Bacteriophages, the most abundant biological entities on Earth, play a central role in regulating microbial populations and driving bacterial evolution. Dominated by tailed double-stranded DNA viruses (Caudovirales), phages orchestrate complex interactions with their hosts that begin with the highly specific process of adsorption, mediated by receptor-binding proteins and host surface-localized receptors. This review delves into the molecular architecture and specificity of phage-host recognition, emphasizing the diverse range of bacterial receptors- including lipopolysaccharides, outer membrane proteins, teichoic acids, pili, flagella, and capsular polysaccharides- found in both Gram-negative and Gram-positive bacteria. In parallel, we explore the multi-layered defense mechanisms employed by bacteria to evade infection, including receptor masking, biofilm-mediated resistance, superinfection exclusion, toxin-antitoxin systems, CRISPR-Cas immunity, and restriction-modification systems. In response, phages have evolved counterstrategies including structural alterations in receptor-binding proteins, anti-CRISPR factors, DNA mimicry, and dual adsorption modules to overcome host immunity. The review integrates recent advances in our understanding of phage-host molecular conflicts, drawing attention to the evolutionary dynamics shaping phage-host coadaptation and highlighting the translational potential of phage biology in therapeutic and biotechnological applications.
Phage Host Interaction, Cell Surface Receptors, Antiphage Defense, Phage Counterdefense
Bacteriophages are viruses that specifically infect bacteria and are found in nearly all microbial ecosystems. Estimates suggest their global numbers reach 1029 -1030 particles, making them one of the most abundant biological entities on Earth. These viruses are environmentally stable and, once inside a susceptible host, replicate efficiently.1 Since their discovery over a century ago, phages have attracted considerable attention for their practical applications. They have been explored as alternatives to antibiotics, as tools for bacterial biocontrol, and as vectors in gene delivery studies. Structurally simple and genetically compact, phages are obligate parasites that rely entirely on the replication and translation machinery of their bacterial hosts to produce progeny. Most phages are highly specific, infecting only one bacterial species or a narrow range of strains.2 Beyond their therapeutic potential, phages play an important ecological role. By infecting and lysing bacteria, they influence host population size, species composition, and metabolic activity, thereby shaping the dynamics of microbial communities. This capacity to modulate bacterial diversity and function makes them key contributors to ecosystem balance and a useful model for studying the molecular basis of host-microbe interactions.3
Most bacteriophages belong to the order Caudovirales and are characterized as tailed viruses with a double-stranded DNA genome. This predominance has been documented in large-scale surveys of the virion.4 Phage replication can proceed through several infection strategies. In the lysogenic pathway, for example, the phage genome, known as a prophage, becomes stably maintained within the bacterial cell. It may integrate into the host chromosome or persist as a plasmid-like element. In either case, the prophage is replicated along with the host genome and can remain dormant for extended periods, while expressing specific regulatory proteins that control both host and phage gene activity. Under stressful environmental conditions, prophages may exit their dormant state, transitioning into the lytic cycle, which culminates in the lysis of the host cell and the release of new virions. Alternatively, pseudo-lysogeny represents a non-canonical phage life strategy wherein the phage neither lyses the host nor integrates into its genome, avoiding stable long-term residence.5,6 A critical and universal step in all phage life cycles is adsorption, which hinges upon the precise recognition of specific host surface proteins. Independent of the subsequent replication strategy, the initial phase of infection invariably involves the identification and attachment of the phage particle to the bacterial cell surface. This adsorption process is a fundamental mechanism shared among viruses infecting organisms across the three domains of life.7,8
Although phages are small and present at low density on the bacterial surface, they usually succeed in attaching to their receptors after very few encounters. The speed at which a bacterial population is cleared depends largely on how quickly virions adsorb to their hosts. In simplified experimental systems, this adsorption rate can be estimated from basic factors such as phage diffusion, bacterial cell size, and the likelihood that a collision results in a stable attachment. When bacterial numbers are high, free phages disappear from the environment more quickly because adsorption events are more frequent. Conversely, when phage concentrations rise, bacteria are infected at a faster rate.9
In the past decade, interest in bacteriophages has grown well beyond microbiology and now includes applications in biotechnology, medicine, agriculture, and even food safety.10-12 For these applications to be both safe and effective, it is essential to understand in detail how phages interact with their bacterial hosts. This involves not only identifying the receptors that allow phage attachment but also clarifying the mechanisms of infection and ways bacteria evolve resistance. In this review, we highlight current knowledge on these processes, with particular emphasis on host cell recognition, receptor specificity, and the molecular strategies bacteria use to defend themselves.
Bacteriophage structure and classification
Bacteriophage morphology plays a central role in shaping their interactions with bacterial hosts. Variations in capsid architecture and tail structures influence how effectively a phage can recognize surface receptors, attach to the cell, and initiate infection. Phages that have been described to date display considerable diversity in their host specificity and infection strategies, ranging from narrow to broad host ranges and from strictly lytic to more complex lifestyles. As a result, members of different taxonomic groups often affect microbial communities in distinct ways, reflecting their structural and functional adaptations to particular ecological environments. The morphology of bacteriophages fundamentally governs their interactions with bacterial hosts. Structural complexities, including the organization of the capsid and tail apparatus, critically determine their capacity to recognize, adhere to, and penetrate bacterial cells. Currently characterized bacteriophages exhibit remarkably diverse life strategies, particularly in terms of host range specificity and lytic capabilities. Consequently, phages belonging to distinct taxonomic groups may exert unique influences on the microbial dynamics within specific ecological niches.13
Phages possess a relatively simple architecture, consisting of a proteinaceous capsid that encases their nucleic acid, which may be either DNA or RNA. Based on their structural morphology, they are broadly categorized into tailed, non-tailed, and filamentous types.14 Among these, tailed phages are the most extensively studied. They typically contain three major parts: a capsid that contains the genome, a tail that acts as a conduit for DNA delivery into the host cell, and a specialized tail tip structure that is responsible for recognizing and binding to bacterial receptors. The assembly of double-stranded DNA (dsDNA) phages occurs in a stepwise manner. First, capsid proteins form a precursor “prohead” around a portal complex. The viral genome is then actively pumped into this prohead by an ATP-driven motor. Once the DNA is fully packaged, the head undergoes maturation to stabilize the enclosed genome. Finally, tail and neck components are attached, giving rise to a complete and infectious viral particle.15
Bacteriophages were initially categorized into four primary morphological groups: Tailed phages (Caudoviridae), Icosahedral or polyhedral form (Microviridae), filamentous phages (Inoviridae), and pleomorphic types (Plasmaviridae)
Among bacteriophages, the tailed types grouped under the order Caudovirales are by far the most widespread. It is estimated that they account for close to 96% of all known phages. These viruses typically carry double-stranded DNA, which is packaged inside an icosahedral head and linked to a tail structure. The tail is more than a passive appendage; it acts as a nanomachine that mediates host recognition, binding, and the transfer of genetic material. Tail morphology has been used as a taxonomic marker, leading to the division of Caudovirales into three families: Myoviridae, with their contractile tails; Siphoviridae, which have long but non-contractile and flexible tails; and Podoviridae, characterized by their short, stubby tails.16 More recently, however, the International Committee on Taxonomy of Viruses (ICTV) revised this system, bringing these families under a broader class, Caudoviricetes, to better reflect genome-based and evolutionary relationships.17 Members of this class are not just abundant but also ecologically significant. Through both lytic and lysogenic interactions, they help regulate microbial populations. Beyond ecology, they are attracting interest in applied fields ranging from therapeutic development to biotechnology and bacterial genetics.
The family Microviridae consists of small, non-enveloped bacteriophages that carry single-stranded DNA within an icosahedral capsid. Within this group, three main lineages have been described. The genus Microvirus infects free-living bacteria such as enterobacteria. In contrast, members of the subfamily Gokushovirinae specialize in intracellular parasitic bacteria, including Chlamydia and Spiroplasma. A third lineage, the Alpavirinae, has so far been identified only in the form of prophages. Once these viruses enter a bacterial cell, they produce a complementary minus-strand of DNA. This strand provides the template needed for transcription of viral mRNAs and subsequent gene expression.18,19
The Inoviridae family comprises non-enveloped, flexible, filamentous bacteriophages that primarily infect Gram-negative bacteria by adsorbing to bacterial pili. Their circular, positive-sense single-stranded genomes range from 5.5 to 10.6 kb, encoding 4-15 proteins. These phages replicate via a rolling circle mechanism and release progeny through extrusion without lysing the host. The family includes 43 species across two genera, with genomes that may remain episomal or integrate with the bacterial chromosome.20
The Plasmaviridae family comprises quasi-spherical, membrane-enveloped bacteriophages that infect cell-wall-less bacteria, specifically Acholeplasma species. These phages possess a circular, supercoiled double-stranded DNA genome approximately 12 kb in length, containing 14 open reading frames encoding at least 15 proteins. With a diameter ranging from 50-125 nm, they replicate within the host cytoplasm. The family has no assigned order and includes a single genus and species (Acholeplasma virus L2).21
Strategies underpinning the phage-host binding and invasion
Unlike typical predator-prey relationships, phage-host interactions exhibit remarkable specificity, with phage replication intricately dependent on the metabolic and replicative systems of the bacterial hosts. Bacteriophage infection begins with adsorption, a critical step involving specific interactions between phage binding proteins and bacterial surface receptors. Attachment of the bacteriophage to the host cell may result from Brownian motion, passive diffusion, or directional fluid-mediated transport.22 There are two types of phage adsorption to the host cell. Reversible attachment enables phage particles to detach and subsequently infect other cells of the same bacterial strain within the culture. Upon successful engagement with primary receptors, this transient interaction transitions into irreversible anchoring, triggering the activation of the phage’s injection apparatus. Adsorption not only initiates infection but also defines the host range by determining which bacterial strains can be targeted. Consequently, it serves as the primary determinant of phage-host specificity.7
The process is initiated by a reversible interaction between the phage and the bacterial cell surface, facilitated by the binding of phage tail fibres or specialised appendages to primary receptors located on the bacterial outer envelope- typically lipopolysaccharides (LPS), outer membrane proteins, teichoic acids, or surface structures such as flagella and pili. This transient association serves as a preliminary host recognition step, enabling the phage to assess the compatibility of the potential host before committing to infection.23,24 The vast majority of bacteriophages are classified under the order Caudovirales and are distinguished by their tail structures, which play a pivotal role in host recognition and the delivery of phage genetic material into the bacterial cell.25
A critical initial phase in the life cycle of most bacteriophages entails the attachment of the virion to a specific recognition site on the host cell membrane, subsequently facilitating the injection of phage genomic DNA into the bacterial cell. A given bacteriophage typically exhibits a narrow host range, often targeting a specific microbial strain. This high degree of host specificity is primarily governed by the adsorption process, which is dictated by the structural and molecular characteristics of receptor molecules present on the surface of the bacterial cell against phage receptor-binding proteins (RBPs).22
Receptor-binding proteins are essential components of bacteriophages as they determine the recognition and attachment to the host. In general, RBPs contain three regions: a conserved N-terminal domain, a hinge-like flexible segment, and a C-terminal domain, which often participates in receptor recognition and, in some cases, enzymatic degradation of host surface proteins.26 The exact architecture of RBPs differs across phages, reflecting variations in phage morphology and strategies of host interaction. For example, in the Podovirus Sf6, which infects Shigella flexneri, short tail fibres (tail spike proteins) first engage with the lipopolysaccharide (LPS) layer, allowing a reversible binding step.27 In contrast, in the well-studied Escherichia coli phage T4, the gp12 protein located at the short tail fibres mediates irreversible binding to the LPS core region.28 A different arrangement is seen in Siphoviridae, such as Lactococcus lactis phage TP901-1, where RBPs form part of a larger structural baseplate complex that coordinates attachment.29
The bacterial cell envelope
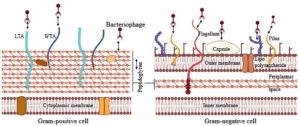
As illustrated in Figure 1, Bacterial surfaces provide a variety of molecular targets that bacteriophages can exploit as receptors. The type of receptor available depends on the bacterium-phage pairing and can range from short peptide sequences to more complex polysaccharide structures.
Figure 1. Schematic representation of bacterial cell surface targets for phage adsorption. Gram-positive (right) and Gram-negative (left) bacterial envelopes are illustrated, highlighting key surface components that function as phage receptors. In Gram-positive bacteria, thick peptidoglycan, teichoic acids, including both lipo and wall teichoic acids, serve as recognition sites. In Gram-negative bacteria, phages exploit outer membrane proteins (OMPs), lipopolysaccharides, pili, and flagella as receptor molecules38 (Source: Biorender.com)
The architecture of the cell envelope differs markedly between Gram-positive and Gram-negative organisms, and these differences shape both bacterial physiology and their interactions with phages. Gram-positive bacteria contain a thick peptidoglycan layer with embedded teichoic acids but lack an outer membrane. By contrast, Gram-negative bacteria have a much thinner peptidoglycan layer situated between the cytoplasmic membrane and an outer membrane rich in lipopolysaccharides (LPS). Such variations not only contribute to differences in antibiotic sensitivity but also strongly influence the way bacteriophages recognize and adsorb to their hosts.30
Surface-localized receptors in Gram-positive bacterial cell wall
Gram-positive bacteria are characterized by a relatively thick cell wall that contains a large amount of peptidoglycan proteins together with teichoic acid (TA). In some species, Teichoic acid may represent more than half of the total cell wall mass. These polymers occur in two major forms: lipoteichoic acids (LTA), which are linked to the cytoplasmic membrane, and wall teichoic acids (WTA), which are covalently associated with the peptidoglycan layer.31 The peptidoglycan itself forms the main structural framework of the wall and lies just outside the cytoplasmic membrane, a feature that is conserved across most bacterial groups. While its primary role is to preserve cell shape and structural integrity, it also provides anchoring sites for various proteins and lipopolysaccharides. Structurally, peptidoglycan comprises linear glycan chains composed of alternating N-acetylglucosamine and N-acetylmuramic acid residues, linked via 1-4 glycosidic bonds, which are further crosslinked by short peptide bridges to form a robust, mesh-like matrix.32 Bam35, a temperate double-stranded DNA bacteriophage that infects Gram-positive Bacillus thuringiensis, uses a sequential entry mechanism involving three distinct stages: rapid receptor binding, peptidoglycan penetration, and interaction with the plasma membrane. Binding to the bacterial surface is dependent on N-acetyl-muramic acid, while peptidoglycan degradation is mediated by a virion-associated hydrolytic activity, and both of these steps occur independently of divalent cations. In contrast, successful penetration through the plasma membrane requires the presence of divalent cations, highlighting a unique ion-dependence at the final entry step.33
In Listeria Phage A500, host recognition occurs via protein-glycopolymer interaction involving the endolysin Ply500, which specifically targets Listeria serovars 4, 5, and 6 through its l-alanoyl-d-glutamate endopeptidase activity. The cell wall binding domain (CBD500) of Ply500 specifically binds 3’-O-acetylated N-acetylglucosamine (GlcNAc) residues in the wall teichoic acid (WTA) polymer. These GlcNAc moieties serve as critical epitopes recognised by a binding site within SH3b-like repeats of the viral domain, ensuring selective cell wall interaction.34 In bacteriophage SPP1, irreversible adsorption to its host, Bacillus subtilis, is facilitated by the host cell membrane protein YueB. However, reversible attachment primarily targets the glucosylated poly (glycerophosphate) chains of the cell wall teichoic acid.35 Similarly, in the broad host range, virulent Listeria phage A511, receptor binding protein Gp108 specifically recognizes N-acetylglucosamine and rhamnose substituents present on the wall teichoic acids of the bacterial cell wall. These examples highlight the pivotal role of teichoic acid modifications in phage host recognition and specificity.36
Upon attachment of the phage to the bacterial surface, a mechanical or chemical signal is transmitted along the tail fiber to the capsid, where the viral genome is encapsulated. The contraction of the phage’s tail sheath triggers the precise penetration of the bacterial cell wall by the internal tail tube, facilitating the release of the viral genome through the portal pore located at a distant capsid vertex. The DNA is subsequently translocated into the host cytoplasm in a sequential manner, leaving the empty phage particle firmly anchored to the bacterial surface.37
Surface-localized receptors in the Gram-negative bacterial cell wall
Bacteriophages targeting Gram-negative bacteria must first overcome the lipopolysaccharide (LPS) barrier, a structurally robust component of the outer membrane stabilized by strong noncovalent interactions. Unlike peptidoglycan, which is susceptible to enzymatic degradation, LPS poses a formidable challenge, prompting many phages to utilize specialized proteins to navigate through it and engage outer membrane protein (Omp) receptors a strategy observed across all tail morphotypes.39
Bacteriophages that target the smooth (S) type of lipopolysaccharide (LPS) exhibit an exceptionally narrow host range due to the high structural variability of the O-antigen across different bacterial taxa. In contrast, phages that recognize rough (R) type LPS or vice versa tend to possess a broad host spectrum, as the conserved nature of the LPS core region is shared among various species and genera of Gram-negative bacteria.40
Recent advancements in the characterization of novel phage isolates infecting Gram-negative bacteria have leveraged lipopolysaccharide profiling to elucidate phage infection strategies, particularly those targeting hosts with impaired or modified LPS biosynthesis.41
The host cell receptors interact with phage proteins either reversibly or irreversibly, and these distinct modes of adsorption are mediated by different sets of viral proteins and host receptors, highlighting a functional specialization in the phage infection process.42 The attachment to the host begins with a selective but reversible interaction with LPS on the bacterial surface. Tail-associated enzymes, particularly those embedded within spike structures, exhibit LPS-degrading activity by targeting and cleaving O-antigen repeats. By breaking down the LPS barrier, the phage gains access to deeper cell surface components, including secondary receptors or the outer membrane itself, where stable and irreversible binding is ultimately established. One of the previously explained examples, where T4 bacteriophage mediates the reversible attachment to the host E. coli surface, is by gp53 proteins located in the long tail fibres, whereas irreversible binding involves gp12 proteins in the short tail fibres, which specifically recognize the heptose region within the host’s lipopolysaccharide.28
Peptidoglycan, although shielded by the outer membrane in Gram-negative bacteria, plays a crucial role during the later stages of phage entry by acting as a secondary barrier and target for enzymatic processing. Escherichia phage T7, a well-studied Podovirus infecting Escherichia coli, follows a multi-step infection pathway involving initial attachment to the outer membrane protein FhuA, followed by localized degradation of the peptidoglycan to enable DNA translocation.43 This process is mediated by a virion-associated lysozyme domain within the tail machinery, which specifically cleaves the β-1,4 glycosidic bonds between N-acetylglucosamine and N-acetylmuramic acid residues. While peptidoglycan recognition in T7 is secondary to outer membrane binding, its efficient hydrolysis is essential for successful DNA delivery into the cytoplasm, highlighting a coordinated interaction between outer membrane receptor engagement and peptidoglycan penetration in Gram-negative hosts.44
The flagella are the initial host contact targets on the bacterial cell, by which the phages enhance their chances of establishing attachment and achieving successful infection. The interaction between the phage head-associated filament and the bacterial flagellum represents a distinct infection strategy, wherein the filament transiently coils around the rotating flagellum, facilitating phage transport towards the cell pole and proximal host receptors.45
In some instances, the phage adsorption process may involve multiple receptors on the bacterial surface, indicating a mechanism that requires sequential or simultaneous interaction with two distinct host receptors. For example, Flagellotropic phage 7-7-1, a member of Myoviridae, infects the soil bacterium Agrobacterium spp. strain H13-3 by targeting actively rotating bacterial flagella. The number, length, and rotational speed of the flagella influence the efficiency of infection. The phage initially binds to the flagellar filament and is subsequently translocated towards the bacterial surface via clockwise rotation. Upon reaching the cell surface, the phage engages with the lipopolysaccharide (LPS), enzymatically degrades it to access the outer membrane, and then injects its genomic DNA into the host cell through a contractile, syringe-like tail structure.46
A distinct class of bacteriophages exhibits plasmid dependency, wherein infection is restricted to bacterial cells that harbor and actively express specific plasmid-encoded genes required for phage adsorption and entry. These typically exploit plasmid-derived surface structures – most notably conjugative pili -as receptors for attachment. A well-characterized example is bacteriophage MS2, which selectively infects Escherichia coli strains carrying the F (fertility) plasmid, which encodes the biosynthesis of F-pili functioning as primary receptors for MS2 binding and subsequent infection.47 The phage maturation (A) protein specifically binds to the pilus shaft, and natural pilus retraction draws the virion towards the cell envelope, enabling direct translocation of the RNA genome into the cytoplasm. In the absence of these plasmid-encoded surface components, the bacterial host remains resistant to phage invasion. Recently, Parra et al. successfully isolated and characterized two novel dsDNA plasmid-dependent bacteriophages designated Lu221 and Hi226, which specifically target Salmonella enterica harbouring the IncN plasmid Pkm101.48 These examples highlight how plasmid-encoded extracellular appendages can be co-opted by bacteriophages to mediate precise host recognition and efficient genome delivery.
Some bacteria develop a thick polysaccharide barrier, such as a capsule, enveloping the cell, which often acts as a key recognition site for bacteriophages. In many phage-host systems, phages employ capsule depolymerases located on tail fibres or the baseplate to degrade this layer, granting access to underlying receptors. For example, Klebsiella phage KP32 encodes two such enzymes, KP32gp37 and KP32gp38, which are highly specific for capsular serotypes K3 and K21, respectively. By targeting and breaking down these serotype-specific polysaccharides, these polymerases limit the narrow host range of phage KP32 to strains that express either the K3 or K21 capsule type.49
Bacterial antiphage defence mechanisms
Bacteria and bacteriophages are locked in a perpetual evolutionary arms race, marked by dynamic cycles of coevolution and coexistence. Within this ongoing interaction, bacteria evolve phage resistance mechanisms to protect their populations, while phages simultaneously adapt through counter-resistance strategies that enable them to bypass these defences and maintain infectivity.50
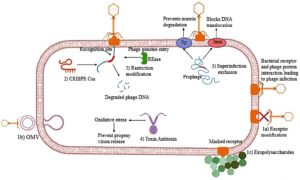
As shown in Figure 2, bacteria orchestrate intricate defense mechanisms that specifically target distinct phases of the bacteriophage life cycle. These defenses encompass the inhibition of phage adsorption, prevention of viral genome injection, and disruption of essential processes, including phage DNA replication, capsid formation, and progeny assembly. Moreover, components derived from these phage resistance systems, most notably the CRISPR-Cas machinery, have laid the foundation for the development of cutting-edge genome editing technologies.
Figure 2. Overview of bacterial anti-phage strategies. Bacterial cells employ multiple protective mechanisms to counteract phage infection at different stages of the viral life cycle. Surface-level defences include 1a) receptor modification, 1b) production of competitive inhibitors, and 1c) Formation of extracellular matrix barriers. 2) CRISPR adaptive immunity, 3) Superinfection exclusion, 4) Toxin-antitoxin systems, and 5) Restriction-modification (RM) systems. Together, these multifaceted defences constitute a layered barrier against phage invasion and proliferation (Source: Biorender.com)
Blocking Phage binding/Interference with phage adsorption
The initiation of bacteriophage infection is universally dependent on the successful adsorption of the virion to specific receptors on the bacterial cell surface. To counter this, certain bacterial strains have evolved sophisticated strategies to impede phage attachment. These adsorption-inhibiting mechanisms generally fall into three categories: direct masking or modification of surface receptors, secretion of extracellular matrix components that hinder access, and production of competitive inhibitors that obstruct phage binding.
Receptor blocking
OmpA is a major outer membrane protein in Escherichia coli that not only facilitates F-mediated conjugation but also serves as a receptor for various bacteriophages. Studies have indicated a potential interaction between the F-plasmid encoded lipoprotein TraT and OmpA of the bacteria, wherein TraT appears to interfere with the binding of OmpA-specific phages, likely through direct interaction. Additionally, notable sequence homology between TraT and phage protein 38-known to recognize bacterial outer membrane proteins-further supports its role in modulating phage-host interactions.51
Competitive inhibitor production
Outer membrane vesicles (OMVs) appear to serve as innate decoys, shielding bacteria from the attack of virulent bacteriophages. In Vibrio cholerae, secreted OMVs have been shown to function as a protective barrier against predation by three lytic phages-ICP1, ICP2, and ICP3. This OMV-mediated defense was demonstrated to be both dose-dependent and reliant on specific receptor interactions, thereby significantly limiting phage infectivity.52 Similarly, hyper-vesiculating mutants of Escherichia coli have been observed to produce OMVs that confer resistance not only against bacteriophage infection but also against antimicrobial peptides such as Polymyxin B and colistin. Notably, when T4 phages were co-incubated with OMVs, they exhibited rapid and irreversible binding, resulting in a marked decrease in phage infectivity due to the sequestration of virions by vesicular structures.53
Extracellular matrix
In Escherichia coli K-12, a quorum-sensing-regulated antiphage defense mechanism has been observed, wherein AHL-mediated signalling in the presence of autoinducers modulates the bacterial response to reduce susceptibility to phage λ and the broad-host-range phage χ.54 Furthermore, quorum-sensing peptides were found to induce abortive infection in E. coli by activating the mazEF toxin-antitoxin system, effectively restricting the propagation of phage P1.55 These findings suggest that quorum-sensing-based regulation of phage sensitivity may represent a widespread strategy within microbial communities.
CRISPR-Cas mediated defense
Among the various strategies bacteria employ to defend against phage infection, the CRISPR-Cas system (Clustered Regularly Interspaced Short Palindromic Repeats and CRISPR-associated proteins) is unique as the only known adaptive and heritable immune mechanism.56 It is prevalent in most archaea and nearly half of bacteria, which defends against lytic phages. This system incorporates short DNA fragments (26-72 bp) from invading phages as spacers, enabling the production of small RNAs that recognize and block future infections by matching viral sequences, thereby generating bacteriophage-insensitive mutants (BIMS).57 In a comprehensive analysis, 44 lysogenic strains of Pseudomonas aeruginosa PA14 were evaluated for their susceptibility to infection by three distinct bacteriophages: JBD18, JBD25, and JBD67. These phages are efficiently neutralized by the native CRISPR-Cas immune system of P. aeruginosa PA14, which prevents successful phage replication. Notably, productive infection by these phages occurred only in mutant PA14 strains in which either the CRISPR loci or the associated cas genes had been deleted, underscoring the essential role of CRISPR-Cas in conferring phage resistance in this bacterial model.58
Blocking phage genome entry
Superinfection exclusion (SIE) is a mechanism by which bacterial strains harbouring a temperate phage genome (prophage) within their chromosome gain resistance to subsequent infection by the same or closely related phages, thereby preventing viral coinfection. The principal strategies by which phages mediate SIE involve either the inactivation of host surface receptors essential for phage binding or the obstruction of phage DNA injection and intracellular translocation.59 In the case of T-even coliphage, rapid superinfection exclusion is established through the action of two phage-encoded proteins: Imm, which blocks DNA translocation and partially restricts its release from superinfecting virions, and Sp protein, which inhibits murein degradation by the phage-associated lysozyme.60
In Pseudomonas aeruginosa, a highly conserved Pf phage protein called PfsE (PA0721) confers resistance to superinfection by phages that utilize the type IV pilus as their receptor by interacting with PilC to block pilus assembly.61 However, the minor capsid protein pVII confers complete protection against pilus-dependent lytic phages by interacting with T4P components PilC and PilJ. In contrast, pIII offers partial superinfection exclusion by interacting with PilJ and TolR/TolA, providing limited defense against pilus-dependent phages.62
Abortive infection
The toxin-antitoxin (TA) systems have emerged as a significant component of bacterial antiviral defense mechanisms by inducing cellular stress in infected bacteria, thereby disrupting the phage replication cycle and preventing the release of progeny virions. It has been increasingly recognized for its widespread presence across sequenced bacterial genomes – some of which encode dozens of distinct TA modules. These systems are generally arranged as two-gene operons comprising a toxin that disrupts essential cellular functions and a specific antitoxin that counteracts its activity under normal physiological conditions.63 In Salmonella typhimurium, the retron Sen-2 element encodes the accessory toxin RcaT, which is normally inactivated by a complex formed between reverse transcriptase (RT) and multicopy single-stranded DNA (msDNA). Upon disruption of msDNA biosynthesis, RcaT is released from inhibition and becomes toxic. Phage infection can activate RcaT through direct modification of msDNA, leading to growth arrest of the host. Conversely, certain prophage-encoded proteins evade this defense by directly neutralizing RcaT activity.64 Collectively, retron-based TA systems serve as abortive infection mechanisms that contribute to phage resistance in bacteria. Another example of DarTG, a recently identified toxin-antitoxin system with two subtypes, DarTG1 and DarTG2, that provide strong protection to Escherichia coli MG1655 against phages RB65 and T5, respectively. Upon infection, the DarT toxin – a DNA ADP-ribosyl transferase -is released and modifies phage DNA, blocking replication and preventing virion production. Some phages, particularly T-even types, have evolved resistance by acquiring mutations in their DNA polymerase or expressing the anti-DarT protein gp61.2, demonstrating phage adaptability against host defenses.65 Another notable recent example is provided by the toxin-antitoxin immunity protein CapRel(SJ46) in E. coli, which, in contrast to the conventional one-to-one recognition paradigm, utilizes a flexible pseudo zinc finger domain to sense two structurally unrelated phage proteins the major phage capsid and small phage protein GP54Bas11. This dual recognition mechanism introduces a robust safeguard for bacteria, as phages must simultaneously mutate both triggers to fully evade detection, thereby limiting their evolutionary escape routes.66
Nucleic acid interference
The restriction-modification (RM) system serves as a critical bacterial defense strategy against foreign genetic elements, such as bacteriophages. It consists of two coordinated enzymatic activities: a restriction endonuclease (REase) and a methyl transferase (MTase). The REase detects and cleaves specific short DNA sequences, typically 4-8 base pairs in length, if these sequences are unmethylated. To safeguard the host genome from inadvertent cleavage, the MTase selectively methylates these same recognition sites on the host DNA, thus protecting it from restriction activity. This system enables bacteria to discriminate between self and non-self-DNA. When the infecting phage DNA enters the cell and lacks the host-specific methylation, it is rapidly targeted and degraded by the REase. The level of immunity conferred by the RM system depends on the compatibility of the phage and the specificity of the RM components present in the host. Collectively, RM systems function as precise molecular sentinels, providing robust protection by promoting the selective destruction of exogenous DNA while maintaining the integrity of the host genome.67,68
In Escherichia coli carrying a Type I RM system (e.g., EcoP1), the restriction endonuclease targets unmethylated DNA at specific recognition motifs. The complementary methyltransferase modifies the host’s DNA at these sites, thereby preventing self-restriction. Phage λ DNA grown in an RM-negative host lacks methylation and is therefore efficiently cleaved upon infection of an RM-positive strain.69 In addition to methylation- based RM system, bacteria can modify their genomes through phosphorothioation, a process in which sulfur replaces a non-bridging oxygen in the DNA backbone and forms the basis of phosphorothioation-dependent RM systems. This chemical signature distinguishes self-DNA from non-self-DNA and directs restriction activity toward invading phage genomes that lack the appropriate modification.70 The Dnd system exemplifies this strategy in E. coli and Salmonella enterica, where DndABCDE introduces phosphorothioate marks at specific motifs in the host chromosome, while the associated DndFGH restriction complex targets and cleaves unmodified phage DNA, conferring resistance to a broad range of bacteriophages.71
Phage counter-defense strategies against bacterial resistance
Phages are also capable of evolving and developing novel strategies to circumvent bacterial defenses, enabling them to sustain their predatory activity. A comprehensive investigation into both microbial defense systems and phage countermeasures is essential for advancing and refining phage therapy.
Phage infectivity is compromised when host receptors are either masked or no longer compatible with the phage’s receptor-binding proteins (RBP). As a result, alterations or loss of these receptors constitute a fundamental strategy by which bacteria develop resistance to phage infection. To overcome bacterial resistance arising from receptor modification or loss, bacteriophages can adapt by altering their receptor-binding proteins (RBPs), which are crucial for host recognition and attachment. Such modifications enable phages to recognize alternative or mutated bacterial surface structures, thereby restoring infectivity. Escherichia coli phage λ, which typically binds to the LamB porin on the bacterial outer membrane. When E. coli develops resistance by mutating or downregulating LamB, V can evolve variants with mutations in its tail fiber protein (encoded by gene J), allowing it to use alternative receptors such as OmpF.72,73 This receptor-switching mechanism demonstrates the phage’s ability to circumvent bacterial resistance through targeted changes in RBPs, thus sustaining the infection cycle and evolutionary battle between phages and their hosts.
Another example is the Escherichia coli podophage K1-5, which encodes two distinct types of tail fiber proteins. This adaptation allows K1-5 to attach to and infect E. coli strains expressing different capsule antigens (K1 and K5), despite antigenic variation in the bacterial population. Thus, by molecular evolution of receptor-binding proteins, bacteriophages can adapt to changes in bacterial surface structures and continue their infection cycle despite bacterial attempts at evading phage adsorption.74
Bacteriophages evade the bacterial CRISPR-Cas adaptive immune system by encoding anti-CRISPR (Acr) proteins within their genomes. These Acr proteins inhibit the assembly and the activity of CRISPR-Cas complexes, thereby blocking target recognition and cleavage, and ultimately allowing successful phage infection and replication within the bacterial host.75 A characterized instance involves Type I-F CRISPR-Cas systems in Pseudomonas aeruginosa and their inhibition by several AcrIF proteins; AcrIF1 binds to the Cas7f backbone of the Csy surveillance complex, thereby blocking the hybridization between the CRISPR RNA (crRNA) and the target DNA. AcrIF2 interacts with the DNA-binding site, specifically blocking the recognition of the protospacer adjacent motif (PAM) on the phage DNA. AcrIF3 acts as a direct inhibitor of the Cas3 helicase-nuclease, an essential enzyme for DNA cleavage in the Type I-F system. Experimentally, engineered bacteriophages carrying these acrIF genes (termed EATPs) have demonstrated the successful suppression of CRISPR immunity in P. aeruginosa, permitting productive infection and replication even in multidrug-resistant (MDR) bacterial strains.76 The inclusion of Acr proteins significantly enhances the ability of therapeutic phages to overcome bacterial immune defenses in both in vitro and in vivo infection models.
Phages can resist bacterial restriction-modification (RM) systems by interfering with the DNA methylation process that bacteria use to distinguish their DNA from foreign invaders. Bacteriophage T7, for example, encodes the OCR (Overcome Classical Restriction) protein that structurally mimics the phosphate backbone of B-form DNA, allowing it to inhibit host restriction enzymes competitively.77 This mimicry not only shields phage DNA from cleavage but also disrupts host methylation by preventing methyltransferase access to DNA, thereby simultaneously blocking both restriction and methylation pathways to ensure efficient phage infection and replication.78,79 Similarly, phages have evolved mechanisms to overcome alternative RM systems, such as those based on DNA phosphorothioation. Notably, phage JSS1 can bypass PT-mediated immunity, allowing it to successfully infect and replicate within PT-positive bacterial hosts.80
The relationship between phages and their bacterial hosts is shaped by a constant balance between how efficiently a virus attaches to the cell surface, the defence strategies used by bacteria, and the countermeasures developed by phages. The remarkable diversity of bacterial receptors and immune mechanisms, together with the adaptive plasticity of phages, exemplifies an ongoing evolutionary arms race that continually shapes microbial ecosystems. The molecular interactions between phages and their host cells are of considerable scientific interest, as unravelling these mechanisms offers opportunities for therapeutic exploitation and biomedical innovations. This ongoing engagement between bacteriophages and bacteria has driven the evolution of highly specialized defense and counter-defense strategies, reflecting a finely tuned molecular dialogue. The diverse bacterial antiphage defense systems- comprising various biomolecules, enzymes, and cellular processes-mirror the vast heterogeneity of phages, suggesting that additional resistance mechanisms remain to be uncovered. Bacteria deploy diverse antiviral mechanisms that act at multiple stages of the phage infection cycle, while phages continually evolve sophisticated tools to circumvent these barriers. Expanding research beyond foundational insight toward a molecular-level understanding, particularly of understudied single-stranded DNA, RNA, and genetically distinct phages, is essential for advancing phage biology and fully elucidating the complexities of bacterial antiviral immunity.
A deep understanding of these antagonistic interactions is essential for optimizing phage therapy, designing synthetic biological tools, and developing next-generation antimicrobial strategies with enhanced specificity and reduced resistance potential.
ACKNOWLEDGMENTS
The authors thank Nitte (Deemed to be) University for the support
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
Both authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
None.
DATA AVAILABILITY
Not applicable.
ETHICS STATEMENT
Not applicable.
- Danis-Wlodarczyk K, Dabrowska K, Abedon ST. Phage therapy: The pharmacology of antibacterial viruses. Curr Issues Mol Biol. 2020;40:81-163.
Crossref - Dion M B, Oechslin F, Moineau S. Phage diversity, genomics and phylogeny. Nat Rev Microbiol. 2020;18(3):125-138.
Crossref - Albrycht K, Rynkiewicz AA, Harasymczuk M, Barylski J, Zielezinski A. Daily Reports on Phage-Host Interactions. Front Microbiol. 2022;13:946070.
Crossref - Ackermann HW. 5500 Phages examined in the electron microscope. Arch Microbiol. 2007;152(2):227-243.
Crossref - Olszak T, Latka A, Roszniowski B, Valvano MA, Drulis-Kawa Z. Phage Life Cycles Behind Bacterial Biodiversity. Curr Med Chem. 2017;24(36):3987-4001.
Crossref - Salmond GPC, Fineran PC. A century of the phage: past, present and future. Nat Rev Microbiol. 2015;13:777-786.
Crossref - Bertozzi Silva J, Storms Z, Sauvageau D. Host receptors for bacteriophage adsorption. FEMS Microbiol Lett. 2016;363(4): fnw002.
Crossref - Dunne M, Hupfeld M, Klumpp J, Loessner MJ. Molecular basis of bacterial host interactions by gram-positive targeting bacteriophages. Viruses. 2018;10(8):397.
Crossref - Abedon ST. Bacteriophage Adsorption: Likelihood of Virion Encounter with Bacteria and Other Factors Affecting Rates. Antibiotics. 2023;12(4):723.
Crossref - Fernandez L, Gutierrez D, Rodriguez A, Garcia P. Application of bacteriophages in the agro-food sector: A long way toward approval. Front Cell Infect Microbiol. 2018;8:296.
Crossref - Ul Haq I, Chaudhry WN, Akhtar MN, Andleeb S, Qadri I. Bacteriophages and their implications on future biotechnology: A review. Virol J. 2012;9:9.
Crossref - Moye ZD, Woolston J, Sulakvelidze A. Bacteriophage applications for food production and processing. Viruses. 2018;10(4):205.
Crossref - Weitz JS, Poisot T, Meyer JR, et al. Phage-bacteria infection networks. Trends Microbiol. 2013;21(2):82-91.
Crossref - Aksyuk AA, Rossmann MG. Bacteriophage assembly. Viruses. 2011;3(3):172-203.
Crossref - Lebedev AA, Krause MH, Isidro AL, et al. Structural framework for DNA translocation via the viral portal protein. EMBO J. 2007;26(7):1984-1994.
Crossref - Fokine A, Rossmann MG. Molecular architecture of tailed double-stranded DNA phages. Bacteriophage. 2014;4(2):e28281.
Crossref - Zhu Y, Shang J, Peng C, Sun Y. Phage family classification under Caudoviricetes: A review of current tools using the latest ICTV classification framework. Front Microbiol. 2022;13:1032186.
Crossref - Roux S, Krupovic M, Poulet A, Debroas D, Enault F. Evolution and diversity of the microviridae viral family through a collection of 81 new complete genomes assembled from virome reads. PLoS One. 2012;7:0040418.
Crossref - Wu Y, Wu Z, Guo L, et al. Diversity and distribution of a prevalent Microviridae group across the global oceans. Commun Biol. 2024;7(1):1377.
Crossref - Knezevic P, Adriaenssens EM, ictv Report Consortium. ICTV virus taxonomy profile: Inoviridae. J Gen Virol. 2021;102(7):001614.
Crossref - Krupovic Mart, ictv Report Consortium. ICTV Virus Taxonomy Profile: Plasmaviridae. J Gen Virol. 2018;99(5):617-618.
Crossref - Moldovan R, Chapman-McQuiston E, Wu XL. On kinetics of phage adsorption. Biophys J. 2007;93(1):303–15.
Crossref - Kemp P, Garcia LR, Molineux IJ. Changes in bacteriophage T7 virion structure at the initiation of infection. Virology. 2005;340(2):307-317.
Crossref - Zeng X, Wang W, Zhu D, et al. Two receptor-targeting mechanisms of lambda-like siphophage Gifsy-1 of Salmonella Typhimurium. PLoS Pathog. 2025;21(7):e1013352.
Crossref - Veesler D, Cambillau C. A Common Evolutionary Origin for Tailed-Bacteriophage Functional Modules and Bacterial Machineries. Microbiol Mol Biol Rev. 2011;75:423-33.
Crossref - Latka A, Lemire S, Grimon D, et al. Engineering the Modular Receptor-Binding Proteins of Klebsiella Phages Switches Their Capsule Serotype Specificity. ASM J. 2021;12(3).
Crossref - Parent KN, Erb ML, Cardone G, et al. OmpA and OmpC are critical host factors for bacteriophage Sf6 entry in Shigella. Mol Microbiol. 2014;92(1):47-60.
Crossref - Ellen T, Gerrit G, Schutz M, et al. The structure of the receptor-binding domain of the bacteriophage T4 short tail fibre reveals a knitted trimeric metal-binding fold. J Mol Biol. 2003;331(2):361-373.
Crossref - Veesler D, Spinelli S, Mahony J, et al. Structure of the phage TP901-1 1.8 MDa baseplate suggests an alternative host adhesion mechanism. PNAS. 2012;109(23):8954-8958.
Crossref - Silhavy TJ, Kahne D, Walker S. The bacterial cell envelope. Cold Spring Harb Perspect Biol. 2010;2(5):a000414.
Crossref - Xia G, Kohler T, Peschel A. The wall teichoic acid and lipoteichoic acid polymers of Staphylococcus aureus. Int J Med Microbiol. 2010;300(2-3):148-154.
Crossref - Vollmer W, Blanot D, De Pedro MA. Peptidoglycan structure and architecture. FEMS Microbiol Rev. 2008;32(2):149-167.
Crossref - Gaidelyte A, Cvirkaite-Krupovic V, Daugelavicius R, Bamford JKH, Bamford DH. The entry mechanism of membrane-containing phage Bam35 infecting Bacillus thuringiensis. J Bacteriol. 2006;188(16):5925-5934.
Crossref - Shen Y, Kalograiaki I, Prunotto A, et al. Structural basis for recognition of bacterial cell wall teichoic acid by pseudo-symmetric SH3b-like repeats of a viral peptidoglycan hydrolase. Chem Sci. 2021;12(2):576-589.
Crossref - Baptista C, Santos MA, Sao-Jose C. Phage SPP1 reversible adsorption to Bacillus subtilis cell wall teichoic acids accelerates virus recognition of membrane receptor YueB. J Bacteriol. 2008;190(14):4989-96.
Crossref - Habann M, Leiman PG, Vandersteegen K, et al. Listeria phage A511, a model for the contractile tail machineries of SPO1-related bacteriophages. Mol Microbiol. 2014;92(1):84-99.
Crossref - Leiman PG, Chipman PR, Kostyuchenko VA, Mesyanzhinov VV, Rossmann MG. Three-dimensional rearrangement of proteins in the tail of bacteriophage T4 on infection of its host. Cell. 2004;118(4):419-429.
Crossref - Dicks LMT, Vermeulen W. Bacteriophage–Host Interactions and the Therapeutic Potential of Bacteriophages. Viruses. 2024;16(3):478.
Crossref - Fernandes S, Sao-Jose C. Enzymes and mechanisms employed by tailed bacteriophages to breach the bacterial cell barriers. Viruses. 2018;10(8):396.
Crossref - Letarov AV. Bacterial Virus Forcing of Bacterial O-Antigen Shields: Lessons from Coliphages. Int J Mol Sci. 2023;24(24):17390.
Crossref - Kulikov EE, Golomidova AK, Prokhorov NS, Ivanov PA, Letarov AV. High-throughput LPS profiling as a tool for revealing of bacteriophage infection strategies. Sci Rep. 2019;9(1):2958.
Crossref - Letarov AV, Kulikov EE. Adsorption of Bacteriophages on Bacterial Cells. Biochemistry (Moscow). 2017;82(13):1632-1658.
Crossref - Perez-Ruiz M, Pulido-Cid M, Roman Luque-Ortega J, Maria Valpuesta J, Cuervo A, Carrascosa JL. Assisted assembly of bacteriophage T7 core components for genome translocation across the bacterial envelope. Proc Natl Acad Sci U S A. 2021;118(34):e202671911.
Crossref - Chen W, Xiao H, Wang L, et al. Structural changes in bacteriophage T7 upon receptor-induced genome ejection. Proc Natl Acad Sci USA. 2021;118(37):1-7.
Crossref - Guerrero-Ferreira RC, Viollier PH, Ely B, et al. Alternative mechanism for bacteriophage adsorption to the motile bacterium Caulobacter crescentus. Proc Natl Acad Sci U S A. 2011;108(24):9963-9968.
Crossref - Gonzalez F, Helm RF, Broadway KM, Scharf BE. More than Rotating Flagella: Lipopolysaccharide as a Secondary Receptor for Flagellotropic Phage 7-7-1. J Bacteriol. 2018;200(19):e00363-1.
Crossref - Spankie TJ, Haywood AL, Dottorini T, Barrow PA, Hirst JD. Interaction of the maturation protein of the bacteriophage MS2 and the sex pilus of the Escherichia coli F plasmid. J Mol Graph Model. 2020;101:107723.
Crossref - Parra B, Cockx B, Lutz VT, Brondsted L, Smets BF, Dechesne A. Isolation and characterization of novel plasmid-dependent phages infecting bacteria carrying diverse conjugative plasmids. Microbiol Spectr. 2024;12(1):e0253723.
Crossref - Majkowska-Skrobek G, Latka A, Berisio R, et al. Phage-borne depolymerases decrease Klebsiella pneumoniae resistance to innate defense mechanisms. Front Microbiol. 2018;9:2517.
Crossref - Koskella B, Brockhurst MA. Bacteria-phage coevolution as a driver of ecological and evolutionary processes in microbial communities. FEMS Microbiol Rev. 2014;38(5):916-931.
Crossref - Riede I, Eschbach M L. Evidence that TraT interacts with OmpA of Escherichia coli. FEBS Lett. 1986;205(2):241-245.
Crossref - Reyes-Robles T, Dillard RS, Cairns LS, et al. Vibrio cholerae outer membrane vesicles inhibit bacteriophage infection. J Bacteriol. 2018;200(15):e00792-17.
Crossref - Manning AJ, Kuehn MJ. Contribution of bacterial outer membrane vesicles to innate bacterial defense. BMC Microbiol. 2011;11:258.
Crossref - Hoyland-Kroghsbo NM, Maerkedahl RB, Svenningsen S Lo. A quorum-sensing-induced bacteriophage defense mechanism. MBio. 2013;4(1):e00362-12.
Crossref - Belitsky M, Avshalom H, Erental A, et al. The Escherichia coli extracellular death factor EDF induces the endoribonucleolytic activities of the toxins MazF and ChpBK. Molecular Cell. 2011;41(6):625-635.
Crossref - Hampton HG, Watson BNJ, Fineran PC. The arms race between bacteria and their phage foes. Nature. 2020;577(7790):327-336.
Crossref - Levin BR, Moineau S, Bushman M, Barrangou R. The Population and Evolutionary Dynamics of Phage and Bacteria with CRISPR-Mediated Immunity. PLoS Genet. 2013;9(3):e1003312.
Crossref - Cady KC, Bondy-Denomy J, Heussler GE, Davidson AR, O’Toole GA. The CRISPR/Cas adaptive immune system of Pseudomonas aeruginosa mediates resistance to naturally occurring and engineered phages. J Bacteriol. 2012;194(21):5728-5738.
Crossref - Leavitt JC, Woodbury BM, Gilcrease EB, Bridges CM, Teschke CM, Casjens SR. Bacteriophage P22 SieA-mediated superinfection exclusion. MBio. 2024;15(2):e0216923.
Crossref - Teklemariam AD, Al-Hindi RR, Qadri I, et al. The Battle between Bacteria and Bacteriophages: A Conundrum to Their Immune System. Antibiotics. 2023;12(2):381.
Crossref - Schmidt AK, Fitzpatrick AD, Schwartzkopf CM, et al. A Filamentous Bacteriophage Protein Inhibits Type IV Pili To Prevent Superinfection of Pseudomonas aeruginosa. mBio. 2022;13(1):e0244121.
Crossref - Wang W, Li Y, Tang K, et al. Filamentous prophage capsid proteins contribute to superinfection exclusion and phage defence in Pseudomonas aeruginosa. Environ Microbiol. 2022;24(9):4285-4298.
Crossref - Harms A, Brodersen DE, Mitarai N, Gerdes K. Toxins, Targets, and Triggers: An Overview of Toxin-Antitoxin Biology. Mol Cell. 2018;70(5):768-784.
Crossref - Bobonis J, Mitosch K, Mateus A, et al. Bacterial retrons encode phage-defending tripartite toxin–antitoxin systems. Nature. 2022;609(7925):144-150.
Crossref - LeRoux M, Srikant S, Teodoro GIC, et al. The DarTG toxin-antitoxin system provides phage defence by ADP-ribosylating viral DNA. Nat Microbiol. 2022;7(7):1028-1040.
Crossref - Zhang T, Cepauskas A, Nadieina A, et al. A bacterial immunity protein directly senses two disparate phage proteins. Nature. 2024;635(8039):728-735.
Crossref - Dupuis M, Villion M, Magadan AH, Moineau S. CRISPR-Cas and restriction-modification systems are compatible and increase phage resistance. Nat Commun. 2013;4:2087.
Crossref - Sneppen K, Semsey S, Seshasayee ASN, Krishna S. Restriction modification systems as engines of diversity. Front Microbiol. 2015;6:528.
Crossref - Weiserova M, Ryu J. Characterization of a restriction modification system from the commensal Escherichia coli strain A0 34/86 (O83:K24:H31). BMC Microbiol. 2008;8:106.
Crossref - Jiang S, Chen K, Wang Y, et al. A DNA phosphorothioation-based Dnd defense system provides resistance against various phages and is compatible with the Ssp defense system. MBio. 2023;14(4):e0093323.
Crossref - He W, Huang T, Tang Y, et al. Regulation of DNA phosphorothioate modification in Salmonella enterica by DndB. Sci Rep. 2015;5:12368.
Crossref - Meyer JR, Dobias DT, Weitz JS, Barrick JE, Quick RT, Lenski RE. Repeatability and contingency in the evolution of a key innovation in phage lambda. Science. 2012;335:428-432.
Crossref - Maddamsetti R, Johnson DT, Spielman SJ, Petrie KL, Marks DS, Meyer JR. Gain-of-function experiments with bacteriophage lambda uncover residues under diversifying selection in nature. Evolution. 2018;72(10):2234-2243.
Crossref - Scholl D, Rogers S, Adhya S, Merril CR. Bacteriophage K1-5 Encodes Two Different Tail Fiber Proteins, Allowing It To Infect and Replicate on both K1 and K5 Strains of Escherichia coli. J Virol. 2001;75(6):2509-2515.
Crossref - Rauch BJ, Silvis MR, Hultquist JF, et al. Inhibition of CRISPR-Cas9 with Bacteriophage Proteins. Cell. 2017;168(1):150-158.e10.
Crossref - Qin S, Liu Y, Chen Y, et al. Engineered Bacteriophages Containing Anti-CRISPR Suppress Infection of Antibiotic-Resistant P. aeruginosa. Microbiol Spectr. 2022;10(5).
Crossref - Walkinshaw MD, Taylor P, Sturrock SS, et al. Structure of Ocr from bacteriophage T7, a protein that mimics B-form DNA. Mol Cell. 2002;9(1):187-194.
Crossref - Stephanou AS, Roberts GA, Cooper LP, et al. Dissection of the DNA Mimicry of the Bacteriophage T7 Ocr Protein using Chemical Modification. J Mol Biol. 2009;391(3):565-576.
Crossref - Kudryavtseva AA, Csefalvay E, Gnuchikh EY, et al. Broadness and specificity: ArdB, ArdA, and Ocr against various restriction-modification systems. Front Microbiol. 2023;14:1133144.
Crossref - He Y, Jiang S, Wang F, et al. Mechanistic insights into JSS1_004-mediated antagonism of the DndBCDE-FGH restriction system and engineering applications. mBio. 2025;16(8):e0138625.
Crossref
© The Author(s) 2026. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.