ISSN: 0973-7510
E-ISSN: 2581-690X
The objective of the study is to determine the antibiotic resistance and biofilm formation of Klebsiella pneumoniae in different surfaces. In this study, 47 K. pneumoniae strains were procured from K.A.P. Viswanatham Government Medical College, Trichy, Tamil Nadu. All the isolates were reconfirmed through biochemical reactions. Maximum resistance was observed against Ampicillin, Cefepime, Cefotaxime, Co-trimoxazole, Aztreonam, and Imipenem. The lowest resistance was noticed against Cefuroxime. Among these 85% strains exhibited multidrug resistance with 13% was carbapenem and 98% ESBL resistant strains. The biofilm formation of all the strains in different surfaces revealed that stainless steel surface found to be adhered high number of cells than in other surfaces. Similarly, the biofilm formation of strains grown with glucose in tryptic soya broth (TSB) enhanced adherence ability. It is concluded that presence of glucose or any sugar substrates enhances the biofilm formation thereby developing high resistance against different antibiotics. This condition is detrimental to human health and causes considerable concern.
Klebsiella pneumoniae, Antibiotic Resistance, Biofilm, Material Surfaces
In 1882, Friedlander was the first to isolate K. pneumoniae from the lungs of pneumonia victims who had died. This encapsulated bacterium was formerly known as Friedlander’s bacillus, but in 1886 it was renamed Klebsiella.1 K. pneumoniae, a ubiquitous Gram-negative encapsulated bacterium which exists on the mucosal surface of mammals. Among Klebsiella spp., K. pneumoniae was considered as a primary pathogen of hospital and community acquired infections and has become an “emergency threat” to public health due to multidrug resistance (MDR). It is a crucial component of nosocomial infections, including pneumonia, bacteremia, septicaemia, urinary tract infections (UTIs), pyogenic liver abscesses, meningitis, wound infections, and burn infections.2 In addition, catheter-associated urinary tract infection (CAUTI) is the hospital acquired infections (HAIs) will increase the patient’s incidence rate.3
K. pneumoniae is described as a bacterium that can adhere to surface contact, develop biofilm, and persist on these surfaces. Biofilm are a major health care problem, reportedly involving 65% of bacterial infections, allowing cells to persist, results in increasing resistance to antibiotics.4 A biofilm is a cluster of microbial cells, which are adhered on the surface and surrounded with a matrix made primarily of polysaccharide components. The polymeric matrix is the adhesion between the polysaccharide cells to protect the cells from the biofilm.5 The biofilm of K. pneumoniae on the internal surface of catheters and other indwelling implants has been recognized as the primary cause of CAUTI.4 It also develops invasive infections by forming biofilm colonization in the gastrointestinal and respiratory tract areas, especially in immunosuppressed patients.
Effective biofilm formations are influenced by the surface type of the material, compared to stainless steel, cell adhesion and biofilm formation in silica gel and polycarbonate are immediate.6 There are other factors that change biofilm formation, including nutrient levels, temperature, and relative humidity (RH) conditions. Biofilm inhibits the penetration of antibiotics and slows the growth rate of bacteria, increases persistent cells, and promotes gene sharing.5,7 In addition, the misuse of antibiotics in the treatment of human illnesses causes a serious threat to human health. Antibiotic resistance genes have been well-documented as an increasing risk that can impart resistance to drugs that have been exposed.8 Recent studies have revealed the interconnection between antibiotic resistance and biofilm formation in K. pneumoniae clinical isolates.5 However, quantitative experimental studies on biofilm formation on various surfaces are lacking. The current study focused on the antibiotic resistance pattern of K. pneumoniae and the biofilm formation of K. pneumoniae on diverse surfaces.
Collection of Strains
As many as K. pneumoniae strains were acquired from K.A.P. Viswanatham government college hospital, Tiruchirappalli, Tamil Nadu, India.
Reconfirmation of Bacterial Isolates
Firstly, the collected strains were grown on tryptic soya agar (TSA) (HiMedia, India) for overnight at 37˚C and transferred to MacConkey agar (HiMedia, India) to study the morphological characterization and finally preceded for the biochemical confirmation analysis.9
Determination of Antibiotic Resistance Pattern
All the confirmed K. pneumoniae isolates were subjected to antibiotic resistance patterns. Following the methods of Clinical Laboratory and Standards Institute (CLSI, 2018) the fresh bacterial isolate was inoculated into a 0.85% NaCl suspension to achieve a turbidity equivalent to 0.5 McFarland standards.10 Using a sterile cotton swab, the culture was swabbed on Mueller Hinton Agar (MHA) (HiMedia, India) and then antibiotic discs were placed. Antibiotics used to determine the resistance pattern of K. pneumoniae were Ampicillin (10µg), Aztreonam (30µg), Cefepime (30µg), Cefotaxime (30µg), Cefuroxime (30µg), Cephalothine, Co – trimoxazole (25µg), Colistin (25µg), Doripenem (10µg), Ertapenem (10µg), Gentamycin(10µg), Imipenem (10µg) and Meropenem (10µg) (HiMedia, India).
Formation of Biofilm on Diverse Surfaces
Analysis of Biofilm Formation on Glass and Polypropylene Tubes in TSB and TSB with Glucose (TSBG)
Qualitative evaluation of biofilm was assessed by the tube method, as formerly reported by Christensen et al. with slight modifications.11 All confirmed bacterial isolates were cultured individually in TSB (HiMedia, India) at 37°C for 24 h. In brief, overnight culture of K. pneumoniae were inoculated in glass or polypropylene tubes filled with 10 ml of TSB (HiMedia, India) and TSBG, incubated at 37°C for 24 h. After incubation, these test tubes were drained and rinsed thrice with sterile water and air dried. Then the tubes were stained using 100 µl of 0.5% crystal violet (CV) (HiMedia, India) and the excess stains were removed by washing the tubes with sterilized water. The tubes were air dried upside down for the observation of biofilm formation. The cell-bound dye in the biofilm was decolorized with 1 ml of 99% ethanol and then transferred to a cuvette. The optical density (OD) was taken at 595 nm in a spectrophotometer (Cary 60 UV – Vis, Agilent Technologies, U.S.A).12
Analysis of Biofilm Formation on Polystyrene Plates in TSB and TSBG
All bacterial isolates were individually cultured in TSB (HiMedia, India) and TSBG at 37°C for 24 h. Each well of a sterile 96-well polystyrene plate (Tarsons, India) was loaded with 90 µl TSB (HiMedia, India) and TSBG respectively, followed by 10 µl of bacterial culture. The plate was then incubated at 37°C for 24 h. At the end of incubation, the medium was removed by inverting the microtitre plate, and then the wells were rinsed thrice using sterile water and air dried. All the wells were stained using 50 µl 0.5% CV (HiMedia, India) for 5 minutes. Then the excess stains were removed by washing with sterile water. The dye associated with the adhering cells was decolorized with 1 ml of 99% ethanol, and in each well, OD was quantified at 595 nm in a spectrophotometer ( Cary 60 UV – Vis, Agilent Technologies, U.S.A).12
Analysis of Biofilm Formation on Stainless Steel in TSB and TSBG
All K. pneumoniae strains were individually cultured in TSB (HiMedia, India) and TSBG. Then the stainless coupons were submerged in the glass tubes loaded with 1 ml of K. pneumoniae strain and 10 ml of sterilized (PBS) (pH – 7.4) respectively and incubated for 2 h at 37°C. The stainless steel coupons with the biofilm were washed with sterile water and air dried after 2h. Then, 1ml of 0.5% CV stain (HiMedia, India) was used to stain the stainless steel coupon. The excess stains were removed using sterile water. The dye associated with the adhering cells were decolorized by using of 1 ml of 99% ethanol, and OD was taken at 595 nm in a spectrophotometer (Cary 60 UV – Vis, Agilent Technologies, U.S.A).12
Total number of K. pneumoniae strains procured from K.A.P. Viswanatham Government College Hospital and their sources are given in Table. A total of 9 strains from blood, 16 strains from pus and 22 strains from urine were obtained and all the strains were reconfirmed.
Table (1):
Distribution of K. pneumoniae in different samples.
S. No. |
Samples |
Number of isolates |
K. pneumoniae distribution (%) |
|---|---|---|---|
1. |
Blood |
9 |
19% |
2. |
Pus |
16 |
34% |
3. |
Urine |
22 |
47% |
Total |
47 |
Antibiotic Resistance Pattern Analysis
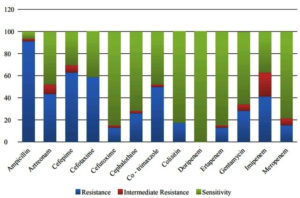
Antibiotic resistance pattern for the K.pneumoniae are presented in Figure 1. The sensitivity of doripenem was detected in all the 47 isolates (100%) and in the test of ESBL antibiotics, the K. pneumoniae strains exhibited its stronger resistance of around 93.1% to Ampicillin, followed by 63% to Cefepime, 58.7% to Cefotaxime, 50% to Co-trimoxazole, 43.4% to Aztreonam, 28.2% to Gentamycin, 26% to Cephalothine, 17.4% to Colistin, 13% to Cefuroxime. In carbapenem antibiotic tested, the isolates expressed their strong resistance against Imipenem 41.3% followed by Meropenem 15.2% and 13% Ertapenem. Among these 65.9% were MDR strains.
Formation of Biofilm on Diverse Material Surfaces
Analysis of Biofilm Formation on Glass and Polypropylene Tubes in TSB and TSBG
Biofilm production of K. pneumoniae on glass surface on TSB and TSBG is presented in Figure 2. All 47 isolates showed a consistent biofilm formation in TSB and TSBG. In TSB significant increase was found in strains 9, 46, and 47 than in TSBG. In strains 7, 8, 11, 14–17, 21, 22, 25 – 27, 30, 31, 35, 36, 38, 39, and 43, TSB seems to have a moderately higher growth rate than TSBG. In TSBG, strains 2, 10, 19, 24, and 45 exhibited a lesser growth. All other strains, with the exception of the ones specified above, remain unchanged. It is observed that when the glucose was supplemented with TSB, 55% of the strains exhibited good growth, while the other 45% of the strains did not exhibit any change over the glass surface.
The biofilm production of K. pneumoniae strains on polypropylene surface is presented in Figure 3. From strains 1 to 14, exhibited a fluctuation in biofilm formation and in strains 15 to 21, showed potential increases in growth rate on TSBG when compared to TSB, whereas strain 22 exhibited no change. Similarly, strains 23 to 29 showed enhanced growth in TSBG. Similarly, strains 30 and 31 showed increased growth on TSB than in TSBG. In strains 32 and 33 significant increased growth is observed. TSB also exhibited increase growth in strains 34 and 35, while strains 36 to 39 exhibited higher growth in TSBG than in TSB, and strain 40 exhibited no change in their growth. In strains 41–43, 45, and 46, there was a potential rise in TSBG and strains 44 and 47 remained unchanged. It was observed that, when the glucose was supplemented in TSB, 68% of the strains exhibited good growth, whereas 27.6% of the strains exhibited lesser growth, and no change is noticed over 4.2% of the K. pneumoniae strains on the polypropylene surface.
Biofilm production of K. pneumoniae on polystyrene surface on TSB and TSBG is presented in Figure 4. All strains exhibited fluctuation in biofilm formation. In TSBG, strains 6, 14, 25, 37, 38, 40, and 46 exhibited a decreased growth, while strains 2, 8, 36, 40, and 41 no much differences was observed. Except for the strains described above, all other strains grew at a faster rate in TSBG than in TSB. It was observed that, when the glucose was supplemented in TSB, 72% of the strains exhibited an increased growth, while 14.8% of the strains exhibited lesser growth and 12.7% of the K. pneumoniae strains on the polystyrene surface exhibited no change.
The biofilm production of K. pneumoniae strains stainless steel surface is presented in Figure 5. Consistent fluctuation was observed except strains 27 and 47, which exhibited a significant, increase in TSBG than in TSB and in strains 13, 21, 29, 35, 39, and 42, lesser growth was noticed in TSBG. Except for the strains mentioned above, all other strains showed normal growth in TSBG than in TSB. It was observed that, when the glucose was supplemented TSB, 59.5% of the strains exhibited increased growth, followed by 8.5% that exhibited a lesser growth and 32% of the strains remained unchanged on stainless steel surface.
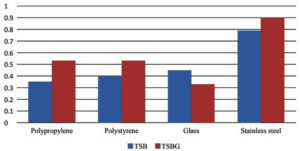
Comparative studies of biofilm formation on various surfaces using TSB and TSBG as examined by the CV staining method over glass, polypropylene, polystyrene, and stainless steel are illustrated in Figure 6. In TSB and TSBG, one strain (47) expressed the greater level of biofilm production over the glass surfaces, whereas one strain (47) in TSB and three strains (41, 43, and 47) in TSBG demonstrated the increased biofilm production over the polypropylene surfaces. Similarly, in TSB four strains (1, 15, 45, and 47) and in TSBG strains 1 and 47 exhibited the increased biofilm production over the polystyrene surfaces. In stainless steel surfaces, however, two strains (4 and 31) in TSB and one strain (27) in TSBG formed the highest biofilm.
As a result, there was no difference among the strains that produced the highest biofilm and those that produced the least biofilm on different surfaces. Stainless steel outperformed various surfaces in terms of influence on biofilm formation. Using TSB, it was found that stainless steel surfaces had increases of 44%, 39%, and 34% when compared to the other examined surfaces (glass, polypropylene, and polystyrene) respectively. While stainless steel surface using TSBG was compared to other evaluated surfaces (glass, polypropylene, and polystyrene), a difference of 26%, 26%, and 46% raise were found respectively. However, the difference was not much enough to be noticed. In all surfaces except the glass surfaces, biofilm growth in TSBG was higher than in TSB.
Gram-negative bacteria are the prominent cause of infection in both community and hospital settings. The rise of microbes resistant to a spectrum of antibiotics used to treat illnesses has now become a major global public health concern throughout the world. In this study, out of 47 tested isolates, 93.1% were resistant to ampicillin. However, 100% of the tested isolates showed sensitivity to doripenem. K. pneumoniae exhibited strong antibiotic resistance against all the antibiotics tested and 65.9% of the isolates were found to be MDR. This in complete agreement with the findings of Cepas et al., who reported MDR of K. pneumoniae strains accounted for 38% of all the K. pneumoniae strains.13 Similarly, Manjula et al. also observed that 90.2% were shown to be MDR K. pneumoniae and also exhibited stronger resistance to penicillin, cephalosporin, fluoroquinolone, aminoglycosides, and sulphonamides which was also supported by Moini et al.14,15 MDR bacteria are posing a significant problem in terms of infection management. So it’s critical to track and optimize antibiotic use through antibiotic stewardship programmes. Since the shortcomings of treatments are prevalent among people who have always obtained single antibiotic therapy, many trials have already shown that using a combination of antibiotics can prevent the future emergence of new resistant strains.
The CV staining technique was used to examine the efficacy of various contact surfaces and glucose supplementation in K. pneumoniae biofilm formation. There was a substantial variation in biofilm production on the material surfaces. In TSBG, the isolates showed higher biofilm formation on stainless, polypropylene, and polystyrene surfaces than on glass surfaces. No association was obtained between the tested isolates and greater biofilm production across diverse material surfaces. In terms of the impact of material surfaces, biofilm production in TSB was higher in stainless steel than in polypropylene, polystyrene, and glass. However, this difference was not significant enough to warrant reporting. In TSBG, higher biofilm formation was found on stainless steel than on polypropylene and polystyrene. Our results indicated that K. pneumoniae cells adhered very rapidly to hydrophilic surfaces more than to hydrophobic surfaces on TSB. Similarly, in TSBG, K. pneumoniae biofilm formation was higher in hydrophilic surface except the glass surfaces. Blackman et al. studied that a certain kind of enrichment medium used has a massive effect on the accumulation of biofilm due to alteration of cell surface components.16 Subsequently, the progression of biofilm and cell adhesion could be affected by glass tubes exposed to disparate solutions. This study also revealed that the different types of media had an impact on biofilm development, which is in accordance with previous findings and in addition, it illustrates that substrate variation is pivotal in the establishment of biofilm.4,17 Cepas et al. examined whether there was a correlation between antibiotic resistance and the ability to produce biofilm in gram-negative bacteria and revealed that there was no clinically important association between general MDR and biofilm formation because MDR strains have not shown a higher predisposition to form biofilm than non-MDR strains.13
Based on the present study, it is recommended that efforts are needed to prevent this resistant strains from spreading and require new therapies for this pathogens. Furthermore, biofilm development on diverse surfaces as evaluated by the CV method varies among K. pneumoniae strains and nutritional circumstances like supplementation of glucose. However, the findings observed by the CV staining approach did not reveal the accurate population of K. pneumoniae in biofilm surfaces, and in addition CV stain is not environmental friendly, light sensitive and hazardous to human health implying that more research is needed to develop an appropriate environmental friendly tool for estimating biofilm formation levels and to investigate biofilm formation across the diverse material surfaces.
ACKNOWLEDGMENTS
The authors would like to express gratitude to Bharathidasan University for allowing to conduct research.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
KS designed the work. LSB performed the experiments, generated data and wrote the manuscript. BAM helped in generating the data and writing the manuscript. KS edited and approved the manuscript for publication.
FUNDING
None.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
This article does not contain any studies on human participants or animals performed by any of the authors.
- Stoodley P, Sauer K, Davies DG, Costerton JW. Biofilms as complex differentiated communities. Annu Rev Microbiol. 2002;56:187-209.
Crossref - Hoiby N, Ciofu O, Johansen HK, et al. The clinical impact of bacterial biofilms. Int J Oral Sci. 2011;3(2):55-65.
Crossref - Nicolle LE. Catheter associated urinary tract infections. Antimicrob Resist Infect control. 2014;3:23.
Crossref - Rode TM, Langsrud S, Holck A, Moretro T. Different patterns of biofilm formation in Staphylococcus aureus under food-related stress conditions. Int J Food Micro. 2007;116(3):372-383.
Crossref - O’Toole G, Kaplan HB, Kolter R. Biofilm formation as microbial development. Annu Rev Microbiol. 2000;54:49-79.
Crossref - Jo SH, Baek SB, Ha JH, Ha SD. Maturation and survival of Cronobacter biofilms on silicone, polycarbonate, and stainless steel after UV light and ethanol immersion treatments. J Food Prot. 2010;73(5):952-956.
Crossref - Kim H, Bang J, Beuchat LR, Ryu JH. Fate of Enterobacter sakazakii attached to or in biofilms on stainless steel upon exposure to various temperatures or relative humidities. J Food Prot. 2008;71(5):940-945.
Crossref - Singh VK, Singh R, Kumar A, Bhadouria R, Singh P, Notarte KI. Antibiotics and Antibiotic Resistance Genes in Agroecosystems as Emerging Contaminants. Sustainable Agriculture Reviews. 2020;50:177–210.
Crossref - Indrajith S, Mukhopadhya AK, Chowdhury G, et al. Molecular insights of carbapenem resistance Klebsiella pneumoniae isolates with focus on multidrug resistance from clinical samples. J Infect Public Health. 2021;4(1):131-138.
Crossref - Wayne PA, Performance standards for antimicrobial disk susceptibility testing. Clinical and Laboratory Standard Institute, 28th Ed. CLSI, USA, 2018;38(3).
- Christensen GD, Simpson WA, Bisno AL, Beachey EH. Adherence of slime-producing strains of Staphylococcus epidermidis to smooth surfaces. Infect Immun. 1982;37(1):318-26.
Crossref - Lee JS, Bae YM, Lee SY, Lee SY. Biofilm Formation of Staphylococcus aureus on Various Surfaces and Their Resistance to Chlorine Sanitizer. J Food Sci. 2015;80(10):M2279-86.
Crossref - Cepas V, Lopez Y, Munoz E, et al. Relationship between biofilm formation and antimicrobial resistance in gram-negative bacteria. Microb Drug Resist. 2019;25(1):72-79.
Crossref - GM N, CM G Nagshetty K, et al. Antibiotic susceptibility pattern of ESbetaL producing Klebsiella pneumoniae isolated from urine samples of pregnant women in Karnataka. J Clin Diagn Res. 2014;8(10):DC08-11.
Crossref - Moini AS, Soltani B, Ardakani AT, et al. Multidrug-resistant Escherichia coli and Klebsiella pneumoniae isolated from patients in Kashan. Iran Jundishapur J Microbiol. 2015;8(10): 27517.
Crossref - Blackman IC, Frank JF. Growth of Listeria monocytogenes as a biofilm on various food-processing surfaces. J Food Prot. 1996;59(8):827-831.
Crossref - Hood SK, Zottola EA. Biofilms in food processing. Food Control. 1995:6(1); 9-8.
Crossref
© The Author(s) 2022. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.