ISSN: 0973-7510
E-ISSN: 2581-690X
Halophilic actinobacteria produce secondary metabolites with diverse biological activities. This study investigates bioactive compounds from termite mound isolates collected across Virudhunagar, Tamil Nadu. Twenty halophilic isolates were screened; only ten tolerated 12% NaCl. Among them, isolate N3 exhibited strong antibacterial activity against Klebsiella pneumoniae MTCC 39 and Staphylococcus aureus MTCC 96, and antifungal activity against Candida albicans MTCC 282. Bioactive metabolites were purified using TLC and identified via GC-MS, revealing diketopiperazine- and peptide-based compounds with antimicrobial, antitumor, and anti-inflammatory potential. 16S rRNA sequencing confirmed isolate N3 as Microbacterium barkeri (98% similarity). The study highlights the potential of halophilic actinobacteria as sources of novel antimicrobial agents and proposes possible coupling pathways between hydroxyl and carbonyl groups within bioactive diketopiperazines.
Actinobacteria, Antibacterial, Anti-tumour, Anti-Inflammatory, Termite Mound
Termite mounds have been traditionally used in many cultures for treating skin rashes and diseases due to their rich mineral content, microbial properties, and potential healing effects. The soil from termite mounds contains antibacterial, antifungal, and anti-inflammatory compounds that may help soothe skin irritations, infections, and wounds. The importance of termite mound applications lies in their natural healing abilities. The clay-like soil is packed with essential minerals like calcium, magnesium, and iron, contributing to skin repair and detoxification. Additionally, termite mounds host beneficial microorganisms that might promote skin microbiome balance, reducing inflammation and infections. Innovation in this practice could involve scientific validation and modern applications of termite mound extracts in dermatology. By isolating bioactive compounds, researchers can develop eco-friendly, sustainable skincare products derived from termite mound materials. This approach could lead to the creation of natural ointments, creams, or masks that harness the mound’s healing potential while ensuring safety and effectiveness. With rising interest in organic and traditional medicine, integrating termite mound-based treatments into modern dermatology could provide a new frontier for sustainable skincare, benefiting communities that have long relied on these natural remedies.
Gram-positive bacteria having a high genomic DNA content of guanine and cytosine (G+C) are known as actinobacteria. They have been observed in a range of ecological contexts. Their component species have been identified as the principal source of several bioactive chemicals, accounting for 70% of all antibiotics found to date.1 Roughly 25% of these were found to be non-Streptomyces species, and the remaining 75% were found to be Streptomyces.2
Each genome of Streptomyces has many biosynthetic gene clusters (BGCs), which are the source of various bioactive substances used in medicine or agriculture.3 The phylum Bacteria has 30 main groupings, of which this is one of the biggest. This phylum has 374 genera, the biggest of which is Streptomyces, 63 families, 18 orders, 14 suborders, and six classes.4 Streptomyces is a Gram-positive bacterium that shares physiological traits with certain fungal species and a GC content between 69% and 78% with other Actinobacteria genera.5
The Streptomycetaceae family and the Streptomycetales order encompass them. Actinobacteria have produced approximately 39% of distinct natural chemicals, with the Streptomyces genus alone contributing over 80%. Many novel Actinobacteria have been identified due to the theory that strains from previously unidentified sources may be great targets for the discovery of bioactive chemicals.6 The objective is to investigate the diversity of actinobacteria in termite nests and explore their bioactive potential as antimicrobial bioactive compound.
Termites are a kind of social insect in the Isoptera order, and are very similar to cockroaches. They inhabit vast colonies with a ruler, their queen, and workers, all of whom have distinct behaviours and functions. Soil termites assist to circulate nutrients in the soil by digesting dead plant materials. Termite populations are generally located in a protective building termed a “termite nest”.7 The subterranean variety is typically found underground at depths ranging from 50 to 100 cm.8
Termite mound soil have rich amount of clay minerals such as kaolinite, smectite and montmorillonite. These minerals may help to eliminate toxins and skin conditions, speed up healing. Termite mound soil has a diverse microbiological life, including bacteria and fungi. Some of these microorganisms can produce bioactive compounds with antibacterial, anti-inflammatory or immunomodulatory properties. The organic matter in termite mound soil may help to improve its therapeutic effects by delivering nutrients and promoting a healthy microbial ecology. This organic compound may include bioactive molecules with therapeutic potential. Alkalinity: Termite mound soil is typically alkaline, which could explain its antibacterial and wound-healing properties. Alkaline environments can inhibit the growth of certain bacteria and fungus, resulting in fewer diseases.9
The excessive use of antibiotics has resulted in growing antimicrobial resistance.10 This greatly increases the demand for new drug and antibiotic research and development. Natural chemicals obtained from microbes, synthetic and semi-synthetic derivatives equivalents have traditionally been crucial sources of antibiotics.11 Current investigations have increasingly directed their focus toward extreme environments for having competitive conditions that promote the production of novel secondary metabolites as a defense or/and signaling strategy.12 Also, the enzymes and products formed under these challenging conditions are more physically and chemically stable when facing high pressure and temperature.13,14 These metabolites encompass a diverse array of compounds, including antimicrobial, antifungal, antitumor, antivirals, and antioxidants critical in modern medicine.15,16 The aim of this research is to find the diversity of halophilic actinobacteria in termite mound soil which is obtained from various areas. It also includes the study of investigation in biotechnological capabilities of actinobacteria which is related to antibacterial, anti- tumour and anti-inflammatory properties when the bioactive compound is tested against the specific pathogens.
Halophilic actinobacteria, thriving in high-salinity environments such as salt lakes and marine or saline soils, are prolific producers of enzymes, antibiotics, and other bioactive secondary metabolites. These unique organisms synthesize compounds rarely found in non-halophilic bacteria, many of which display potent antibacterial properties effective against drug-resistant pathogens. Their adaptation to extreme salinity drives the evolution of specialized metabolic pathways that enable the production of antimicrobial and stress-tolerant molecules, making them promising candidates for broad-spectrum antibiotic development.
This study aimed to isolate and characterize halophilic actinobacteria from termite mounds in Virudhunagar through morphological, biochemical, and Gram-staining analyses. The isolates were screened for antimicrobial activity against Klebsiella pneumoniae, Staphylococcus aureus, and Candida albicans. Secondary metabolites were purified and characterized using TLC and GC-MS, and the most potent isolate (N3) was identified via 16S rRNA sequencing. The study further explores the mechanisms and therapeutic potential of the identified bioactive compounds as antimicrobial, antitumor, and anti-inflammatory agents.
Collection of termite mound samples
Samples of termite mounds were obtained from three different sites in Nathampaati labelled as (N), Pudhupatti (P), and K. Pudur labelled as (S). Each termite nest sample (200 g per sample from each location) was packaged in plastic bags and stored in an icebox for up to 24 hours before being moved to the laboratory. The samples were ground into fine particles. The dried materials were mixed in sterile distilled water and thoroughly combined before being assessed using a pH meter.
Media preparation
To prepare the media, the following ingredients were used: Glucose – 1.8 g, KCl – 1.2 g, MgSO4 – 50 mg, CaCl2 – 78 mg, CaCo3 – 50 mg, NaCl – 12 g, KH2PO4 – 20 mg, LB broth – 2 g, and Agar – 1.8 g for every 100 ml of distilled water. The amount of each ingredient was measured using a weighing machine. After that, the ingredients were dissolved in purified water. The medium’s pH was brought to 7.2 as needed. After that, agar was added and cooked until it totally dissolved. Once all the components had dissolved, the media was autoclaved. To prevent contamination, autoclave sterilization was carried out at 121 °C for 15 minutes at a pressure of 15 pounds.
Isolation of Halophilic Actinobacteria from Termite Mound
Halophilic Actinobacteria were isolated from a termite mound by diluting dried N, P and S samples with water in a 1:1 ratio. The samples were then pretreated using a dry bath at 55 °C for 10 minutes. Subsequently, the samples were spread onto LB broth with 12% NaCl media. For twenty-four hours, all plates were incubated at 55 °C. The individual colonies that were produced were then adhered to a different plate.
Colony morphology and storage
Colonial morphology is utilized as the initial step in identifying colonies, with emphasis on size, colour, and Gram’s staining. Glycerol stock is employed for the extended preservation of isolates at -86 °C to facilitate future studies.
SEM analysis of cell morphology
SEM study of cell cultures entails comparing the surface morphologies of normal and malignant cells to light microscopy images of the same cells in their live condition. Different research groups have developed unique sample preparation protocols to visualize specific biological samples with the required resolution. Most SEM protocols include fixation steps with paraformaldehyde and/or glutaraldehyde, followed by osmium tetroxide. Specimens are then dehydrated with ethanol or acetone, dried using critical point drying, and coated with metal. These protocols can achieve a resolution of 5 nm, sufficient for investigating a cell’s ultrastructure. However, most morphometric descriptors discussed in this review paper do not necessitate such high resolution.
Biochemical tests
Traditional tests include simple biochemical tests like the Indole Test, Methyl Red Test, Voges-Proskauer Test, Citrate Utilisation Test, Catalase Test, Starch Hydrolysis, and Carbohydrate Fermentation, as well as staining and microscopy methods like gram staining, endospore staining, and Grams staining.48
Methods for identification of isolates by 16S rRNA analysis
Isolates were cultured on suitable growth media under optimal conditions for halophilic actinobacteria. Pure cultures were obtained through repeated streaking. To ensure high-quality DNA, genomic DNA was extracted using a commercial DNA extraction kit or a conventional bacterial DNA extraction methodology, such as the CTAB method. Universal primers, such as 27F: 5′-AGAGTTTGATCMTGGCTCAG-3′ and 1492R: 5′-TACGGYTACCTTGTTACGACTT-3′, were used to amplify the 16S rRNA gene by PCR. PCR conditions comprised a 5-minute initial denaturation at 94 °C. 30 cycles of 94 °C (30 seconds) denaturation, 55 °C (30 seconds) annealing, and 72 °C (1 minute) extension. Final extension: 10 minutes at 72 °C. Electrophoresis on a 1% agarose gel stained with ethidium bromide and seen under UV light was used to validate the amplified products. A gel extraction or PCR cleaning kit was used to purify the PCR results, and an automated DNA sequencer was used to sequence them. The 16S rRNA sequences were aligned and compared with reference sequences in the NCBI GenBank database using BLAST to ascertain sequence similarity utilizing further sequence alignment and BLAST analysis. Phylogenetic trees were constructed using software like MEGA (Molecular Evolutionary Genetics Analysis) to analyse evolutionary relationships. Isolates were identified based on sequence similarity (>97%) and clustering with known species in phylogenetic trees.
Extraction of secondary metabolite
The 20 isolates were chosen to extract Secondary Metabolite based on the antimicrobial study. 20 ml of LB broth with 12% NaCl was prepared, and the isolates were then inoculated into individual tubes. The culture broths of these isolates were centrifuged at 6000 rpm for 10 minutes to obtain cell-free supernatant. The cell-free supernatant was collected, and a solvent mixture of ethyl acetate, toluene and acetone was added at a ratio of 7:1:3. After thorough mixing, the solution was stored at -20 °C for 24 hours. Following this period, the solution was separated into two layers, with the upper layer being carefully transferred into separate tubes. Finally, the extract was air-dried.
Thin layer chromatography
The chemicals included in the isolates’ extract were visualised using thin layer chromatography. Silicone gel (20 × 20 cm, 0.25 mm Alugram® SIL G/UV 254, Macherey and Nagel, Duren) and mobile phases [hexane:toluene:acetone] in a 7:1:3 ratio were used to perform the chromatography on aluminium plates. Under a UV lamp, the chemicals were seen.
Antibacterial studies on microbial bioactive compound
The work area was disinfected using a disinfectant and an open burner. A total of 20 microbial bioactive compounds were tested against K. pneumoniae MTCC 39 using the disc diffusion method. Plates of Meat Extract agar were prepared. A sterile cotton swab was immersed in the K. pneumoniae MTCC 39, and excess liquid was removed by pressing the swab against the tube wall. The agar plate was rotated in order to swab the whole surface. On the agar surface, the inoculum was dispersed uniformly. To ensure proper absorption of the inoculum by the medium, the plates were allowed to dry for approximately five minutes. Seven microliters of the bioactive substance were put onto a sterile disc that was set on the agar surface. This step needed to be completed within 15 minutes of inoculation. Each disc was gently pressed onto the agar to ensure proper attachment. After then, the plates were typically incubated at 37 °C for the whole night. After incubation, the antibiotic discs’ surrounding zones of inhibition were examined, and each zone’s diameter was quantified.47
Bioautography studies on bioactive compound
Ten metabolites were chosen based on the disc diffusion method. Subsequently, 10 µl of each metabolite was applied to TLC plates and incubated at 37 °C for one to two hours. Following this, Petri dish was placed in an airtight container, and water was added to the dish to maintain a moist environment. The TLC plates were then transferred to the Petri dish using forceps, and the bacterial culture of S. aureus MTCC 39 was spread on the plate. We observed the zones of inhibition surrounding each metabolite and measured the width of each zone after a 24-hour incubation period at 37 °C. The disc diffusion method was used for the confirmation, and N3 was compared. Additionally, the technology of Gas Chromatography-Mass Spectrometer (GC-MS) was used to identify the bioactive chemicals.17
Antibiotic sensitivity test
The isolates were chosen based on their activity and subsequently subjected to an antibacterial susceptibility test. A total of 10 isolates were tested against various antibiotics using the Kirby-Bauer disk-diffusion method. The antibiotics were prepared at a concentration of 100 mg/ml, with 50 µl of each antibiotic added to the media and poured onto the plates. The bacterial isolates were then patched onto the plates. Following a 24-hour incubation period, the results were examined. Similarly, 100 µl of antibiotic was added in the subsequent steps. The antibiotic disks used included Tetracycline, Erythromycin, Streptomycin, Penicillin, Ampicillin, Norfloxacin, Gentamicin, Amoxicillin, Amikacin, and Levofloxacin. After being left to dry at room temperature, the plates were incubated for 18 to 24 hours at 37 °C. The data were evaluated for sensitivity or resistance following the incubation period.
MIC (Minimum Inhibitory Concentration)
The minimum amount of an antimicrobial substance that, upon incubation, prevents a bacterium from growing visibly. A series of test tubes or wells with varying concentrations of the antimicrobial agent. Bacteria are added, and the tubes are incubated 37 °C. The MIC is the concentration in the first tube where no visible growth (turbidity) is observed.
MBC (Minimum Bactericidal Concentration)
An antimicrobial substance kills 99.9% of the bacteria at its lowest concentration. Identifies whether the antimicrobial is bactericidal (kills bacteria) or bacteriostatic (inhibits growth but does not kill). Samples from tubes that show no signs of growth are plated on an agar medium once the MIC has been established. The lowest concentration that prevents colony formation on the plate is the MBC.
Differences Between MIC and MBC
MIC focuses on stopping bacterial growth, while MBC confirms bacterial death. For bactericidal agents, the MIC and MBC values are typically close, while for bacteriostatic agents, MBC is much higher than MIC.
Impact
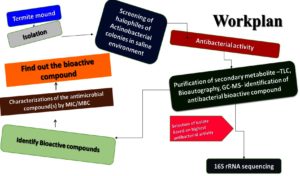
Termites can thrive in a wide range of environments, spanning from dense forests to urban settings and across the diverse ecosystems of South India. These highly organized insects, belonging to the Blattodea order, are critical in maintaining the ecosystem by decomposing organic matter and shaping the environment. Their population growth is aided by the warm and humid climate of South India, where they contribute to soil fertility, ecosystem stability, and nutrient circulation. The saliva of termites at the heart of their colonies is a remarkable example of biological adaptability. Despite its seemingly insignificant nature, termite saliva contains a wealth of proteins, enzymes, and other bioactive compounds that support the colony’s activities and impact its interactions with the surrounding environment. Research on termite saliva sheds light on a complex web of ecological relationships and potential applications, and termites play essential roles in natural and human-influenced ecosystems for experimental design in workflow studies (Figure 1). Our research highlights the potential for New Drug Development, particularly concerning the N3 isolate derived from bioactive compounds, which includes antitumor, antimicrobial, and anti-inflammatory properties, as well as the pressing challenge of antibiotic resistance. This emphasizes the urgent necessity to identify new antibiotics. The development of new antibacterial remedies for work plans may benefit from the unique bioactive chemicals found in halophilic actinobacteria (Figure 1). These microorganisms flourish in saline environments, which remain largely untapped in both pharmaceutical and industrial sectors. Their unique characteristics make them a sustainable choice for antibiotic production, featuring distinctive metabolites, resilience to harsh conditions, a broad spectrum of bioactivity, and the promise of innovative drug discovery, thus alleviating competition for resources essential for food production.
Figure 1. Shows the antimicrobial bioactive compound from spore halophilic actinobacterial isolate of the Termite mound and screening the experimental design of the workflow chart
Isolation and collection of halophilic actinobacteria from termite mound
The termite mound samples were collected from three specific locations: Nathampatti (N), Pudhupatti (P), and K. Pudur (S) in the Virudhunagar district (Figures 2a-c). Soil sampling distances in the surrounding areas were chosen to evaluate the variation in soil nutrients as the distance from the termite mounds increased. A total of 20 isolates from various actinobacteria were gathered from termites in Nathampatti (N), Pudhupatti (P), and K. Pudur (S) regions for examination (Figure 2). The culture medium used for bacterial growth (LB with 12% NaCl) was controlled to confirm no interference with the tested compounds produced by the actinobacteria.
Figure 2. Shows different locations of Termite mound from Virudhunagar district (A) Termite mound soil collected from Nathampatti, (B) Termite mound soil collected from Pudhupatti, (C) Termite mound soil collected from K. Pudur
Media preparation and isolation of halophilic actinobacterial colonies
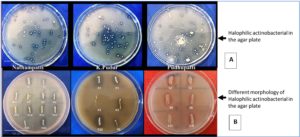
Actinobacterial isolates were typically grown on YMA medium, but for this study, a modified LB medium was used with the following composition: KCl (1.2%), NaCl (12%), starch (1%), glucose (1.8%), and oatmeal milk (0.5%). The isolates were characterized both macroscopically and microscopically. Macroscopic characterization involved observations of colony shape, including elevation, pigmentation, and the colour of the aerial and substrate mycelium. Using a light microscope set to 1000x magnification, Grams staining was used for microscopic characterization in order to view the mycelium’s structure and form. These methods were employed to identify variations in cultural characteristics. Based on colony morphology and color, halophilic bacterial organisms were selected and transferred to separate plates (Figure 3). The authors isolated three strains: N1-34 from Nathampatti, P1-10 from Pudhupatti (Virudhunagar) and S1-8 from K. Pudur.
Figure 3. (A) Halophilic colonies on the modified agar plate in different locations of Virudhunagar district, (B) screening the different morphology of halophilic colonies on the patch with the 12% NaCl with agar plate
Morphological characterization
Actinobacterial colonies with comparatively different colours were rough, starchy, and irregularly shaped, and they were cultivated on 12% NaCl agar media. Isolates based on the antimicrobial study; 24 colonies were selected. The isolates from Nathampatti are N2, N3, N4, N5, N7, N9, N10, N11, N16, N25, N26, and N28. The isolates from K. Pudur are S1, S2, S3, S6, S11, and S21. The isolates from Pudhupatti are P1, P2, P3, P5, P6, and P10. Li et al.5 have classified the colony features of actinobacteria into two types. The first group may generate aerial mycelium and has a velvety, colourful colony surface. The colony surface of the second group was colourless and smooth, and they did not generate aerial mycelium. Among the colonies, zone creation was chosen for more research.
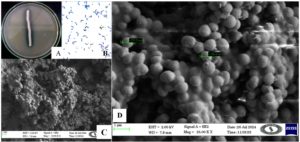
Characterization of colony morphology of halophilic actinobacterial isolates N3
Milky and sticky white halophilic colonies were observed (Figures 4a-d). In microscopic observation, gram-positive small cocci are joined together and look like a rod. SEM analysis is similar to the microscopic observation that cocci-shaped bacteria are joined together to form rod-shaped. Actinobacteria exhibit phenotypic characteristics that are crucial for describing a genus or species. Actinobacteria physiological and biochemical assays require careful consideration of a number of factors. These include selecting strains from closely related taxa and the type strain of the appropriate genera for comparison based on phylogenic information from 16S rRNA analyses and Indole, MR activity, VP broth study, citrate utilization, catalase, and different sugar, such as sucrose, fructose, lactose, starch hydrolysis (Figures 5a-b, Table 1). Methodology is also a concern, and it is advisable to use established approaches and relevant tests. Evidence of their reliability compared to established methods must be provided if new methods are employed.
Table (1):
Studies on biochemical test and different sugar hydrolysis of the different actinobacterial isolates from Termite mound
No. |
Isolates |
Indole |
MR |
VP |
Citrate utilization Test |
Catalase |
Sucrose |
Fructose |
Lactose |
Starch hydrolysis |
|---|---|---|---|---|---|---|---|---|---|---|
1 |
N2 |
– |
+ |
– |
– |
+++ |
++ |
++ |
– |
– |
2 |
N3 |
– |
– |
– |
+ |
+++ |
– |
++ |
– |
+ |
3 |
N6 |
– |
+ |
– |
– |
+++ |
+ |
++ |
– |
– |
4 |
N7 |
– |
+ |
– |
+ |
+++ |
++ |
+++ |
– |
– |
5 |
N11 |
– |
+ |
– |
– |
+++ |
++ |
+++ |
– |
– |
6 |
P1 |
– |
+ |
– |
– |
+++ |
+ |
++ |
– |
– |
7 |
P2 |
– |
+ |
– |
– |
+ |
– |
++ |
– |
– |
8 |
S3 |
– |
+ |
– |
– |
++ |
++ |
+++ |
– |
– |
9 |
S6 |
– |
+ |
– |
– |
++ |
++ |
+++ |
– |
– |
– sign indicate negative response, + indicates weak positive activity, ++ indicates moderate positive activity, and +++ indicates strong positive activity
Figure 4. (A) Colony morphology of Actinobacterial isolate N3 (Microbacterium barkeri M17) in 12% agar plate with clear halophilic zone formed, (B) Gram’s staining of N3 600x view, (C) SEM images of N3 Actinobacterial isolate for 1 µm size of the cells, and (D) SEM images of N3 Actinobacterial isolate for 1 µm and measure single cell size =738.4 nm
Also, testing should be performed in duplicate or triplicate since the phenotypic characteristics of actinobacteria might be influenced by culture conditions and other variables. It is also crucial that the trials include appropriate positive and negative controls. It is impossible to draw similar conclusions about the Actinobacterial phenotype from only seeing the organism. In their phenotypic examination, strains of the most closely related taxa have to be compared using the same techniques. The type strain of the relevant genus’ type species must be included in the comparisons.49
16S rRNA sequence analysis
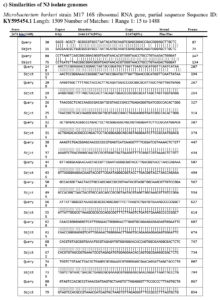
The actinobacterial strains exhibiting the most significant biological activities underwent additional characterization utilizing molecular features. Following the 16S rRNA gene sequencing, the acquired sequences were deposited into the NCBI Gene Bank Nucleotide Database (Figure 5 a-c). Subsequent analysis of the sequences through NCBI’s BLASTN algorithm revealed that the strains were affiliated with two distinct bacterial species. The high degree of sequence similarity observed with a specific bacterial species already present in the Gene Bank led to this conclusion. Specifically, Isolate N3 displayed sequence similarity with the Microbacterium barkeri strain.
Figure 5. (a) Studies on N3 isolate (Microbacterium barkeri M17) for 16S rRNA gene sequencing, deposited into the NCBI GenBank Nucleotide Database, and (b) Phylogenetic tree of molecularly characterized isolates, (c) Identification and Sequences of gene identification
Antibacterial activity of cell free extract
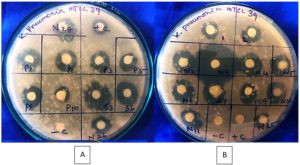
Antibacterial metabolites were extracted thrice from the cell-free supernatant using various solvents, among which ethyl acetate proved most effective. The concentrated extract formed a greenish-yellow residue, later diluted with methanol for testing. Sterilized discs containing 10 µL of the extract were placed on Klebsiella pneumoniae MTCC 39–inoculated LB agar plates. After 24 hours of incubation, notable inhibition zones were observed (Figures 6a,b), confirming the strong antibacterial activity of the N3 isolate. The antibacterial assay (Table 2) includes mean inhibition zones ± standard deviations, calculated from three independent replicates (n = 3). Among the isolates, N3 exhibited the largest and most consistent inhibition zone (0.71 ± 0.02 cm), comparable to the positive control streptomycin (0.43 ± 0.02 cm). This indicates the reproducible and potent antibacterial activity of the N3 isolate’s secondary metabolites against test pathogens. The bioactive compounds effectively inhibited pathogenic bacteria, suggesting their potential in developing new antimicrobial agents. Overall, halophilic actinobacteria from termite mounds demonstrated significant antibacterial potential, underscoring their value as promising sources of novel bioactive metabolites to combat antibiotic resistance.
Table (2):
Selection of 10 different actinobacterial isolates from 20 isolates and their antibacterial efficiency of different isolates from Termite mound against Klebsiella pneumoniae MTCC 39 (106 cells/mL)
| No. | Name of the actinobacterial isolate | The antibacterial activity of the inhibition in cm. | ||
|---|---|---|---|---|
| Before incubation | After incubation | Results | ||
| 1 | Control (Ethyl acetate alone) | Nil | Nil | Nil |
| 2 | Positive control1 mg/L Streptomycin | 0.192 | 0.43 ± 0.02 | ++ |
| 3 | P3 (Positive control) | 0.184 | 0.71 ± 0.03 | +++ |
| 4 | N2 | 0.225 | 0.27 ± 0.01 | + |
| 5 | N3 | 0.223 | 0.71 ± 0.02 | +++ |
| 6 | N6 | 0.232 | 0.24 ± 0.01 | + |
| 7 | N7 | 0.215 | 0.23 ± 0.02 | + |
| 8 | N11 | 0.259 | 0.29 ± 0.02 | + |
| 9 | N26 | 0.256 | 0.31 ± 0.02 | + |
| 10 | P1 | 0.241 | 0.31 ± 0.01 | + |
| 11 | P2 | 0.241 | 0.28 ± 0.01 | + |
| 12 | S3 | 0.184 | 0.27 ± 0.01 | + |
| 13 | S6 | 0.185 | 0.29 ± 0.02 | + |
| 14 | Termite mound soil | 0.188 | 0.40 ± 0.02 | ++ |
– sign indicate negative response, + indicates weak positive activity, ++ indicates moderate positive activity, and +++ indicates strong positive activity
Figure 6. (A) Shown antibacterial activity studies on Nathampatti halophilic actinobacterial isolates from Termite mound against Klebsiella pneumoniae MTCC 39 (10⁶ cells/mL). (B) Pudhupatti and K. Pudhur halophilic actinobacterial isolates from Termite mound against Klebsiella pneumoniae MTCC 39 (10⁶ cells/mL)
Mechanism of action
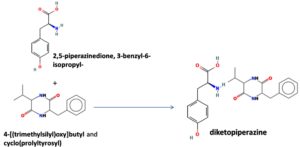
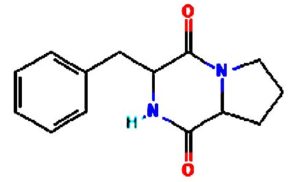
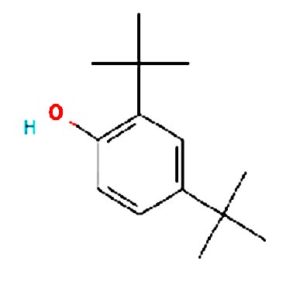
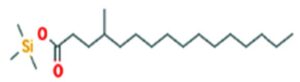
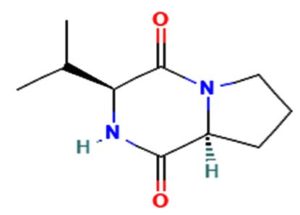
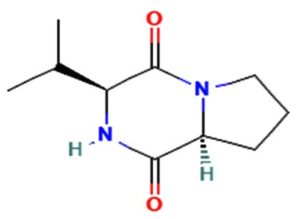
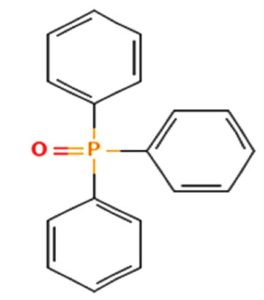
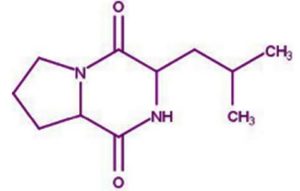
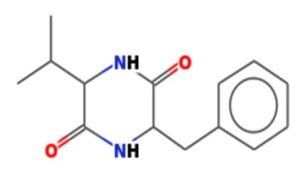
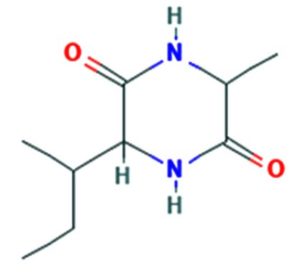
The merging of compounds such as 4-[(trimethylsilyl)oxy] butyl and cyclo(prolyltyrosyl) with 3-benzyl-6-isopropyl- 2,5-piperazinedione to create a new compound involves chemical synthesis. Interestingly, the known compounds above were functional group of analysis in the form of 4-[(trimethylsilyl)oxy] butyl contains a silyl ether with a hydroxyl functionality, which can act as a nucleophile after deprotection. A cyclodipeptide structure containing amino acid residues with reactive groups such as hydroxyl (-OH) in tyrosine and possibly amine (-NH2) or carboxyl (-COOH) groups at termini. 2,5-Piperazinedione, 3-benzyl-6-isopropyl-: A diketopiperazine (DKP) structure with potential reactive sites at amide or carbonyl positions. And then potential reaction pathways were involved in the hydroxyl group of 4-[(trimethylsilyl)oxy] butyl can react with the carbonyl group of the diketopiperazine or the carboxyl group in cyclo(prolyl tyrosyl). A coupling reagent like EDCI or DCC could facilitate the linkage between amino (-NH2) or carboxyl groups of the peptide and the diketopiperazine.
The biological properties of the merged compound depend on the resultant structure and functional groups. The Enzyme Inhibition of the diketopiperazine moiety is known to inhibit proteases by mimicking peptide substrates. If the new compound forms a cyclic structure, it might interact with enzymes that recognize similar cyclic peptides, blocking their active sites. Amphiphilic properties from the trimethylsilyl or diketopiperazine groups may allow the compound to disrupt bacterial or fungal membranes. And compounds with diketopiperazine or tyrosine-like residues can generate reactive oxygen species (ROS), leading to oxidative stress in microbial cells. Tyrosine-containing peptides can interfere with kinases or phosphatases in signaling pathways, potentially modulating cell growth or immune responses. DNA or RNA Binding the cyclic and aromatic nature of diketopiperazine and peptide residues may facilitate binding to nucleic acids, affecting transcription or translation (Figure 7).
Figure 7. Defense mechanism of action for two chemical and formation of diketopiperazine and antibiotic function
The compound Pyrrolo[1,2-a] pyrazine-1,4-dione, hexahydro-3- is a cyclic dipeptide and does not fall under the category of β-lactam antibiotics. Instead, it is a non-β-lactam compound. The structure of this compound is derived from diketopiperazines (cyclic dipeptides), characterized by a six-membered ring structure that includes two amide bonds. Unlike β-lactams, which have a four-membered lactam ring, this structure does not contain such a feature. The activity of cyclic dipeptides, including this compound, is known for diverse biological properties such as antibacterial, antifungal, anticancer, and anti-inflammatory activities. It is do not typically target bacterial cell wall synthesis like β-lactam antibiotics (e.g., penicillin or cephalosporins). However, the Pyrrolo[1,2-a] pyrazine-1,4-dione, hexahydro-3- is a non-β-lactam compound with potential bioactive properties.
Commercial Antibiotic analysis of 10 different actinobacterial isolates of antibiotic sensitive analysis
The efficacy of the ten chosen actinobacterial isolates varied regarding their antibacterial properties, as demonstrated by the formation of inhibition zones (Table 3). The isolates examined in this study are N2, N3, N6, N7, N11, N26, P1, P2, S3, and S6. Antibiotic susceptibility was assessed on these selected isolates for further investigation. Among the ten isolates, only one actinobacterial isolate, N3, exhibited inhibition of 2.078 cm, displaying not only inhibitory effects but also 90% sensitivity to all synthetic antibiotic compounds, except Amoxicillin, which displayed resistance (Table 3). The term “sensitivity” refers to the ability of the cultured bacterial pathogen’s growth to be hindered, or not, by exposure to specific antibiotics. The outcomes of the antibiotic sensitivity test are classified as susceptible, intermediate, and resistant. Sensitivity analysis, also known as susceptibility testing, helps doctors choose the best antibiotic to eradicate an infectious microbe. Microorganisms that infiltrate the body and produce an illness include bacteria and fungus. Sensitivity analysis is a technique used to determine how sensitive bacteria are to antibiotics. It also assesses how well the medication gets rid of the germs. The test results can be used to determine which medications have the best chance of curing the illness.
Table (3):
Selected antibacterial and their efficiency of different antibiotic sensitivity assays
| Name of the Antibiotics | Name of the Actinobacterial isolates | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| N2 | N3 | N6 | N7 | N11 | N26 | P1 | P2 | S3 | S6 | |
| Tetracycline | S | S | S | R+ | S | S | S | R+ | R+ | R++ |
| Ampicillin | S | S | S | S | S | S | S | S | S | S |
| Streptomycin | S | S | R++ | S | S | S | R++ | S | S | S |
| Gentamycin | S | S | R++ | R+ | S | S | R++ | R++ | R+ | S |
| Penicillin | R++ | S | S | R++ | S | R+++ | R++ | R++ | R+++ | R++ |
| Norfloxacin | R++ | S | S | R++ | S | R+++ | R++ | R++ | R+++ | R++ |
| Amoxicillin | R++ | R+ | S | R++ | R+++ | R+++ | R++ | S | R+++ | R++ |
| Amikacin | R++ | S | S | R+ | S | S | R++ | R+ | R++ | R+ |
| Erythromycin | S | S | R++ | S | S | S | S | S | R++ | S |
| Levofloxacin | S | S | S | S | S | S | S | S | S | S |
S indicates sensitivity; R+, low levels of resistance, R++, indicate moderate resistance, R+++ indicate high resistance
Thin layer chromatography on a microbial bioactive compound by screening of antimicrobial-producing actinobacteria
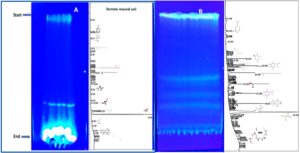
The study of antibacterial activity from halophilic actinobacteria isolated from termite mounds aims to explore their potential as sources of novel bioactive compounds. Twenty different actinobacterial isolates were screened, and bioactive compounds were extracted using solvent extraction. The crude extracts were dried at room temperature and further analysed. TLC method was employed to separate and identify active compounds. These isolates displayed significant antibacterial activity, demonstrating their potential to combat pathogenic bacteria. Termite mounds provide a unique microenvironment fostering diverse and resilient actinobacteria, making them valuable in the search for new antimicrobial agents to address antibiotic resistance challenges.
The ethyl acetate, toluene, and acetone crude extracts were separated using TLC plates for strains N6, N26, N5, P1, and P6, resulting in 2 fractions. Strain N9 yielded 5 fractions, while N-10,3,4,11,7,28,25, 2; P-3,5, and P-2,10; S-6 yielded 3 fractions. However, S3 & N4 yielded no fraction (Figure 8). In order to separate or purify combinations of chemical and biological substances, bioactive compounds are frequently screened using TLC method. Its simplicity, economy, ease of use, and minimal solvent requirements make it useful. Because it is carried out under ambient settings, it is also regarded as the best separation technique for high throughput analysis.18 Both medium polar and non-polar bands were produced by the TLC screening of the bioactive chemical pattern arrangement library from the extracellular extracts of the twenty actinobacterial cultures.
Figure 8. Thin layer chromatography on a cell free extract from 20 different halophilic actinobacterial isolates from microbial bioactive compounds pattern in TLC plate
Studies on Bioautography of 10 different halophilic actinobacterial isolates
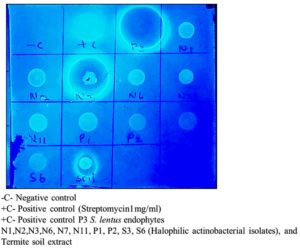
TLC bioautography is a method used for drug screening that involves combining TLC separation with the assessment of biological activity (Figure 9). Research has shown that this technique can confirm the presence of active natural products, including antibacterial, antifungal, antitumor, antioxidant compounds, and enzyme inhibitors. Bioautography is a process that allows for the isolation of potential hits or leads through the use of a suitable chromatographic method followed by a biological detection system. In this study, various actinobacterial secondary metabolites were analysed alongside Termite mound metabolite. Following the application of the bacterial culture S. aureus MTCC 96 on the plate and subsequent incubation, positive results were obtained from the positive control (+C), and Figure 8 have clear zones were observed from P3, N3, and also Termite mound soil. However, in other isolates, very little inhibition was found in the plate, while inhibition was not observed in 100% of the cases.
Figure 9. Studies on Bioautography of 10 different halophilic actinobacterial isolates with termite mound soil extract
GC-MS analysis of N3 isolate (Microbacterium barkeri M17), and their bioactive compounds and function
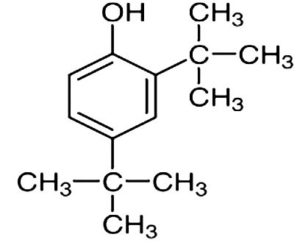
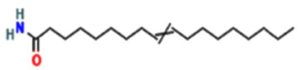
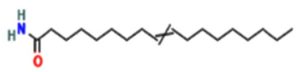
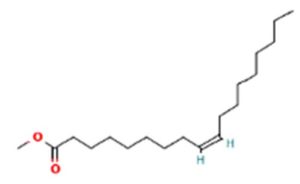
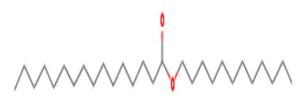
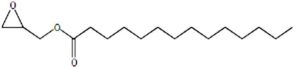
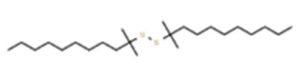
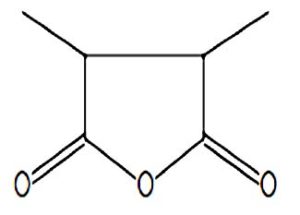
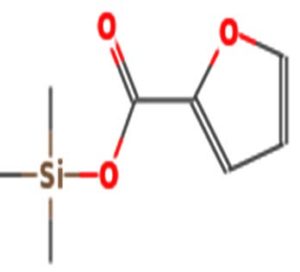
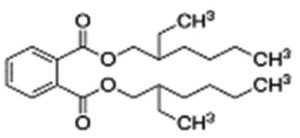
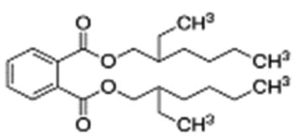
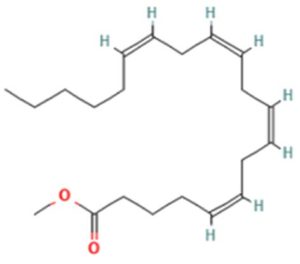
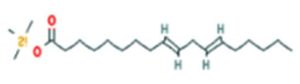
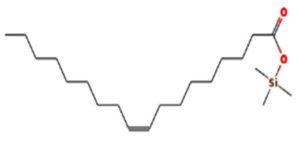
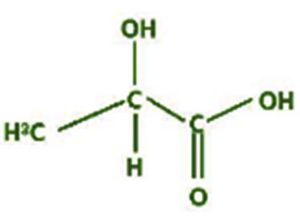
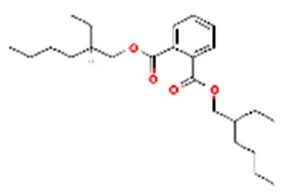
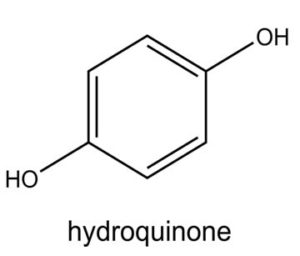
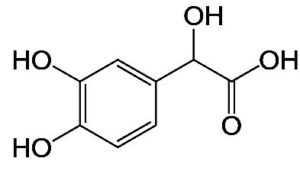
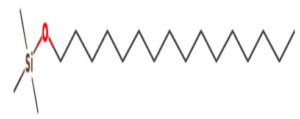
A profile of the fundamentals and significance of Thin Layer Chromatography (TLC) in research in general, and termite mound and N3 (Microbacterium barkeri M17) Actinobacteria isolate in particular, was attempted in this work (Figure 9). For many years, ordinary chemistry labs have employed thin layer chromatography, a straightforward, affordable, and user-friendly planar chromatographic method, to regularly separate chemical and biological substances. Analyte spots on the TLC plate are often visualized using chemical and optical techniques. Additionally, it may be used widely to detect contaminants in a chemical. The evaluation of TLC and its use in estimating bioactive chemicals from termite mounds and N3 (Microbacterium barkeri M17) Actinobacteria isolates is highlighted in the study (Figure 10). The authors have to find an important bioactive compound in the Termite mound, namely (Nattampatti termite mound and N3 secondary metabolite, and the comparison of bioactive compounds found in the library. However, it is crucial to assess the bioactive characteristics of various compounds, including antitumor compounds such as Pyrrolo[1,2-a] pyrazine-1,4-dione, hexahydro-3-, Cyclo (L-prolyl-L-valine), Triphenylphosphine oxide, Methyl gamma-linolenate, Elaidic acid-TMS, and Cyclo (alanylleucyl). Additionally, antimicrobial compounds like 2,4-Di-tert-butylphenol, Dibutyl phthalate, Palmitic Acid-TMS, Hexadecanoic acid, 4-[(trimethylsilyl)oxy]butyl, Cyclo (prolyl-tyrosyl), Methyl oleate, Linoleic acid-TMS, Methyl arachidonate, 2-Furoic acid-TMS, and 2,5-Piperazinedione, 3-benzyl-6-isopropyl- need to be examined. Furthermore, the anti-inflammatory compounds 9-Octadecenamide and Acetamide, N-(2-methoxy-5-methylphenyl) should also be evaluated (Table 4).
Table (4):
GC-MS analysis of N3 isolates (Microbacterium barkeri M17), and their bioactive compounds and compare the Termite mound soil extract name, structure and function
Figure 10. (A) TLC profile of bioactive compounds extracted from termite mound soil, showing distinct separation patterns under solvent system used. (B) TLC profile of crude bioactive compounds obtained from the N3 isolate (Microbacterium barkeri ), indicating the presence of prominent metabolite bands corresponding to antimicrobial activity
Efficacy of MIC and MBC on N3 isolates (Microbacterium barkeri M17)
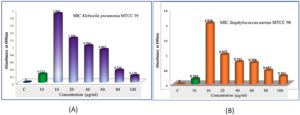
Studying the Minimum Inhibitory Concentration (MIC) and Minimum Bactericidal Concentration (MBC) of bacterial metabolites is critical for understanding their antimicrobial efficacy and potential therapeutic applications. Actinobacteria metabolites for bioactive compounds derived from termite mound Actinomyces species are promising for antimicrobial drug discovery. MIC and MBC studies validate their efficacy. Many bacterial metabolites, such as cyclo(L-prolyl-L-valine) or benzoic acid derivatives, show activity against pathogens like Staphylococcus aureus MTCC96, K. pneumoniae MTCC 39, for MIC/MBC testing bridges the gap between discovery and therapeutic application (Figures 11a-b). Combination Therapies are evaluating MIC and MBC in combination with existing antibiotics can lead to synergistic therapies. Studying MIC and MBC of bacterial metabolites is a cornerstone of antimicrobial research. It provides quantitative insights into their efficacy, helps optimize their use as drugs, and ensures that their therapeutic potential is harnessed effectively. Actinomyces-derived compounds like cyclo(L-prolyl-L-valine) is particularly relevant for discovering new, potent antimicrobial agents.
Figure 11. Minimum inhibitory concentration (MIC) evaluation of crude bioactive compounds from the N3 isolate (Microbacterium barkeri M17).(A) MIC assay against Staphylococcus aureus MTCC 96 at different concentrations of the crude extract.(B) MIC assay against Klebsiella pneumoniae MTCC 39 at varying concentrations of the crude bioactive compounds
Standard Curve and MIC studies of streptomycin on pathogenic organisms
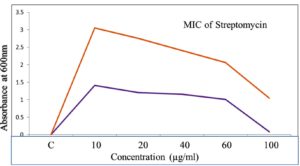
The standard curve of streptomycin was established to evaluate its antimicrobial potency against pathogenic organisms, including Staphylococcus aureus MTCC 96 and Klebsiella pneumoniae MTCC 39. This approach enables the determination of minimum inhibitory concentration (MIC) values, which signify the lowest concentration of streptomycin required to inhibit microbial growth effectively (Figure 12). The MIC studies revealed that streptomycin exhibited significant antibacterial activity against both pathogens, highlighting its efficacy as a therapeutic agent. S. aureus MTCC 96, a Gram-positive bacterium, and K. pneumoniae MTCC 39, a Gram-negative pathogen, are commonly associated with severe infections and demonstrate varying susceptibility profiles. Quantitative analysis based on the standard curve provided a precise correlation between streptomycin concentration and microbial growth inhibition. These findings underscore the importance of MIC studies in assessing antibiotic effectiveness, optimizing dosage regimens, and addressing resistance concerns in clinical settings.
Figure 12. Studies on standard curve of Streptomycin shows that MIC studies on pathogenic organism of Staphylococcus aureus MTCC96, K. pneumoniae MTCC 39
TLC Patterns and MIC of Microbacterium barkeri M17 Against Pathogenic Organisms
Studies on the TLC patterns and MIC of the Microbacterium barkeri M17 isolate (N3) revealed significant antimicrobial activity against three pathogenic organisms: Staphylococcus aureus MTCC 96, Klebsiella pneumoniae MTCC 39, and C. albicans MTCC 282. A total of eight TLC patterns were analysed, demonstrating diverse bioactive compounds. Among these, the 4th and 5th patterns exhibited prominent zones of inhibition against all tested pathogens, indicating the presence of potent antimicrobial metabolites. The zone of inhibition was measured and compared with positive controls, streptomycin (10 mg/mL) and ciprofloxacin (10 mg/mL), which served as benchmarks for evaluating the effectiveness of the bioactive compounds (Table 5). The 4th and 5th TLC patterns consistently produced inhibition zones comparable to or exceeding those of the controls, underscoring the therapeutic potential of the compounds extracted from Microbacterium barkeri. The MIC studies further confirmed the inhibitory effects of these compounds, highlighting their capability to suppress the growth of both Gram-positive and Gram-negative pathogens. The synergistic impact of the identified compounds may offer promising avenues for developing novel antimicrobial agents. These findings emphasize the role of Microbacterium barkeri M17 as a valuable source for bioactive compounds, with the potential to combat multidrug-resistant pathogens effectively.
Table (5):
Studies on the TLC pattern and MIC of N3 isolates (Microbacterium barkeri M17) against three different pathogens
| No. | Experiments | Different pathogens | ||
|---|---|---|---|---|
| S. aureus MTCC 96 | K. pneumoniae MTCC 39 | C. albicans MTCC 282 | ||
| Zone of inhibition (in mm) | ||||
| 1 | Control | 0.0 | 0.0 | 0.0 |
| 2 | Streptomycin (10 mg/ml) | 2.211 ± 0.003 | 1.999 ± 0.002 | 0.985 ± 0.003 |
| 3 | Ciprofloxacin (10 mg/ml) | 2.981 ± 0.004 | 2.000 ± 0.001 | 1.031 ± 0.001 |
| 4 | TLC pattern-1 | 0.001 ± 0.003 | 0.000 ± 0.003 | 0.000 ± 0.003 |
| 5 | TLC pattern-2 | 0.001 ± 0.003 | 0.000 ± 0.003 | 0.000 ± 0.003 |
| 6 | TLC pattern-3 | 0.003 ± 0.003 | 0.000 ± 0.003 | 0.000 ± 0.003 |
| 7 | TLC pattern-4 | 1.111 ± 0.006 | 1.000 ± 0.007 | 0.677 ± 0.001 |
| 8 | TLC pattern-5 | 0.996 ± 0.001 | 0.888 ± 0.002 | 0.502 ± 0.001 |
| 9 | TLC pattern-6 | 0.011 ± 0.003 | 0.010 ± 0.003 | 0.005 ± 0.003 |
| 10 | TLC pattern-7 | 0.000 ± 0.000 | 0.000 ± 0.000 | 0.000 ± 0.000 |
| 11 | TLC pattern-8 | 0.000 ± 0.000 | 0.000 ± 0.000 | 0.000 ± 0.000 |
‘± ‘sign indicate minimum three replicates
TLC Patterns and MIC/MBC Analysis of Microbacterium barkeri M17
The Microbacterium barkeri M17 (N3 isolate) was evaluated for antimicrobial activity through TLC patterns (4th and 5th) and MIC/MBC assays against Staphylococcus aureus MTCC 96 and Klebsiella pneumoniae MTCC 39. The 4th and 5th patterns showed significant zones of inhibition for all tested pathogens, comparable to the positive controls, streptomycin and Ciprofloxacin (1 mg, 5 mg, and 10 mg/mL). MIC studies confirmed the bioactivity of these compounds, while synthetic antibiotics demonstrated negative MBC results. In contrast TLC pattern 4th and 5th also get positively (Table 6). These findings emphasise the therapeutic potential of bioactive chemicals derived from Microbacterium barkeri as antimicrobials against multidrug-resistant bacteria.
Table (6):
Studies on the TLC pattern and MIC/MBC of N3 isolates (Microbacterium barkeri M17)
| Treatment | MIC | MBC | ||||
|---|---|---|---|---|---|---|
| Zone of inhibition in mm | ||||||
| (1 mg/mL) | (5 mg/mL) | (10 mg/mL) | (1 mg/mL) | (5 mg/mL) | (10 mg/mL) | |
| Control | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| Streptomycin | 0.987 | 1.385 | 2.451 | – | – | – |
| Ciprofloxacin | 0.444 | 0.688 | 1.301 | – | – | – |
| TLC pattern-4 | 0.201 | 0.833 | 1.007 | + | + | + |
| TLC pattern-5 | 0.806 | 0.998 | 1.081 | + | ++ | ++ |
– sign indicate negative response; + sign indicate positive response; ++ indicates strong positive activity
Actinobacteria with halophilic characteristics were obtained using LB broth with 12% NaCl media from the termite mound. After screening 30 isolates, one organism was selected for identification. The identification process involved assessing colony color, morphology, biochemical and carbohydrate fermentation tests, gram staining, and SEM analysis. The 16S rRNA gene sequencing technique was used to further characterise the isolate. Twenty of the 26 microorganisms that Hussaini et al.9 recovered were gram-positive, and six were Gram-negative. Bacillus species, Citrobacter freundii, Enterococcus cloacae, Enterococcus faecalis, E. coli, K. pneumoniae, Pseudomonas species, S. aureus, S. lentus, and S. ureae were among the isolates that were detected. The samples were spread on sterile soya medium plates and nutrient agar plates and then incubated at 37 °C for 24 hours. Biochemical assays, colony morphology, and gramme staining were used to identify the bacterium. Devi et al.50 isolated a total of 132 isolates, including 70 bacteria, 58 actinomycetes, and 4 different fungal strains from termite mound soil samples collected from various areas of Himachal Pradesh. The isolation process involved using nutrient agar, potato dextrose agar, and actinomycetes isolation agar.
The microorganisms were distinguished based on their morphology and biochemical features. Similarly, Rajni Devi et al.51 identified Staphylococcus saprophyticus, Bacillus methylotrophicus, and Bacillus sp. through the standard plate count technique (Wollun, 1982)52 using various media like nutrient agar, Jensen’s Agar, and Pikoskaya’s agar. The identification of bacterial isolates was carried out by assessing their morphological and biochemical characteristics following the standard procedure outlined in Bergey’s Manual of Systematic Bacteriology. The efficient isolates underwent molecular characterization through the 16S rRNA gene sequencing method, and the obtained sequences were analysed using the online NCBI Blast Program.
In this current investigation on antibacterial properties, it was found that out of 30 isolates, 20 were able to inhibit K. pneumoniae. The secondary metabolite was obtained through solvent extraction, and subsequent screening revealed that the N3 metabolic compound exhibited significant inhibition on S. pneumoniae using disc diffusion and bioautography techniques. Sawadogo et al.53 conducted a study on the antimicrobial activity of 60 isolates against various pathogenic bacteria and fungi using the paper disc diffusion method. Among the isolates tested, MBm2, MBm8 (bacteria), and MBm26 (actinobacterium) displayed superior antibacterial and antifungal activity against all pathogens. Krishanti et al.54 reported that dual culture methods assessed antibacterial and antifungal activity on ISP2 agar plates. Initial antibacterial screening revealed that six isolates inhibited bacterial infections, whereas four isolates had antifungal activity against three fungal species.
The Actinomycetes isolate exhibited the highest inhibitory activity against E. coli, indicating its potential to inhibit the growth of Gram-negative bacteria rather than Gram-positive bacteria. Similarly, Sujada et al.6 found that the isolate displayed broad-spectrum antimicrobial activity against various microorganisms. The antimicrobial substances produced by the isolate showed significant inhibitory effects comparable to streptomycin or benomyl against Bacillus cereus, Enterococcus faecalis, Staphylococcus aureus, MRSA, Proteus mirabilis, and Fusarium sp. Furthermore, the antifungal activities of AMHU-5 against Cryptococcus neoformans, Aspergillus flavus, Rhizoctonia solani AG-2, and Sclerotium solani were superior to those derived from PDB. In this investigation, 20 different halophilic isolates were collected from termite mounds, and several tests were performed, such as gram staining, biochemical examinations, and evaluations of antibiotic resistance. All isolates were screened under high salt conditions of 12% NaCl. Among them, only 10 isolates grew in the medium, and others did not grow. Interestingly, N3 was chosen due to its antibacterial highest level of inhibiting zone (K. pneumoniae MTCC 39, S. aureus MTCC 96) and antifungal (C. albicans MTCC 282) properties. The research included the purification and analysis of bioactive compounds using TLC and GC-MS for both Termite mound soil and N3 secondary metabolite and the comparison of bioactive compounds found in the library. However, the crucial for assessing the bioactive characteristics of antitumor compounds Pyrrolo[1,2-a] pyrazine-1,4-dione, hexahydro-3-, Cyclo (L-prolyl-L-valine), Triphenylphosphine oxide, Methyl gamma-linoleate, Elaidic acid-TMS, Cyclo(alanylleucyl), antimicrobial compounds 2,4-Di-tert-butylpheno, dibutyl phthalate, Palmitic Acid-TMS, Hexadecanoic acid, 4-[(trimethylsilyl)oxy]butyle, Cyclo(prolyltyrosyl), Methyl oleate, Linoleic acid-TMS, Methyl arachidonate, 2-Furoic acid-TMS, 2,5-Piperazinedione, 3-benzyl-6-isopropyl-, and anti-inflammatory compounds, 9-Octadecenamide, Acetamide, N-(2-methoxy-5-methylphenyl). Additionally, the 16S rRNA sequence confirmation was performed, revealing that the isolates of N3 belonged to the Microbacterium barkeri of 98% similarity species.
In summary, this study demonstrates the potential of Microbacterium barkeri M17 and other halophilic Actinobacteria as key players in the search for next-generation antimicrobial agents. Their remarkable ability to produce diverse and potent bioactive compounds establishes them as a valuable resource for pharmaceutical role of potential reaction pathways were involved in the hydroxyl group of 4-[(trimethylsilyl)oxy] butyl can react with the carbonyl group of the diketopiperazine or the carboxyl group in cyclo (prolyltyrosyl). A coupling reagent like EDCI or DCC could facilitate the linkage between amino (-NH2) or carboxyl groups of the peptide and the diketopiperazine and biotechnological applications.
This study shows that termite mound environments harbor halophilic actinobacteria with remarkable antimicrobial potential. Among the isolates examined, the N3 strain—identified as Microbacterium barkeri—stood out for its strong antibacterial and antifungal activity. Chemical profiling through TLC and GC-MS revealed that this isolate produces a rich mix of bioactive metabolites, including diketopiperazines, peptides, and fatty acid derivatives known for antimicrobial, anti-inflammatory, and antitumor properties. The MIC and MBC results further confirmed the potency of these compounds. Overall, the findings highlight that termite mound–derived halophilic actinobacteria are a promising and largely untapped source of new bioactive compounds, offering valuable prospects for future antimicrobial drug development.
ACKNOWLEDGMENTS
The authors are thankful to the Vice Presidents, Chancellor, and Vice-Chancellor of Kalasalingam University, Krishnankoil, Tamil Nadu, for their support.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
NA and HN conceptualized and designed the study. NA performed data collection, analysis and wrote the manuscript. HN supervised the study and revised the manuscript. Both authors read and approved the final manuscript for publication.
FUNDING
None.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
Not applicable.
- Butler MS, Buss AD. Natural products-the future scaffolds for novel antibiotics? Biochem Pharmacol. 2006;71(7):919-929.
Crossref - Berdy J. Bioactive microbial metabolites. J Antibiot. 2005;58(1):1-26.
Crossref - Tran PN, Yen MR, Chiang CY, Lin HC, Chen PY. Detecting and prioritizing biosynthetic gene clusters for bioactive compounds in bacteria and fungi. Appl Microbiol Biotechnol. 2019;103(8):3277-3287.
Crossref - Sharma M, Dangi P, Choudhary M. Actinobacteria: source, identification, and their applications. Int J Curr Microbiol App Sci;2014:3(2):801-832
- Olanrewaju OS, Babalola OO. Streptomyces: implications and interactions in plant growth promotion. Appl Microbiol Biotechnol. 2019;103:1179-1188.
Crossref - Tiwari K, Gupta RK. Rare actinomycetes: a potential storehouse for novel antibiotics. Critical Reviews in Biotechnology. 2012;32(2):108-132.
Crossref - Berlanga M, Paster BJ, Guerrero R. The taxophysiological paradox: changes in the intestinal microbiota of the xylophagous cockroach Cryptocercus punctulatus depending on the physiological state of the host. International Microbiology. 2009;12(4):227-236.
Crossref - Haľková B, Tuf I, Tajovský K, Mock A. Subterranean biodiversity and depth distribution of myriapods in forested scree slopes of Central Europe. ZooKeys. 2020;930:117-137.
Crossref - Duran-Bautista EH, Silva-Olaya AM, Llanos-Cabrera MP, Yalanda-Sepulveda K, Suárez JC. Soil-feeding termites build islands of soil physical and chemical fertility in pastures in Colombian Amazon. PLOS One. 2025;20.
Crossref - Berdy J. Thoughts and facts about antibiotics: where we are now and where we are heading. J Antibiot. 2012;65(8):385-395.
Crossref - Demain AL, Fang A. The Natural Functions of Secondary Metabolites. In: Fiechter, A. (eds) History of Modern Biotechnology I. Advances in Biochemical Engineering/Biotechnology, vol 69. Springer, Berlin, Heidelberg 2000;69:1-39.
Crossref - Machado H, Sonnenschein EC, Melchiorsen J, Gram L. Genome mining reveals unlocked bioactive potential of marine Gram-negative bacteria. BMC Genomics. 2015;16(1):158.
Crossref - Shah MP, Dey S (Eds.). Trends in biotechnology of Polyextremophiles. Springer. 2024.
Crossref - Barzkar N, Sukhikh S, Babich O. Study of marine microorganism metabolites: New resources for bioactive natural products. Front Microbiol. 2024;14:1285902.
Crossref - Eltokhy MA, Saad BT, Eltayeb WN, et al. Metagenomic nanopore sequencing for exploring the nature of antimicrobial metabolites of Bacillus haynesii. AMB Expr. 2024;14(1):52.
Crossref - Palacios-Rodriguez AP, Espinoza-Culupu A, Duran Y, Sanchez-Rojas T. Antimicrobial activity of Bacillus amyloliquefaciens BS4 against Gram-negative pathogenic Bacteria. Antibiotics. 2024;13(4):304.
Crossref - Joel EL, Bhimba BV. Fungi from mangrove plants: antimicrobial & anticancer potentials. International Journal of Pharmacy and Pharmaceutical Sciences, 2012;4(3):139-142. https://innovareacademics.in/journal/ijpps/Vol4Issue3/4237.pdf
- Cheng SC, Huang MZ, Shiea J. Thin layer chromatography/mass spectrometry. J Chromatogr A. 2011;1218(19):2700-2711.
Crossref - Sommerauer C, Rebernik P, Reither H, Nanoff C, Pifl C. The noradrenaline transporter as site of action for the anti-Parkinson drug amantadine. Neuropharmacology. 2012;62(4):1708-1716.
Crossref - Dolezal M, Zitko J. Pyrazine derivatives: a patent review (June 2012 – present). Expert Opin Ther Pat. 2015;25(1):33-47.
Crossref - Jia T, Zhao M, Pan L, Deng C, Zou J, Zhang X. Effect of phenolic antioxidants on the thermal oxidation stability of high-energy-density fuel. Chemical Engineering Science. 2021; 247:117056.
Crossref - David I, Orboi MD, Simandi MD, et al. Fatty acid profile of Romanian’s common bean (Phaseolus vulgaris L.) lipid fractions and their complexation ability by β-cyclodextrin. PLoS One. 2019;14(11):e0225474.
Crossref - Bailly C. Insights into the Bioactivities and Mechanism of Action of the Microbial Diketopiperazine Cyclic Dipeptide Cyclo(L-leucyl-L-prolyl). Marine Drugs. 2025;23(10).
Crossref - Al-Ostoot FH, Sherapura A, Malojirao VH, et al. Modulation of DNA damage response by targeting ATM kinase using newly synthesized di-phenoxy acetamide (DPA) analogs to induce anti-neoplasia. Pharmacol Rep. 2021;73(5):1344-1360.
Crossref - Liu X, Yang J, Song X, et al. A potential novel treatment strategy for breast cancer: The regulation of apoptosis and metastasis by mitochondria-targeted lupeol-triphenylphosphine derivatives. Bioorganic chemistry. 2025;167:109214.
Crossref - Azman AS, Othman I, Fang CM, Chan KG, Goh BH, Lee LH. Antibacterial, Anticancer and Neuroprotective Activities of Rare Actinobacteria from Mangrove Forest Soils. Indian J Microbiol. 2016;57(2):177-187.
Crossref - Abdel Raoof GF, Al-Abd A. Phytochemical Characterization and Anticancer Potential of Avocado Seed Oil. Chemistry & biodiversity. 2025;e01833
Crossref - Alqahtani FY, Aleanizy FS, Mahmoud AZ, et al. Chemical composition and antimicrobial, antioxidant, and anti-inflammatory activities of Lepidium sativum seed oil. Saudi J Biol S. 2019;26(5):1089-1092.
Crossref - Javid S, Purohit MN, Yogish Kumar H, Ramya K, et al. Semisynthesis of Myristic Acid Derivatives and their Biological Activities: A Critical Insight. Journal of Biologically Active Products from Nature. 2020;10(6): 455-472.
Crossref - Musthafa KS, Balamurugan K, Pandian SK, Ravi AV.
2,5-Piperazinedione inhibits quorum sensing-dependent factor production in Pseudomonas aeruginosa PAO1. J Basic Microbiol. 2012;52(6):679-686.
Crossref - Mitra S, Das R, Emran T, et al. Diallyl Disulfide: A Bioactive Garlic Compound with Anticancer Potential. Front Pharmacol. 2022;13.
Crossref - Sessa C, Valota O, Geroni C. Ongoing phase I and II studies of novel anthracyclines. Cardiovasc Toxicol. 2007;7(2):75-79.
Crossref - Sarova D, Kapoor A, Narang R, Judge V, Narasimhan B. Dodecanoic acid derivatives: Synthesis, antimicrobial evaluation and development of one-target and multi-target QSAR models. Med Chem Res. 2010;20(6):769-781.
Crossref - Korbecki J, Bajdak-Rusinek K. The effect of palmitic acid on inflammatory response in macrophages: an overview of molecular mechanisms. Inflamm Res. 2019;68(11):915-932.
Crossref - Kaviyarasan S, Kavitha K, Indra N, Meenakshisundram P, Thirukumaran K. Isolation, characterization and metabolic profiling of seed endophyte B. licheniformis against Sarocladium oryzae in rice. Plant Science Today. 2025;12(sp1).
Crossref - Malak AY, Mohamed MK, Abd El-Mawla MAA, Zaher MA. Cytotoxic and antimicrobial effects of selected Egyptian Asteraceae species as well as GC-MS metabolite profiling of Senecio cruentus lipophilic fraction. Bulletin of Pharmaceutical Sciences Assiut University. 2023;46(1):39-49.
Crossref - Pal P. Recent advances in syntheses and antibacterial activity of novel furan derivatives. Intech open. 2024.
Crossref - Horie Y, Nomura M, Ramaswamy BR, Harino H, Yap CK, Okamura H. Effects of non-phthalate plasticizer bis (2-ethylhexyl) sebacate (DEHS) on the endocrine system in Japanese medaka (Oryzias latipes). Comp Biochem Physiol C Toxicol Pharmacol . 2023;264:109531.
Crossref - Hammock BD, McReynolds CB, Wagner K, et al. Movement to the clinic of soluble epoxide hydrolase inhibitor EC5026 as an analgesic for neuropathic pain and for use as a nonaddictive opioid alternative. J Med Chem. 2021;64(4):1856-1872.
Crossref - Kusumah D, Wakui M, Murakami M, Xie X, Yukihito K, Maeda I. Linoleic acid, a-linolenic acid, and monolinolenins as antibacterial substances in the heat-processed soybean fermented with Rhizopus oligosporus. Biosci Biotechnol Biochem. 2020;84(6):1285-1290.
Crossref - Ceci C, Lacal PM, Tentori L, De Martino MG, Miano R, Graziani G. Experimental evidence of the antitumor, antimetastatic, and antiangiogenic activity of ellagic acid. Nutrients. 2018;10(11):1756.
Crossref - Abd Alsaheb RA, Aladdin A, Othman NZ, et al. Lactic acid applications in pharmaceutical and cosmeceutical industries. J Chem Pharm Res. 2015;7(10):729-735.
- Giner RM, Rios JL, Manez S. Antioxidant activity of natural hydroquinones. Antioxidants. 2022;11(2):343.
Crossref - Parcheta M et al. Mandelic acid derivatives. Materials, 2022;15(15):5413.
Crossref - Mou Y, Meng J, Fu X, et al. Antimicrobial and antioxidant activities and effect of 1-hexadecene addition on palmarumycin C2 and C3 yields in liquid culture of endophytic fungus Berkleasmium sp. Dzf12. Molecules. 2013;18(12):15587-15599.
Crossref - Roy RN, Laskar S, Sen SK. Dibutyl phthalate, the bioactive compound produced by Streptomyces albidoflavus 321.2. Microbiol Res. 2006;161(2):121-126.
Crossref - Clinical and Laboratory Standards Institute. (2020). Performance Standards for Antimicrobial Susceptibility Testing; 30th ed. CLSI Supplement M100. Wayne, PA: Clinical and Laboratory Standards Institute. Retrieved from https://clsi.org/standards/products/microbiology/documents/m100/
- Vos P, Garrity GM, Jones D. et al. (Eds.). Bergey’s manual of systematic bacteriology: Volume 3: The Firmicutes (Vol. 3). Springer Science & Business Media. 2011. https://link.springer.com/book/10.1007/978-0-387-68572-4
- Li Q, Chen X, Jiang Y, Jiang C. Cultural, physiological, and biochemical identification of actinobacteria. Intech open. 2016.
Crossref - Devi R, Thakur R, Gupta MK. Isolation and molecular characterization of bacterial strains with antifungal activity from termite mound soil. Int J Curr Microbiol Appl Sci. 2018;7(4):1-7.
- Devi R, Thakur R. Screening of bacteria for PGP traits. J Pharmacogn Phytochem, 2018;7(2):1681–1686.
Crossref - Wollum II. AG. Cultural methods for soil microorganisms. In: Page AL, Miller RH, Keeney DR, 2nd eds. Methods of Soil Analysis, Part 2: Chemical and Microbiological Properties. American Society of Agronomy and Soil Science Society of America. 1982:718-802.
Crossref - Sawadogo JB, Traore S, Hien SET, Pale D, Maiga Y, Dianou D, Ouattara AS. Int J Curr Microbiol App Sci. 2023;12(9):288-301.
Crossref - Krishanti NPRA, Zulfiana D, Wikantyoso B, Zulfitri A, Yusuf S. Antimicrobial Production by an Actinomycetes Isolated from the Termite Nest. J Trop Life Sci. 2018;8(3).
Crossref
© The Author(s) 2026. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.