ISSN: 0973-7510
E-ISSN: 2581-690X
The anthropogenic use of mercury (Hg) has led to the wide outlook to the problem and health concerns caused by it. Metagenomic approach has opened the galore to the ease of access to the gene pool. The permissible limit of Hg is 0.001ppm but there are some sites in India having very high concentration of Hg (Ulhas estuary, Mumbai, having 107 ppm Hg). Library of clones containing mer operon, was constructed using culture independent technique. Out of 150 clones studied, UC07 has shown growth at 70ppm Hg. It has also shown antibiotic resistance towards vancomycin (30 mcg), Kanamycin (30 mcg) and Norfllaxin (10 mcg). Clone has the capability to volatilize 97.94% of 5ppm Hg which gradually decreased to 47% with the increase in Hg to 70ppm. Validation study using actual site (Ulhas soil) showed the effective conversion of Hg up to 99.9% by UC07. The clone contains both merA and merB genes, so it can be effectively utilized for organic and inorganic contaminated site. This is the first study of validation of clone using Hg contaminated site. The results are quite promising which shows that UC07 can be utilized effectively for bioconversion of both organic as well as inorganic form of Hg contamination.
Mercury, metagenomic, mercury resistance, mer operon, antibiotic resistance, mercury volatilization
Mercury (Hg) is naturally occurring element found in environment and amongst the top most toxic elements in the plethora of 6 million substances and found in earth crust at varied array from 21 (lower crust) to 56 (upper crust) ppb1 in the elemental form and mixture of minerals (hypercinnabar, metacinnabar and cinnabar etc.)2 It is one of the most widely used heavy metal due to its characteristic feature to form amalgams with other metals (dental amalgams or gold extraction) and also in agriculture as bactericides, fungicides, insecticides and herbicides.
These anthropogenic usages of mercury have led to pollute the water bodies and surrounding soil by direct and indirect discharge of mercury, which in turn has turn hazardous to the organisms living in such habitat and is also one of the major concerns to human health as it could lead to neurotoxic diseases. The need to study on mercury elevated because of its usage as topical disinfectant and antiseptics in the community and hospitals. It has been estimated that 800 tons of Hg(II) is released in environment due to weathering of 1010 tons of rocks globally every year, furthermore 200 million tons of mercury is estimated to be present in oceanic reservoir. The concentration of dissolved Hg (II) in water varies from 0.03 to 2.0 sg/liter (i.e., up to 10-8 M), in the midst of such surroundings, it is not astonishing that bacteria have evolved with the mechanism to overcome such toxic compound. The cluster of genes responsible for mercury resistance was discovered as mercury operon (mer operon) consisting of two major genes merA and merB. merA genes, encoded in structural gene found on mer loci or transposon 501. Also, evolutionary studies depict the presence of merA gene in some microorganism due to horizontal gene transfer. Gene merA encodes for enzyme known as “mercury reductase” that converts the inorganic form of mercury (Hg2+) to volatile and less toxic form of mercury (Hgo) and merB genes encodes for organomercury lyase that breaks down organic form of mercury (CH3 Hg + ) to Hg2+ . These enzymes are found in the cytoplasm of the cell. In addition, other genes are also present like genes for transport (merT)3. Depending upon the presence of gene in bacteria two type of resistance mechanisms are found, termed as Broad spectrum- where the organism carries both the genes merA and merB conferring resistance against HgR and the other as Narrow spectrum: where only merA gene is present providing resistance to bacteria against only Hg2+ . As per various researches it has been found that merB gene is present upstream to merA gene conferring resistance to both organic and inorganic form of mercury3,4. The increase in Hg pollution has augmented antibiotic resistance5. The presence of antibiotic resistance is thought to be linked with the presence of mercury found in environment6.
There are few sites in India like Ulhas estuary, Kodai kanal and Panipat industrial area which have surpassed the permissible limit of Hg (1×10-3 ppm) as per Indian standards and WHO and has entered the food chain. One of such site is Ulhas estuary containing 107ppm of Hg concentration7. This study was undertaken with the objective to obtain a clone (containing mer operon) which has the ability to withstand high concentration of Hg as well as efficiently convert its organic form to in-organic toxic form (Hg2+) and further transformation into non-toxic volatile form (Hgo).
Sample collection and analysis of Hg concentration in soil
Soil sample was collected from Ulhas estuary, Thane, Maharashtra. The soil was taken from 10 cm beneath the surface in the clean container and was immediately stored at 4°C till further use. CV-AAS analysis of the soil was done by digesting it by mixing it with equal volumes of water and nitric acid and heated till the volume reduced to one third of original volume. Later the content was filtered through Whatman filter paper and reading was taken for Hg volatilization.
Metagenomic library construction of (mer operon)
Metagenomic DNA was isolated from the soil sample by using MOBIO kit by given protocol. DNA quantification was done by diluting the DNA up to 500 folds and taking UV absorption at 260nm. Partial restriction digestion was performed to get the DNA in range of 5-10kb for cloning. 100ml digestion mixture was setup in sterile micro-centrifuge tubes which had 80ml of DNA, 9ml of sterile double distilled water, 10ml of 10X buffer solution and 1ml of Sau 3A1 restriction enzyme. The reaction was setup in the ice and immediately transferred to 16°C cold water bath for 4h. Reaction was stopped by heating the mixture at 70°C. The partially digested metagenomic DNA was subjected to gel electrophoresis in 1% agarose andviewed under UV light. DNA fragments in the range of 5-10kb were eluted using HiYield Gel/PCR DNA Mini Kit (Real Biotech Corporation, Taiwan). Partially digested DNA (400ng) in range of 5-10kb was ligated to pUC 19 vector (predigested with BamHI) and dephosphorylated using T4 DNA ligase enzyme at 4°C for 16h in micro-centrifuge tubes the reaction was terminate by heating the reaction mixture at 70°C. The ligated DNA product was introduced into the host bacteria i.e. E.coli DH10B by electroporation at 200W, 25mf and 2.5 kV, which was carried out by Micropulser II (Bio-Rad). The reaction mixture was mixed with 1ml of Luria Broth (LB), 100ml was plated on Luria Bertani (LB) agar plates supplemented with ampicillin, X-Gal and IPTG. The plates containing transformants were kept in Incubator at 37°C for 16h.
Screening and Selection of mercury resistant clones
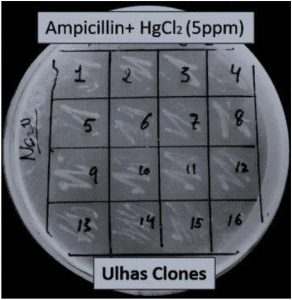
White colonies were picked and streaked on the LB agar plates supplemented with 5ppm of Mercury Chloride (HgCl2) and kept in incubator at 37°C for 18-72h. The clones which were able to grow at 5ppm of Hg were inoculated in LB supplemented with 5-90ppm of HgCl2 and the ones showed growth at maximum Hg concentration were chosen for acclimatization. The cultures were inoculated in LB with 5ppm HgCl2 and incubated at 37°C for 48h. It was further transferred to 10, 20, 30 till 80ppm of HgCl2 . OD was taken at 600nm.
The clone which has shown growth at highest concentration of HgCl2 was selected for further studies. Since the clone was able to grow in presence of HgCl2, indicated that it carries merA gene.It was also examined for presence of merB gene by inoculating it in LB with CH3HgCl2 (5ppm and 10ppm) and incubating at 37°C at 180rpm for 24 – 48 h. OD was taken at 600nm. The clones were also inoculated in LB supplemented with 5ppm-80ppm HgCl2 respectively for cfu count and incubated at 37°C for 18-24h. Overnight grown culture (100µl) was plated onto LB agar and incubated at 37°C for 16h. CFU was calculated by using online CFU athttp://openCFU.source-forge.net/8.
Quantification and validation of Hg volatilization
Since the selected clone indicated the presence of both merA and merB genes, it was checked for its ability to convert in-organic toxic form of Hg to non-toxic volatile form. The clone was grown in LB supplemented with 10, 20 up to 80 ppm of HgCl2 and, incubated at 37°C at 180rpm. The culture was withdrawn on 2nd , 4 th and 6 th day. Culture was centrifuged at 6000rpm for 10 minutes and supernatant was used for quantification of Hg volatilization by CV-AAS (Analytical Jena make (Vario-6) cold vapor HG-AAS).
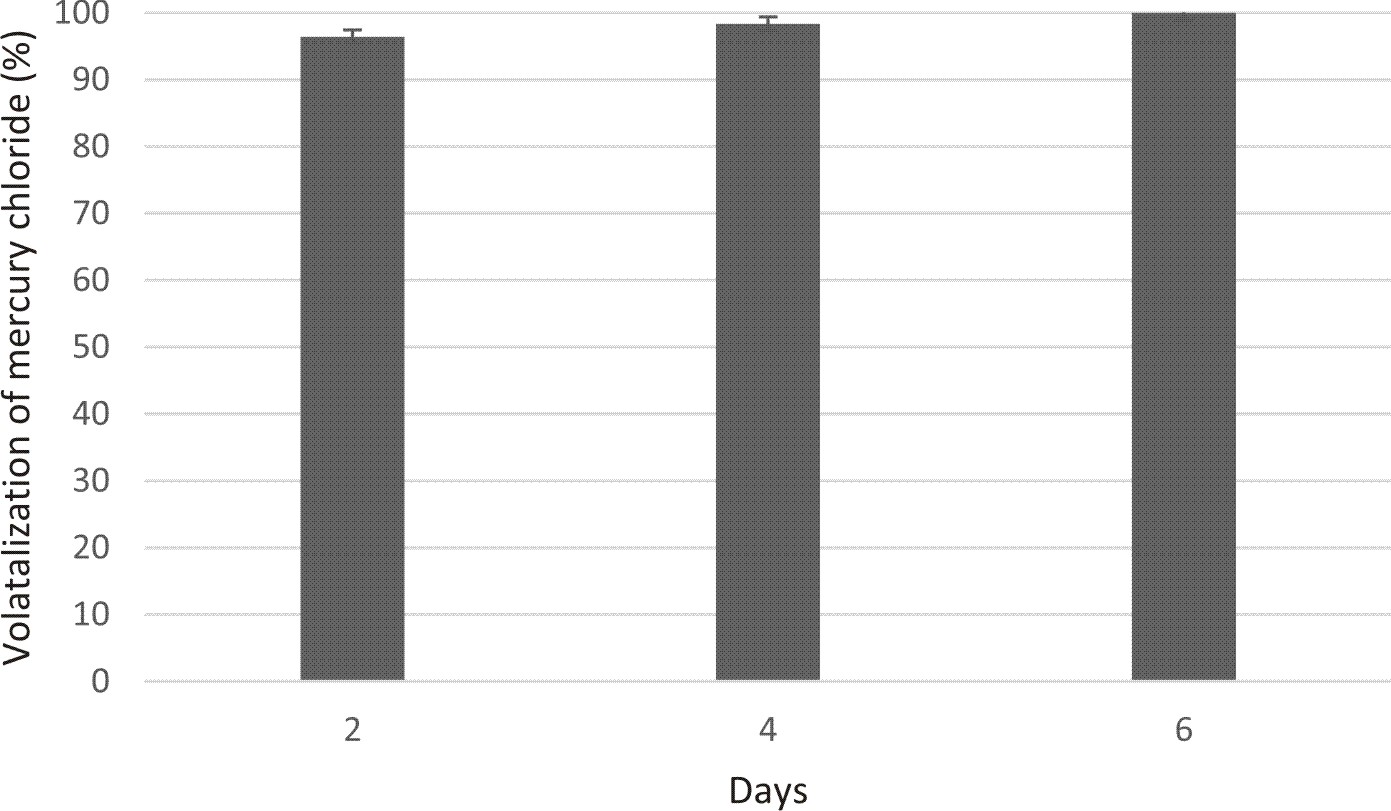
For validation, the Ulhas soil sample (without autoclaving) was mixed with sterile soil (3:2wt/wt) to make 10 kg of mixture with Hg concentration to approx. 60 ppm. 500ml (1 OD) of overnight grown culture of selected clone was sprayed on the diluted soil. Samples were withdrawn on 2nd , 4th and 6th day respectively. CV-AAS was done to check the ability of clone to transform toxic form of mercury to non-toxic form as an application to original site.
Antibiotic resistance
Antibiotic resistance was checked through disk diffusion method against Streptomycin (10 and 25 mcg), Norflaxin (10mcg), Tetracyclin (30mcg), Ampicillin (10, 20 and 30mcg), Kanamycin (30mcg), Vancomycin (30mcg), Gentamycin (30mcg). Zone of inhibition (in cm) was measured after 18h of incubation.
Identification of the mercury determinants
(a) Genomic DNA Isolation
Culture was inoculated into 10 mL of nutrient broth and incubated overnight at 37°C. Culture was centrifuged at 6,000rpm at room temperature (RT) for 10 minutes followed by subsequent washing of pellet 2-3 times by 1X TE buffer. Pellet was thoroughly mixed with 40µl of 10% SDS and 400µl of 1X TE buffer and incubated at 37°C for 30 minutes. Centrifugation was done at 7000rpm for 10 minutes at RT and supernatant was collected into fresh micro-centrifuge tubes and then phenol chloroform-based extraction was done. DNA was concentrated using absolute ethanol by leaving it at -20°C overnight and was thoroughly centrifuged at 10,000 rpm for 15 minutes at RT followed by subsequent washing with 70% ethanol and air dried. Finally the DNA was re-suspended in 50µl of 1x TE buffer.
(b) Amplification of merA and merB genes
Amplification of merA and merB genes were done using merA F; R and merB F; R primers sets and PCR Master Mix (Fermentas Life Sciences, USA) in 50µl reaction using an automated thermocycler (Eppendorf AG 22331). The amplified PCR product along with 10kb and 1kb DNA ladder (Merck millipore, India) was visualized on agarose gel. The PCR products ~1.6kb for merA and ~500bp for merB were eluted and purified using HiYield Gel/PCR DNA Mini Kit (Real Biotech Corporation, Taiwan) and sequencing was done.
Construction of phylogenetic tree
For phylogenetic analysis, merA and merB gene sequences were compared with merA gene and merB sequences respectively, available in Gene Bank using BLAST Search tool. While studying the genes, ten best sequences showing maximum similarity (»99%) with individual gene were selected for the construction of phylogenetic tree. Multiple sequence alignment was done using CLUSTAL OMEGA9 .The statistical testing was done by resampling the dataset for 1000 times in program SEQBOOT. To estimate evolutionary distance between the sequences DNADIST program was used to calculate pairwise distance. The resultant distance matrix was used to draw tree with neighbor joining method using program NEIGHBOR10. Finally, the phylogeny tree was viewed in Tree View, version 1.6.6.
Metagenomic DNA isolation and restriction digestion
The concentration of the DNA isolated from Ulhas estuary site was 700ng/ml (260nm). DNA fragment in the range of 5 to 10Kb was gel eluted and ligated onto dephosphorylated plasmid digested with BamH1 restriction enzyme and metagenome was digested by using Sau3A1 (Fig. 1).
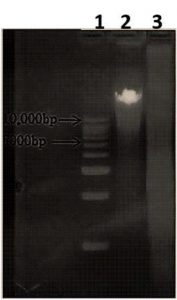 Fig. 1. Metagenome of Ulhas soil sample. Lane 1: 10 kb DNA ladder, 2: Metagenomic DNA (Ulhas); 3: Restricted digested with Sau3A1
Fig. 1. Metagenome of Ulhas soil sample. Lane 1: 10 kb DNA ladder, 2: Metagenomic DNA (Ulhas); 3: Restricted digested with Sau3A1Metagenomic Library construction, screening and selection of clones
More than 150 clones were selected and screened for Hg tolerance (Fig. 2). Ninety-eight of them were able to grow on the LB agar plates supplemented with 5ppm of HgCl2.
Table 1
Ulhas clones showing the growth at different concentrations of Hg.
Sl.No |
Concentration of HgCl2 |
Number of tolerant clones |
|---|---|---|
1 |
05 |
98 |
2 |
10 |
84 |
3 |
20 |
32 |
4 |
30 |
16 |
5 |
40 |
4 |
6 |
50 |
– |
7 |
60 |
– |
8 |
70 |
– |
9 |
80 |
– |
10 |
90 |
– |
Table 2
CFU count of Ulhas clones at different Hg concentration
| Sl.No | Mercury Chloride Concentration (ppm) | CFU | |||
|---|---|---|---|---|---|
| UC07 | UC26 | UC79 | UC126 | ||
| 1 | Control (0) | 802 | 604 | 508 | 617 |
| 2 | 5 | 1024 | 965 | 722 | 712 |
| 3 | 10 | 921 | 567 | 684 | 525 |
| 4 | 20 | 662 | 410 | 312 | 338 |
| 5 | 30 | 427 | 368 | 293 | 216 |
| 6 | 40 | 415 | 307 | 301 | 267 |
| 7 | 50 | 397 | – | 63 | 52 |
| 8 | 60 | 274 | – | – | – |
| 9 | 70 | 131 | – | – | – |
| 10 | 80 | – | – | – | – |
Table 3 : Acclimatization of Ulhas clones at different Hg concentration
| Sl.No | Mercury Chloride Concentration (ppm) | Clones | |||
|---|---|---|---|---|---|
| UC07 | UC26 | UC79 | UC126 | ||
| 1 | 10 | √ | √ | √ | √ |
| 2 | 20 | √ | √ | √ | √ |
| 3 | 30 | √ | √ | √ | √ |
| 4 | 40 | √ | √ | √ | √ |
| 5 | 50 | √ | √ | √ | |
| 6 | 60 | √ | |||
| 7 | 70 | √ | |||
| 8 | 80 | ||||
(√) showing growth of clones
The clones (UC07, UC26, UC79, UC126) were grown overnight at different concentrations of HgCl2 (5 to 80 ppm) and number of viable cells were calculated (Table 2). The clones were acclimatized gradually by growing them at higher Hg concentration, only UC07 was able to grow till 70ppm Hg conc. hence selected for further studies (Table 3). UC07 was able to grow in existence of HgCl2, indicated the clone carrying merA gene. It was also checked for presence of merB gene by using CH3HgCl2 as a substrate. UC07 was able to grow at 5ppm and 10ppm CH3HgCl2 showing the OD, 1.67 and 1.45 at 600nm respectively, confirming the presence of merB gene as well.
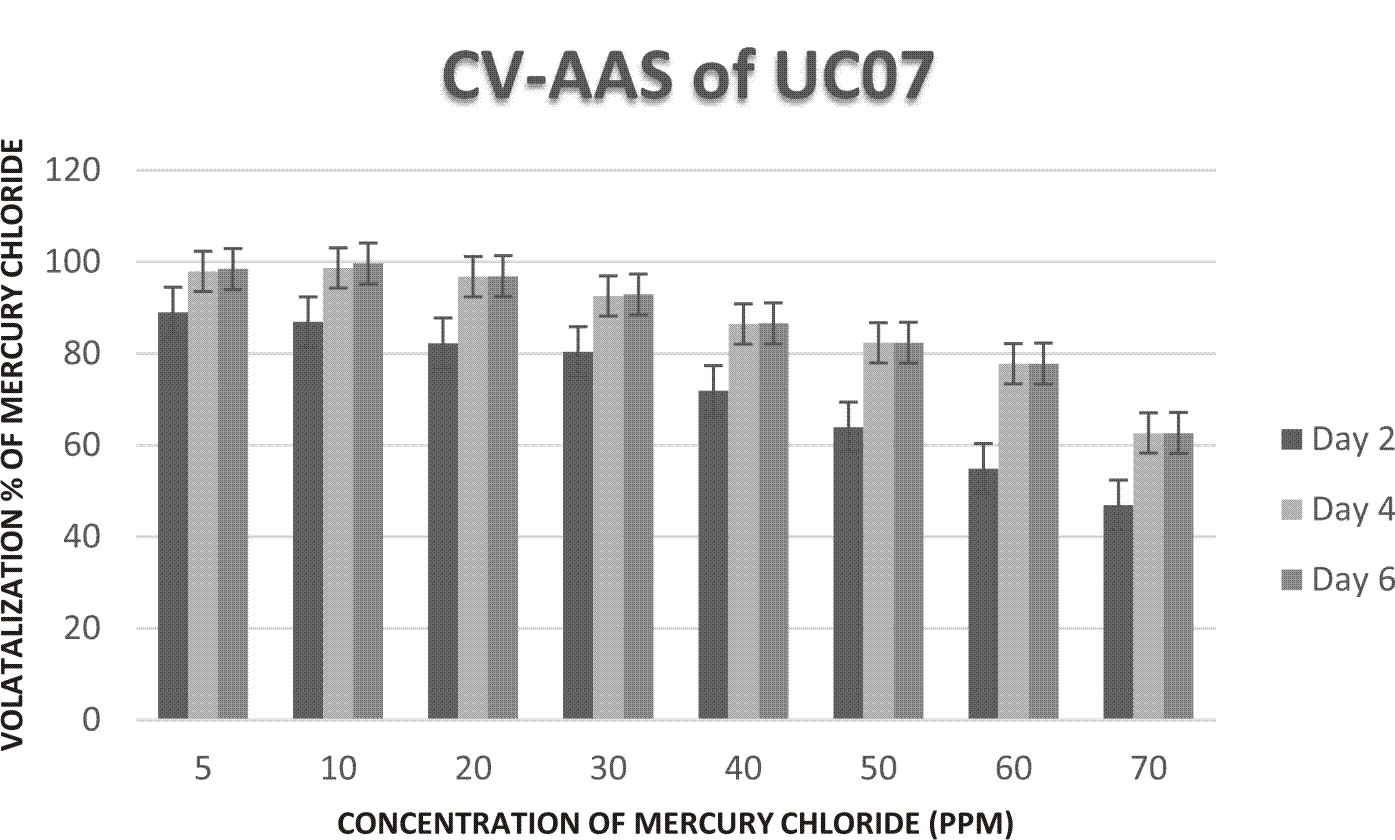
Quantification and validation of Hg volatilization
The quantification of Hg volatilization was done by CV-AAS for the samples withdrawn on day 2nd, 4th and 6th. It was observed that UC07 was able to effectively volatilize Hg supplemented in media. The conversion was found to be maximum at 5ppm (97.94%) on day 6th whereas on day 4th (93.23%) and day 2nd (86.34%) (Fig. 3). In all the Hg conc. (ppm) taken, the maximum bioconversion can be considered on 6 th day.
Antibiotic resistance
Antibiotic activity of UC07 was checked by disk diffusion method. On the basis of the measurement of the zone of inhibition (in cm) it was observed that UC07 was resistance to Vancomycin (30 mcg), Kanamycin (30 mcg) and Norflaxin (10 mcg), whereas it has shown susceptibility towards Streptomycin (25 mcg) (Table 4). 3.5 Identification and deposition of merA and merB gene sequences: The merA gene that codes for mercury reductase enzyme and merB gene coding for methylmercury lyase of UC07 were sequenced. The partial sequences were checked for chimera using DECIPHER’s Find Chimeras web tool (http://decipher.cee.wisc.edu/FindChimeras.html). The presence of both the genes merA (Accession No: MG772720) and merB (Accession No: MG755404) gave the indication of mercury volatilization which was supported by further experiments.
Phylogenetic analysis of merA and merB genes
The sequence obtained for merA and merB genes were globally aligned using BLAST and top ten matches showing similarity »99% were used for phylogenetic tree construction (Fig. 5 and Fig. 6). The presence of merA and merB genes within the phylogenetic tree with high boot strap value (993-1000) confirms that UC07 contain both the genes.
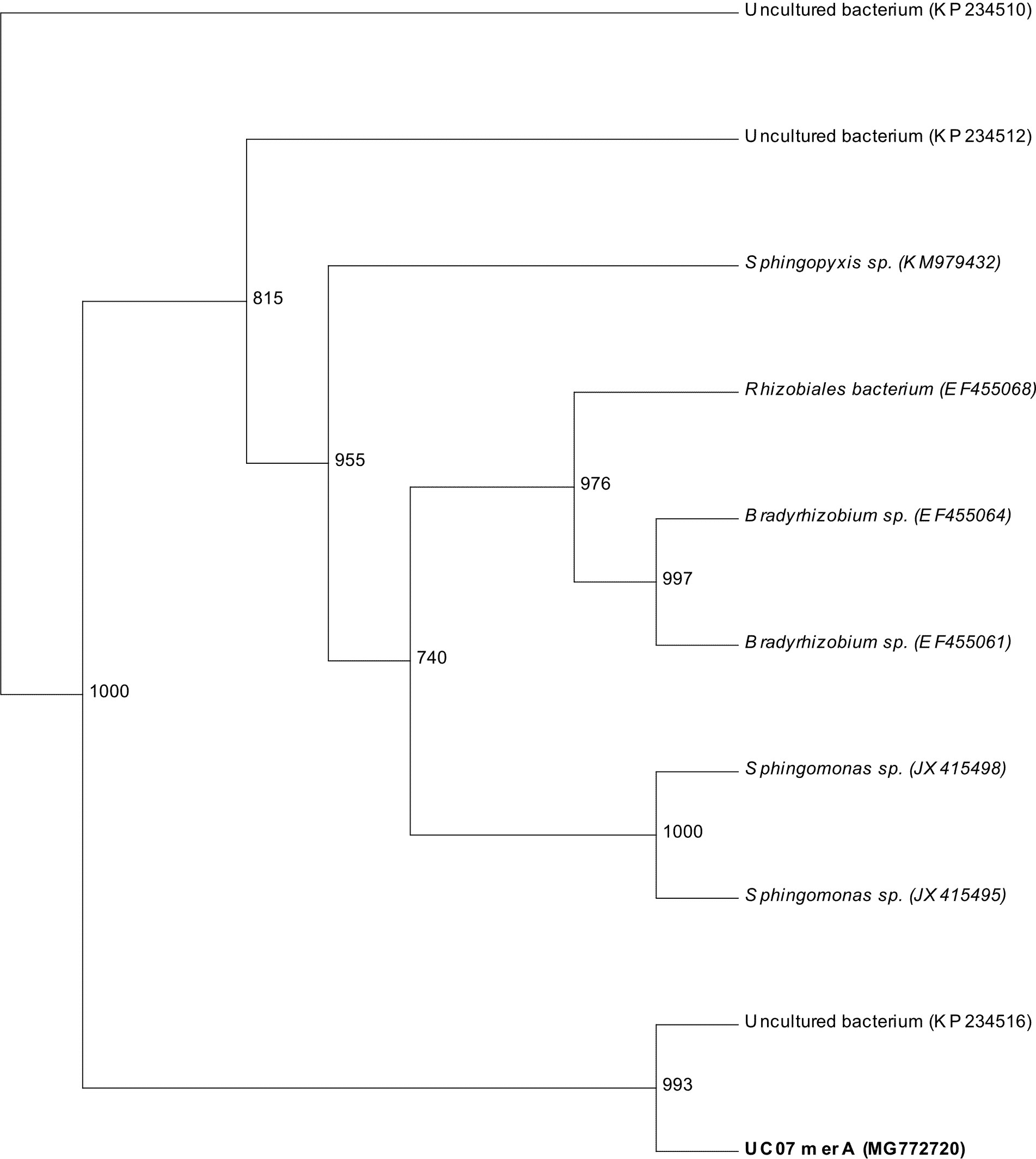 Fig. 5. phylogenetic tree based on partial sequences of merA genes was produced applying the Neighbor-Joining method. The bootstrap values, calculated from 1000 replicates, show the percentage of replicate trees in which the associated ratio clustered together.
Fig. 5. phylogenetic tree based on partial sequences of merA genes was produced applying the Neighbor-Joining method. The bootstrap values, calculated from 1000 replicates, show the percentage of replicate trees in which the associated ratio clustered together.
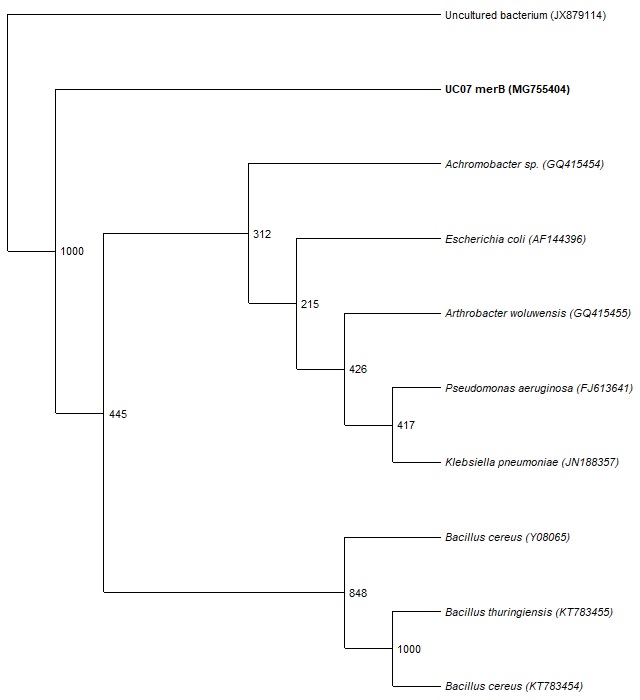 Fig. 6. phylogenetic tree based on partial sequences of merA genes was produced applying the Neighbor-Joining method. The bootstrap values, calculated from 1000 replicates, show the percentage of replicate trees in which the associated ratio clustered together.
Fig. 6. phylogenetic tree based on partial sequences of merA genes was produced applying the Neighbor-Joining method. The bootstrap values, calculated from 1000 replicates, show the percentage of replicate trees in which the associated ratio clustered together.Table 4
Antibiotic sensitivity assay of UC07
| Antibiotics | Disc Conc. | Resistant | Intermediate | Susceptible |
|---|---|---|---|---|
| Vancomycin | 30 mcg | + | ||
| Kanamycin | 30 mcg | + | ||
| Norflaxin | 10 mcg | + | ||
| Getamycin | 30 mcg | + | ||
| Tetracyclin | 30mcg | + | ||
| Streptomycin | 10 mcg | + | ||
| 25 mcg | + |
*Resistant (0-4mm), Intermediate (5-8mm) and Suseptible (9-12mm)
Extensive use of mercury by various industries has become major cause of concern as it is dumped without proper discard procedure has found to enter the water bodies and as well as human food chain. Various researchers have found that there are many bacteria which can convert toxic form of (Hg2+ ) to non-toxic form of mercury (Hgo). These microorganisms are of great use as they play important role in biological degradation of mercury in its habitat3. Metagenomic approach gives the new insight of isolating the genomes of the bacteria which are un-cultivable i.e. cannot be cultured in lab11,12. Among various clones, UC07 was found to be the best as it can grow in 70ppm of HgCl2. UC07 can be used for broad spectrum volatilization as it was able to utilize both HgCl2 and CH3HgCl2. The gene product converts the dangerous methylmercury and other organic mercury derivatives to ionic mercury; which can then be reduced to Hgo13. It has shown antibiotic resistance towards vancomycin (30 mcg), Kanamycin (30 mcg) and Norfllaxin (10 mcg). Hgr loci and antibiotic resistance are commonly found on the same plasmids and transposons14, also microorganisms found in mercury pollutant sites have been discovered with antibiotic resistance5. Results indicated that maximum volatilization in case of media study was seen on day 4 (97.94%) whereas it was ~100% on day 6 in validation study using Ulhas soil. The Hg concentration in Ulhas soil was reduced down to 60ppm which may have favored the conditions for the autochthonic microbes enhancing the volatilization. The PCR and sequencing confirmed the presence of merA and merB genes within UC07 and the tree constructed using these sequences revealed the close relationship of both merA and merB already present in database. Since the clone can be used for broad spectrum bioconversion of Hg, it has also shown efficient volatilization on actual site (Ulhas soil). This clone can thus be maintained and explored for efficient volatilization of Hg.
Hg is one of the sixth most toxic elements found among the plethora of different element found on the earth. Its exposure can give rise to serious health problems and sometimes even death. Due to alarming situation it is important to find new strategies for its control. Bio conversion is one of the most widely chosen approaches in order to reduce the toxic waste. Ulhas soil, Mumbai, India was used to construct metagenomic library of mer operon. The clone obtained, was able to grow at 70ppm Hg. It has shown promising results in bioconversion of Hg in actual site and can be used effectively for volatilization of both organic and inorganic form of Hg.
We are thankful to Department of Science and Technology, Government of India, for providing the funds to carry out the study, Dr. Rakesh Sharma, IGIB for providing his help and also thankful to Amity University, Sector-125, Noida, UP, for providing infrastructure.
On behalf of all authors, the corresponding author states that there is no conflict of interest.
- Wedepohl KH. The composition of the continental crust. Geochimica et cosmochimica Acta,1995; 59(7):1217-32.
- Barkay T, Miller SM, Summers AO. Bacterial mercury resistance from atoms to ecosystems. FEMS Microbiol Rev., 2003; 27(2-3):355-84.
- Jaiswal G, Porwal S. Role of mercury resistance (mer) operon in bioreme-diation of mercury contamination. J. of Biol and Chem Res., 2014; 2: 986-97.
- Nascimento AM, Chartone-Souza E. Operon mer: bacterial resistance to mercury and potential for bioremediation of contaminated environments. Genet and Mol Res., 2003; 2(1):92-101.
- McArthur JV, Tuckfield RC. Spatial patterns in antibiotic resistance among stream bacteria: effects of industrial pollution. Appl. and Environ. Microbiol., 2000; 66(9):3722-6.
- Sant’ana YX, Chartone-Souza E, and Ferreira MD. Drug resistance and colicinogeny of Salmonella typhimurium strains isolated from sevrage-contamined surface water and humans in Belo Horizonte, Brazil Revista de microbiologia, 1989; 20(1):41-9.
- Porwal S, Singh R. Cloning of merA Gene from Methylotenera mobilis for Mercury Biotransformation. Ind. J. of Microbiol., 2016; 56(4):504-7.
- Geissmann Q. OpenCFU, a new free and open-source software to count cell colonies and other circular objects. PloS one, 2013; 8(2):e54072.
- McWilliam H, Li W, Uludag M, Squizzato S, Park YM, Buso N, Cowley AP, Lopez R. Analysis tool web services from the EMBL-EBI. Nucleic acids research, 2013; 41(W1), W597-W600.
- Felsenstein J, (1993) Phylip (Phylogeny Inference Package) version 3.57c. Department of Genetics, University of Washington, Seattle. Distribution: http://evolution.genetics.washington.edu /phylip.html, 2018.
- Sharma R, Ranjan R, Kapardar RK, Grover A. 'Unculturable'bacterial diversity: An untapped resource. Curr. Sci., 2005; 10:72-7.
- Kalia VC. Extending genomic limits through metagenomic exploration. J. Cosmol., 2010; 13:3625-7.
- Lal D, Lal R. Evolution of mercuric reductase (merA) gene: a case of horizontal gene transfer. Microbiology, 2010; 79(4):500-8.
- Wireman J, Liebert CA, Smith T, Summers AO. Association of mercury resistance with antibiotic resistance in the gram-negative fecal bacteria of primates. Appl. and Environ. Microbiol., 1997; 63(11):4494-503.
© The Author(s) 2019. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.