ISSN: 0973-7510
E-ISSN: 2581-690X
Bacillus cereus JCM44 was isolated from the soil samples collected from the Kanyakumari district of Tamil Nadu, using a minimal agar medium. It was characterized by its ability to biotransform chitin into chitosan at 30°C for 48 h. The Chitosan (CS) was recovered by using 0.1N Sodium hydroxide and 2% acetic acid. It was confirmed by iodine solution with dark purple colouration and qualitatively identified by acetone and ethanol. CS formation has been identified from X-ray diffraction (XRD) and Fourier Transform Infrared Spectroscopy (FTIR). Scanning Electron Microscopy (SEM) analysis elucidated the formation of an average size of 349 nm to 1µm with the morphology of a non-porous, smooth membrane with irregular shape and crystallite. The Zeta potential study of the bacterially synthesized chitosan showed their excellent stability with a surface charge of -2.28 mV. Bacterially Synthesized Chitosan (BSC), with a polydispersity index (PDI) value of 0.626 indicates the aggregation of particles. The peaks of higher wavenumber confirm the formation of blend in Bacterially Synthesized Chitosan (BCS) at OH and NH was denoted. The XRD analysis of the BSC detected the crystallinity of the samples when compared to the commercial chitosan.
Bacillus cereus JCM44, Chitosan, SEM, DLS, XRD, FT-IR
Chitosan (CS) is a degradable biopolymer and it’s a derivative of chitin.1 Though, the supply of marine resources is variable, seasonal and very expensive.2 CS has distinctive properties with various purposes in biomedical and is mostly applied as elicitors in agriculture-based sectors.3-4 However, CS is a polysaccharide generated by the deacetylation of chitin, using harsh chemicals may decrease the quality of CS and high in its processing rate. Initially, various fungi such as Phycomyces blakesleeanus, Mucor rouxii are for the production of a deacetylated product of chitin.5-6 Simple nutrients are used to culture such microbes, which helps to recover the CS without difficulty.7-9
Fermentation technologies have been used, recently for culturing organisms that contain CS. This approach could be an attractive method for the production of biopolymer CS. In most fungi, CS is found in the outer surface (cell wall), where it likely shows an essential part as a glycan. Chitin deacetylase (CDA) has a role in biotransformation of chitin to chitosan by the hydrolysis of acetamido group.10 Some of the investigators studied the production and the isolation of CS from microbes broadly as an alternative to marine shell waste-derived product, as in Aspergillus sp.11 Most of the fungal strains have low CDA-producing capacity and their fermentation requirements are much more difficult. Therefore, new appropriate potential strains of bacteria which produce CDA should be required to replace the fungal strains. Numerous bacterial species which produce CS were reported early, including Bacillus thermoleovora.12
The problems of chemical processing should be transformed using various microorganisms,13-15 and proteolytic enzymes16-17 used to remove the proteins and mineral content. The production of microbial enzymes either in pure or partially purified form is an important aspect of bio-based industries. The applications of microbial enzymes in the industry are very diverse.18 Therefore, the use of specific microbial CDAs to catalyze the conversion of chitin to CS was considered interesting due to certain advantages over chemical reactions. In microbial fermentation, deproteinization follows through the activity of microbial proteases and demineralization by the acid produced by microorganisms during fermentation.19 In this present study, CS was produced by fermentation using Bacillus cereus JCM44 and the characteristics of synthesized CS were analyzed.
Materials
Chitosan (Shrimp shell) with the deacetylation degree of ≥ 75%, chitin
(CAS 1398-61-4, degree of deacetylation: 65-70%) and all the chemicals with analytical grade were purchased and used from Himedia Laboratories Pvt. Ltd. The purchased chemicals were used as it is without any further purification.
Isolation of CDA producing bacteria
Coastal soil samples of 5 to 10 cm depth were collected from different locations in Kanyakumari district, Tamil Nadu, India. One gram of soil sample was taken and dissolved in 9mL of sterile distilled water and serially diluted up to 10-9. The Minimal medium (MM) containing (w/v) 1% chitin was added as a sole source of carbon, 0.2% Sodium Nitrate (NaNO3), 0.1% Dipotassium hydrogen Phosphate (K2HPO4), 0.1% Potassium Dihydrogen Phosphate (KH2PO4), 0.05% Magnesium Sulphate (MgSO4) and 2% Agar were used for the isolation of bacterial isolates and incubated at 35°C for 24 to 48 h.20,21 Using a diagnostic strip test, the bacterial isolates producing CDA were screened for CDA activity. A potent bacterial isolate (JCM44) was selected based on quantitative screening. Physiological and biochemical tests of JCM44 like catalase, oxidase, indole, Methyl Red-Voges Proskauer (MR-VP) and citrate utilization tests were performed. 16S rDNA sequencing was done by 27F (5’- AGAGTTTGATCMTGGCTCAG -3’) and 1492R (5’- TACGGYTACCTTGTTACGACTT -3’).22 The PCR product was purified, sequenced and submitted to National center for Biotechnology Information (NCBI). An accession number was obtained.
Chitin to CS – Biotransformation (JCM44)
For the biotransformation of Chitin to CS the medium containing (1g of yeast extract, 0.4g of Ammonium Sulfate (NH4)2SO4, 0.15g of KH2PO4 with a pH of 8.0 and chitin (50 mg) was added along with it. The medium was taken in a conical flask (250 mL) and (1%) inoculum of the positive isolate JCM44 was added along with it and one of the flasks without inoculum was taken as control. The flasks were incubated for 2 days. Flasks were taken after incubation for CS recovery.23
CS recovery
The incubated flasks were taken for CS recovery. The broth (fermented) was taken and centrifuged (12000 rpm) for 15 min. The pellet attain was taken and 10mL of (0.1N) sodium hydroxide was added along with it. After the content was mixed, it was autoclaved for 15 min. The tubes were taken and allowed for cooling. In the alkaline action, all the cell debris got solubilized. Again the tubes were centrifuged (12000 rpm) for 15 min and the pellet was collected. Acetic acid (2% 10 mL) was added to the pellet and the pellet was left overnight at room temperature for the solubilization of CS. Again the tubes were centrifuged (12000 rpm) for 15 min the very next day. The pellet was collected and (1N) sodium hydroxide was added along with it. A white precipitate was formed which shows the presence of CS.21
Qualitative Estimation of CS
Centrifuged (5000 rpm) the precipitate again for 15 min. and washed using distilled water twice to attain pH 7. Again the precipitate was resuspended in 0.5mL dis. H2O and it was kept for drying at 55°C for 2 to 4 h. The confirmatory test was done to the attained precipitate. To confirm the precipitate as CS, iodine solution (a few drops) was added to the precipitate, it turns the precipitate dark brown and to acidify, 1% Sulphuric Acid (a few drops) was added along with it. It changes its colour from dark brown to purple, which demonstrates the presence of CS.21
Quantitative Estimation of CS
The clean Petri plates were taken and weighed. The precipitated CS was washed thoroughly with distilled water (pH 7) and resuspended in 1 mL of dis. H2O (pH 7). 1mL of chitosan suspension was poured on the clean Petri plates and kept for drying at 55°C for (2–6) h. The precipitate CS was weighted after drying.21
Characterization of Bacterially synthesized chitosan (BSC)
Scanning Electron Microscope (SEM)
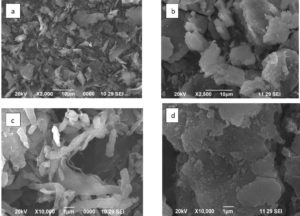
The extracted BSC was analyzed with Scanning Electron Microscope (SEM) (Jeol JSM-Model SEM 6390, Japan). The dried samples of BSC images were observed and compared with the commercial chitosan at 10,000x magnification at an accelerated voltage of 20 kV.
Dynamic Light Scattering (DLS)
Zeta potential and particle size were measured for BSC, (Zetasizer, Nano ZS, Malvern Instruments Ltd.,) based on a DLS technique. The stability of the BSC was measured at a stable measurement temperature of 25°C, and a scattering angle of 90°. Photon Correlation Spectroscopy (PCS) was used for the size evaluation of BSC.
The polydispersity index (PDI)
The homogeneity dispersion of BSC is a measure of PDI (0 to 1). The homogeneity is shown by values closer to 0 (zero) and non-uniformity values (> 0.5).
X-ray Diffraction Method (XRD)
XRD data of the diffraction pattern of BSC was obtained with Shimadzu-model XRD 6000, operated at a voltage of 40 kV and a current of 30 mA with Cu, Ka radiation in a 2θ configuration from 10-60°.
Fourier-transform infrared spectroscopy (FT-IR) analysis
The IR spectra of BSC were analyzed with FT-IR Spectrophotometer, Perkin Elmer Make-Model spectrum, RX over the frequency series of 4000 – 400 cm−1. The results were compared to commercial (Shrimp shell) chitosan.
Isolation and Identification
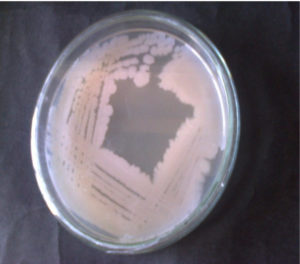
In this study, the highest CDA-producing bacterial isolate JCM44 Figure 1a. The morphology of strain JCM44 was milky-white with a smooth surface, rod and Gram-positive. CDA-producing bacterial isolate JCM44 was exposed to a strip test where p-nitroacetanilide was used as an indicator. The bacterial isolate JCM44 showed a positive result by producing yellow colour in chitin agar broth supplied with indicator p-nitroacetanilide Figure 1b. Also, the isolate was positive for catalase, oxidase, MR-VP, citrate utilization test and negative for indole. 16S rDNA sequencing was done to isolate JCM44 and deposited in the NCBI with accession number MG917734.1. The sequence has high homology to Bacillus cereus and with greater than 99% of sequence identity and a phylogenetic tree was constructed. Various researchers have found that marine bacteria were reported that major producers of CDA including Bacillus sp.21,24,25
Production of CS
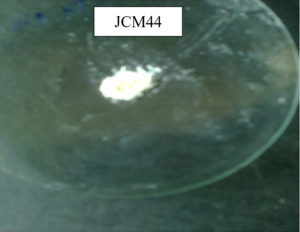
The selected bacterial isolate Bacillus cereus JCM44 has shown the highest chitin degradation was cultivated in a production medium containing chitin Figure 2 (a). A white colour powder was obtained after the incubation, centrifugation and treatment with acetic acid and NaOH indicating the CS released from raw chitin. The resulting precipitate was further confirmed by its reaction with iodine solution Table. Once the isolate was proven to release CS from the chitin, its yield was also determined. The yield of chitosan by Bacillus cereus JCM44 was 60mg/L as maximum Figure 2(b). Early reports of CS production from certain fungi shows, 0.12 g/L of CS in liquid medium,26 78.3 mg/L of CS by A. niger, 1.8g/L of CS by Absidia coerulea.27,28 In this study, the production of CS by the bacterial isolate Bacillus cereus JCM44 is not much more when compared to fungal isolates but by enhancing the medium conditions of CDA production will be increased for the production of CS. Hence, this bacterial isolate can be used on an industrial scale for the biotransformation of chitin to CS evidencing it to be a hopeful isolate for a biologically approachable process.
Table : Qualitative Estimation of BCS.
Flask Type |
Fermented broth of isolate Bacillus cereus JCM44 |
Control flask (Not inoculated) |
|---|---|---|
Chitosan recovered in Neutralization (2% acetic acid) with (1N) NaOH |
10ml of 2% Acetic acid White precipitate observed |
10ml of 2% Acetic acid No precipitate observed |
Iodine reaction |
Dark purple coloration |
NIL |
Figure 2 (a). Biotransformation of chitin to chitosan using Bacillus cereus (JCM44) and its recovery
SEM
The SEM analysis of BSC showed that it has a non-porous, smooth membrane with irregular shape and crystallite in nature; this structural modification may be due to aggregation of the particles, this was due to their high surface charge Figure 3 (b&d). This was in agreement with the previous data reported in CS from B. portentous.29 The morphology of the BSC was different from that of commercial CS, Figure(3a) which has only the fibrous exterior could be owing to the extraordinary molecular lining. Rough layers of flakes and pores were seen in lower magnification (×2,500) Figure 3b. At ×10,000 magnification, BSC has pores with smooth and round surfaces; crumbling flasks have seen in Figure 3d. Next to Figure 3(d) when compared to the commercial CS Figure 3c in the previous reports.30-34 SEM images showed larger pores in insects and commercial chitin due to the separation of chitin and CS synthesis from various parts of the insects.
DLS
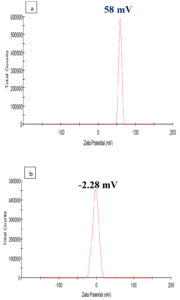
DLS is used to determine the stability and particle size in the suspension based on surface charge. The stability of the BSC particles synthesized was related to the commercial CS in the present study and was measured by using Zetasizer and is shown in Figure 4 (a & b). The BSC particles and commercial CS were diluted to dis. H2O and dimensions of the particles were achieved at the temperature of 25°C in triplicates. The Zeta potential of -2.28 mV of BSC was perfect for the physical constancy of any suspension when compared to the commercial CS which has a zeta potential of +58.9 mV. The results conclude that the BSC is more stable than the commercial chitosan. A similar trend was observed by35 works, which presented the zeta potential value (4mV) of EA@CSNP, which shows the nanoparticle’s optimistic surface charge with less stability. Due to the presence of drugs on the polymer surface, the Zeta potential values should be decreased. For the physical stability, the particle charge shows > + 30mV and < -30mV is ideal of any suspension. Similar trends were found for chitosan nanoparticles (CSNPs) synthesis by36 shown + 30mV to -5mV which might be the occurrence of alkali on the particles.
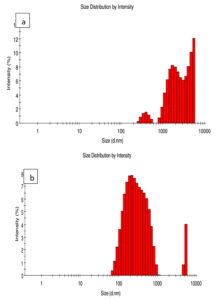
Polydispersity Index (PDI)
The PDI describes the degree of “non-uniformity”, and simplifies thoughtful of scattering and accumulation of the particles in the medium. The dimension scattering of BSC and commercial CS is shown in Figure 5 (a & b). PDI values reflect the dimension scattering. PDI values will be shown based on the higher and lower range of particles. The PDI value of BSC is shown as 0.0626 and the PDI value for commercial CS was 0.534; this shows the heterogeneous nature of the particles. It was reported that PDI more than 0.5 is indicative of aggregation of CS particles.37 This reveals the aggregation of the particles also size distribution of BSC is smaller than the commercial CS. Similar particle size distribution for CSNPs with heterogeneous dispersion was reported.38
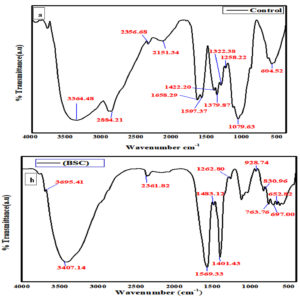
FT-IR analysis
FT-IR spectra were done to detect the structural groups of BSC. The CS from Bacillus cereus JCM44, as related to the physical property with the commercial CS used as a control Figure 6(a&b). The IR studies had shown strong absorption bands of 3695, 3407, 2361, 1569, 1262, 928, 830, 763, 697 and 652 cm-1. In Commercial CS IR studies had shown, the bands at 3364, 2884, 2356, 2151, 1653, 1597, 1422, 1379, 1322, 1258, 1079, 604 cm-1 that between 3364 and 697 cm-1. Absorption bands at 3695 cm-1 of BSC specify O-H and NH bond indicates the reduced hydrogen bonding. Absorption bands at 2361, 1401 (–C≡C–, C-H) and 1569 cm-1 (N-O) reveal alkenes and nitro groups. Absorption bands at 697 cm-1 indicated alkane C-H bending. The absorption bands show, commercial CS at the various peaks 1597, 1422, 1379, 1322, 1258, 1079, and 604 cm-1, which indicates the stretching absorption peaks (aromatic amine and alkane groups). Also, the IR spectral peaks of commercial CS of 3364, 2884, 2356, 2151, and 1653 cm-1 were comparable to BSC. This demonstrates the confirmation of CS. Similar reports have been reported in39 and also similar results of IR stretching peaks observed in polysaccharide (Chitosan) prepared by processing prawn waste (shell) were revealed by.40
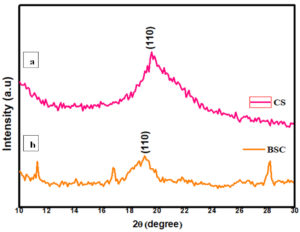
XRD
The crystalline nature and phase purity of the prepared and commercially obtained sample were analyzed using powder XRD diffraction. The high intense peak was observed at 19.3°and 19.1° of 110 planes for the CS and BSC, respectively, which correspond to the characteristic peak of chitosan. Comparatively, a slight decrement in the intensity of the peak for BSC sample was observed which might be due to the low crystalline nature of the synthesized chitosan. Thus, this confirms the formation of CS Figure 7 (a & b). In addition, a few peaks other than the characteristic peak of the chitosan were also observed. The presence of such peaks reveals the trace amount of materials in the sample which were used to prepare BSC. Similar reports were showed in fungal CS and Shrimp shell chitosan where the same major characteristic peaks were observed at 19.9° and 19.8-20.7°.41-44
It can be inferred from the present study that CDA secreted by Bacillus cereus JCM44 appears to be a prospective strain for the biotechnological application for the production of CS. The BSC was characterized by FT-IR, XRD, SEM, DLS (Zeta potential) and polydispersity index (PDI) analysis and compared with commercial CS. The BSC was irregular in shape with a smooth surface. FTIR reveals the presence of OH, NH groups, alkenes and amine groups of chitosan. The crystalline nature of synthesized chitosan was confirmed with XRD analysis. The Zeta potential and the Polydispersity index showed the stability of the synthesized bacterial chitosan and it revealed that the particle was heterogeneous which indicates the aggregation of the particles. Significant production of CS by enhancing bacterial isolates helps the financial, feasible and ecologically approachable process. Such synthesized CS is biocompatible, having the assets to be recycled for bio-based requirements. The vital data attained contributes to our knowledge of the capabilities and possibilities of Bacillus cereus JCM,44 and successfully reduced the expenses of fabrication of CS.
ACKNOWLEDGMENTS
The authors would like thank Department of Biotechnology, Periyar University, Salem, Tamil Nadu, India for strengthening the instrumentation facility.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
SI designed the research work, carried out all the experiments and wrote the manuscript.
IAP designed the research work, supervised, validated and reviewed the manuscript. Both authors read and approved the final manuscript for publication.
FUNDING
This work was supported by the University Grants Commission (UGC)-MANF Fellowship (Awarded on F1-17.1/2013-14/MANF-2013-14-CHR-TAM-25806 /(SA-III/Website) dt.06-Feb-2014, Government of India.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
This article does not contain any studies on human participants or animals performed by any of the authors.
- Raval R, Simsa R, Raval K. Expression studies of Bacillus licheniformis chitin deacetylase in E. coli rosetta cells. Int J Biol Macromol. 2017;104(Part B):1692-1696.
Crossref - Crestini C, Kovac B, Giovannozzi-Sermanni G. Production and isolation of chitosan by submerged and solid-state fermentation from Lentinus edodes. Biotechnol Bioeng. 1996;50(2):207-210.
Crossref - McGahren WJ, Perkinson GA, Growich JA, Leese RA, Ellestad GA. Chitosan by fermentation. Process Biochem. 1984;19(3):88-90.
- Shahidi F, Arachchi JKV, You JJ. Food applications of chitin and chitosan. Trends Food Sci Tech. 1999;10(2):37-51.
Crossref - Muzzarelli RAA, Chitin Oxford: Pergamon Press. 1977:82-252.
- Knorr, D., Beaumont, M. & Pandya, Y. Potential of acid soluble and water soluble chitosan in biotechnology. In G. Skjak-Braek, T. Anthonsen & P. Sanford (Eds) Chitin and chitosan London, New York: Elsevier Appl. Sci. 1989: 101-118.
- White SA, Farina PR, Fulton I. Production and isolation of Chitosan from Mucor rouxii. Appl Environ Microbial. 1979;38(2):323-328.
Crossref - Shimahara K, Takiguchi Y, Kobayashi T, Uda K, Sannan T. Screening of mucoraceae strains suitable for chitosan production. In Chitin and Chitosan, ed. G. Skjak-Braek, Anthonsen T, Sandford P. Appl Sci. 1989:319-332.
- Synowiecki J. Use of krill chitin as an enzyme support. In Chitin in Nature and Technology, ed. Muzzarelli R A A, Jeuniaux C, Gooday G. Plenum Press, New York, 1986:417-420.
- Ma Q, Gao X, Tu L, et al. Enhanced Chitin Deacetylase Production Ability of Rhodococcus equi CGMCC14861 by Co-culture Fermentation With Staphylococcus sp. MC7. Front Microbiol. 2020;11:592477.
Crossref - Natarajan K, Riyaz AB, Sathish R. Production and Evaluation of chitosan from Aspergillus Niger MTCC strains. Iran J Pharm Res. 2011;10(3):553-558.
Crossref - Toharisman A, Suhartono MT. Partial Purification and Characterization of Chitin Deacetylase Produced By Bacillus Thermoleovorans LW-4-11, Scientific Repository, IPB Bogor Agricultural University. 2008.
- Wang SL, Chio SH. Deproteinization of shrimp and crab shell with the protease of Pseudomonas aeruginosa K-187. Enzyme Microb Technol. 1998;22(7):629-633.
Crossref - Cira LA, Huerta S, Hall GM, Shirai K. Pilot scale lactic acid fermentation of shrimp wastes for chitin recovery. Process Biochem. 2002;37(12):1359-1366.
Crossref - Shirai K, Guerrero I, Huerta S, et al. Effect of initial glucose concentration and inoculation level of lactic acid bacteria in shrimp waste ensilation. Enzyme Microb Technol. 2001;28(4-5):446-452.
Crossref - Gagne N, Simpson BK. Use of proteolytic enzymes to facilitate the recovery of chitin from shrimp wastes. Food Biotechnol. 1993;7(3):253-263.
Crossref - Broussignac P. Chitosan a natural polymer not well known by the industry. Chim Ind Genie Chim. 1968;99:1241-1247.
- Akila RM. Fermentative production of Fungal chitosan, a Versatile Biopolymer (perspective and its applications). Adv Appl Sci Res. 2014;5(4):157-170.
- Rao MS, Munoz J, Slevens WF. Critical factors in chitin production by fermentation of shrimp biowaste. Appl Microbiol Biotechnol. 2000;54(6):808-813.
Crossref - Zhou G, Zhang H, He Y, Li He. Identification of a chitin deacetylase producing bacteria isolated from soil and its fermentation optimization. Afr J Microbiol Res. 2010;4(23):2597-2603.
- Kuldeep K, Vikrant D, Vikas S, Uma B. Isolation and Characterization of Chitosan-Producing Bacteria from Beaches of Chennai, India. Enzyme Res. 2012;6:421683.
Crossref - Lane D. 16S/23S rRNA sequencing. In: Stackebrandt, E, Goodfellow, M. (Ed.), Nucleic Acid Techniques in Bacterial Systematics. John Wiley and Sons, Inc., New York, 1991:115-147.
- Vadake RS. Biotransformation of Chitin to Chitosan, United States Patent 5739015, 1998.
- Sherin I, Arulselvi PI. Isolation and evaluation of potential CDA producing bacterial strain Bacillus cereus (JCM44) isolated from coastal areas of south India. Res J Biotechnol. 2020;15(6):1-9. URL:http://www.scopus.com/inward/record.url?eid=2-s2.0-85090902919&partnerID=MN8TOARS
- Ischaider, Hasnah N, Seniwati D. Production and application of chitin deacetylase from Bacillus licheniformis HSA3-1a as Biotermicide. Marina Chim Acta. 2014;1(1):8-12.
Crossref - Jin Hu K, Hu JL, Ho KP, Yeung KW. Screening of fungi for chitosan producers, and copper adsorption capacity of fungal chitosan and chitosanaceous materials, Carbohydr Polym. 2004;58(1):45-52.
Crossref - Muzzarelli RAA, Ilari P, Tarsi R, Dubini B, Xia W, Chitosan from Absidia coerulea. Carbohydr Polym. 1994;25(1):45-50.
Crossref - Davoust N, Persson A, Effects of growth morphology and time of harvesting on the chitosan yield of Absidia repens. Appl Microbiol Biotechnol. 1992;37(5):572-575.
Crossref - Kucukgulmez A, Celik M, Yanar Y, Sen D, Polat H, Kadak AE. Physicochemical characterization of chitosan extracted from Metapenaeus stebbingi shells. Food Chem. 2011;126(3):1144-1148.
Crossref - Ifuku S, Nomura R, Morimoto M, Saimoto H. Preparation of chitin nanofibers from mushrooms. Materials. 2011;4(8):1417-1425.
Crossref - Kaya M, Seyyar O, Baran T, Erdogan S, Kar M. A physicochemical characterization of fully acetylated chitin structure isolated from two spider species: with new surface morphology. Int J Biol Macromol. 2014;65:553-558.
Crossref - Kaya M, Baran T. Description of an new surface morphology for chitin extracted from wings of cockroach (Periplaneta americana). Int J Biol Macromol. 2015f; 75:7-12.
Crossref - Kaya M, Erdogan S, Mol A, Baran T. Comparison of chitin structures isolated from seven orthopteran species. Int J Biol Macromol. 2015b;72:797-805.
Crossref - Kaya M, Bulut E, Mujtaba M, et al. Gender influences differentiation of chitin among body parts. Arch Insect Biochem Physiol. 2016;93(2):96-109.
Crossref - Arya G, Vandana M, Acharya S, Sahoo SK. Enhanced anti proliferative activity of Herceptin (Her2) conjugated germcitabin- loaded chitosan nanoparticle in pancreatic cancer therapy. Nanomedicine. 2011;7(6):859-570.
Crossref - Ilaiyaraja N, Devi A, Khanum F. Chlorogenic acid loaded chitosan nanoparticles with sustained release property, retained antioxidant activity and enhanced bioavailability. Asian Journal of Pharmaceutical Sciences. 2015;10(3):203-211.
Crossref - Bathool A, Vishakante GD, Khan MS, Shivakumar HG. Development and characterization of atorvastatin calcium loaded chitosan nanoparticles for sustained drug delivery. Adv Mat Lett. 2012;3(6):466-470.
Crossref - Sujima Anbu A, Sahi SV, Venkatachalam P. Synthesis of Bioactive Chemicals Cross-linked Sodium Tripolyphosphate (TPP) – Chitosan Nanoparticles for Enhanced Cytotoxic Activity against Human Ovarian Cancer cell Line (PA-1). J Nanomed Nanotechnol. 2016;7(6):1-9.
Crossref - Puvvada YS, Vankayalapati S, Sukhavasi S. Extraction of chitin from chitosan from exoskeleton of shrimp for application in the pharmaceutical industry. Int Curr Pharm J. 2012;1(9):258-263.
Crossref - Sneha P, Aiswarya J, Changam Sheela S, Sanjay M Cherian. Extraction And Purification Of Chitosan From Chitin Isolated From Sea Prawn (Fenneropenaeus Indicus). Asian J Pharm Clin Res. 2014;7(4):201-204.
- Duan W, Chen C, Jiang L, Li GH. Preparation and characterization of the graft copolymer of chitosan with poly[rosin-(2-acryloyloxy)ethyl ester]. Carbohydr Polym. 2008;73(4):582-586.
Crossref - Ramya R, Sudha P.N, Mahalakshmi J. Preparation and Characterization of Chitosan Binary Blend. International Journal of Scientific and Research Publications . 2012;2(10):1-9.
- Yen M-T, Mau J-L. Physico-chemical characterization of fungal chitosan from shiitake stipes. LWT – Food Sci Technol. 2007;40(3):472-479.
Crossref - Prashanth KVH, Kittur FS, Tharanathan RN. Solid state structure of chitosan prepared under different N-deacetylating conditions. Carbohydr Polym. 2002;50(1):27-33.
Crossref
© The Author(s) 2023. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.