detailed as we characterized CTX-M genes further and found that they are consistent mostly of CTX-M15.
The CTX-M15 enzyme belongs to the CTX-M-1 cluster and is derived from CTX-M-3 by a single Asp-240-Gly substitution27 and has the highest catalytic activity against ceftazidime among all the CTX-M clusters, this explains the high resistance level to Ceftazidime among this group of our isolates. CTX-M15 has been reported from different parts of the world, it has been described in many genera of enterobacteriacae across the world, and is the most reported variant in much of Europe28. We identified CTX-M15 in a number of Enterobacterial species, including E.coli, Klebsiella spp., Proteus spp., Enterobacter spp., Citrobacter freundii and Morganella morganii. All these strains were resistant to multiple antimicrobial agents. CTX-M15 variant was also detected in fecal commensal isolates in this study, two E.coli isolates from the community and six isolates including E.coli and Klebsiella spp from the fecal samples of hospitalized patients. The CTX-M-15 variant which occurs mainly in E.coli seems to be epidemic in most countries. It is reported from many countries within the African continent with other ESBL-enzymes. From Tanzania in a pediatric hospital29 and an intensive care unit30, from Cameroon31. Another report from sub-Sahara Africa was from Central African Republic32, Malawi26. And also North Africa from Tunisia33.
Conjugation studies from this study have shown the possibility of horizontal gene transfer between various enterobacterial species carrying resistance genes to many antimicrobial groups including b-lactams. This could explain the acquisition of CTX-M15 and other ESBLs genes by different species of enterobacteriaceae. Other studies has reported similar findings, from Nigeria34, and Malawi26, showed that ESBL genes were associated with large conjugative plasmids that carry other resistance determinants and were responsible of resistance transfer between different bacterial species.
Only one isolate (E.coli) from this study was encoding blaCTX-M9, reported and sequenced in an E.coli from Spain35. and also later reported in an E.coli causing community acquired UTI infections36.
Variable blaSHV genes encoding ESBLs have been detected in this study from clinical as well as commensals. blaSHV5 and blaSHV5a were detected in 9.7% of the ESBL producing isolates, both genes were isolated from Enterobacter spp and from K. pneumoniae. SHV5a was detected in an E.coli isolate from the community commensal collection as well. SHV5 has been reported by different studies from Thailand37, Poland38, Australia39 and was isolated from a healthy student in Lebanon40. From the African continent SHV-5 has been reported from South-Africa in two different studies41,42.
blaSHV12 was detected in two (2.7%) isolates (Klebsiella pneumoniae and an Enterobacter spp); both isolates were CTX-M15 producers. blaSHV12 is the most commonly identified genotype worldwide. within the African continent SHV-12 was reported from Tanzania29,31, Senegal43, Cameroon32 and from the Central African Republic33.
blaSHV genes with specific hydrolytic activities to certain antimicrobials were also detected. SHV-38 was detected in Klebsiella and an Enterobacter isolates, its hydrolytic activity includes only ceftazidime and imipenem. It has been reported from France45. SHV-26 was also detected in this study in two K. pneumoniae isolates, they were resistant to co-amoxyclav. SHV-26 is a b-lactamase with reduced susceptibility to clavulanic acid that renders bacteria intermediately resistant to an inhibitor containing b-lactam. Other blaSHV genes were also reported by this study including blaSHV-28 among two isolates K. pneumoniae and E.coli.
And SHV-11 which infrequently reported and is not considered as an extended spectrum b-lactamase44. The prototypes to TEM and SHV-1/SHV-1a b-lactamase enzymes were detected as well among the majority of the isolates.
This part of the study has illustrated a great variety of beta-lactamases genes, which reflects the magnitude of the problem of antimicrobial resistance in Sudan, and the wide misuse of the expanded-spectrum beta-lactams. Over all 69% of the isolates were carrying blaCTX-M gene as well as blaSHV gene. 31% were carrying only blaSHV genes and were all ESBL-producers.
Detection of CTX-M15 and the variety of SHV enzymes among the majority of isolates would render all third generation cephalosporins ineffective for the treatment of enterobacterial infections. Finding of Amp-C within these isolates was an alarming sign of the development of resistance to cephamycins which have not yet been introduced into clinical practice in Sudan. All the above would limit the treatment options for the physicians who would be faced with only one option. Imipenem has recently been introduced in The Sudan. However all the isolates showed complete susceptibility to it and it could be one of the options for the treatment of multi-resistant infections. However already a resistance gene (blaSHV-38) that confers resistance to ceftazidime as well as to imipenems has been detected among these isolates.
Resistant bacteria may also spread and become major infection-control problems. Not only within health-care institutions, but also in the community. Clinically important bacteria, such as extended-spectrum b-lactamase (ESBL)-producing E.coli, are increasingly observed in the community as has been reported in several studies37,46.
In this study, multiple drug resistance to three or more antimicrobials is highly prevalent among the clinical isolates found in variable enterobacterial species. A wide range of gram-negative bacterial species were found to be harboring resistance to the majority of commonly used b-lactam antibiotics as well as to other antimicrobial groups. Infection with multi- resistant pathogens will complicate the treatment of nosocomial gram–negative bacterial infections within the Sudanese hospitals and will increase the mortality rates due to ineffective treatment of patients complicated by the limited options available for the doctors. This will result in further nosocomial dissemination of multi-resistant isolates due to selective pressure applied by the use of these drugs.
This work was supervised by late professor C.A.Hart, who passed away before publishing this work. A special thanks to the professors John E. Corkill for his close supervision and guidance and Ahmed Abd Elmageed for his support.
This research is fully sponsored by Ahfad University for Women, Sudan. I hereby declare that this work does not include any conflict of interest or affiliation or involvement in any organization. Ethics of human and animal experimentation not applied.
- Bush, K.; Jacoby, G. A.; Medeiros, A. A. A functional classification scheme for beta-lactamases and its correlation with molecular structure. Antimicrobial Agents& Chemotherapy, 1995; 39(6): 1211-33.
- Burriel, A. R.; Mastoraki, F.; Kritas, S. Antimicrobial resistance of animal and human strains of Enterobacteriaceae in Greece. New Microbiology Journal , 2003; 26(1): 27-32.
- Rahal, J. J. Extended-spectrum beta-lactamases: how big is the problem? Clin Microbiology& Infection. 2000; 6(2): 2-6.
- Canton, R. & Coque,T.M. The CTX-M beta-lactamase pandemic. Current Opinion in Microbiology. 2006; 9: 466-475.
- Pitout, J. D. & Laupland, K. B. Extended-spectrum beta-lactamase-producing Enterobacteriaceae: an emerging public-health concern. Lancet Infectious Diseases. 2008; 8: 159-166.
- Endimiani, A & Paterson, D.L. Optimizing therapy for infections caused by Enterobacteriaceae producing extended-spectrum b-lactamases. Seminars in Respiratory and Critical Care Medicine 2007 ; 28(6): 646-55.
- Pitout, J.; Nordmann. P.; Laupland, K. B. & Poirel, L. Emergence of Enterobacteriaceae producing extended-spectrum beta-lactamases (ESBLs) in the community. Journal of Antimicrobials & Chemotherapy., 2005; 56: 52-59.
- Kritu, P.; Prakash, G.; Shiba, K.R.; Reena, K.M.; RAM, N.S.; Ganesh. R. Antibiogram typing of gram negative isolates in different clinical samples of a tertiary hospital. Asian Journal of pharma-ceutical and clinical research. 2013; 6(1): 153-156.
- Meeta, S.; Sati, P.; Preeti, S. Prevalence and antibiogram of extended spectrum beta-lactamase (ESBL) producing gram negative bacilli and further molecular characterization of ESBL producing Escherichia coli and Klebsiella spp. Journal of Clinical Diagnosis and Research. 2013; 7(10): 2168-2172.
- Rupinder. B.; Geeta, W.; Shikha, J. Prevalence of extended spectrum b-lactamses in multidrug resistant strains of gram negative Bacilli. Journal of Academia and Industrial Research. 2013; 1(9): 558-560.
- Majda, Q.; Najma, A.; Summyia, B. Evaluation of extended spectrum beta-lactamase mediated resistance in Escherichia coli and Klebsiella in urinary tract infection at a tertiary care hospital. Biomedical Sciences, 2013; 29: 78-81.
- Paterson, D. L. & R. A. Bonomo. Extended-spectrum beta-lactamases: a clinical update. Clinical Microbiology Reviews, 2005; 18(4): 657-86.
- Peirano, G.; Richardson, D.; Nigrin, J.; McGeer, A.; Loo, V.; Toye, B.; Alfa, M.; Pienaar, C.; Kibsey, P.; Pitout, J.D.D. High prevalence of ST131 isolates producing CTX-M-15. Antimicrobial agents chemotherapy, 2010; 54(3): 1327-1330.
- Sibhghatulla, S.; Jamale, F.; Shazi, S.; Syed, M.; Danish, R.; Mohamed, A. K. Antibioitc resistance and extended spectrum beta-lactamases: Types, epidemiology and treatment. Saudi Journal of Biological Sciences, 2015; 22: 90-101.
- Ahmed, A.; Osman, H.; Mansour, A.M.; Musa, H.A; Ahmed, A.B; Karrar, Z.; Hassan H.S. Antimicrobial agent resistance in bacterial isolates from patients with diarrhea and urinary tract infection in the Sudan. American Journal of Tropical Medicine and Hygine, 2000; 63(5-6): 259-63.
- Ibrahim,M.E.; Bilal,N.E.; Hamid, M.E. Increased multi-drug resistant Escherichia coli from hospitals in Khartoum state, Sudan. African Health Sciences. 2012; 12(3): 368–375.
- Mekki, A.H.; Hassan, A.N; Elsayed, D.M. Extended spectrum beta lactamases among multidrug resistant Escherichia coli and Klebsiella species causing urinary tract infections in Khartoum. Journal of Bacteriology research , 2010; 11(3):18-21.
- Muna,Y. The Prevalence of Extended Spectrum b-lactamase and Amp-C. producing bacteria in a Sudanese tertiary hospital. Sudan.Medical Journal, 2015; 50(3).
- Yousif, M.; Alobaid, A.; Mohamed, E.; Ahmed, M.; Khalid, E.; Babiker, A.; Kamal, E. Phenotype and Genotype of Extended-spectrum-beta-Lacta-mases in Sudanes patients with UTI. Basic Research Journal of Microbiology, 2016; 3(4) ISSN 2354-4082
- Forbes, B.A.; Sahm, D.F.; Weissfield, A.S. Overview of bacterial identification- Methods and strategies. Baily and Scotts Diagnostic Microbiology, 12th edition. Mosby Elsevier, 2007.
- Andrews, J M. BSAC standardized disc susceptibility testing method (version 3). Journal of Antimicrobial Chemotherapy, 2014; 53: 713–28.
- Andrews, J M. Determination of Minimum Inhibitory Concentrations. Journal of Anti-microbial Chemotherapy, 2002; 49(6): 1049.
- Carter, MW; Oakton, KJ; Warner, M.; Livermore, DM. Detection of Extended-Spectrum beta-lactamases in Klebsiellae with the Oxoid combination disk method. Journal of Clinical Microbiology. 2000; 38: 4228-4232.
- Kim, J.; Lim,Y .M.; Jeong,Y.S.; Seol, S. Y. Occurrence of CTX-M-3, CTX-M-15, CTX-M-14, and CTX-M-9 extended-spectrum beta-lactamases in Enterobacteriaceae clinical isolates in Korea. Antimicrobial Agents and Chemotherapy, 2005; 49(4): 1572-5.
- Schwaber, M. J.; Raney,P.M.; Rasheed,J.K.; Biddle, J.W.; Williams, P.; McGowan, J.E.; Tenover, Jr. & F. C. 2004. Utility of NCCLS guidelines for identifying extended-spectrum beta-lactamases in non-Escherichia coli and Non-Klebsiella spp. of Enterobacteriaceae. Journal of Clinical Microbiology, 2004; 42(1): 294-8.
- Gray, K. J.; Wilson, L.K.; Phiri, A.; Corkill, J.E.; French, N.; Hart.C.A. Identification and characterization of ceftriaxone resistance and extended-spectrum beta-lactamases in Malawian bacteraemic Enterobacteriaceae. Journal of Antimicrobial Chemotherapy, 2006 ; 57(4): 661-5.
- Brigante, G.; Luzzaro, F.; Perilli, M.; Lombardi, G.; Coli, A.; Rossolini,G.M.; Amicosante,G.; Toniolo, A. Evolution of CTX-M-type beta-lactamases in isolates of Escherichia coli infecting hospital and community patients. International Journal of Antimicrobial Agents, 2005; 25(2): 157-62.
- Karisik, E.; Ellington, M.J.; Pike, R.; Warren, R.E.; Livermore,D.M.; Woodford,N. Molecular charac-terization of plasmids encoding CTX-M-15 {beta}-lactamases from Escherichia coli strains in the United Kingdom. Journal of Antimicrobial Chemotherapy, 2006; 58(3): 665-8.
- Blomberg, B.; Jureen, R.; Manji, K.P.; Tamim, B.S.; Mwakagile,D.S.; Urassa, W.K.; Fataki,M.; Msangi, V.; Tellevik,M.G.; Maselle,S.Y.; Langeland,N. High rate of fatal cases of pediatric septicemia caused by gram-negative bacteria with extended-spectrum beta-lactamases in Dar es Salaam, Tanzania. Journal of Clinical Microbiology, 2005; 43(2): 745-9.
- Ndugulile, F.; Jureen, R.; Harthug, S.; Urassa,W.; Langeland, N. Extended spectrum beta-lactamases among Gram-negative bacteria of nosocomial origin from an intensive care unit of a tertiary health facility in Tanzania. BMC Infectious Diseases, 2005; 5: 86.
- Gangoue-Pieboji, J.; Bedenic, B.; Koulla-Shiro, S.; Randegger, C.; Adiogo, D.; Ngassam, P.; Ndumbe, P.; Hachler, H. Extended-spectrum-beta-lactamase-producing Enterobacteriaceae in Yaounde, Cameroon. Journal of Clinical Microbiology, 2005; 43(7): 3273-7.
- Frank, T.; Arlet, G.; Gautier, V.; Talarmin,A.; Bercion, R. Extended-spectrum beta-lactamase-producing Enterobacteriaceae, Central African Republic. Emerging Infectious Diseases, 2006; 12(5): 863-5.
- Lavollay, M.; Mamlouk, K.; Frank,T.; Akpabie,A.; Burghoffer, B.; Ben Redjeb,S.; Bercion, R.; Gautier, V.; Arlet, G. Clonal dissemination of a CTX-M-15 beta-lactamase-producing Escherichia coli strain in the Paris area, Tunis, and Bangui. Antimicrobial Agents & Chemotherap, 2006; 50(7): 2433-8.
- Soge, O. O.; Queenan A.M.; Ojo,K.K.; Adeniyi,B.A.; Roberts,M.C. CTX-M-15 extended-spectrum (beta)-lactamase from Nigerian Klebsiella pneumoniae. Journal of Antimicrobial Chemother, 2006; 57(1): 24-30.
- Sabate, M.; Tarrago, R.; Navarro, F.; Miro E.; Verges, C.; Barbe,J.; Prats, G. Cloning and sequence of the gene encoding a novel cefotaxime-hydrolyzing beta-lactamase (CTX-M-9) from Escherichia coli in Spain. Antimicrobial Agents and Chemotherapy, 2000; 44(7): 1970-3.
- Rodriguez-Bano, J.; Navarro, M.D.; Romero, L.; Martinez-Martinez, L.; Muniain, M.A.; Perea, E.J.; Perez-Cano, R.; Pascual, A. Epidemiology and clinical features of infections caused by extended-spectrum beta-lactamase-producing Escherichia coli in non-hospitalized patients. Journal of Clinical Microbiology, 2004; 42(3): 1089-94.
- Chanawong, A.; M’Zali, F.H.; Heritage, J.; Lulitanond, A.; Hawkey,P.M. SHV-12, SHV-5, SHV-2a and VEB-1 extended-spectrum beta-lactamases in Gram-negative bacteria isolated in a university hospital in Thailand. Journal of Antimicrobial Chemotherapy, 2001; 48(6): 839-52.
- Gniadkowski, M.; Palucha, A.; Grzesiowski, P.; Hryniewicz,W. Outbreak of ceftazidime-resistant Klebsiella pneumoniae in a pediatric hospital in Warsaw, Poland: clonal spread of the TEM-47 extended-spectrum beta-lactamase (ESBL)-producing strain and transfer of a plasmid carrying the SHV-5-like ESBL-encoding gene. Antimicrobial Agents and Chemotherapy, 1998; 42(12): 3079-85.
- Mulgrave, L.; Attwood, P.V. Characterization of an SHV-5 related extended broad-spectrum beta-lactamase in Enterobacteriaceae from Western Australia. Journal of Pathology, 1993; 25(1): 71-5.
- Moubareck, C.; Daoud, Z.; Hakime, N.I.; Hamze, M.; Mangeney,N.; Matta,N.; Mokhbat, J.E.; Rohban, R.; Sarkis,D.K.; Doucet-Populaire, F. Countrywide spread of community- and hospital-acquired extended-spectrum beta-lactamase (CTX-M-15)-producing Enterobacteriaceae in Lebanon. Journal of Clinical Microbiology, 2005; 43(7): 3309-13.
- Pitout, J. D.; Thomson, K.S.; Hanson,N.D.; Ehrhardt, A.F.; Moland,E.S.; Sanders,C.C. beta-Lactamases responsible for resistance to expanded-spectrum cephalosporins in Klebsiella pneumoniae, Escherichia coli, and Proteus mirabilis isolates recovered in South Africa. Antimicrobial Agents and Chemotherapy, 1998; 42(6): 1350-4.
- Essack, S. Y.; Hall,L.M; Livermore,D.M. Klebsiella pneumoniae isolate from South Africa with multiple TEM, SHV and AmpC beta-lactamases. International Journal of Antimicrob Agents, 2004; 23(4): 398-400.
- Cardinale, E.; Colbachini, P.; Perrier-Gros-Claude,J.D.; Gassama, A.; Aidara-Kane,A. Dual emergence in food and humans of a novel multiresistant serotype of Salmonella in Senegal: Salmonella enterica subsp. enterica serotype 35:c:1,2. Journal of Clinical Microbiology, 2001; 39(6): 2373-4.
- Nuesch-Inderbinen, M. T.; Kayser.F.H.; Hachler.H. Survey and molecular genetics of SHV beta-lactamases in Enterobacteriaceae in Switzerland: two novel enzymes, SHV-11 and SHV-12. Antimicrobial Agents and Chemotherapy, 1997; 41(5): 943-9.
- Poirel, L.; Heritier,C.; Podglajen,I.; Sougakoff,W.; Gutmann.L.; Nordmann, P. Emergence in Klebsiella pneumoniae of a chromosome-encoded SHV beta-lactamase that compromises the efficacy of imipenem. Antimicrobial Agents and Chemotherapy, 2003; 47(2): 755-8.
- Tenover, F. C. Mechanisms of antimicrobial resistance in bacteria. American Journal of Medicine, 2006; 119(6 Suppl 1): S3-10; S62-70.
© The Author(s) 2019. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.


