ISSN: 0973-7510
E-ISSN: 2581-690X
Pseudomonas aeruginosa (PA) is part of a group of common nosocomial pathogens that exhibit multidrug resistance, thus proving to be a significant threat to healthcare. This study analyzes the ability of four commonly used antibiotics to observe eradication of the PA biofilm growth. Ceftazidime (CAZ), Tobramycin (TOB), Ofloxacin (OFLX), Meropenem (MEM), were tested against overnight cultures of PA strain PA01. The minimal inhibitory concentrations (MIC) of planktonic cells for all the four antibiotics were determined using broth microdilution while the minimal bactericidal concentrations (MBCs) were determined by colony count after antibiotic treatment and regrowth. Biofilm growth inhibition was performed by treating cells with antibiotic at the time of inoculation while eradication was determined by adding antibiotics 24 hours after inoculation, allowing mature biofilm formation, followed by the measurement of absorbance. PA planktonic cells exhibited highest susceptibility to MEM compared to overnight grown PA biofilm which demonstrated resistance to CAZ, complete sensitivity to ofloxacin, and minimal sensitivity to TOB and MEM. PA biofilm displayed dose-dependent sensitivity to TOB, MEM and OFLX, and a significant level of resistance to CAZ during inhibition phase. However, in the eradication phase, PA showed significant resistance to TOB followed by CAZ while PA biofilm showed sensitivity at higher concentrations of MEM. Our study exhibits that PA strain PA01 is resistant to ceftazidime in both planktonic and biofilm phases. While ofloxacin proved to be the most effective even at lower concentrations when compared with other antibiotics, tobramycin was most effective at higher concentrations for eradicating and inhibiting PA biofilms.
Pseudomonas aeruginosa, biofilm, CAZ: ceftazidime, OFLX: ofloxacin, MEM: meropenem, TOB: tobramycin
Pseudomonas aeruginosa is a species of great medical significance due to its status as an important opportunistic human pathogen.1 It is important to note that these Gram-negative bacteria single-handedly account for 10% of all hospital-acquired infections and are also responsible for killing thousands of people annually.2 These high mortality rates can be attributed to the appearance of drug-resistant strains and biofilm formation.3 Costerton et al have defined biofilm as a three-dimensionally structured group comprising of different microorganisms, typically enclosed in a self-produced polymeric matrix, that adhere to a biotic or an abiotic surface.4 It has been shown that biofilm formation starts with the adherence of single planktonic cells to a conducive surface. These initial adherent cells grow into microcolonies which then further proliferate into mature biofilms. The matured biofilms produce components of the extracellular matrix (ECM) which are critical in maintaining the distinctive structures of cellular aggregates.5 The ECM is crucial in biofilm persistence because of its role in protecting the colonies from environmental fluctuations and acting as a barrier against antimicrobial agents from the host or otherwise. Most importantly, compared to their planktonic equivalents, bacteria encased in biofilm can be more resistant to antibiotics by almost 1000-fold. This level of resistance is often enough to render the relevant antibiotic therapy ineffective.6,7 Several factors are plausible contributors to enhanced bacterial resistance: production of ECM, different cell surface properties of bacterial embedded in biofilm, metabolic shift, and slow growth.8 In P. aeruginosa, the efflux pump MexAB-OprM is largely responsible for resistance to antibiotics. The MexAB-OprM system, a tripartite pump is one of the largest multi drug resistant efflux pumps with high levels of expression of the RND (Resistance nodulation division) family in P. aeruginosa. The pump extrudes antimicrobials across the outer membrane, which in turn helps to bestow resistance to beta-lactams that target the cell wall synthesis.9 Over the years, antibiotic therapy has been the mainstay of treatment however, due to adaptive resistance, the commonly used antibiotics are becoming no more effective in completely eradicating PA.
Currently, eight categories of antibiotics are primarily used to treat P. aeruginosa infections. These categories comprise of penicillin with β-lactamase inhibitors (BLI) (ticarcillin and piperacillin in combination with clavulanic acid or tazobactam), monobactams (aztreonam), aminoglycosides (amikacin, netilmicin, gentamicin, tobramycin), carbapenems (meropenem, imipenem), cephalosporins (cefepime, ceftazidime), polymyxins (colistin, polymyxin B), Fosfomycin and fluoroquinolones (levofloxacin, ciprofloxacin, ofloxacin).11 Fluroquinolones monotherapy has also shown effectiveness against PA in vitro, indicating its potential as an alternative to beta lactams.10
P. aeruginosa develops resistance to aminoglycosides by producing inactivating enzymes, fluoroquinolones by producing DNA gyrase and topoisomerase IV which causes target modifications. For carbapenems, P. aeruginosa produces Class B Metallo-β-lactamase (IMP, VIM, SPM, GIM), Class A serine carbapenemase (KPC), and Class D carbapenemase (OXA-types: OXA-40) to inactivate the antibiotics. Other mechanisms include Efflux systems (overexpression).11 In addition to the production of carbapenemases, the over production of major efflux pumps such as MexAB-OprM, MexXY-OprM, and MexCD-OprJ also contributed to the carbapenem resistance in PA.15 Whereas the beta lactam resistance in Pseudomonas aeruginosa is mainly driven by the over production of AmpC beta lactamase present in the chromosome.12
With increasing resistance of PA01 to these antibiotics, we therefore, investigated four antibiotics from different categories for both the planktonic and biofilm phase and determined their resistance pattern and their effectiveness in inhibiting PA01 biofilm formation and eradication.
Objectives
- To optimize the growth conditions and process for measurement of Pseudomonas aeruginosa PA01 biofilms.
- To study the antibiotic sensitivity of Pseudomonas aeruginosa PA01 planktonic cells and biofilms.
- To study the effect of various antibiotics on Pseudomonas aeruginosa PA01 on Planktonic phase, to establish minimum inhibitory concentrations (MIC) and Minimum Bactericidal concentrations (MBC) for each antibiotic.
- To evaluate the extent of inhibition and eradication of Pseudomonas aeruginosa PA01 biofilm biomass by various antibiotics.
Bacterial strains, growth media, and conditions
The Pseudomonas aeruginosa strain, PAO1 utilized in this study was preserved by our laboratory at −80°C. Tryptic Soy Agar (TSA) and Tryptic Soy Broth (TSB) media were employed to culture the planktonic form of Pseudomonas aeruginosa bacteria whereas, M63 media supplemented with arginine alone or glucose and casamino acid were used for formation of pseudomonas aeruginosa biofilm.
Materials
Antibiotic discs of ceftazidime (CAZ), Meropenem (MEM), tobramycin (TOB), piperacillin (PRL) and ofloxacin (OFLX) were purchased from Oxoid limited, UK. The powdered form of those antibiotics was purchased from Acros organic and Alfa aesar. The powdered antibiotics were dissolved in bioreagent water according to the manual of antimicrobial susceptibility testing published by American society of Microbiology (2005). The final stock concentrations of MEM, OFLX, TOB, and CAZ in stock solution were 0.25 ug/ml, 0.5 ug/ml, 1 ug/ml, 2 ug/ml, 4ug/ml, 8 ug/ml, 16 ug/ml, 32 ug/ml, 128 ug/ml, 128 ug/ml, 64 ug/ml, respectively.
Optimization of growth media for biofilm growth (M63 supplemented with arginine vs glucose and casamino)
To determine the optimized media for biofilm production, we tested two media as described by Microtiter Dish Biofilm Assay 2011.
- M63 minimal media enhanced with magnesium sulfate, glucose, and casamino acid.
- M63 minimal media augmented with magnesium sulfate and arginine.
Hundred-micromilliter (100 µmL) of the media was added to each 96 well dish. This was triplicated and incubated for overnight at 37°C. The biomass of biofilm was determined by a plate reader at 540 nm after CV staining.
Antibiotics disc diffusion testing on Pseudomonas aeruginosa
Antibiotic diffusion assay for both fresh and overnight plates were determined as described by the Manual of Antimicrobial Testing except that CFU counting was used to standardize the culture.13 This assay includes steps such as selection of PAO1 colonies, preparation of liquid culture suspension, inoculating entire TSA plates, adding antimicrobial disks within 15 minutes of plate inoculation, overnight incubation, and measurement of zone of inhibition.13 For planktonic phase, overnight plates were used to prepare liquid cultures and incubated at 37°C for 3.5 hours to reach the log phase. TSA plates were inoculated entirely with bacterial culture and antibiotic discs were placed on the agar plates after which it was incubated for overnight at 37°C. In the biofilm phase, plates inoculated entirely with the bacteria were incubated at 37°C overnight before antibiotics discs were place on the agar plates. These plates were re-incubated at 37°C overnight. The antibiotic discs used were Ofloxacin (OFX) 5 ug/ml, Meropenem (MEM) 10 ug/ml, Tobramycin (TOB) 10 ug/ml, Piperacillin (PRL) 100 ug/ml and Ceftazidime (CAZ) 30 ug/ml. In diameter of the inhibition zones were computed and designated as susceptible, intermediate, or resistant.
Minimal Inhibitory Concentration (MIC)
The minimal inhibitory concentration (MIC) was determined by broth microdilution method described by the CLSI 2015. Using McFarlan 0.5 solution as a standard for turbidity of the overnight broth culture, 10 ul of P. aeruginosa was dispensed per well in a 96-well plate. The various concentrations of antibiotics were added, and the volume adjusted with TSB to 100 ul in each well. The plates were incubated overnight at 37°C. MIC was established as the lowest concentration with absence of any visible bacterial growth.
Minimal Bactericidal Concentration (MBC)
The MBC of planktonic P. aeruginosa were determined for the four antibiotics according to the Manual of Antimicrobial Susceptibility Testing published by American society of Microbiology (2005). Using an overnight PAO1 plate incubated, 3 loop full of cells in 150 ml of TSB were incubated at 37°C on a shaker. Ten-milliliter (10 mL) of culture were taken during the logarithmic phase into 36 sterile tubes (triplicated) and standardized to OD600 before treating with the various concentrations of antibiotics. (Ofloxacin: 0.5 ug/ml, 1ug/ml and 32 ug/ml, Ceftazidime: 4 ug/ml, 8 ug/ml, 16 ug/ml, Meropenem: 1ug/ml, 2 ug/ml, 3 ug/ml, Tobramycin: 0.5 ug/ml, 1ug/ml, 2 ug/ml). The antibiotic-treated culture was serial diluted 10-1 to 10-7 and plated. The plates were incubated at 37°C overnight. CFUs and MBC established for each plate.
Eradication assay and biofilm inhibition
The biofilm inhibitory assay carried out in microplates.14 4 μl bacterial suspension from 1: 100 overnight culture and 196 μl TSB were dispensed per well in a 96-well microplate and exposed to different concentrations of the 4 antibiotics. After overnight incubation at 37°C, biofilm biomass was determined by a plate reader after CV staining and dissolved with acetic acid at 540 nm. For biofilm eradication, 100 μl bacterial suspension from 1:100 overnight culture was dispensed per well in microplates. After static incubation at 37°C overnight, the supernatants were removed, and the wells treated with the various concentrations of the antibiotics. The plates were incubated for overnight at 37°C and biofilm biomass determined as previously described.
Optimization of biofilm growth media
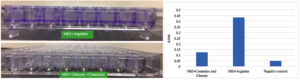
When glucose and casamino acid was replaced with arginine as a sole carbon and energy source, more robust biofilm production was noted (Fig 1a). It was found that arginine supplemented M63 media showed 3.5 folds increase in P. aeruginosa biofilm production capacity compare to glucose and casamino supplemented M63 media (Fig 1b). Arginine supplemented M63 media can be recommended to produce robust P. aeruginosa biofilm compared to casamino supplemented M63 media.
Fig. 1. Biofilm-forming capacity Pseudomonas aeruginosa in M63 minimal media with arginine vs glucose and casamino. Formation of biofilms in 96-well microtiter plates incubated at 37°C for 24 hours were stained utilizing crystal violet. 1(a) Representative images of biofilm formation after CV staining. 1(b) Graphical representation of Pseudomonas aeruginosa biofilm-forming capacity as measured by spectrophotometric determination
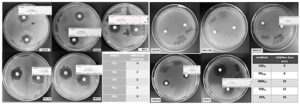
The planktonic cells of P. aeruginosa showed most sensitive to meropenem followed by piperacillin, ofloxacin, tobramycin and ceftazidime. The planktonic cells did not show complete resistance to any of the antibiotic tested (Fig 2a). Whereas the overnight grew P. aeruginosa showed maximum sensitivity to ofloxacin followed by tobramycin and meropenem, and complete resistant to ceftazidime and piperacillin (Fig. 2b). As hypothesized, P. aeruginosa plates after 24 hours might have started producing biofilms making difficult for these antibiotics to penetrate to kill the bacteria.
Fig. 2. Antibiotics disc susceptibility assay on pseudomonas aeruginosa planktonic cells (2a) and biofilm (2b) (Scale added to measure the zone of inhibition)
Effect of different antibiotics on Pseudomonas aeruginosa biofilm inhibition and eradication
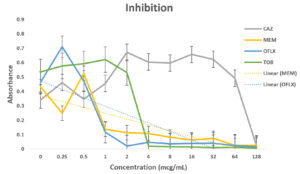
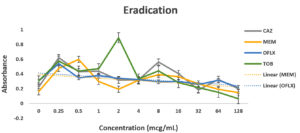
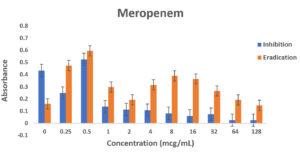
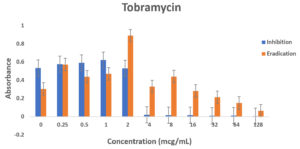
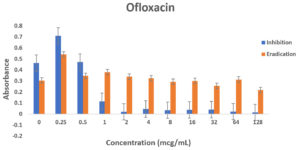
Pseudomonas aeruginosa showed dose dependent sensitivity to OFLX, MEM, TOB and resistance to CAZ in inhibition phase (Fig. 3a). P. aeruginosa exhibited a significant resistance to all antibiotics except for ofloxacin during eradication phase. In eradication phase, P. aeruginosa biofilms displayed sensitivity at higher concentration of tobramycin, (32ug/ml). The highest level of resistance observed for CAZ, and TOB at lower concentrations, respectively during eradication phase (Fig 3b).
Fig. 3. Antibiotic activity of different antibiotics on PA01. The effects of antibiotics on biofilm inhibition (n = 6) where the antibiotic is applied at the time of inoculation (a) and biofilm eradication (n = 6) where the antibiotic is applied after mature biofilm formation (b) at various concentrations. The absorbance of 540 nm (A540 nm) was utilized for data collection of residual biofilms
Pseudomonas aeruginosa infection is one of the significant threats among acutely ill and immunocompromised patients in healthcare centers. The strain PA01 is a clinical isolate that has been turned into a laboratory strain. The genome sequence for the strain is publicly available and an ordered transposon mutant library is also available, making it very amenable to study.16
Table (1):
Determination of the Minimal Inhibitory Concentration (MIC) and Minimal Bactericidal Concentration (MBC).
Antibiotics |
MIC (ug/ml) |
MBC (ug/ml) |
|---|---|---|
Ofloxacin |
2.0 |
2.0 |
Ceftazidime |
4.0 |
4.0 |
Meropenem |
1.0 |
3.0 |
Tobramycin |
2.0 |
2.0 |
Arginine is known to stimulate a more robust biofilm in PA01.17 Our arginine supplemented M63 media showed 3.5 folds increase in P. aeruginosa biofilm growth compared to glucose and casamino supplemented M63 media. The planktonic cells of Pseudomonas aeruginosa showed highest susceptibility to MEM then OFLX, TOB and CAZ. Whereas, P. aeruginosa 24 hours mature biofilm demonstrated resistance to CAZ, minimal sensitivity to TOB and MEM; and absolute sensitivity to OFLX.
Remarkably, the MIC and MBC concentrations for OFLX, CAZ, and TOB were the same (2 ug/ml, 4 ug/ml, 2 ug/ml respectively), whereas MEM showed MIC 1ug/ml and MBC 3ug/ml.
Dose-dependent sensitivity to MEM, TOB and OFLX, and CAZ in inhibition phase was observed. Significant level of resistance to tobramycin (till 128 ug/ml), OFLX and CAZ was established in the eradication phase. Again, P. aeruginosa biofilm was sensitive at higher concentration of MEM, (32ug/ml) during the eradication phase. A recent study indicated that PA resistance to carbapenems are contributed to processes such as production of carbapenemase, efflux pumps.15
Biofilm formation is time-dependent. The results show that less concentration of the same antibiotic is needed to treat an early P. aeruginosa infection compared to the 24-hour mature infection. For the treatment of severe P. aeruginosa infections in critically ill patients, administering proper antibiotic regimen empirically and earlier has presented positive outcomes and it may assist in decreasing mortality.18 The results obtained indicates that Pseudomonas aeruginosa strain PA01 is resistant to ceftazidime in biofilm phase and as planktonic bacteria. Importantly, ofloxacin showed the most effectiveness even at lower concentrations when compared with other antibiotics, whereas tobramycin was most effective at higher concentrations for eradicating and inhibiting PA biofilms. However, the side effects incidents could increase when higher concentrations of antibiotics are administered. In a recent study, it was concluded that PA biofilms will not be eradicated with low dose tobramycin. 19 Overall, ofloxacin is the most preferred antibiotic for both eradication and inhibition of PA biofilms. Pseudomonas aeruginosa strain PA01 biofilms exhibited least eradication with ceftazidime at all the concentrations.
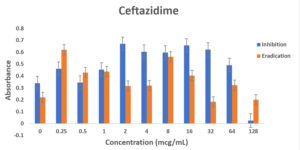
Fig. 4. Biofilm forming capacity (n=6) of P. aeruginosa against different concentration of CAZ (a), MEM (b), TOB (c) and OFLX (d) during inhibition and eradication phase. The data collection are performed at absorbance 540 nm
Even though wild-type strain of PA is still found to be susceptible to ceftazidime, AmpC overexpression with or without point mutation may decrease the susceptibility to β-lactams with exception of carbapenems. This might be the reason why in the eradication experiment, as the concentration of ceftazidime was increasing, AmpC was overexpressed exhibiting the resistance.12,20 Tobramycin has shown a similar effect in both the inhibition and eradication experiments, (Fig 3a and 3b as well as (Fig. 3a and 3b as well Fig 4c), though as the antibiotic concentration increases, the resistance decreases. In our study, ofloxacin was determined to be the most consistent best option for treating P. aeruginosa infection as shown in Fig 3a and 3b and Fig 4d.
Clinicians are now shifting to newer combination antibiotic therapies to treat resistant strains of P. aeruginosa such as ceftolozane/tazobactam, ceftazidime/avibactam, etc. Recent study indicates that the continuous infusion of ceftolozane/tazobactam may prove to be effective and safe to treat infections caused by PA in the outpatient settings.21 Evaluation of safety and efficiency of these novel drug combinations require additional data from research study.
This study shows that Pseudomonas aeruginosa strain PA01 is resistant to ceftazidime in planktonic and biofilm state. It is also demonstrated here that PA01 biofilm formation and eradication was inhibited by ofloxacin. Tobramycin is most effective at higher concentrations for both inhibition and eradication of PA biofilms. Further genomic experiments will be warranted to identify genes that are overexpressed or suppressed during the antibiotics treatment to understand PA01 biofilm resistance to ceftazidime.
ACKNOWLEDGMENTS
The authors special appreciation to Dr. Zvi Loewy, Jaspreet Kaur, Calvin Wang for their assistance.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
PB conceptualized and supervised the study, designed the methodology and participated in final review and editing. DP, YH, PS, YH, MB, BS applied methodology and did the investigation. DP contributed to software work and wrote the draft. PS did the validation and grammatical editing and performed the formal data analysis.
FUNDING
None.
ETHICS STATEMENT
Not applicable.
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the manuscript and available from the corresponding author on reasonable request.
- Sousa AM, Pereira MO. Pseudomonas aeruginosa Diversification during Infection Development in Cystic Fibrosis Lungs-A Review. Pathogens. 2014;3(3):680-703.
Crossref - Mena KD, Gerba CP. Risk Assessment of Pseudomonas aeruginosa in Water. Rev Environ Contam Toxicol. 2009;201:71-115
Crossref - Bassetti M, Vena A, Croxatto A, Righi E, Guery B. How to manage Pseudomonas aeruginosa infections. Drugs in Context. 2018;7:212527.
Crossref - Costerton JW, Stewart PS, Greenberg EP. Bacterial biofilms: A common cause of persistent infections. Science. 1999;284(5418):1318-1322.
Crossref - Hentzer M, Wu H, Andersen JB, et al. Attenuation of Pseudomonas aeruginosa virulence by quorum sensing inhibitors. EMBO Journal. 2003;22:3803-3815.
Crossref - Chu M, Zhang MB, Liu YC, et al. Role of berberine in the treatment of methicillin-resistant Staphylococcus aureus infections. Sci Rep. 2016;6(1):24748.
Crossref - Drilling A, Morales S, Boase S, et al. Safety and efficacy of topical bac- teriophage and ethylenediaminetetraacetic acid treatment of Staphylococcus aureus infection in a sheep model of sinusitis. Int. Forum Allergy Rhinol. 2014;4(3):176-186.
Crossref - Stryjewski ME, Corey GR. Methicillin-resistant Staphylococcus aureus: An evolving pathogen. Clin Infect Dis. 2014;58(Suppl 1):S10-S19.
Crossref - Tsutsumi K, Yonehara R, Ishizaka-Ikeda E, et al. Structures of the wild-type MexAB-OprM tripartite pump reveal its complex formation and drug efflux mechanism. Nature Communications. 2019;10(1):1520.
Crossref - Wu PF, Lin YT, Wang FD, Yang TC, Fung CP. Is fluoroquinolone monotherapy a useful alternative treatment for Pseudomonas aeruginosa bacteremia? Infection. 2018;46(3):365-373.
Crossref - Fujii A, Seki M, Higashiguchi M, Tachibana I, Kumanogoh A, Tomono K. Community-acquired, hospital-acquired, and healthcare-associated pneumonia caused by Pseudomonas aeruginosa. Respir Med Case Rep. 2014;12:30-33.
Crossref - Torrens G, Hernandez SB, Ayala JA, et al. Regulation of AmpC-Driven β-Lactam Resistance in Pseudomonas aeruginosa: Different Pathways, Different Signaling. mSystems. 2019;4(6):e00524-19.
Crossref - Cavalieri SJ. Manual of Antimicrobial Susceptibility Testing. United States: American Society for Microbiology. 2009.
- Qu L, She P, Wang Y, et al. Effects of norspermidine on Pseudomonas aeruginosa biofilm formation and eradication. Microbiology Open. 2016;5(3):402-412.
Crossref - Hassuna NA, Darwish MK, Sayed M, Ibrahem RA. Molecular Epidemiology and Mechanisms of High-Level Resistance to Meropenem and Imipenem in Pseudomonas aeruginosa. Infection and Drug Resistance. 2020;13:285-293.
Crossref - Liberati NT, Urbach JM, Miyata S, et al. An ordered, nonredundant library of Pseudomonas aeruginosa strain PA14 transposon insertion mutants. Proc Natl Acad Sci U S A. 2006;103(8):2833-2838.
Crossref - Otoole GA. Microtiter Dish Biofilm Formation Assay. J Vis Exp. 2011;(47):2437.
Crossref - Pena C, Suarez C, Tubau F, et al. Carbapenem-resistant Pseudomonas aeruginosa: factors influencing multidrug-resistant acquisition in non-critically ill patients. Eur J Clin Microbiol Infect Dis. 2009;28(5):519-522.
Crossref - Mangiaterra G, Cedraro N, Vaiasicca S, et al. Role of Tobramycin in the Induction and Maintenance of Viable but Non-Culturable Pseudomonas aeruginosa in an In Vitro Biofilm Model. Antibiotics. 2020;9(7):399.
Crossref - Davies DG, Parsek MR, Pearson JP, Iglewski BH, Costerton JW, Greenberg EP. The involvement of cell-to-cell signals in the development of a bacterial biofilm. Science. 1998;280(5361):295-298.
Crossref - Jones BM, Huelfer K, Bland CM. Clinical and Safety Evaluation of Continuously Infused Ceftolozane/Tazobactam in the Outpatient Setting. Open Forum Infectious Diseases. 2020;7(2):ofaa014.
Crossref
© The Author(s) 2021. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.