ISSN: 0973-7510
E-ISSN: 2581-690X
The goal of this research is on antibiotic resistance and biofilm formation of Escherichia coli on different surfaces. 37 E.coli isolates were obtained from K.A.P. Viswanatham Government Medical College, Tiruchirappalli, Tamil Nadu, India. Biochemical assays were used to re-confirm all the isolates. Ampicillin, Cefepime, Cefotaxime, Co-trimoxazole, Tetracycline and Levofloxacin showed substantial levels of resistance. Meropenem, Tigecycline, and Colistin showed the least amount of resistance. 75.6% of the E.coli strains were multidrug resistant (MDR). Biofilm formation of E.coli was higher in TSBG than in TSB in all (polystyrene, polypropylene, glass and stainless steel) surfaces. It is evident that the presence of glucose or any sugar substrate promotes biofilm development, resulting in notable antibiotic resistance. This situation is hazardous to human health.
E.coli, Antibiotic Resistance, Biofilm, Surfaces
E. coli is a gram negative, facultative anaerobic, versatile bacterium, mostly found in the lower intestine of humans and endothermic animals. They colonize in the host and in the environment.1 They are the first bacterial species to colonize the gut right after birth.2 Most strains of E. coli are usually beneficial and this commensal E. coli hardly causes disease excluding immunocompromised host, and conditions such as in peritonitis.3 It causes pneumonia, appendicitis, gastrointestinal infections, septicemia, and meningitis.4 Antibiotic resistance is a potentially significant threat to human health worldwide, over the last few decades.2 It is expected that, by 2050, E. coli infections might contribute to 30% of 10 million multi-drug infections annually.5 World Health Organization (WHO) had declared that antimicrobial resistance (AMR) is one among the top ten public health threats globally and death due to AMR will increase from 7,00,000 to 10 million per year globally.6,7. Antimicrobial resistance occurs when there is a high increase of resistance to antibiotics by the bacteria. E. coli has the ability to colonize various environments and this led to an advantage of acquiring antibiotic resistance from other bacteria within its environment.2
Another major issue in healthcare is formation of biofilm, which contributes to 65% of bacterial infections. E. coli is a complex microbe which also leads to diarrhoea, biliary and urinary tract infections in humans and in animals.8,9 Few of these infections are suspected to prevail as biofilm-associated infections.10 Moreover, E. coli biofilms play a major role in other serious human diseases like prostatitis, and catheter-associated infections and otitis media.11-13 Biofilms are aggregates in which cells are ingrained within a matrix of extracellular polymeric substance adhere to each other or to an inert or living surface.4,14 It leads to an increase in antibiotic resistance by allowing cells to persist. Antibiotic resistance determinants are transferrable to other isolates of E. coli, also to other bacteria within the gastrointestinal tract and to acquire resistance from other organisms.15 A detailed study of biofilm helps in developing methodologies to withstand biofilm formation. Various studies disclose the connection between biofilm formation and antibiotic resistance. Studies on quantification of biofilm formation on various surfaces are still not sufficient. The present study was undertaken to screen antibiotic susceptibility, and biofilm formation of E. coli on various surfaces.
Collection of Strains
37 clinical strains were collected from K.A.P. Viswanatham Government Medical College, Tiruchirappalli, Tamil Nadu, India. All bacterial isolates were subjected to a series of confirming tests.
Confirmation of Bacterial Isolates
The collected strains were grown on tryptic soya agar (TSA) (Himedia, India) for overnight at 37°C and transferred to MacConkey agar to study the morphological characterization and finally proceeded for the biochemical confirmation analysis.16
Antibiotic Susceptibility Test
All confirmed E. coli strains were tested for their antibiotic resistance patterns by Kirby-Bauer disc diffusion method on Muller-Hinton agar, as per the guidelines of Clinical and Laboratory Standards Institute (CLSI) (2018).17 The antibiotics used in this study were Ampicillin (AMP, 10mcg), Cefepime (CPM, 30 mcg), Cefotaxime (CTX, 30 mcg), Co-Trimoxazole (COT, 25 mcg), Tetracycline (TE, 30 mcg), Levofloxacin (LE, 5 mcg), Gentamycin (GEN, 10 mcg), Imipenem (IPM, 10 mcg), Ertapenem (ETP, 10 mcg), Meropenem (MRP, 10 mcg), Tigecycline (TGC, 30 mcg), Colistin (CL, 10 mcg), and Doripenem (DOR, 10 mcg). Overnight grown bacterial cultures were inoculated into 0.85% NaCl and the suspension was adjusted to equivalent of 0.5 McFarland Standard. Sterile cotton swabs were dipped in the suspension and swabbed on the Muller Hinton Agar plate (MHA). The antibiotic discs were then placed aseptically with sterile forceps on swabbed MHA plates. The plates were then incubated for 24 h at 37°C. The guidelines provided by Clinical Laboratory Standards Institute (CLSI, 2018) were used to evaluate the patterns as “Sensitive”, “intermediate resistant” or “resistant”.17
Quantification of Biofilm Production on Various Surfaces
Biofilm formation on glass and polypropylene tubes in tryptic soya broth (TSB) and tryptic soya broth with 1% glucose (TSBG).
The study was carried out using tube method. Each of the selected E. coli strains was cultured individually in 10 ml of TSB and TSBG (Hi-media, India) in glass and polypropylene tubes for 24 h at 37°C. The glass tubes were decanted, rinsed multiple times with sterile water, and air dried. Then the tubes were stained with 100 μl of 0.5% Crystal Violet (CV) (Hi-media, India). Excess stain was destained by rinsing with sterile distilled water. Tubes were dried in an inverted position and noticed for biofilm formation. The excess stain on biofilm cells were removed by washing with 1 ml of 99% ethanol, and then 100 μl was transferred to a cuvette.18 The Optical Density (OD) of each well was estimated at 595 nm in a spectrophotometer (Cary – 60 UV-Vis, Agilent Technologies, USA).
Biofilm Formation on Polystyrene Plates in Tryptic Soya Broth (TSB) and Tryptic Soya Broth with 1% Glucose (TSBG)
Each of the E. coli strains was inoculated individually in TSB and TSBG (Himedia, India), and grown for 24 h at 37°C. Sterile 96-well polystyrene plates (Tarsons, India) were loaded with 90 μl of TSB and TSBG, and inoculated with 10 μl of bacterial culture. Plates were then incubated at 37°C for 24 h. After incubation, the plate was kept upside down and the medium was discarded. The wells were rinsed thrice with sterile water and dried. Wells were then stained using 50 μl of 0.5% crystal violet for 5 min. The wells were washed with distilled water to remove excess stains. The stained adherent cells were decolorized using 50 μl of 99% ethanol. The OD of individual wells was estimated at 595 nm in a spectrophotometer18 (Cary – 60 UV-Vis, Agilent Technologies, USA).
Biofilm Formation on Stainless Steel in TSB and TSBG
All E. coli strains were inoculated in TSB and TSBG (Hi-media, India), and grown for 24 h at 37° C. In a glass tube with stainless steel coupons, 1 ml of E. coli was inoculated into 10 ml of sterile phosphate buffer saline (pH – 7.4), and incubated for 2 h at 37°C. After 2 h of incubation, the coupons were air dried and submerged into TSB and TSBG. The cells adhered on the stainless steel coupons were incubated at 37°C for 24 h. After 24 h of incubation, the coupons were washed thrice with sterile water and dried. Then, 1 ml of 0.5% crystal violet was used to stain the coupons. Excess stain was removed by washing with sterile water. Then it was decolorized using 1 ml of 99% ethanol, and the OD was taken at 595 nm in a spectrophotometer18 (Cary – 60 UV-Vis, Agilent Technologies, USA).
The confirmational test of E. coli was performed on 37 clinical strains. Table shows the percentage of distribution of E. coli from various sources, among which Urine had high number (59.4%) followed by, Pus (29.7%), Blood (5.4%), Sputum (2.7%), and Vaginal swab (2.7%).
Table:
Distribution of E.coli collected from different sources.
Source |
Total number of isolates (n=37) |
% of E.coli distribution (%) |
|---|---|---|
Urine |
22 |
59.4 |
Pus |
11 |
29.7 |
Blood |
02 |
5.4 |
Sputum |
01 |
2.7 |
Vaginal swab |
01 |
2.7 |
Antibiotic Resistance Pattern
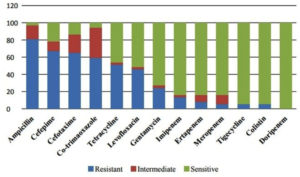
The antibiotic resistant patterns of E. coli from different clinical samples are presented in Figure 1. From the results, it is clear that all the strains showed resistance to at least one antibiotic except Doripenem for which all strains were sensitive. Among the antibiotics tested, E. coli isolates exhibited maximum resistance against Ampicillin (81%), followed by Cefepime (67.5%), Cefotaxime (64.86%), Co-Trimoxazole (56.7%), Tetracycline (51.3%), Levofloxacin (51.3%), Gentamycin (24.3%), Imipenem (10.8%), Ertapenem (8.1%), Meropenem (5.4%), and Tigecycline (5.4%) and Colistin (5.4%).
Biofilm Formation of E. coli on Different Surfaces
Quantification of Biofilm Formation on Glass and Polypropylene Tubes in Tryptic Soya Broth (TSB) and Tryptic Soya Broth with Glucose (TSBG)
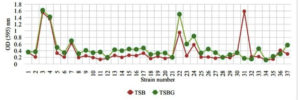
Biofilm formation on glass surface with TSB and TSBG is presented in Figure 2. From strains 1 to 16, strains 1 and 2 showed slow growth, whereas strains 3 and 4 showed a high growth. Strains 5 and 6 exhibited declined growth and strain 7 had slight increase in growth. From strain 8 to strain 21, there was a fluctuation in growth both in TSB and TSBG. Strain 22 showed good growth both in TSB and TSBG. With a decline in strain 23, strain 24 showed a slight increase in the growth. Strains 25 to 37 showed growth fluctuations both in TSB and TSBG. Strains 3 and 31 exhibited highest biofilm formation in TSB whereas, strains 3 and 22 showed highest biofilm production in TSBG on glass surface.
In all 37 strains, growth was greater in TSBG than in TSB, except the strain 31. It was observed that addition of glucose to TSB enhanced the growth on glass surface.
Polypropylene Surface
Biofilm formation on polypropylene surface with TSB and TSBG is presented in Figure 3. All 37 strains exhibited growth both in TSB and TSBG. In TSB and TSBG, strains 1, 3, 7, 8, 9, 10, 11, and 12 showed no much growth, whereas strains 2, 5, 6, 13, 14, 15, and 16 showed a mild increase in their growth. From strain 17 to strain 37, it is evident that strains 18, 19, 21, 22, 25, and 26 showed poor growth whereas strains 17, 20, 23, 24 showed a better growth than the other strains. Strain 27 exhibited a moderately high growth and there was a sudden decline in growth for the strains 28, and 29. Again, there was a high growth in strains 30, and 32, in which strain 31 had a slight decline in growth. In all 37 strains, growth was higher in TSBG than in TSB, but the difference of growth in E. coli was not much difference between TSB and TSBG. It was observed that addition of glucose to TSB enhanced the growth of biofilm on polypropylene surface.
Polystyrene Surface
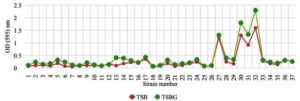
Biofilm formation on polystyrene surface with TSB and TSBG is presented in Figure 4. Strain 1 remained unchanged. From strains 2 to 25, there was less growth on biofilm with fluctuations both in TSB and TSBG, except strain 19. Strain 26 remained unchanged and strain 27 showed highest biofilm formation in polystyrene surface in TSBG. In TSB, strains 27 to 37 showed no appreciable increase in growth with mild fluctuations.
In all 37 strains, growth was higher in TSBG than in TSB. But the difference of growth in E. coli on polystyrene surface was not much between TSB and TSBG, except for the strain 27. It was observed that addition of glucose to TSB enhanced the growth on polystyrene surface.
Stainless steel Surface
Formation of biofilm on stainless steel surface with TSB and TSBG is presented in Figure 5. Both in TSB and TSBG, strains 1 to 5 showed minimum growth. Strains 6 and 7 exhibited highest growth level both in TSB and TSBG. From strains 8 to 23, there was continuous fluctuation both in TSB and TSBG. Strains 24 to 33 showed a substantial increase in growth, whereas in strains 28 and 29 high growth was noted. From strains 34 to 37 there was a decline in growth of biofilm. In TSB, strain 7 had the highest growth. In TSBG, strains 6 and 7 had the maximum level of growth. In all 37 strains, growth was higher in TSBG than in TSB. But the difference of growth in E. coli on stainless steel surface was not much between TSB and TSBG. It was observed that addition of glucose to TSB enhanced the growth on stainless steel surface.
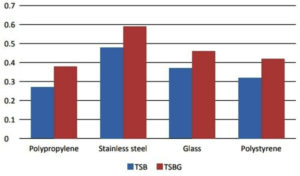
Comparative studies of biofilm formation on different surfaces (polystyrene, polypropylene, glass and stainless steel) using TSB and TSBG (Himedia, India) as quantified by crystal violet staining method are presented in Figure 6. In TSB and TSBG on polypropylene surface, one strain (strain 32) was found to form high biofilm formation. Three strains (strains 3, 22 and 31) showed highest biofilm formation in glass surface in TSB whereas, in TSBG, two strains (strains 3, and 22) showed highest biofilm formation. Only one strain (strain 19) expressed the high level of biofilm formation both in TSB and TSBG in polystyrene surface. However, strains 6 and 7 were found to be high in biofilm formation both in TSB and TSBG in stainless steel surface.
As a result, there was no association between the strains that produced maximum level of biofilm on different surfaces. From this data, it is noted that 60% of the strains which produced highest biofilm was from urine source and 40% of the strains were from pus source. In terms of the influence of the surface on biofilm production, steel outperformed other surfaces followed by glass, polystyrene and polypropylene. This difference, however, was not big enough to be noticed. Overall, biofilm formation in TSBG was higher than in TSB in all surfaces. It is known from this study, that the presence of glucose or any sugar substrate promotes biofilm development, resulting in significant resistance to antibiotics. The results of this work indicate that the ability of biofilm formation on various surfaces was found to be different among strains.
Out of 37 tested E. coli isolates, 81% were resistant to ampicillin. For Doripenem, 100% of the strains were sensitive. From this study, 75.6% of the isolates were found to be multi-drug resistant E. coli. Majority of the MDR E. coli strains were also resistant to fluoroquinolones, sulfonamides, and tetracyclines. AMR, according to the (WHO) World Health Organization (2021), is occurring all throughout the planet, jeopardising humans’ ability to manage infectious diseases, as well as compromising along with other health and medical advancements.7 Mukherjee et al. have reported in their study that multi-drug resistant E. coli showed 97.5% resistance to ampicillin and also, 37 out of 40 strains (92.5%) were multi-drug resistant.19 Antibiotic resistance pattern varies between different geographic regions. Odonkor and Addo et al. concluded from their study that the prevalence of multi-drug resistant E. coli was 49.48%.20 Similarly, Singh et al. reported that inappropriate usage of antibiotics leads to multi drug resistance by various pathogens.21
In the present study, MDR E. coli strains exhibited high resistance to ampicillin. This has been linked to the over usage of ampicillin drug in humans. Third-generation cephalosporins are used to cure many body site infections caused by gram negative bacteria.22 The current study exhibited maximum level of resistance to third generation cephalosporins such as Cefepime (67.5%) and Cefotaxime (64.86%). Multi-drug resistance of E. coli to three or more unrelated families of antibiotics was already reported by others,23-25 and it’s a major health threat. Combination of antibiotics has been proven in some studies to prevent the formation of new resistant strains.
Biofilm formation of E. coli on various types of materials was quantified using the crystal violet staining method in TSB and TSBG. It was observed from this study that biofilm formation of E. coli was greater in TSBG than in TSB in all surfaces. Hence, glucose was found to enhance the biofilm production in TSB. Also, there was no association observed between the strains producing highest biofilm. In terms of the influence of the surface on biofilm production, stainless steel surface had the highest biofilm than rest of the surfaces. Wilks et al. had stated that E. coli cells survived for 28 days at 4°C and in room temperature on stainless steel surface.26 Our study also showed that E. coli attached effectively to hydrophilic surface like stainless steel than hydrophobic surface. The exposure of glass tubes to different growth media might have brought changes in their cell surface chemistry and minimized the adhesion process of E. coli cells to glass surface, which is hydrophilic. Also, Stainless steel lacks chemical properties that destroys the bacteria (Packiyam et al.)27 Hence, the capable pathogens can colonize and multiply on stainless steel surfaces. Thus, stainless steel has the potential to serve as an infection reservoir (Solheim et al.)28 Alotaibi and Bukhari had reported that hydrophobicity, temperature, pH and culture medium, and also concentration of microorganism are factors that may affect adhesion process of biofilm formation.29 Overall, biofilm formation in TSBG was higher than in TSB in all surfaces. It is concluded from this study, that the presence of glucose or any sugar substrate enhances biofilm development, which may result in remarkable antibiotic resistance. In certain gram negative bacteria, the acquisition of particular antimicrobial resistance can compromise or improve biofilm formation.30 Multidrug resistant strains, on the other hand, do not appear to form more biofilm than non-MDR. Biofilm production was found in both MDR and non – MDR strains. From this study, 28 strains (75.6%) were found to be multi drug resistant (MDR) and 09 strains (24.3%) were non-multi drug resistant (Non-MDR). Cepas et al. reported that there was no relationship between multi drug resistance and biofilm formation.30 Therefore, MDR strains of E. coli did not tend to have more biofilm production than the non-MDR. Hence, understanding the pathogenesis associated with biofilm formation aids in therapeutic development.
Increase in multiple resistance to antimicrobial agents is a major challenge in the development of therapeutic approaches. Hence, more studies are needed to clarify the association between antimicrobial resistance and biofilm formation and quantify the levels of bacterial biofilm on various surfaces.
ACKNOWLEDGMENTS
The authors would like to thank Bharathidasan University for allowing us to conduct our research work.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
KP supervised and designed part of the work. KS designed and monitored the work. BAM performed the experiment and generated the data. LSB helped in generating data. BAM wrote the manuscript. LSB helped in manuscript writing. KS edited and approved the manuscript.
FUNDING
None.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
Not applicable.
- Braz VS, Melchior K, Moreira CG. Escherichia coli as a multifaceted pathogenic and versatile bacterium. Front Cell Infect Microbiol. 2020;10:548492.
Crossref - Galindo-Mendez M. Antimicrobial Resistance in Escherichia coli. E. Coli. Infections-Importance of Early Diagnosis and Efficient Treatment, 2020;1-20.
Crossref - Kaper JB, Nataro JP, Mobley HL. Pathogenic Escherichia coli. Nat Rev Microbiol. 2004;2(2):123-140.
Crossref - Kumar A, Leite AFV, Maekawa LS, et al. Management of E. coli Sepsis. In E. Coli Infections-Importance of Early Diagnosis and Efficient Treatment. IntechOpen. 2020.
Crossref - Reid CJ, Wyrsch ER, Chowdhury PR, et al. Porcine commensal Escherichia coli: a reservoir for class 1 integrons associated with IS26. Microb Genom. 2017;3(12):e000143.
Crossref - De Kraker ME, Stewardson AJ, Harbarth S. Will 10 million people die a year due to antimicrobial resistance by 2050? PLoS Med. 2016;13(11):e1002184.
Crossref - World Health Organization. Antimicrobial Resistance: a top ten global public health threat. EClinical Medicine.2021;41:101221.
Crossref - Johnson JR, Stell AL. Extended virulence genotypes of Escherichia coli strains from patients with urosepsis in relation to phylogeny and host compromise. J Infect Diseases. 2000;181(1):261-272.
Crossref - Moulin-Schouleur M, Schouler C, Tailliez P, et al. Common virulence factors and genetic relationships between O18: K1: H7 Escherichia coli isolates of human and avian origin. J Clin Microbiol. 2006;44(10):3484-3492.
Crossref - Reisner A, Krogfelt KA, Klein BM, Zechner EL, Molin S. In vitro biofilm formation of commensal and pathogenic Escherichia coli strains: impact of environmental and genetic factors. J Bacteriol. 2006;188(10):3572-3581.
Crossref - Naves P, Del Prado G, Huelves L, et al. Measurement of biofilm formation by clinical isolates of Escherichia coli is method-dependent. J Appl Microbiol. 2008;105(2):585-590.
Crossref - Ren D, Zuo R, Barrios AFG, et al. Differential gene expression for investigation of Escherichia coli biofilm inhibition by plant extract ursolic acid. Appl Environ Microbiol. 2005;71(7):4022-4034.
Crossref - Sharma D, Misba L, Khan AU. Antibiotics versus biofilm: an emerging battleground in microbial communities. Antimicrob Res Infect Control. 2019;8(1):1-10.
Crossref - Piperaki ET, Syrogiannopoulos GA, Tzouvelekis LS, Daikos GL. Klebsiella pneumoniae: virulence, biofilm and antimicrobial resistance. Pedi Infect Disease J. 2017;36(10):1002-1005.
Crossref - Rasheed M U, Thajuddin N, Ahamed P, Teklemariam Z, Jamil K. Antimicrobial drug resistance in strains of Escherichia coli isolated from food sources. Rev Inst Med Trop Sao Paulo. 2014;56(4):341-346.
Crossref - Indrajith S, Mukhopadhyay AK, Chowdhury G, et al. Molecular insights of carbapenem resistance Klebsiella pneumoniae isolates with focus on multidrug resistance from clinical samples. J Infect Public Health. 2021;14(1):131-138.
Crossref - Wayne, PA. Clinical Laboratory Standards Institute (CLSI) Performance standards for antimicrobial disk susceptibility tests. Document M100S, 28th Ed., CLSI Vol – 38 No.3, Fourteenth informational supplement, USA. 2018
- Lee JS, Bae YM, Lee SY, Lee SY. Biofilm formation of Staphylococcus aureus on various surfaces and their resistance to chlorine sanitizer. J Food Sci. 2015;80(10):M2279-M2286.
Crossref - Mukherjee M, Basu S, Mukherjee SK, MajuMder M. Multidrug-resistance and extended spectrum beta-lactamase production in uropathogenic E. coli which were isolated from hospitalized patients in Kolkata, India. J Clin Diagn Res: JCDR. 2013;7(3):449.
Crossref - Odonkor ST, Addo KK. Prevalence of multidrug-resistant Escherichia coli isolated from drinking water sources. Int J Microb. 2018;2018:7204013.
Crossref - Singh VK, Singh R, Kumar A, Bhadouria R, Singh P, Notarte KI. Antibiotics and Antibiotic Resistance Genes in Agroecosystems as Emerging Contaminants. Sustainable Agriculture Reviews. 2021;50:177-210. Springer.
Crossref - Abd El-Baky RM, Ibrahim RA, Mohamed DS, Ahmed EF, Hashem ZS. Prevalence of virulence genes and their association with antimicrobial resistance among pathogenic E. coli isolated from Egyptian patients with different clinical infections. Infect Drug Res. 2020;13:1221-1236.
Crossref - Oteo J, Lazaro E, de Abajo FJ, Baquero F, Campos J, Spanish members of EARSS. Spanish members of EARSS. Antimicrobial-resistant invasive Escherichia coli, Spain. Emerg Infect Dis. 2005;11(4):546-553.
Crossref - Bartoloni A, Pallecchi L, Benedetti M, et al. Multidrug-resistant commensal Escherichia coli in children, Peru and Bolivia. Emerg Infect Dis. 2006;12(6):907-913.
Crossref - Sahm DF, Thornsberry C, Mayfield DC, Jones ME, Karlowsky JA. Multidrug-resistant urinary tract isolates of Escherichia coli: prevalence and patient demographics in the United States. Antimicrob Agents Chemother. 2001;45(5):1402-1406.
Crossref - Wilks SA, Michels H, Keevil CW. The survival of Escherichia coli O157 on a range of metal surfaces. Int J Food Microbiol. 2005;105(3):445-454.
Crossref - Packiyam R, Kananan S, Pachaiyappan S, Narayanan U. Effect of storage containers on coliforms in household drinking water. Int J Curr Microbiol App Sci. 2016;5(1):461-477.
Crossref - Solheim HT, Sekse C, Urdahl AM, Wasteson Y, Nesse LL. Biofilm as an environment for dissemination of stx genes by transduction. Appl Environ Microbiol. 2013;79(3):896-900.
Crossref - Alotaibi GF, Bukhari MA. Characterization and evaluation of biofilm formation by. Klebsiella pneumonia MBB9 isolated from epilithic biofilms of the porter brook river, Sheffield. Edel J Biomed Res Rev. 2021;3:14-24.
Crossref - Cepas V, Lopez Y, Munoz E, et al. Relationship between biofilm formation and antimicrobial resistance in gram-negative bacteria. Microb Drug Res. 2019;25(1):72-79.
Crossref
© The Author(s) 2022. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.