ISSN: 0973-7510
E-ISSN: 2581-690X
The objective of this study was to find an alternative to chemical control of pathogenic fungi in wheat, using microorganisms that are safe and that can be isolated from the same biotopes of the pathogens. Lactic acid bacteria isolated from soft wheat grains were screened for their antifungal activity against Fusarium graminearum Schwab, Aspergillus flavus Link and Aspergillus parasiticus Speare, using two techniques (overlay and co-culture) on De Man, Rogosa, and Sharpe agar. The overlay method showed that out of forty-six lactic acid bacteria, five isolates showed an inhibition of radial growth range from 1% to 73.89%. According to the co-culture method, the most efficient biological agent for wheat mold growth isolate was LAB001 with an average rate of inhibition of 31.18% against A. flavus, 42.26% against A. parasiticus and 55.53% against F. graminearum. Lactic acid bacteria LAB001 was identified as Enterococcus faecium with 99.6% of similarity. E. faecium LAB001 can be considered as promising isolate for the biocontrol of pathogenic molds in small grain cereals.
Biocontrol; lactic acid bacteria; pathogenic molds; small grain cereals.
Soft wheat occupies a central place in the culinary and eating habits of the Algerian population. Although more than a million and a half hectares of wheat have been cultivated on average since 1961 (FAO, 2018), grain yields remain below the demand of the national market. Therefore, Algeria remains dependent on countries with large grain production capacities such as France and Canada for importation of this strategic food. According to FAO (2018), Algeria imported around 2.53.100 tons of wheat in 1965 and around 7.454.603 tons in 2011, despite a tripling of national production.
Massive importation and long-term storage of wheat grains can promote the spread of complex fungi, with damage ranging from the loss of nutritional quality and sanitation of the grain to the accumulation of various mycotoxins. Fusarium graminearum schwab, Aspergillus flavus Link and Aspergillus parasiticus Speare are widely recognized pathogen of cereals that is ubiquitous in almost all countries of the world (Battilani et al., 2016; Merhej et al., 2011; Franco et al., 2011; Miedaner et al., 2010; Pitt and Hocking, 2009; Parry et al., 1995). In addition, these species can infect wheat with mycotoxins such as trichothecenes B, including deoxynivalenol (DON), nivalenol (NIhV), and their derivatives, fumonisine and aflatoxin (B1, B2, G1 and G2), at a high rate.
Owing to these concerns, the Algerian government is required to prepare a policy to inhibit or eliminate these molds. In this context, using biocontrol of plant pathogenic species generally regarded as safe (GRAS) method and is considered a good alternative to chemical fungicides. Indeed, the antagonistic agents selected in this study, lactic acid bacteria, are known for their ability to inhibit the growth of molds and have been used for millennia in the preservation of foodstuffs (Sevgi and Tsveteslava, 2015). Thus, their GRAS status and probiotic potential makes lactic acid bacteria promising candidates for biological control of molds. Lactic acid bacteria are able to live in and adapt to extremely diverse environments and substrates. Indeed, these microorganisms have well-adapted systems that protect them against various stresses they may encounter (Romeo et al., 2001). The main purpose of this study was to screen novel indigenous lactic acid bacteria strains from soft wheat for antifungal effects against three toxigenic fungi, especially F. graminearum, A. Flavus and A. parasiticus.
Fungal isolates
Three strains of F. graminearum, A. flavus and A. parasiticus were isolated from soft wheat grains imported from France during 2015, stored in cooperative grain and dried vegetable silos in the Bechar district, southwestern Algeria (31°37’00’’ north, 2°13’00’’ south), in 2015 at ambient temperature and humidity. The method used for grains sampling was the one described by Codex CAC / GL 50-2004. We first took random and primary samples of similar sizes and then, we formed the composite sample by mixing the primary samples. Finally, once in the laboratory, we dispatched the composite sample at fifty final samples (200g).
Using a solution of 1.5° sodium hypochlorite, grain surfaces were disinfected for 10 min, followed by rinsing twice with sterile distilled water. This was done to eliminate all external pollutions of fungal or bacterial origins. Once dried, 10 grains per plate were incubated at 25°C in darkness for 7 days on Dichloran Chloramphenicol Peptone Agar (DCPA) selective medium for F. graminearum, and on Potato Dextrose Agar (PDA) for Aspergillus flavus and A. parasiticus strains (Pitt and Hocking, 2009). The selected strains were stored in PDA slant tubes at +4°C and in sporal solution in Eppendorf tubes containing Potato Dextrose Broth (PDB) with 25% glycerol at “20°C.
Isolation of lactic acid bacteria
The lactic acid bacterial strains used in this study are indigenous strains isolated from the soft wheat grains as fungal isolates. The isolation technique applied was the one proposed by Chen et al. (2005) and amended by Dalie et al. (2010). 1g of wheat sample (neither disinfected nor crushed) was added to 5ml of De Man, Rogosa, and Sharpe (MRS) broth and incubated at 30°C under anaerobic condition. After 72h of incubation, each mixture was diluted in 9ml of sterile peptone water (0.2% w/v) up to the dilution 10-10. 100µl of each dilution were spread onto MRS agar and incubated at 30°C for 72 hours. Ten percent of the purified isolates were Gram-stained and catalase tested. Only gram-positive and catalase-negative isolates were selected and stored in MRS broth with 30% glycerol at “20°C (Boudra and Niderkorn, 2004).
lactic acid bacteria and its antifungal activity
Two techniques were followed to study antifungal activity. The first was assayed using the streaking or overlay methods (Ström et al., 2002; Magnusson and Schnurer. 2001). Each strain of lactic acid bacteria was inoculated in 2 cm lines on 15 ml of MRS agar plates and incubated in anaerobic condition for 48h at 30°C. The plates were then, overlaid with 10ml of PDA (0.8% w/w agar) containing 106 spores/ml of each strains of F. graminearum, A. flavus and A. parasiticus. After 72h of aerobic incubation at 25°C, the inhibition zone was measured around the bacterial streaks and was presented as a percentage of no fungal growth of the plate area per bacterial streak. This assay was done in duplicate.
The second method was examined using co-culture on MRS agar. According to Florianowicz, (2001): 200µl of young bacterial culture (12 to 16h) were inoculated in depth of 15 ml of MRS agar. After the solidification of the culture medium, one sterile disk was deposited in the center of the plate and then saturated with 10µl of sporal suspension (106 spores/ml). Cultures were incubated at 25°C for one week. Through measuring the diameter of the colonies in two perpendicular directions, linear growth was determined every two days. Antifungal activity was expressed in terms of colony growth inhibition as follows: Inhibition (I) or antifungal activity (A.A.F) = 100 × [1″ (DE / DT)], in which DE is the diameter of a fungal colony in the presence of lactic acid bacteria, and DT is the diameter of a fungal colony without lactic acid bacteria (as a control).
Physiological characterization of the isolate showing the strongest antifungal effect
Physiological characterization of the lactic acid bacterial strain showing the strongest antifungal effect was carried out in MRS broth at different temperatures (10°C, 30°C, 37°C, and 45°C), with different pH values (4.4 and 9.6), and with different concentrations of NaCl (6.5% and 18%) at 30°C for 48 hours of incubation. 100 ml of MRS broth were inoculated with lactic acid bacteria LAB001 culture (12 to 16h) at the rate of 1% (v/v) and incubated for 48 hours. A heat resistance test was studied through exposing the isolate to a temperature of 60°C for 30 min, then incubating the sample at 30°C for 24 hours. Bacterial growth was determined by measuring the optical density at 600 nm (Dalie et al., 2010).
Biochemical characterization of the isolate showing the strongest antifungal effect
Various tests were carried out including tests of the fermentation type, growth on blue milk Sharmen, arginine hydrolysis, citrate use, dextran and acetone production, and fermentation profile performed within API20 Strep gallery (BioMérieux, REF 20600, France), according to the instructions provided by the manufacturer. The bacterial suspension was dispensed into API20 Strep strips wells and coated with paraffin oil. Then, the strips were incubated at 30°C for 24 h to 48h. The results were red based on colors reactions according to a chart provided by the manufacturer. An identification was made using the APIWEB TM Database.
Statistical analysis
Two independent experiments were performed for all assays of antifungal activity and mean values ± standard deviation (SD) were calculated.
Screening of lactic acid bacterial strains for their antifungal effects
Overlay method
Forty-six presumptive colonies of lactic acid bacteria were isolated on MRS agar medium in which 99% cocci and 1% rods. According to the percentages of antifungal inhibition, the LAB strains were randomly classified into five levels: 80 to 100% (Level 1), 50 to 79% (Level 2), 25 to 49% (Level 3), 15 to 24% (Level 4) and 0 to 14% (Level 5). According to the results summarized in Table 1, lactic acid bacteria strain LAB001 showed the greatest antifungal activity (Figure 1), followed by strain LAB003, which showed fairly good inhibitory activity. Isolates LAB002 and LAB005 showed very low activity, whereas the other 43 isolates showed no inhibitory activity against the molds strains tested. Therefore, isolates LAB001 and LAB003 were selected for the second assay.
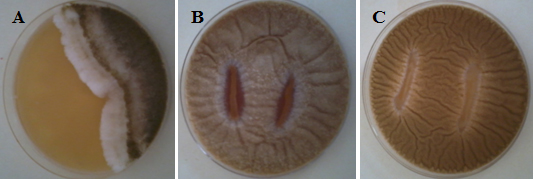
Fig. 1. Plates from Antifungal activity of some LAB by the overlay agar method. (A) LAB001 Vs A. flavus showing clear zones of fungal inhibition (B) LAB002 Vs A. parasiticus. (C) LAB029 Vs A. parasiticus.
Table (1):
Screening of antifungal activity of lactic acid bacteria isolates in overlay assay.
| Fungal species | Strains | Inhibitory activity rate (%) | |||
|---|---|---|---|---|---|
| LAB isolates | |||||
| LAB001 | LAB002 | LAB003 | LAB005 | ||
| F. graminearum | S 1 | 68.59±1.89 | 10.65±2.29 | 55.32±5.12 | 6.03±0.09 |
| S 2 | 73.89±1.46 | 8.25±0.82 | 43.65±3.75 | 6.29±0.26 | |
| S 3 | 69.39±2.05 | 13.96±1.19 | 51.35±1.05 | 4.38±2.51 | |
| A. flavus | S 4 | 56.95±5.89 | 0.00±0 | 28.34±2.79 | 5.75±0.35 |
| S 5 | 69.44±4.20 | 5.5±0.64 | 33.35±4.72 | 4.5±1.41 | |
| S 6 | 59.72±0.40 | 4.78±0.52 | 40.56±4.71 | 1±1.41 | |
| A. parasiticus | S 7 | 51.66±2.85 | 3.89±0.50 | 25.89±2.96 | 4.75±2.45 |
| S 8 | 50.56±0.79 | 6.50±2.83 | 24.45±2.29 | 3.5±0.54 | |
| S 9 | 47.22±1.57 | 2.25±3.18 | 20.00±3.10 | 2±0.71 | |
* Result is given as mean value ±SD.
Co-culture method
Results shown in Figure 2 and Figure 3 clearly showed the significant antagonistic effect of the isolate LAB001 after seven days of co-culture against the studied mold species. The maximum effect was observed against the first strain of F. graminearum (62.83%) and A. parasiticus (66.77%). On the other hand, the inhibitory activity of LAB003 on the radial growth oscillates between a minimum of 7% against A. flavus and a maximum of 54.83% against F. graminearum. As a result, the most resistant fungus was A. flavus, and the most sensitive was F. graminearum.Based on these results, the strain LAB001 was considered the most effective biological agent.
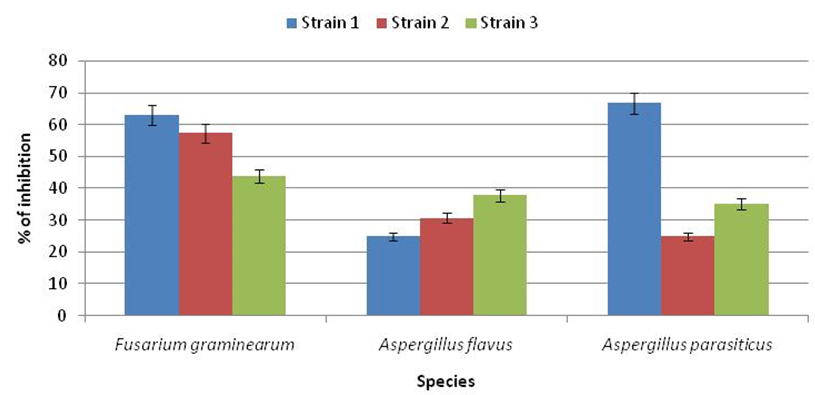
Fig. 2. Rate of inhibition of radial growth of mold strains by LB001 after 7days
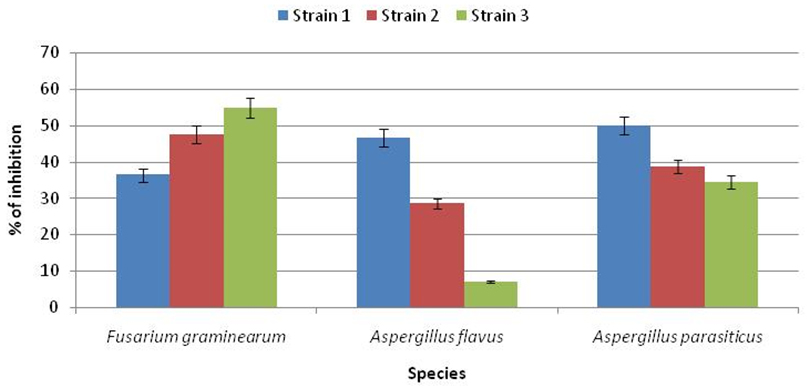
Fig. 3. Rate of inhibition of radial growth of mold strains by LB003 after 7days
Characterization and identification of isolate LB001
The results (Table 2) show that strain LAB001 is able to grow within a range of temperatures [10°C to 45°C], and an optimum temperature of 37°C. In addition, it can resist a temperature of 60°C for 30 minutes. Furthermore, this strain is capable of growing at pH values ranging from 4.4 to 9.6 or in culture media containing up to 6.5% NaCl. Biochemically, isolate LAB001 is homofermentative, meaning it is able to grow on milk “blue Sharmen” and reduce blue methylene at 0.1% and 0.3% (of blue methylene). It is capable of hydrolyzing arginine, esculine, hippurate and degrading citrate via the enzyme citratase. The isolate LAB001 can ferment several sources of carbon especially, glucose, lactose, amidon, arabinose, mannitol, ribose and trehalose, with production of a-Galactosidase, ß-Glucuronidase, ß-Galactosidase, Pyrrolidonyl-arlamidase, Leucine-amino-peptidase. According to the APIWEB TM Database identification system, strain LAB001 shows 99.6% similarity with Enterococcus faecium.
Table (2):
Physiological and biochemical characteristics of isolate LAB001.
Characteristics |
Profile of the isolate LAB001 |
|---|---|
Cell shape |
Coccus |
Cell association |
Chain |
Type of fermentation |
Homofermentation |
Temperature: |
|
10°C |
+ |
30°C |
+ |
37°C |
+ |
45°C |
+ |
60°C |
+ |
Concentration of NaCl: |
|
6.50% |
+ |
18% |
– |
pH: |
|
4.4 |
+ |
9.6 |
+ |
Type of fermentation : |
Homofermentation |
Growth on Methylene blue milk : |
|
0.10% |
+ |
0.30% |
+ |
Hydrolysis of : |
|
Arginine |
+ |
Esculin |
+ |
Hippurate |
+ |
Fermentation of : |
|
Glucose |
+ |
Lactose |
+ |
Amidon |
+ |
Arabinose |
+ |
Mannitol |
+ |
Raffinose |
– |
Ribose |
+ |
Sorbitol |
– |
Trehalose |
+ |
Inulin |
– |
Glycogen |
– |
Production of : |
|
Acetoin |
ND |
±-Galactosidase |
+ |
ß-Glucuronidase |
+ |
ß-Galactosidase |
+ |
Alkaline phosphatase |
– |
Pyrrolidonyl-arlamidase |
+ |
Leucine-amino-peptidase |
+ |
(+): positive; (-): negative; ND: not determined
Several scientific studies have focused on lactic acid bacteria of plants for biological control against pathogenic microorganisms and their associated toxins (Matel and Cornea, 2014; Oliveira et al., 2015b; Gerbaldo et al., 2012; Sathe et al., 2007; Cabo et al., 2002; Clevland et al., 2001). In this study, we hypothesized that lactic acid bacteria could inhibit the development of fungi isolated from the same substrates. We speculated that this would result in vitro interactions that more closely mimic in vivo interactions (Kerry, 2000). This approach has been used in several studies of biological control organisms (Dalie et al., 2010 ; Bleve et al., 2006; Okigbo, 2005). In the present study, a screening for antifungal activity against mold strains responsible for wheat grains contamination was done for forty-six lactic acid bacteria strains that were isolated. The rates of inhibition of fungal growth through overlay and co-culture methods showed that the most efficient isolate was LAB001 which exerted an important antifungal activity on all mold strains that were examined. These methods were also reported by other authors (Oliveira et al., 2015a; Dalie et al., 2010; Ström et al., 2002; Magnusson et al., 2002) as a good assay to evaluate the antifungal potential of lactic acid bacteria. On the other hand, we observed that the antifungal effect was LAB and fungi strain-dependent.
The isolate that showed the most promising antifungal activity was identified phenotypically as Enterococcus faecium (isolate LAB001). This isolate showed resistance to various stresses that might be encountered in the environment (e.g., 10°C<T<45°C, heat resistance at 60°C for 30 minutes; 4.4<pH<9.6). This is advantageous for biocontrol and suggests that E. faecium can be used under various experimental conditions in future studies. We suggest that selection of the most effective strain is fundamental for bio-preservation. Selection should be based first, on the safety of the organism, and second on its adaptability to storage conditions that will allow it to reduce toxigenic fungal contaminants rapidly. In this sense, several scientists reported that E. faecium is considered a healthy agent used as a natural starter culture, contributing to the development of the organoleptic properties of Mediterranean cheese (Sarantinopoulos et al., 2002; Monero et al., 2003; Favaro et al., 2015) and may play an important role in the preservation of food and its quality (Calo-Mata et al., 2008; Chen et al., 2010).
Moreover, E. faecium is known to produce high levels of enterocin (Vera Pingitore et al., 2012; Laukova , 2012 ; Galvez et al., 2011) or enterolysin A (Dortu and Thonart, 2009). Its action against several microorganisms, i.e., Listeria monocytogenes, Staphylococcus spp., fungi, and yeasts, and other organisms, has been studied widely (Monero et al., 2002; Hosseini et al., 2009; Galvez et al., 2011, Bourouni Chahad et al., 2012; Favaro et al.; 2014).
According to Rehaiem (2014), some active enterococci strains have been suggested as safe candidates due to the growing interest for the usage of probiotics, along with the currently most commonly used strains of Lactobacillus and Bifidobacterium. Furthermore, Calo-Mata et al. (2008) declared that the unlimited use of the E. faecium (NCIMB) for calves and piglets has been approved by the Commission of the European Communities [Commission Regulation (EC) No. 1333/2004], and in the preparation of chickens for fattening [Commission Regulation (EC) No. 1810/2005].
In this study, LAB was isolated from the same biotopes as mold pathogens. Screening of bacterial isolates for their antifungal activity was accomplished using two techniques (overlay and co-culture) in MRS agar. The results showed that the most effective isolate was E. faecium strain LAB001, with 99.6% similarity and based on physiological and biochemical characteristic. Additional studies on the reduction and/or eradication of mold mycotoxins in wheat grains should be conducted.
- Battilani, P., Toscano, P., Van der Fels-Klerx, H.J., Moretti, A., Camardo Leggieri, M., Brera, C., Rortais, A., Goumperis, T., & Robinson, T. Aflatoxin B1 contamination in maize in Europe increases due to climate change. Nature. Sci. Rep, 2010; 6 (24328), 1-7.
- Bleve, G., Grieco, F., Cozzi, G., Lagrieco, A., & Visconti, A. Isolation of epiphytic yeasts with potential for biocontrôle of Aspergillus carbonarius and Aspergillus niger on grape. Int. J. Food Microbiol, 2006; 108: 204-209.
- Boudra, H., & Niderkorn, V., Conservation of strains of fermentative bacteria by cryopreservation. The Research Center of Clermont-Theix. Champanelle. France 2004.
- Bourouni Chahad, O., El Bour, M., Calo-Mata, P., Boudabous, A., & Barros-Velazquez, J. Discovery of novel biopreservation agents with inhibitory effects on growth of food-borne pathogens and their application to seafood products. Research in Microbiology, 2012; 163: 44-54.
- Cabo, M.L., Braber, A.F., & Koenraad, P.M.F.J. Apparent antifungal activity of several lactic acid bacteria against Penicillium discolor is due to acetic acid in the medium. J. Food Prot, 2002; 65: 1309-1316.
- Calo-Mata, P., Arlindo, S., Boehme, K., De Miguel, T., Pascoal, A., & Barros-Velazquez J. Current Applications and Future Trends of Lactic Acid Bacteria and their Bacteriocins for the Biopreservation of Aquatic Food Products. Food Bioprocess Technol, 2008; 1: 43-63.
- Chen, Y.S., Yanagida, F., & Shinohara, T. Isolation and identification of lactic acid bacteria from soil using an enrichment procedure. Let. Appl. Microbial, 2005; 40: 195-200.
- Chen, Y.S., Miyashita, M., Suzuki, K.I., Sato, H., Hsu, J.S., & Yanagida, F. Lactobacillus pobuzihii sp. nov., isolated from pobuzihi (fermented cummingcordia). International Journal of Systematic and Evolutionary Microbiology, 2010; 60: 1914–1917.
- Cleveland, J., Monteville, T., Nes, F., & Chikindas, M. Bacteriocins: Safe, natural antimicrobials for food preservation. Int. J. Food Microbiol, 2001; 71(1), 1-20.
- Dalie, D.K.D., Deschamps, A.M., Atanasova-Penichon, V., & Richard-Forget, F. Potential of Pediococcus pentosaceus (L006) isolated from Maize leaf to suppress Fumonisin-producing Fungal Growth. J. Food Prot, 2010; 6(9),1129-1137.
- Dortu, C., & Thonart, P. The bacteriocins from lactic acid bacteria: characteristics and interests for the bioconservation of food products. Biotechnol. Agron. Soc. Environ, 2009; 13(1): 143-154.
- FAOSTAT. Consulted online: http://faostat3.fao.org/download/T/TP/F. Cited the 08 january 2018.
- Favaro, L., Basaglia, M., Casella, S., Hue, I., Dousset, X., Gombossy de Melo Franco B.D., & Todorov, S.D. Bacteriocinogenic potential and safety evaluation of non-starter Enterococcus faecium strains isolated from home made white brine cheese. Food Microbiology, 2014; 38: 228-239.
- Favaro, L., Lucia, A., Penna, B., & Todorov, S.D. Bacteriocinogenic LAB from cheeses-Application in biopreservation? Trends in Food Science & Technology, 2015; 41(1): 37-48.
- Florianowicz, T. Antifugal activity of some microorganisms against Penicillium expansum. Eur. Food. Res. Technol, 2001; 212: 282-286.
- Franco, T.S., Garcia, S., Hirooka, E.Y., Ono, Y.S., & Santos, J.S. Lactic acid bacteria in the inhibition of Fusarium graminearum and deoxynivalenol detoxification. J. App. Microbiol, 2011; 111; 739-748.
- Galvez, A.A., Dauphin, R.D., Destain, J., Campos, D., & Thonart, P. The entrérocoques: advantages and disadvantages in biotechnology. Biotechnol. Agron. Soc. Environ, 2011; 16(1): 67-76.
- Gerbaldo, G.A., Barberis, C., Pascual, L., Dalcero, A., & BarBeris, L. Antifungal activity of two Lactobacillus strains with potential probiotic properties. FEMS Microbiol Lett, 2012; 332: 27-33.
- Hosseini1, S.V., Arlindo, S., Bohme, K., Fernandez-No, C., Calo-Mata, P., & Barros-Velazquez, J. Molecular and probiotic characterization of bacteriocinproducing Enterococcus faecium strains isolated from nonfermented animal foods. Journal Applies Microbiology, 2009; 107: 1392-1403.
- Kerry, B.R. Rhizosphere interactions and exploitation of microbial agents for the biological control of plant-parasitic nematodes. Ann. Rev. Phytopathol, 2000; 38: 423-991.
- Laukova, A. Potential Applications of Probiotic, Bacteriocin-Producing Enterococci and Their Bacteriocins. In: Lahtinen S., Ouwehand A.C., Salminen S. & Wright A.V. (Taylor & Francis Groyp Ed), Lactic Acid Bacteria : Microbiological and Funcional Aspects. 2012; (pp. 40-53).
- Magnusson, J., & Schnurer, J. Lactobacillus coryniformis subsp. coryniformis strain Si3 produces a broad-spectrum proteinaceous antifungal compound. App. Env. Microbiol, 2001; 67: 1-5.
- Matei, A., & Cornea, C.P. Antifungal activity of some lactic acid bacteria isolated from materials of vegetal origin. Biotechnologies, 2014; 18: 42-47.
- Merhej, J., Richard-Forget, F., & Barreau, C. Regulation of trichothecenes biosynthesis in Fusarium: recent advances and new insights. Appl Microbiol Biotechnol, 2011; 91: 519-528.
- Miedaner, T., Bolduan, C., & Melchinger, A.E. Aggressivness and mycotoxin production of eight isolates each of Fusarium graminearum and Fusarium verticillioides for ear rot on susceptible and resistant early maize inbred lines. Eur. J. Plant Pathol, 2010; 127: 113-123.
- Monero, M.R.F., Leisner, J.J., Tee, L.K., Ley, C., Radu, S., Rusul, G., Vancanneyt, M., & De Vuyst, L. Microbial analysis of Malaysian tempeh, and characterization of two bacteriocins produced by isolates of Enterococcus faecium. Journal of Applied Microbiology, 2002; 92: 147–157.
- Monero, M.R.F., Callewaert, R., Devreese, B., Van Beeumen, B., & De Vuyst, L. Isolation and biochemical characterisation of enterocins produced by enterococci from different sources. Journal of Applied Microbiology, 2003; 94: 214–229.
- Okigbo, R.N. Biological control of postharvest fungal rot of yam (Dioscorea spp.) with Bacillus subtilis. Mycopathol, 2005; 159: 307-314.
- Oliveira, P., Brosnan, B., Furey, A., Coffey, A., Zannini, E., & Arendt, E.K. Lactic acid bacteria bioprotection applied to the malting process. Part I: strain characterization and identification of antifungal compounds, Food Control, 2015a; 51: 433-443.
- Oliveira, P., Brosnan, B., Jacob, F., Furey, A., Coffey, A., Zannini, E., & Arendt E.K. Lactic acid bacteria bioprotection applied to the malting process. Part II: Substrate impact and mycotoxin reduction, Food Control, 2015b; 51: 444-452.
- Parry, D.W., Jenkinson, P., & McLeod, L. Fusarium ear blight (scab) in small grain cereals-a review. Plant pathol, 1995; 44: 207-238.
- Pitt, J.I., & Hocking, A.D. Fungi and food spoilage. Springer Editor. 2009; 519 p.
- Rehaiem, A., Ben Belgacem, Z., Reza Edalatian, M., Martínez, B., Rodríguez, A., Manai, M., & Guerra, N.P. Assessment of potential probiotic properties and multiple bacteriocin encoding-genes of the technological performing strain Enterococcus faecium MMRA. Food Control, 2014; 37: 343-350.
- Romeo, Y., Bouvier, J., & Gutierrez, C. The response to osmotic stress of lactic acid bacteria Lactococcus lactis and Lactobacillus plantarum. Lait, 2001; 81: 49-55.
- Sarantinopoulos, P., Leroy, F., Leontopoulou, E., Georgalaki, M.D., Kalantzopoulos, G., Tsakalidou, E., & De Vuyst, L. Bacteriocin production by Enterococcus faecium FAIR-E 198 in view of its application as adjunct starter in Greek Feta cheese making. International Journal of Food Microbiology, 2002; 72: 125-136.
- Sathe, S.J., Nawani, N.N., Dhakephalkar, P.K., & Kapadnis, B.P. Antifungal lactic acid bacteria with potential to prolong shelf-life of fresh vegetable. J. appl. Microbiol, 2007; 103: 2622-2628.
- Sevgi, E., & Tsveteslava, I.I. Antifungal activity of lactic acid bacteria, isolated from Bulgarian wheat and rye flour. J. Life Sciences, 2015; 9: 1-6.
- Ström, K., Sjögren, J., Broberg, A., & Schnürer, J. Lactobacillus plantarum MiLAB 393 produces the antifungal cyclic dipeptides cyclo (L Phe-L Pro) and (L Phe-trans-4-OH-L Pro) and phenyl lactic acid. Appl. Environ. Microbial, 2002; 68: 4322-4327.
- Vera Pingitore, E., Todorov, S.D., Sesma, F., & Gombossy de Melo Franco, B.D. Application of bacteriocinogenic Enterococcus mundtii CRL35 and Enterococcus faecium ST88Ch in the control of Listeria monocytogenes in fresh Minas cheese. Food Microbiology, 2012; 32: 38-47.
© The Author(s) 2018. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.


