ISSN: 0973-7510
E-ISSN: 2581-690X
J Pure Appl Microbiol, 2019, 13 (2): 689-695 | Article Number: 5538
https://dx.doi.org/10.22207/JPAM.13.2.05 | © The Author(s). 2019
Received: 29/03/2019 | Accepted: 04/05/2019 | Published: 15/05/2019
Ricinus communis is a plant that is characterized by its different applications, in addition to being a wild plant favors its inclusion in agroecological practices because it does not represent high costs for producers. The aim of the work was to evaluate the bacterial isolates of R. communis with respect to its inhibitory activity, chitinolytic, DNase, solubilization of phosphate, biofilm formation and growth promoting effect in Allium sativum and Medicago sativa. The collection of R. communis leaves was carried, which were washed and disinfected with sodium hypochlorite solution in order to isolate bacteria in nutritive agar. The isolates obtained were identified by PCR amplification with the oligonucleotides RM and RB and the amplified products were sequenced for identification. The following tests were carried out on each of the isolates: growth inhibition vs Fusarium oxysporum, chitinolytic activity, DNase, solubilization of phosphate, biofilm formation and growth evaluation in A. sativum and M. sativa. Ten isolates were obtained that were identified as Pseudomonas 50%, Enterobacter 30% and Bacillus 20%. Growth inhibition evaluations against F. oxysporum showed that Bacillus showed significant inhibitory activity. The total isolates were negative for chitinolytic activity, 50% were positive for DNase, 40% positive for solubilization of phosphate and for biofilm formation 25% promoted weak formation and 10% moderate. Both plant models showed an increase in the biomass of fresh weight and dry weight of their roots when they were inoculated with the bacterial consortium at 50%, 75% and 100%.
Castor waste, Pseudomonas, Enterobacter, Bacillus, metabolic activity, inhibition, Fusarium oxysporum.
Ricinus communis is a plant of the Euphorbiaceae family considered a wild species and in several countries it is used as a raw material in the production of biofuel, since it has a high production of seed to obtain oil1,2. Other work related to R. communis has focused on technological development for non-food purposes, economic valuation of consumption and production of raw materials and recently studies have been carried out with clinical application of extracts derived from this plant3-6.
The identification of the microbiota present in the residues of R. communis is very promising, due to the contribution of knowledge regarding the metabolic capacity of each one of the isolated strains that favor the protein value of the agroindustrial residues and the enzymatic capacity, facilitating the composting process. Establishing a wide range of relationships between microorganisms in ecosystems, cataloged as agents of biological control and promoters of plant growth 7-9.
In order to minimize negative effects and increase production in agroecosystems research is carried out with various phytosanitary products, plant extracts with insecticidal and antimicrobial action, in addition to products that are used in agroecological practices, however there is still little information that contributes to the definition of suitable strategies for their application in agroecological production systems10.
It has been reported that castor bean cake is a nutrient-rich residue and as a consequence it harbors a considerable number of microorganisms, being Bacillus sp., The genus that is isolated most frequently and that is characterized by forming endospores that make it resistant to the environment In addition, it is a producer of hydrolytic enzymes and antagonists of phytopathogenic fungi. Endophytic bacteria have also been described that inhabit plant tissues actively colonizing them and exerting beneficial functions for the host, as is the case of Acinetobacter, Enterobacter and Pseudomonas11,12.
In relation to that R. communis presents a diverse microbiota, its isolation and characterization will allow the knowledge of microorganisms with potential application in the agroecological practice. The objective of the present investigation was to evaluate the bacterial isolates of R. communis with respect to its inhibitory activity, chitinase, DNase, solubilization of phosphate, biofilm formation and growth promoting effect in A. sativum and M. sativa.
Collection of Ricinus communis and bacterial isolation
Ricinus communis leaves were collected at the margin of Atoyac River in Puebla-Mexico (19.030426-98.224988, subhumid temperate climate, average annual temperature 17.5°C, average precipitation of 1270 mm), transported in paper bags at room temperature for processing in the Research center of microbiological sciences of science institute of Benemerita Universdad Autonoma de Puebla.
The collected plant material was washed with sterile water and disinfected with 0.125% sodium hypochlorite solution (NaClO) for 10 minutes. After disinfection, fragments of 1 cm2 were placed on nutritious agar allowing to incubate for 24 hours at 37°C. From the bacterial development around the fragments of the leaves, isolation was carried out on nutritive agar.
Isolations identification
For identification, the DNA extraction of the isolates was performed using the PureLink Genomic DNA Kit (Invitrogen, USA), amplified by the Polymerase Chain Reaction (PCR) with universal oligonucleotides RM and RB of the 16S gene (5′- AGA GTT TGA TYM TGG CTC AG-3 ‘) and RM (5′- GGA CTA CCA GGG TAT CTA ATC C-3’). The PCR reaction consisted of a final volume of 25 μL per sample (1X PCR buffer, 0.2 mM dNTPs, 1 mM MgCl2, 0.5 μM primers, recombinant DNA polymerase taq (Invitrogen, USA) 1U and 100 ng of bacterial DNA). Using a mini MS thermal cycler (BioRad, USA) and with the following amplification program: initial denaturation at 95°C for 5 minutes, 35 cycles of 92°C for 1 minute, 57°C for 30 seconds, and 72°C for 1 minute. The PCR products were analyzed by agarose gel electrophoresis stained with 1% BIOTIUM. The PCR products were sequenced in biomolecular detection center of Benemerita Universidad Autonoma de Puebla; for which vials containing approximately 20 μL of the products of each PCR reaction were sent. The sequences were analyzed using the CLC main Workbench program 6.1 QIAGEN and the BLAST program 2.2.27 comparing them with the nt NCBI database.
Fusarium oxysporum growth inhibition evaluation
Each isolate was planted on potato dextrose agar, making a line parallel to 2 cm to the end of the plate and incubating for 24 hours at 30°C. A sample with a diameter of 0.5 cm of F. oxysporum was placed with one week of growth at the other end of the plate, the test was carried out in triplicate and had a control where only F. oxysporum was sown. Inhibition was observed from the radial growth of the fungus, estimated based on the difference between the growth of the bacterial isolation compared with the strain and the growth in the control culture. The evaluation was made when the controls were completely invaded.
Isolates chitinase activity evaluation
The stria isolates were seeded in medium supplemented with colloidal chitin as the sole carbon source and incubated for 24 hours at 30°C, the formation of hydrolysis rings around the colony was interpreted as a positive test.
Isolates DNase activity evaluation
The isolates were sown by stria on DNAase Test agar incubated for 24 hours at 30°C, then 10 mL of 1 N HCl was added to each sample and allowed to interact for 30 minutes, the formation of a halo around the bacterial growth was interpreted as a test positive.
Isolates phosphate solubilizing activity evaluation
From the isolations a massive seeding was carried out in the SRS medium, being incubated for 8 days at 37°C, then colonies were selected that grew acidifying the culture medium and forming a transparent halo around the colony, indicating the phosphate solubilizing activity.
Isolates biofilm formation evaluation
Each of the isolates was cultured in nutritious broth to adjust the inocula to 1×106 CFU/mL. From the 24-hours cultures, 200 μL were placed in each well (in triplicate), including the negative control (the medium without inoculation), and incubated for 24 hours at 30°C. Once the incubation time had elapsed, the content of the plate was eliminated, three washes being carried out with sterile distilled water. Violet cristal (1%, 200 μL) was added to each well, allowing it to interact for 5 minutes, in order to stain the bacteria present in the biofilm. Excess dye was removed with distilled water, adding 200 μL of 99% ethanol to each well and reading the optical density at 630 nm using a PoweamWHYM201 plate reader.
The cut-off point (DOC) and the classification of the strains were established according to the recommendations of Stephanovic et al., (2004)13, classifying them as: not producing biofilm, weak, moderate or strongly producing producers. The DOC was established by the average of the optical densities obtained in the negative control plus 3 standard deviations. It was considered as non-biofilm producer those strains with a DOC less than 1 DOC, weak producing strains to those with a DOC greater than 1 DOC and lower than 2 DOC, moderate those that present a DOC between 2 DOC and 4 DOC and strongly producers to those that present a DOC superior to 4 DOC.
Preparation and evaluation of the bacterial consortium in the growth of Allium sativum and Medicago sativa
Obtained and identified the isolates a bacterial consortium was prepared, taking a colony of each isolated genus and placed in test tubes separately and containing 1 mL of nutritive broth incubated at 24 hours at 37°C. Subsequently, the three samples were transferred and mixed in a bottle with 100 mL of nutrient broth, incubated for 24 hours at 37°C, in order to adjust the inoculum to 1×106 CFU/mL, with the following concentrations 25%, 50%, 75% and 100%.
For the experimental design four treatments were established with six replicates, each sample was made up of 100 grams of Eco-substrate plus A. sativum or M. sativa seed, which were inoculated with 25%, 50%, 75% and 100% of consortium and including a control group (samples not inoculated). The samples were inoculated with 1 mL at the different percentages and were evaluated at 20 days of development, under controlled temperature and photoperiod conditions. After this period, the roots were washed with distilled water, separated from the aerial part of the plant, registering its fresh weight and dry weight.
Statistical analysis
The data referring to the percentage of F. oxysporum growth inhibition and of the biomass related to the fresh weight and dry weight of the roots of A. sativum and M. sativa in the different treatments were analyzed with ANOVA (P<0.05) software SPSS 17.0 for Windows.
From the leaves of R. communis ten bacterial isolates were obtained, from which by means of PCR amplification and sequencing, the genus Pseudomonas (50%) RCMS-01, Enterobacter (30%) RCMS-02 and Bacillus (20%) RCMS-03, with coverage and identity greater than 95%, for 100% of the isolates (Table 1). The use of bacterial consortium has been increased by producers due to their successful effects as biostimulators in plants, pathogen suppressors and pest controllers, favoring agro-productive behavior in different crops14,15.
Table (1):
Identification by sequencing of bacterial isolates
Code |
Description |
Query cover (%) |
Perc. Ident (%) |
|---|---|---|---|
RCMS-01 |
Pseudomonas sp |
100 |
98.96 |
RCMS-02 |
Enterobacter sp |
100 |
98.63 |
RCMS-03 |
Bacillus sp |
100 |
98.56 |
The isolates obtained from Ricinus communis were assigned codes, the five Pseudomonas isolates were grouped in the RCMS-01 code, the three Enterobacter isolates in RCMS-02 and the two Bacillus isolates in RCMS-03. RCMS = Research center of microbiological sciences.
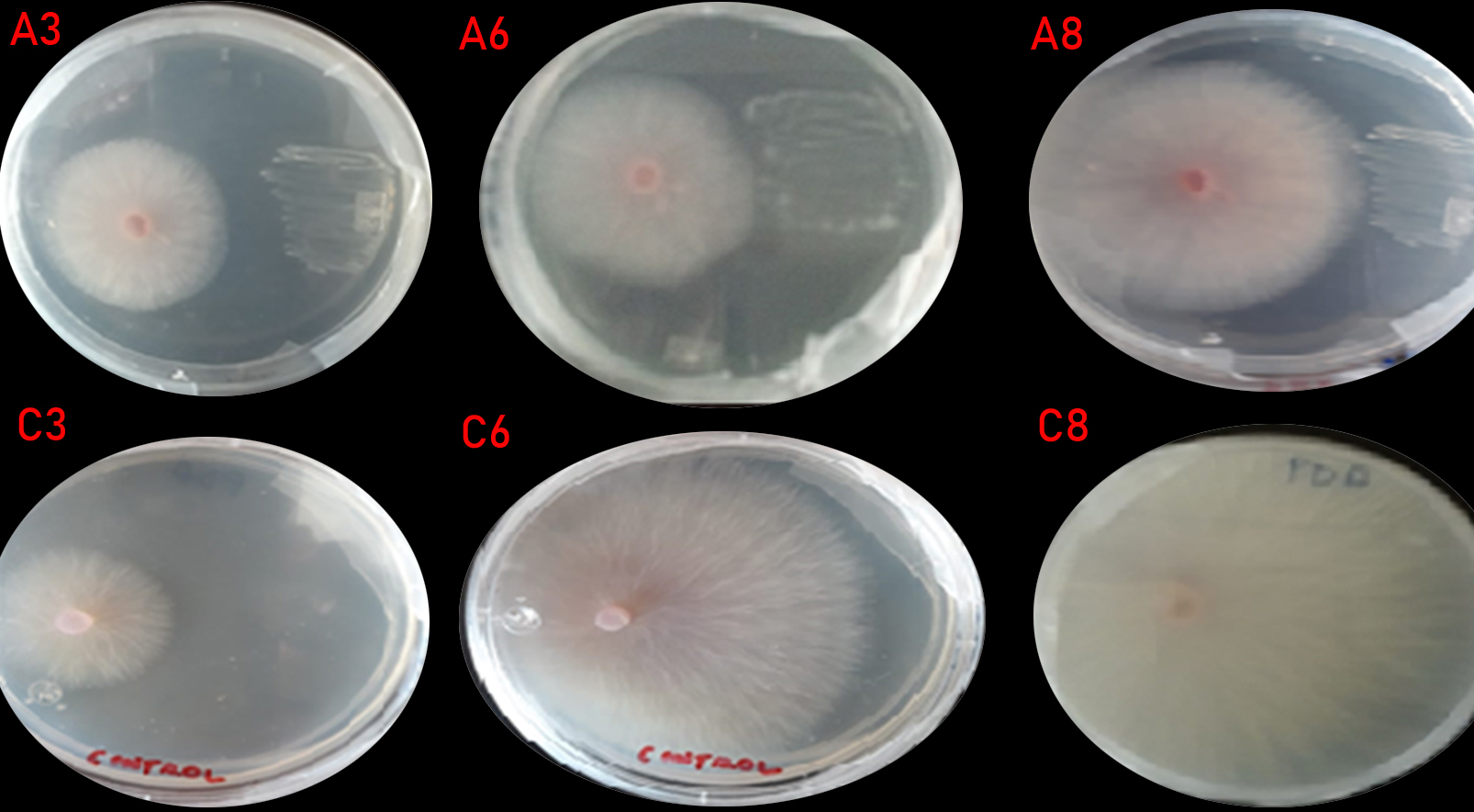
Regarding the evaluation of F. oxysporum growth inhibition in presence of isolates, Bacillus RCMS-03 significantly inhibited the fungus growth with respect to the values obtained in control group and the other two isolated genus, showing significant difference (P <0.05) (Table 2 and Figure 1). According to Orietta and Larrea (2001)16 the mechanism exerted by the bacteria is due to the production of secondary metabolites, which can be volatile or diffusible, but in any case, without coming into physical contact with the pathogen, can inhibit or restrict their growth.
Fig. 1. Panel (A3, A6 and A8) shows Fusarium oxysporum inhibition growth in presence of Bacillus isolation, Panel (C3, C6 and C8) shows Fusarium oxysporum growth without presence of isolation.
Table (2):
Percentage values of Fusarium oxysporum growth in bacterial isolates presence
Day/Isolation |
Control |
Pseudomonas RCMS-01 |
Enterobacter RCMS-02 |
Bacillus RCMS-03 |
|---|---|---|---|---|
1 |
15.6 |
13.2 |
12 |
13.2 |
2 |
28.9 |
21.6 |
22.8 |
21.6 |
3 |
42.1 |
27.7 |
34.9 |
25.3 |
4 |
55.4 |
31.3 |
36.1 |
27.7 |
5 |
67.4 |
38.5 |
38.5 |
31.3 |
6 |
83.1 |
44.5 |
43.3 |
34.9 |
7 |
95.1 |
51.8 |
46.9 |
38.5 |
8 |
100 |
60 |
49.3 |
39.7 |
a |
ab |
ab |
b |
Different letters between columns indicate significant difference (P <0.05)
Evaluation of chitinase activity, the total isolates were negative. In the same way Cabra et al., (2014)8, found that the qualitative tests performed on 16 strains for the detection of chitinase yielded negative results concluding that none of the strains evaluated has the capacity to produce chitinolytic enzymes in microorganisms isolated from residues of R. communis.
Fifty percent of the isolates showed DNase activity, this metabolic capacity being indicative of pathogenicity in most of the bacteria that express it, including among them the microorganisms of genus Pseudomonas17.
The determination of solubilization of phosphate was carried out qualitatively, the appearance of haloes around the seeding in the SRS medium indicated a positive test in 40% of the isolates. It has been reported that isolates of Enterobacter and Bacillus in different anatomical sites of R. communis plant show deaminase activity, production of indole acetic acid and solubilization of phosphate18.
Of the isolations obtained in the present work, 25% were weak producers of biofilm and 10% were moderate producers. This represents an important metabolic process, since it has been demonstrated that the expression of biofilm and the production of surfactin favors the biological control capacity of Bacillus subtilis19. In addition, the establishment of biofilms favors the permanence and interaction of the bacteria in the plant tissues, which has applications in agriculture, since it can be used in the practice of biological control of pathogens, access to new niches, continue colonizing the host and promote the stimulation of plant growth20-22.
From the ten isolates obtained (five isolates of Pseudomonas RCMS-01, three isolates of Enterobacter RCMS-02 and two isolates of Bacillus RCMS-03) the bacterial consortium (1×106 CFU / mL) was formed with the following concentrations 25%, 50%, 75% and 100%.
Plant growth results derived from the application of the bacterial consortium in A. sativum and M. sativa, showed a significant increase in biomass (P <0.05), increasing the fresh and dry weight of the roots with concentrations of 50%, 75 % and 100% with respect to the control (Table 3). This shows the effect of modifying the nutrient accumulation and the plant’s own activities, which were stimulated with the treatments applied.
Table (3):
Biomass values in Allium sativum and Medicago sativa roots
Treatments |
Fresh weight A. sativum |
Dry weight A. sativum |
Fresh weight M. sativa |
Dry weight M. sativa |
|---|---|---|---|---|
Control |
0.11 ± 0.01 a |
0.024 ± 0.01 a |
0.022 ± 0.004 a |
0.0020± 0.0003 a |
25% |
0.39 ± 0.05 b |
0.029 ± 0.01 a |
0.027 ± 0.002 a |
0.0029 ±0.0005 b |
50% |
0.42 ± 0.04 b |
0.123 ± 0.01 b |
0.032 ± 0.005 b |
0.0029 ±0.0004 b |
75% |
0.43 ± 0.08 b |
0.105 ± 0.01 b |
0.034 ± 0.002 b |
0.0030 ±0.0003 b |
100% |
0.54 ± 0.11 c |
0.118 ± 0.01 b |
0.045 ± 0.007 c |
0.0034 ±0.0004 c |
Different letters between rows indicate significant difference (P <0.05)
With respect to genus isolated in the present work, their role in beneficial interactions in plants, usually cataloged as agents for biological control and microorganisms promoting plant growth, has been documented. In the interaction of these microorganisms may be synergistic effects that potentiate the benefits or conversely antagonistic effects23. Therefore, it is suggested to consider agroecological practices because they provide the basic principles to study, design and manage agroecosystems that are productive and conservators of natural resource24.
In conclusion, isolates belonging to genus Pseudomonas (50%), Enterobacter (30%) and Bacillus (20%) were obtained. The consortium evaluated showed significant inhibition in F. oxysporum growth. Total isolates did not present chitinase activity, DNase activity 50% of isolates were positive, and 40% for phosphate solubilization were positive. In the capacity to form biofilm, 25% was characterized by weak formation and 10% by moderate formation. The bacterial consortium formed from the isolates of R. communis promoted an increase in biomass in A. sativum and M. sativa, increasing the fresh and dry weight of the plant. The potential of these bacterial isolates is related to their metabolic pathways and adaptability, which allows their inclusion as a sustainable alternative in different agroecological practices.
Acknowledgments
The Authors are grateful to CONACYT for the scholarship granted to the first author to develop his masters studies in Sustainable Management of Agroecosystems and Vicerrectoria de Investigacion y Estudios de Posgrado (VIEP), Science Institute of Benemerita Universidad Autonoma de Puebla.
Conflicts Of Interest
The authors declare that there are no conflicts of interest.
Authors’ Contribution
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
Funding
This Project was supported by grant 100233566-VIEP2019 from the Vice Presidency of Research and Graduate Studies, Autonomous University of Puebla, Mexico.
Data Availability
All datasets generated or analyzed during this study are included in the manuscript and/or the Supplementary Files.
Ethics Statement
This article does not contain any studies with human participants or animals performed by any of the authors.
- Solis B.J.L., Muסoz O.A., Escalante E.J.A., Zamarripa C.A. Crecimiento de variedades y componentes del rendimiento de higuerilla (Ricinus communis L.) en Montecillo, estado de Mexico. Revista Mexicana de Ciencias Agricolas, 2016; 7: 311-323.
- Rodriguez H.R., Zamarripa C.A. Competitividad de la higuerilla (Ricinus communis) para biocombustible en relacion a los cultivos actuales en el estado de Oaxaca, Mexico. Revista Mexicana de Agronegocios, 2013; 32: 306-318.
- Alvarado-Montero H.R., Monge-Montero C., Vargas M.S., Giselle L., Mata-Segreda J.F. Absence of a molecular structural effect on thermo-dynamic properties of several biodisel materials. Cuadernos de Investigacion UNED, 2018; 10: 362-367.
- Sandoval G.M.A., Almendarez-Hernandez M.A., Nieto G.A., Troyo D.E., Ortega R.A., Beltran M.L.F. Estudios Sociales Revista de Alimentacion Contemporanea y Desarrollo Regional, 2018; 28: 1-24.
- Sousa N.L., Cabral G.B., Vieria P.M., Baldoni A.B., Aragao F.J.L. Bio-detoxification of ricin in castor bean (Ricinus communis L.) seeds. Scientific Reports, 2017; 13: 1-9.
- Badaro M.M., Salles M.M., Leite V.F.M., Arruda C.N.F., Oliveira V.C., Nascimento C.D., Paranhos H.F.O., Silva-Lovato C.H. Clinical trial for evaluation of Ricinus communis and sodium hypochlorite as denture cleanser. Journal of Applied Oral Science, 2017; 25: 324-334.
- Cabra-Cendales T., Meneses-Cabezas D.C., Galeano-Vanegas N.F. Identificacion de microorganismos asociados a residuos de higuerilla (Ricinus communis). Revista Colombiana de Quamica 2015; 44: 10-15.
- Cabra C.T., Rodriguez G.C.A., Villota C.C.P. Capacidad antagonica y quinolitica de microorganismos aislados de residuos de higuerilla (Ricinus communis). Biotecnologia en el Sector Agropecuarios y Agroindustrial, 2014; 12: 56-61.
- Izzeddin A.N., Medina T.L. Efecto del control biologico por antagonistas sobre fitopatogenos en vegetales de consumo humano. Salus/Revista de la Facultad de Ciencias de la Salud, 2011; 15: 8-12.
- Mamprim A.P., Angeli A.L.F., Silva P.F.G., Andressa F.M., Castilho M.C., Barbosa P.R. Efecto de productos fitosanitarios sobre parametros biologicos de Bauveria bassiana (Bals.) Vuill. (Hypocreates: Cordycipitaceae). Revista de Proteccion Vegetal, 2014; 29: 128-136.
- Kumar R., Singh S., Singh O. Bioconversion of lignocellulosic biomass: biochemical and molecular perspective. Journal Indian of Microbiolgy Biotechnology, 2008; 35: 377-391.
- Riveron A.M.B., Coimbra D.L.G. Pesquisa da existencia de bacterias Gram negativas endofiticas potencialmente aptogenicas para o homem em hortalicas provenientes de cutlivos organicos. Revista Venezolana de Endocrinologia y Metabolismo, 2015; 13: 14-24.
- Stephanovic S., Cirkovic I., Ranin, L., Svabic-Vlahovic M. Biofilm formation by Salmonella spp. and Listeria monocytognes on plastic surfaces. Letters Applied Microbiology, 2004; 38: 428-443.
- Olivera V.C., Leiva L., Calero A., Melendrez J.F. Empleo de microorganismos nativos multi-proposito (MNM) en el comportamiento agro-productivo de cultivos hortםcolas. Agrotecnia Cuba, 2015, 39: 34-40.
- Almoguea F.M., Terrero M.W., Rodriguez M.R. Impacto del curso los microorganismos eficientes y su uso en la agricultura en docentes de especialidades agropecuarias, provincia Cienfuegos. Revista Conrado, 2016; 12: 40-46.
- Orietta F., Larrea V. Microorganismos anta-gonistas para el control fitosanitario. Manejo Integrado de Plagas (Costa Rica), 2001; 62: 96-100.
- Ma L.S., Hachani A., Lin J.S., Filloux A., Lai E.M. Agrobacterium tumefaciens deploys a superfamily of type VI secretions DNase effectors as weapons for interbacterial competition in plants. Cell Host & Microbe, 2014; 16: 94-104.
- Prasad M.N.V., Rajkumar M., Annapurna D. Bioremediation and bioeconomy “Potential of castor bean (Ricinus communis L.) for phytoremediation of metalliferous waste assisted by plant groth-promoting bacteria: possible cogeneration of economic products”. Elsevier, 2016. Pp. 149-175.
- Bais H.P., Fall R., Vivanco J.M. Biocontrol of Bacillus subtilis against infection of Arabidopsis roots by Pseudomonas syringae is facilitated by biofilm formation and surfactin production. Plant Physiology, 2004; 134: 307-319.
- Andrews J.S., Rolfe S.A., Huang W.E., Scholes J.D., Banwart S.A. Biofilm formation in environmental bacteria is influenced by different macro-molecules depending on genus and species. Environmental Microbiology, 2010; 12: 2496-2507.
- Rojas B.M.M. Quorum sensing en la associacion beneficiosa de las bacterias con las plantas. Revista Colombiana de Biotecnologםa, 2011; 13: 135-143.
- Arruebarrena-Di Palma A., Pereyra C., Moreno-Ramorez L., Xiqui-Vazquez M.L., Baca B.E., Pereyra M.A., Lamattina L., Creus C.M. Denitrification derived nitric oxide modulates biofilm formation in Azospirillum brasilense. FEMS Microbiology Letters, 2013; 338: 1-9.
- Cano M.A. Interaccion de microorganismos beneficos en plantas: micorrizas, Trichoderma spp., Pseudomonas spp., una revision. Revista UDCA Actualidad & Divulgacion Cientafica, 2011; 14: 15-31.
- Sanchez P.M., Prager M.M., Naranjo R.E., Sanclemente O.E. El suelo, su metabolismo, ciclaje de nutrientes y practicas agroecologicas. Agroecologia, 2012, 7: 19-34.
© The Author(s) 2019. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.