ISSN: 0973-7510
E-ISSN: 2581-690X
This work attempts to assess the antimicrobial potential of actinobacteria isolated from limestone mining sites which hitherto, is an under-explored niche for exploring novel bioactive metabolites. Actinobacteria were selectively isolated from Mawsmai, Meghalaya, India, a limestone mining area, using different pretreatment methods. Forty-seven isolates were obtained, which were identified based on their morphological, biochemical and chemotaxonomical characteristics. Streptomyces was the dominant cultivable genera which constituted 76% of the isolates cultivated. All the isolates were screened for antimicrobial activity against three Gram-negative viz. Escherichia coli, Pseudomonas aeruginosa and Klebsiella pneumoniae, and three Gram-positive bacteria viz. Staphylococcus aureus, Bacillus subtilis and Micrococcus luteus and besides, two candidal species viz. Candida albicans and C. tropicalis. 19% of the total isolates showed antibacterial activity against at least one of the test bacterial strains used. The identity of the four bioactive isolates viz. LD-21, LD-29, LD-34 and LD-39 was confirmed as Streptomyces sp. on the basis of their 16S rDNA sequence and 16S rRNA secondary structure analysis. These isolates showed antibacterial activity against at least two Gram-positive bacteria and all the four harbored at least one of the three biosynthetic gene clusters viz. type-I and type-II polyketide synthases and non-ribosomal peptide synthetase which are related to synthesis of bioactive metabolites.
Streptomyces, Antimicrobial, Biosynthetic gene clusters, 16S rRNA secondary structure.
Actinobacteria is a phylum of Gram-positive or Gram-variable bacteria with diverse morphology ranging from cocci to highly differentiated mycelia1. They have high G+C content in their DNA ranging from 50-70 mol %1. They are widely distributed in all kinds of terrestrial and aquatic environments1. Most of them are saprophytic, however, some are pathogenic to plants and animals1. Actinobacteria has been one of the important sources of naturally-derived antibiotics for the last 75 years and will remain so in the years to come especially for the discovery of novel antibiotics2. Antibiotics have also been discovered from various other natural sources like plants, animals, fungi and bacteria3. Naturally-derived antibiotics, even though are more difficult to discover due to complexity in identifying new scaffolds4, are superior over the synthetic ones because they are products of natural selection and have gone through the long process of evolution and have established compatibility with biological targets4. More than two-thirds of clinically used antibiotics were either natural products or their semi-synthetic derivatives5. However, in recent years, there has been some lag in discovering new classes of natural product antibiotics and this has put a pressure on the pharmaceutical industry due to the emergence of resistance amongst the pathogens. Reasons like rediscovery of known compounds, complexity of the natural antibiotic scaffolds and their suitability as drugs, their unstability, very low yield and difficulty in purification have constrained pharmaceutical companies to shift their interests to synthetic antibiotics in the 21st century4 to stay ahead of emerging antibiotic-resistance. Recent advancement in new generation genome sequencing technology, bioinformatics, analytical chemistry, metagenomics and genome-scale metabolic models4–7 has now impelled the researchers to return to natural products for drug discovery. Search for microbial taxa from under-explored ecological niches increases the possibility of discovering novel molecules from nature5. Limestone-rich areas mainly dominated by a CaCO3 environment, are under-explored ecological niches for mining of microorganisms for various metabolic properties. Recent studies have revealed that both hypogean and epigean limestone actinobacteria are potential antibiotics producers8–14. The state of Meghalaya located in the north-eastern part of India falls under the Indo-Burma Biodiversity Hotspot15. Therefore, mining for antibiotics producing actinobacteria from under-explored ecological niche of limestone mines of diversity-rich state of Meghalaya enhances the possibility of discovery of novel metabolites. This paper, therefore, reports on diversity of actinobacteria having antibacterial activities from limestone mining areas located at Mawsmai, Meghalaya, India.
Sample collection
The sampling site was from a limestone mining area at Mawsmai near Sohra in Meghalaya, India (N25o15.364′ E91o43.885′). Soil samples from around the mining area were aseptically collected in sterilized polythene bags and brought to the lab and stored at 4±1oC until further use.
Determination of soil physico-chemical characteristics
Soil and air temperatures (in oC) were measured in three different locations using a soil thermometer and an ordinary thermometer respectively. Moisture content (%) of the soil was determined gravimetrically16. pH was determined using a digital pH meter17. Organic carbon was determined following the method of Anderson and Ingram (1993)17. Total nitrogen was determined by the Kjeldahl Method18. Available Phosphorus was determined by the molybdate blue method16. Potassium was determined by the flame photometer method18.
Soil pretreatment and isolation of actinobacteria
The collected soil samples were air-dried at room temperature for one week and mixed equally into one composite sample. The air-dried samples were then pretreated by (i) dry heating at 120oC for 1 hour, (ii) wet heating at 70oC for 15 minutes in a water bath, (iii) 1.5% phenol at 25±1oC for 30 minutes, (iv) 0.2% humic acid at 25±1oC for 30 minutes and (v) combination of i & iii. The unpretreated air-dried soil samples were also used as inoculum for isolation. 1g pretreated and unpretreated soil samples were suspended in 9ml sterilized distilled water and serially diluted. All the undiluted and serially diluted soil sample suspensions were used as inocula for isolation. Five selective media were used for isolation viz. Actinomycete Isolation Agar (AIA), Bennett’s Agar (BA), Starch Casein Agar (SCA), Streptomyces Agar (SA) and Humic acid Vitamin Agar (HVA)19. The above media were amended with Nystatin (50µg ml-1) and Rifampicin (20µg ml-1). 100µl of the soil sample suspensions were inoculated on the above selective media and incubated at 28±1oC for up to four weeks. Based on colony morphology, colonies were picked from the culture plates, sub-cultured and maintained in Bennett’s agar medium. The recovery of actinobacterial colonies from different pretreated samples and media were recorded as actinobacterial population number in cfu g-1 soil. All observations were recorded in triplicates and expressed as sample mean ± standard error. Multiple comparisons of sample means were done using ANOVA with Duncan’s test at p £ 0.05. Statistical analysis was performed with XLSTAT software.
Morphological characterization
The selected colonies were grown in ISP3 (oat meal agar) medium by coverslip culture method20 and incubated at 28±1oC for 7 days. The coverslip cultures were then placed on a glass slide and observed under the microscope (Olympus CX21i) at 400X magnification for presence of aerial and substrate mycelia, spore chains and other reproductive structures.
Biochemical and chemotaxonomic characterization
Biochemical tests viz., casein, xanthine, hypoxanthine and tyrosine hydrolysis were performed following the methods of Berd (1973)21. Chemotaxonomic analyses viz. cell wall chemotype of 2,6-diaminopimelic acid (DAP) and whole cell sugars were performed by thin-layer chromatography following the methods of Hasegawa et al. (1983)22. The scheme for preliminary identification of actinobacteria was followed according to Berd (1973)21 and Staneck and Roberts (1974)23.
Phylogenetic analysis of 16S rDNA sequence
Genomic DNA of the selected bioactive isolates were extracted by enzymatic method24 followed by PCR of 16S rDNA. The primers used for amplification were 27F (5’-AGAGTTTGA-TCMTGGCTCAG-3’) and A3R (5’-CCAGCCCCACCTT-CGAC-3’)25. PCR reactions were carried out in an Eco96TM thermal cycler (Himedia, India) in 25µl reaction mixture containing PCR buffer (Promega) with 1.5mM MgCl2, 0.2mM of each dNTP (Promega), 0.5µM of each primer (IDT), 0.625U of Taq DNA polymerase (Promega) and 5µl of extracted DNA as template DNA. Amplification parameter was an initial denaturation of 95oC for 5 minutes, 35 cycles of 95oC for 45 seconds, 56oC for 60 seconds, 72oC for 90 seconds and final extension of 72oC for 7 minutes. The amplified products were analyzed by agarose gel electrophoresis (with 1.2% w/v agarose added with 1µl/10ml v/v LabSafeTM Nucleic Acid Stain, G-Biosciences). A 100bp DNA ladder was used for size reference. The amplified products were purified and sequenced at Xcelris Labs Ltd., Ahmedabad, India. The partial 16S rDNA sequences obtained were matched with type strains from EzBioCloud database26 for determining sequence similarities. Phylogenetic relationships of the selected bioactive isolates along with thirty-two related type strains were determined using MEGA version 727 and phylogenetic tree was constructed using the Maximum Likelihood method based on the Tamura 3-parameter model28. A discrete Gamma distribution was used to model evolutionary rate differences among sites (+G). The rate variation model allowed for some sites to be evolutionarily invariable (+I). Bootstrap analysis29 was done with 1000 replicates. The partial 16S rDNA sequences were deposited in GenBank.
Phylogenetic analysis of predicted 16S rRNA secondary structure
The use of 16S rRNA secondary structures for phylogenetic analysis of actinobacteria was understood from Ghosh et al. (2019)30. The 16S rDNA sequences obtained were converted into 16S rRNA sequences and subsequently the 16S rRNA secondary structures with their minimum free energy (MFE) were predicted using the RNAfold server31 at the ViennaRNA Web Services (http://rna.tbi.univie.ac.at/). The obtained 16S rRNA secondary structures of the selected bioactive isolates along with their MFEs were then compared with the related type strains from EzBioCloud database (which were used for phylogenetic analysis based on 16S rDNA sequence) to confirm their identity. Consensus 16S rRNA secondary structures were also generated using the RNAalifold server32 at the ViennaRNA Web Services between the bioactive isolates and the related type strains with the closest MFEs to analyze for conserved regions. The selected 16SrDNA sequences with the closest MFEs were aligned using ClustalX 2.1 (http://www.clustal.org/download/2.1/) and sequence overhangs trimmed before using the aligned file for generating consensus 16S rRNA secondary structure.
Screening of isolates for antimicrobial activity
Isolates were screened for antimicrobial activity against Escherichia coli MTCC 1669, Pseudomonas aeruginosa MTCC 4673, Klebsiella pneumoniae MTCC 10309, Staphylococcus aureus MTCC 9886, Bacillus subtilis MTCC 1305, Micrococcus luteus MTCC 1538, Candida albicans MTCC 7253 and C. tropicalis MTCC 184. The above test microorganisms were obtained from Microbial Type Culture Collection and Gene Bank, CSIR-Institute of Microbial Technology, Chandigarh, India. Culture filtrates used for antimicrobial assays were prepared by first culturing the isolates in 5ml Bennett’s broth supplemented with 0.05% humic acid, pH 7.533 in 15ml tubes and incubated at 28±2oC for 15 days. The broth cultures were then filtered using an ordinary filter paper followed by sterilization using membrane filters (PVDF 0.2µm). Uninoculated media were used as negative-controls while amikacin (30µg), vancomycin (30µg) and amphotericin-B (20µg) discs were used as positive-controls. Antimicrobial screening was performed by agar-well diffusion method34. Antimicrobial activity was assessed as zone of inhibition (mm) by subtracting the diameter of the agar-well from the diameter of inhibition. All the culture filtrates were prepared and tested in triplicates.
Detection of biosynthetic gene clusters
PCR was performed on the genomic DNA of the selected bioactive isolates for checking the presence of three biosynthetic gene clusters (BGCs) related to synthesis of bioactive meta-bolites viz. type I polyketide synthase (PKS-I), type II polyketide synthase (PKS-II) and Non-ribosomal peptide synthetase (NRPS)8,35. Three pairs of primers viz. K1F (5’-TSAAGTCSAACATCGGBCA-3’) – M6R (5’-CGCAGGTTSCSGTACCAGTA-3’), KSaF (5’-TSGCSTGCTTGGAYGCSATC-3’) – KSaR (5’-TGGAANCCGCCGAABCCGCT-3’) and A3F (5’-GCSTACSYSATSTACACSTCSGG-3’) – A7R (5’-SASGTCVCCSGTSCGGTAS-3’) were used respectively8. PCR was performed in final volume of 25µl containing PCR buffer (Promega) with 1.5mM MgCl2, 0.2mM each dNTP (Promega), 0.4µM of each primer (IDT), 0.5U of Taq DNA polymerase (Promega), 10% DMSO and 5µl of extracted DNA as template DNA. Multiplex PCR for PKS-I and PKS-II and simple PCR for NRPS were performed and amplifications were carried out in an Eco96TM thermal cycler (Himedia, India) in the following cycling parameter: initial denaturation of 95oC for 5 mins, 35 cycles of 95oC for 30 secs, 55oC (for PKS-I and PKS-II) / 57oC (for NRPS) for 2 mins, 72oC for 4 mins and final extension of 72oC for 10 mins.
Soil physico-chemical characteristics
Soil and air temperatures of the study site at the time of sample collection (July) were 21oC and 19oC respectively akin to a sub-tropical climate as Mawsmai, Meghalaya, India falls under such climatic zone36. Moisture content of the soil was 16.64±3.85%. The average soil pH was slightly alkaline (7.12±0.9). Soil organic carbon was 2.46±0.05%. Total soil nitrogen, available phosphorus and potassium were 0.08±0.01%, 3.39±0.2µg g-1 and 1.35±0.65µg g-1 respectively. Moisture content of the soil was higher than that reported by Lamare and Singh (2017)37 who conducted study on changes in soil quality in a limestone mining area in East Jaintia Hills from the same state of Meghalaya, India. They reported soil moisture content which ranged from 5.55% to 6.86%. The soil moisture content from our study site was higher because samples were collected on a rainy day. Soil moisture content from limestone mining sites is usually low due to presence of a mixture of granular, aggregated and blocky soil structure lowering the water holding capacity and also as a result of low organic matter37. Lamare and Singh (2017) also reported slightly alkaline nature of limestone soil37 and such soils are generally characterized by alkaline pH38. Soil organic carbon from our study site was also higher than that reported by Lamare and Singh (2017) who recorded 0.23% to 0.47% of soil organic carbon from intensively mined areas and the reason was probably our soil samples was a composite of samples collected from intensively mined and sparsely mined areas. Total soil nitrogen, though slightly higher than that reported by Lamare and Singh (2017)37, was low due to loss of vegetation cover due to mining activities accompanied with loss of nitrogen-fixing microorganisms37. Available phosphorus and potassium were comparatively much lesser to that reported by Lamare and Singh (2017)37. Other workers have also reported low levels of phosphorus and potassium from limestone mining areas39,40. Lamare and Singh (2017) however, reported no significant changes in phosphorus and potassium levels of limestone-mined and unmined areas37. Generally, soil organic carbon, nitrogen, available phosphorus and potassium are relatively lower in concentration in limestone mining areas41.
Isolation of actinobacteria
Highest actinobacterial population number was recorded from un-pretreated and humic acid pretreated soil samples i.e. respectively 5.6±2.13×102 and 4.0±1.56×102 cfu g-1 soil. Statistically insignificant difference was recorded in population number of these two methods at p<0.05 (Fig.1). The differences in population numbers between wet heat pretreated and humic acid pretreated samples were also statistically insignificant at p<0.05 (Fig.1). The three pretreatment methods were significantly better than the other pre-treatment methods. Minimum number of Actinobacteria was recovered using dry heating and phenol pretreatment methods. This may be because most actinobacteria are heat killed when samples are exposed to 100oC for a duration of one hour42 and phenol has been reported to reduce the growth of actinobacteria43. Among the selective media used, humic acid vitamin agar (HVA) and Streptomyces agar (SA) statistically gave the highest actinobacterial population number at 3.78±1.95×102 and 3.11±1.88×102 cfu g-1 soil respectively and were significantly better than the other three media used at p<0.05 (Fig.1). Humic acid present in HVA serves the sole carbon and nitrogen sources and has been observed to activate spore germination in many actinobacterial species19. SA on the other hand is one of the routinely used media for selective isolation and maintenance of actinobacteria. The un-pretreated samples recorded highest actinobacterial population when inoculated on HVA (10.67±0.88×102 cfu g-1soil) and SA media (10.00±0.58×102 cfu g-1 soil). The difference was, however, statistically insignificant at p<0.05 (Fig.1). The actinobacterial population number recorded from humic acid pretreated sample inoculated on HVA medium (9.00±0.58×102 cfu g-1 soil) was also not significantly different from the unpretreated sample inoculated on SA medium (10.00±0.58×102 cfu g-1 soil).
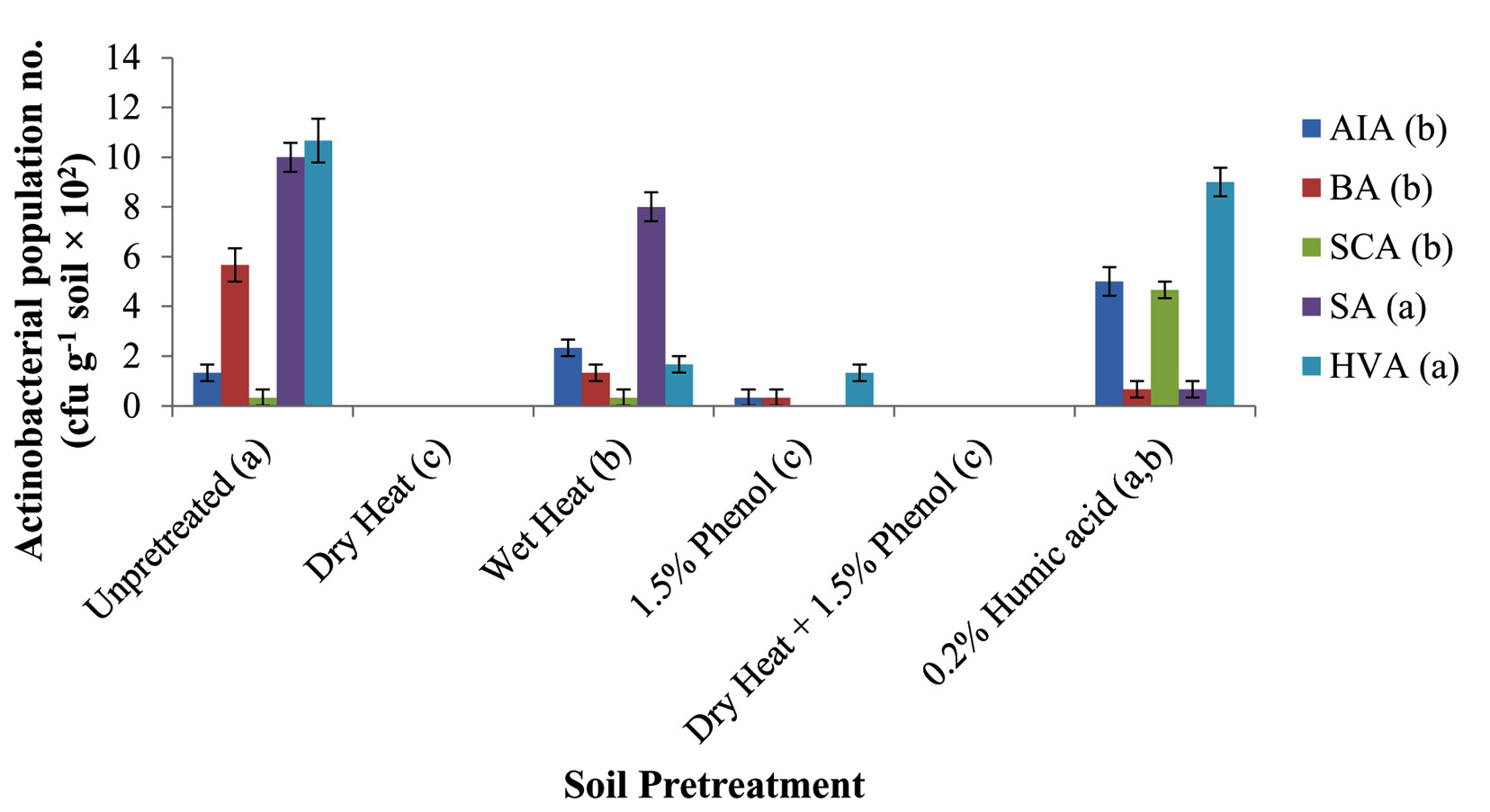 Fig. 1. Recovery of actinobacteria from different pretreated soil samples on different selective media. (AIA, Actinomycete Isolation Agar; BA, Bennett’s Agar; SCA, Starch Casein Agar; SA, Streptomyces Agar; HVA, Humic acid Vitamin Agar. Different lower case letters within parenthesis were significantly different among different pretreatments and among different selective media at p<0.05. Vertical bars represent standard error of mean.)
Fig. 1. Recovery of actinobacteria from different pretreated soil samples on different selective media. (AIA, Actinomycete Isolation Agar; BA, Bennett’s Agar; SCA, Starch Casein Agar; SA, Streptomyces Agar; HVA, Humic acid Vitamin Agar. Different lower case letters within parenthesis were significantly different among different pretreatments and among different selective media at p<0.05. Vertical bars represent standard error of mean.)Diversity of actinobacteria
On the basis of micromorphological characteristics, forty-seven isolates were identified as actinobacteria and all of them showed Gram-positive reaction. Based on biochemical and chemotaxonomic characteristics21,23, thirty-six isolates belonged to Streptomyces genera and the remaining eleven isolates were found to be non-streptomycetes, out of which nine were identified as Nocardia, one as Actinomadura and one as Rhodococcus (Table 1). From the cultivated isolates, it showed a dominance of Streptomyces genera which comprised 76% of the total number of isolates. Recent reports on actinobacteria from limestone niches have also shown Streptomyces to be the dominant genera8,13,44,45. A number of novel actinobacterial strains have also been reported from other limestone niches. Nimaichand et al. (2012)46 reported Streptomyces manipurensis sp. nov., a novel actinomycete, isolated from a limestone deposit site in Manipur, India. Li et al.(2014)47 reported Streptomyces canchipurensis sp. nov., isolated from a limestone environment at Hundung, Manipur, India. Cao et al. (2015)48 reported Lentzea guizhouensis sp. nov., a novel lithophilous actinobacterium isolated from limestone from the Karst area of Guizhou, China. Hezbri et al. (2015)49 reported Geodermatophilus sabuli sp. nov., a g-radiation-resistant actinobacterium isolated from a limestone collected in the Sahara desert of Tunisia. Take et al. (2018)50 reported Streptomyces boninensis sp. nov., isolated from the soil of a limestone cave in the Ogasawara Islands of Japan. Dominance of Streptomyces from the cultivated isolates of this work signifies the adaptation of this genus to thrive on a wide range of nutrients including the synthetic media used during isolation and the genus can also grow faster than the other slower-growing actinobacterial genera thereby establishing its micro niche much earlier than the others9.
Table (1):
Biochemical and chemotaxonomic profile of actinobacterial isolates from the limestone mining site at Mawsmai, Meghalaya
| Isolate | Biochemical profile | Chemotaxonomic profile | Identified genera | ||||
|---|---|---|---|---|---|---|---|
| Casein hydrolysis | Xanthine hydrolysis | Hypoxanthine hydrolysis | Tyrosine hydrolysis | Cell wall DAP | Whole cell sugars | ||
| LD-01 | + | – | + | + | LL | Ara | Streptomyces |
| LD-02 | – | + | + | + | LL | Nd | Streptomyces |
| LD-03 | – | + | + | + | LL | Nd | Streptomyces |
| LD-04 | + | + | + | + | LL | Nd | Streptomyces |
| LD-05 | + | + | + | + | LL | Nd | Streptomyces |
| LD-06 | + | + | + | + | LL | Nd | Streptomyces |
| LD-07 | + | + | + | + | LL | Nd | Streptomyces |
| LD-08 | + | + | + | + | LL | Nd | Streptomyces |
| LD-09 | – | – | + | + | meso | Gal, Ara | Nocardia |
| LD-10 | – | – | + | – | meso | Gal, Ara | Nocardia |
| LD-11 | – | – | + | – | meso | Gal, Ara | Nocardia |
| LD-12 | + | + | + | + | LL | Gal | Streptomyces |
| LD-13 | – | – | + | + | meso | Gal, Ara | Nocardia |
| LD-14 | + | + | + | + | LL | Nd | Streptomyces |
| LD-15 | + | + | + | + | LL | Nd | Streptomyces |
| LD-16 | + | – | + | – | meso | Gal, Ara | Nocardia |
| LD-17 | – | – | – | + | meso | Gal, Ara | Nocardia |
| LD-18 | – | + | + | + | LL | Gal | Streptomyces |
| LD-19 | + | + | + | + | LL | Gal | Streptomyces |
| LD-20 | – | – | – | – | meso | Gal, Xyl | Nocardia |
| LD-21 | + | + | + | + | LL | Nd | Streptomyces |
| LD-22 | + | + | + | + | LL | Nd | Streptomyces |
| LD-23 | + | + | + | + | LL | Gal, Xyl | Streptomyces |
| LD-24 | + | + | + | + | LL | Nd | Streptomyces |
| LD-25 | + | + | + | + | LL | Xyl | Streptomyces |
| LD-26 | + | + | + | + | LL | Nd | Streptomyces |
| LD-27 | + | – | + | + | LL | Ara | Streptomyces |
| LD-28 | + | + | + | + | LL | Nd | Streptomyces |
| LD-29 | + | + | + | + | LL | Nd | Streptomyces |
| LD-30 | – | + | + | + | LL | Nd | Streptomyces |
| LD-31 | – | + | + | + | LL | Nd | Streptomyces |
| LD-32 | + | + | + | + | LL | Nd | Streptomyces |
| LD-33 | + | + | + | + | LL | Nd | Streptomyces |
| LD-34 | + | + | + | – | LL | Nd | Streptomyces |
| LD-35 | + | + | + | + | meso | Nd | Actinomadura |
| LD-36 | – | – | + | – | LL | Nd | Streptomyces |
| LD-37 | + | + | + | + | LL | Nd | Streptomyces |
| LD-38 | + | + | + | + | LL | Nd | Streptomyces |
| LD-39 | + | + | + | – | LL | Nd | Streptomyces |
| LD-40 | + | + | + | + | LL | Nd | Streptomyces |
| LD-41 | + | +/- | – | + | LL | Nd | Streptomyces |
| LD-42 | + | + | + | + | LL | Nd | Streptomyces |
| LD-43 | + | + | + | + | LL | Nd | Streptomyces |
| LD-44 | – | – | – | – | meso | Ara | Rhodococcus |
| LD-45 | + | + | + | + | LL | Nd | Streptomyces |
| LD-46 | – | – | + | + | meso | Nd | Nocardia |
| LD-47 | – | – | + | + | meso | Nd | Nocardia |
DAP, 2,6-diaminopimelic acid; LL and meso are two different isomers of DAP; Ara, Arabinose; Gal, Galactose; Xyl, Xylose; +, positive; -, negative; +/-, uncertain; Nd, Not detected
Antimicrobial activity
Antimicrobial screening results from the agar well diffusion assay revealed that 19% of the isolates showed antibacterial activity against at least one of the bacterial strains tested. The screening results also showed stronger antibacterial activity against Gram-positive than Gram-negative bacteria. The isolates, however, did not show anti-candidal activity. Other recent reports on antimicrobial activity of actinobacteria from limestone niches also showed stronger anti-Gram-positive than anti-Gram-negative bacteria. Nimaichand et al. (2015)8 reported on antimicrobial activity of actinobacteria from limestone deposit sites in Hundung, Manipur, India where eighteen actinobacterial isolates showed antibacterial activity against Gram-positive Bacillus subtilis and five isolates showed antibacterial activity against Gram-negative Escherichia coli. They however, also reported on isolates with anti-fungal activity including anti-candidal activity. Maciejewska et al. (2016)9 also reported on antibacterial activity of moonmilk actinobacterial isolates from a limestone cave in Belgium where the isolates displayed stronger antibacterial activity against Gram-positive bacteria (94% of the phylotypes) than Gram-negative bacteria (71% of the phylotypes). They also reported on anti-fungal activity of the moonmilk isolates where 94% showed activity against a range of fungal strains including Candida albicans. Four isolates from this study viz. LD-21, LD-29, LD-34 and LD-39 showed clear antibacterial activity against at least two Gram-positive bacteria (Fig.2). Isolate LD-21 showed the highest antibacterial activity against Staphylococcus aureus (26±0.58 mm). Antibacterial activity of culture filtrate of LD-21 is shown in Fig.3. The other three isolates viz. LD-29, LD-34 and LD-39 showed moderate anti-bacterial activity against S. aureus with zone of inhibition of 8.33±0.33, 9.33±0.33 and 8.67±0.33 mm respectively. Isolates LD-21 and LD-39 showed low but clear antibacterial activity against Bacillus subtilis with zone of inhibition of 2.67±0.33 and 3.67±0.67 mm respectively. All the above four isolates also showed low but clear antibacterial activity against Micrococcus luteus with zone of inhibition of 2.67±0.33, 1.67±0.33, 4.67±0.88 and 2.33±0.88 mm respectively. All the above four bioactive isolates also showed antibacterial activity against Gram-negative bacterium Pseudomonas aeruginosa but the zone of inhibition of their culture filtrates on the agar well diffusion assay were not as apparent as the positive control (Amikacin disc 30µg) (data not shown) and there were no antibacterial activity against the other Gram-negative bacteria tested. The above four bioactive isolates also possessed at least one of the three biosynthetic gene clusters (BGCs) viz. PKS-I, PKS-II and NRPS (Table 2). Based on the antibacterial activity profile and BGCs, isolates LD-29, LD-34 and LD-39 seemed to have similar metabolic pathways and metabolite clusters. The antimicrobial screening results from this study recovered bioactive isolates with tendency of antimicrobial activity only towards Gram-positive bacteria and failed to recover isolates with broad-spectrum antimicrobial activity. Yet this find is still significant considering the present era where more emphasis are being undertaken to develop narrower spectrum antimicrobial drugs (due to emergence of antimicrobial drug resistance) and recent advancements in rapid and more accurate diagnostic techniques5. It would therefore be interesting to test the above bioactive isolates against more Gram-positive indicator strains and multi-drug resistant clinical isolates. Optimization of the physico-chemical conditions for growth of the bioactive isolates for maximum production of bioactive metabolites51 is currently under way in the hope of finding novel bioactive metabolites.
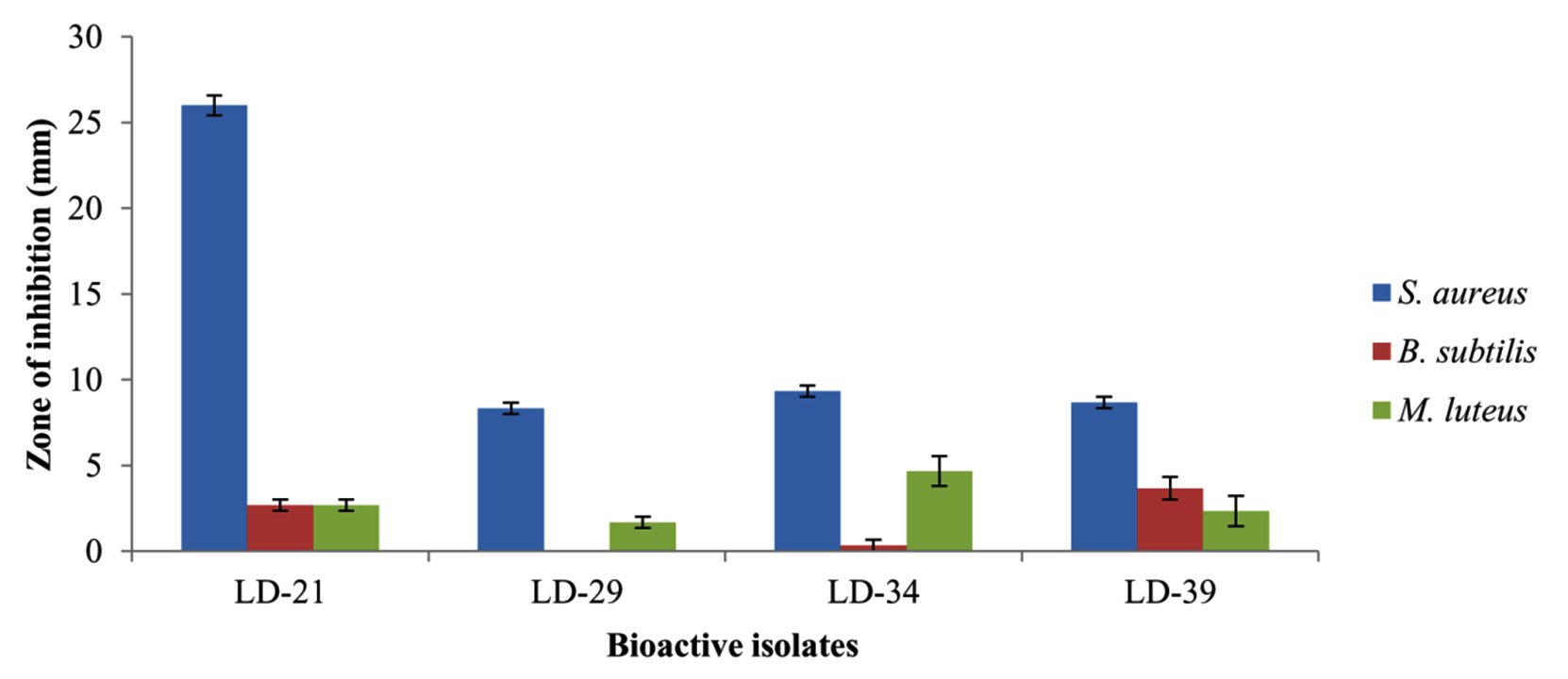 Fig. 2. Antibacterial activity of culture filtrates prepared from the bioactive isolates by agar-well diffusion assay. (All the above bioactive isolates were identified as Streptomyces. S. aureus, Staphylococcus aureus MTCC 9886; B. subtilis, Bacillus subtilis MTCC 1305; M. luteus, Micrococcus luteus MTCC 1538. Vertical bars represent standard error of mean.)
Fig. 2. Antibacterial activity of culture filtrates prepared from the bioactive isolates by agar-well diffusion assay. (All the above bioactive isolates were identified as Streptomyces. S. aureus, Staphylococcus aureus MTCC 9886; B. subtilis, Bacillus subtilis MTCC 1305; M. luteus, Micrococcus luteus MTCC 1538. Vertical bars represent standard error of mean.)Identification of the bioactive isolates
Based on morphological, biochemical and chemotaxonomic profiles (Table 1), all the four bioactive isolates were found to belong to Streptomyces genera. Phylogenetic analysis of their 16S rDNA sequences also confirmed to be Streptomyces. Isolate LD-21 was closest to Streptomyces niveus NRRL2466T (DQ442532) with 99.47% sequence similarity and the other three isolates LD-29, LD-34 and LD-39 were closest to Streptomyces camponoticapitis 2H-TWYE14T (KP784807) with sequence similarity of 99.26%, 99.41% and 99.48% respectively. The generated phylogenetic tree of the bioactive isolates along with the related type strains is shown in Fig.4. GenBank accession numbers of the partial 16S rDNA sequences of the bioactive isolates LD-21, LD-29, LD-34 and LD-39 were assigned from MK659890 to MK659893 respectively. Therefore, the bioactive isolates are now identified as Streptomyces LD-21 (MK659890), Streptomyces LD-29 (MK659891), Streptomyces LD-34 (MK659892) and Streptomyces LD-39 (MK659893). The minimum free energy (MFE) in kcal mol-1 of the predicted 16S rRNA secondary structures of the bioactive isolates also placed them under Streptomyces genera (Table 3). However, on the basis of MFEs, the Streptomyces species with the closest match have changed. The consensus 16S rRNA secondary structures of the bioactive isolates and their closely-related type strains (based on MFE) also showed high degree of conserved regions (Fig.5) confirming that the bioactive isolates belonged to Streptomyces genera. It was also noted that the consensus 16S rRNA secondary structure of Streptomyces LD-21 looked different from the other three bioactive isolates Streptomyces LD-29, Streptomyces LD-34 and Streptomyces LD-39 while these three were similar in their consensus 16S rRNA secondary structures (Fig. 5) which agrees with the maximum likelihood tree (Fig. 4). Thus, it could be inferred that Streptomyces LD-29, Streptomyces LD-34 and Streptomyces LD-39 were phylogenetically closer to one another and they could even belong to the same strain and probably share very similar metabolic pathways and metabolite clusters as already mentioned which is evident from the antibacterial activity of their culture filtrates (Fig.2) and their BGC profiles (Table 2).
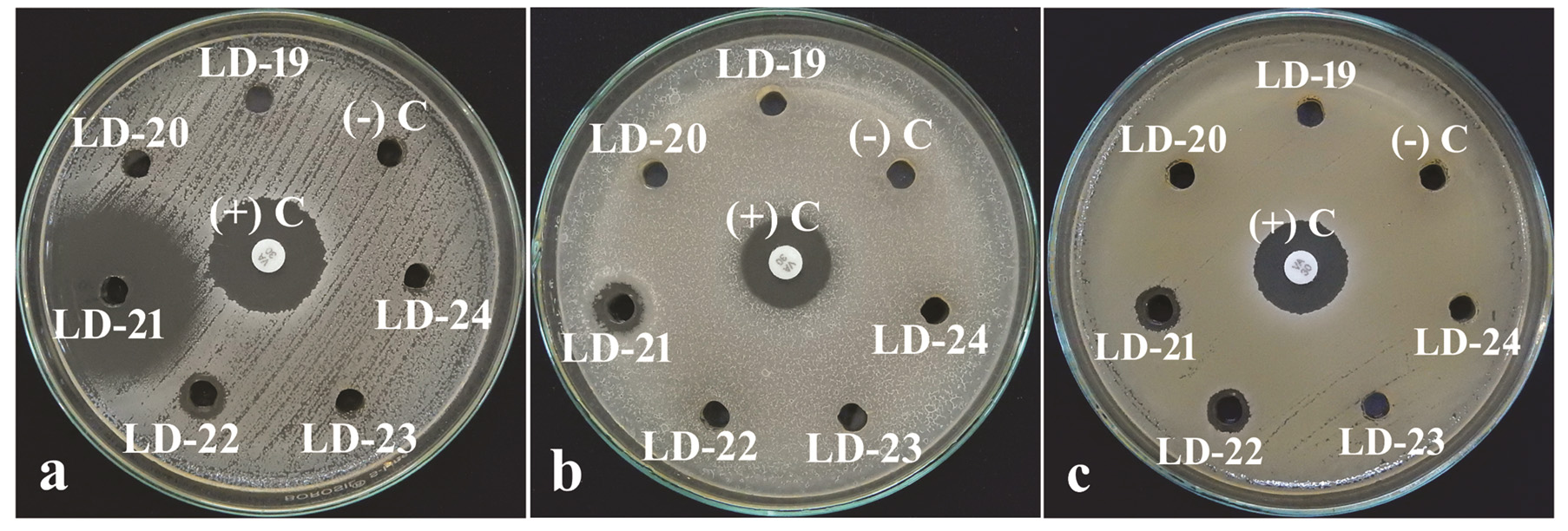 Fig. 3. Antibacterial activity of culture filtrate of bioactive isolate Streptomyces LD-21 and other isolates on a lawn of (a) Staphylococcus aureus, (b) Bacillus subtilis and (c) Micrococcus luteus. (+) C: Positive Control (Vancomycin disc 30µg); (-) C: Negative Control (uninoculated broth).
Fig. 3. Antibacterial activity of culture filtrate of bioactive isolate Streptomyces LD-21 and other isolates on a lawn of (a) Staphylococcus aureus, (b) Bacillus subtilis and (c) Micrococcus luteus. (+) C: Positive Control (Vancomycin disc 30µg); (-) C: Negative Control (uninoculated broth).Table (2):
Detection of biosynthetic gene clusters (BGCs) in bioactive isolates by PCR
| BGC | PCR product length (bp) | Bioactive isolates | |||
|---|---|---|---|---|---|
| LD-21 | LD-29 | LD-34 | LD-39 | ||
| PKS-I | 1200-1400 | – | + | + | + |
| PKS-II | 600 | – | – | – | – |
| NRPS | 700-800 | + | + | + | + |
PKS-I, Type I polyketide synthase; PKS-II, Type II polyketide synthase; NRPS, Non-ribosomal peptide synthetase; +, PCR product detected; -, PCR product not detected
Table (3):
Minimum free energy (MFE) in kcal mol-1 of the predicted 16S rRNA secondary structures of the bioactive isolates along with thirty-two related type strains from EzBioCloud database. Five outgroups have also been included for comparison. The MFEs have been arranged in ascending order.
Strain |
MFE
(kcal mol-1) |
|---|---|
Micromonospora viridifaciens DSM 43909T (X92623)$ |
-553.1 |
Nonomuraea sp. GW 12687T (FN356742)$ |
-552.1 |
Streptomyces niveus NRRL 2466T (DQ442532) |
-542.7 |
Actinomadura darangshiensis DLS-70T (FN646682)$ |
-539.9 |
Streptomyces variegatus NRRL B-16380T (JYJH01000110) |
-537.9 |
Streptomyces flavovariabilis NRRL B-16367T (JNXD01000071) |
-537.9 |
Streptomyces pulveraceus LMG 20322T (AJ781377) |
-536.3 |
Streptomyces gelaticus NRRL B-2928T (DQ026636) |
-535.9 |
Streptomyces sanglieri NBRC 100784T (AB249945) |
-535.1 |
Streptomyces narbonensis NBRC 12801T (AB184157)a |
-534.7 |
Streptomyces LD-21 (MK659890)a# |
-534.3 |
Kitasatospora cinereorecta NBRC 15395T (AB184646) |
-532.9 |
Streptomyces badius NRRL B-2567T (AY999783)b |
-532.8 |
Streptomyces setonii NRRL ISP-5322T (MUNB01000146)b |
-532.8 |
Streptomyces parvus NBRC 3388T(AB184756)b |
-532.8 |
Streptomyces camponoticapitis 2H-TWYE14T (KP784807)b |
-532.8 |
Streptomyces pluricolorescens NBRC 12808T (AB184162)b |
-532.8 |
Streptomyces rubiginosohelvolus NBRC 12912T (AB184240)b |
-532.8 |
Streptomyces pratensis ch24T (JQ806215)b |
-532.8 |
Streptomyces anulatus NRRL B-2000T (DQ026637)b |
-532.8 |
Streptomyces LD-39 (MK659893)b# |
-532.3 |
Streptomyces drozdowiczii NBRC 101007T (AB249957)b |
-531.8 |
Streptomyces humidus NBRC 12877T (AB184213) |
-531.7 |
Streptomyces phaeoluteigriseus DSM 41896T (MPOH01000466)c |
-530.8 |
Streptomyces LD-29 (MK659891)c# |
-530.7 |
Streptomyces globisporus NBRC 12867T(AB184203) |
-530.4 |
Streptomyces fulvissimus DSM 40593T (CP005080) |
-530.4 |
Streptomyces microflavus NBRC 13062T (AB184284) |
-530.4 |
Streptomyces luridiscabiei NRRL B-24455T (LIQV01000394) |
-530.4 |
Streptomyces cyaneofuscatus NRRL B-2570T (JOEM01000050) |
-530.2 |
Streptomyces griseolus NRRL B-2925T (JOFC01000069) |
-530.2 |
Streptomyces halstedii NBRC 12783T (AB184142) |
-530.2 |
Streptomyces sindenensis NBRC 3399T (AB184759) |
-529.8 |
Streptomyces brevispora BK160T (FR692104) |
-529.3 |
Streptomyces griseus subsp. griseus KCTC 9080T (M76388)d |
-528.5 |
Streptomyces LD-34 (MK659892)d# |
-528.5 |
Streptomyces laculatispora BK166T (FR692106) |
-528.3 |
Streptomyces fulvorobeus NBRC 15897T (AB184711) |
-528 |
Mycolicibacterium fortuitum subsp. acetamidolyticum NCH E11620 (NR_104775)$ |
-526.1 |
Nocardia brasiliensis ATCC 19296T (X80591)$ |
-524.8 |
Streptomyces yanii NBRC 14669T (AB006159) |
-522.3 |
a,b,c,d, Bioactive isolates along with their closest related type strains on the basis of MFEs (boldfaced); #, Bioactive isolates from this work; $, Outgroups
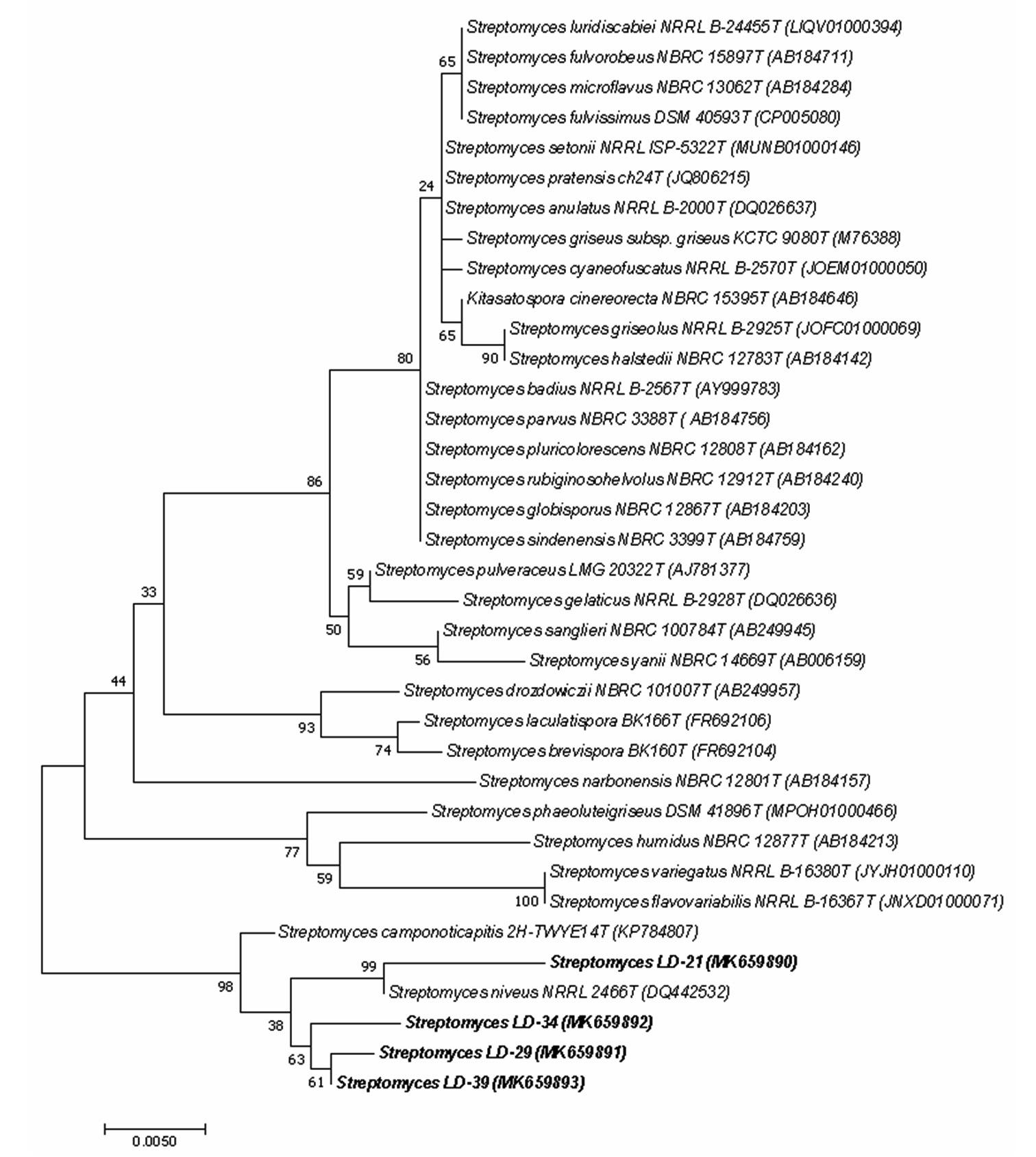 Fig. 4. Maximum Likelihood Tree of the bioactive isolates (boldfaced) Streptomyces LD-21, Streptomyces LD-29, Streptomyces LD-34 and Streptomyces LD-39 along with related type strains based on 16S rDNA sequence. (Alpha-numeric characters within parenthesis are GenBank accession numbers of the corresponding strains. The numbers at the nodes are bootstrap percentages based on 1000 replicates. Horizontal bar equals 0.5% sequence divergence.) (Alpha-numeric characters within parenthesis are GenBank accession numbers of the corresponding strains. The numbers at the nodes are bootstrap percentages based on 1000 replicates. Horizontal bar equals 0.5% sequence divergence.)
Fig. 4. Maximum Likelihood Tree of the bioactive isolates (boldfaced) Streptomyces LD-21, Streptomyces LD-29, Streptomyces LD-34 and Streptomyces LD-39 along with related type strains based on 16S rDNA sequence. (Alpha-numeric characters within parenthesis are GenBank accession numbers of the corresponding strains. The numbers at the nodes are bootstrap percentages based on 1000 replicates. Horizontal bar equals 0.5% sequence divergence.) (Alpha-numeric characters within parenthesis are GenBank accession numbers of the corresponding strains. The numbers at the nodes are bootstrap percentages based on 1000 replicates. Horizontal bar equals 0.5% sequence divergence.)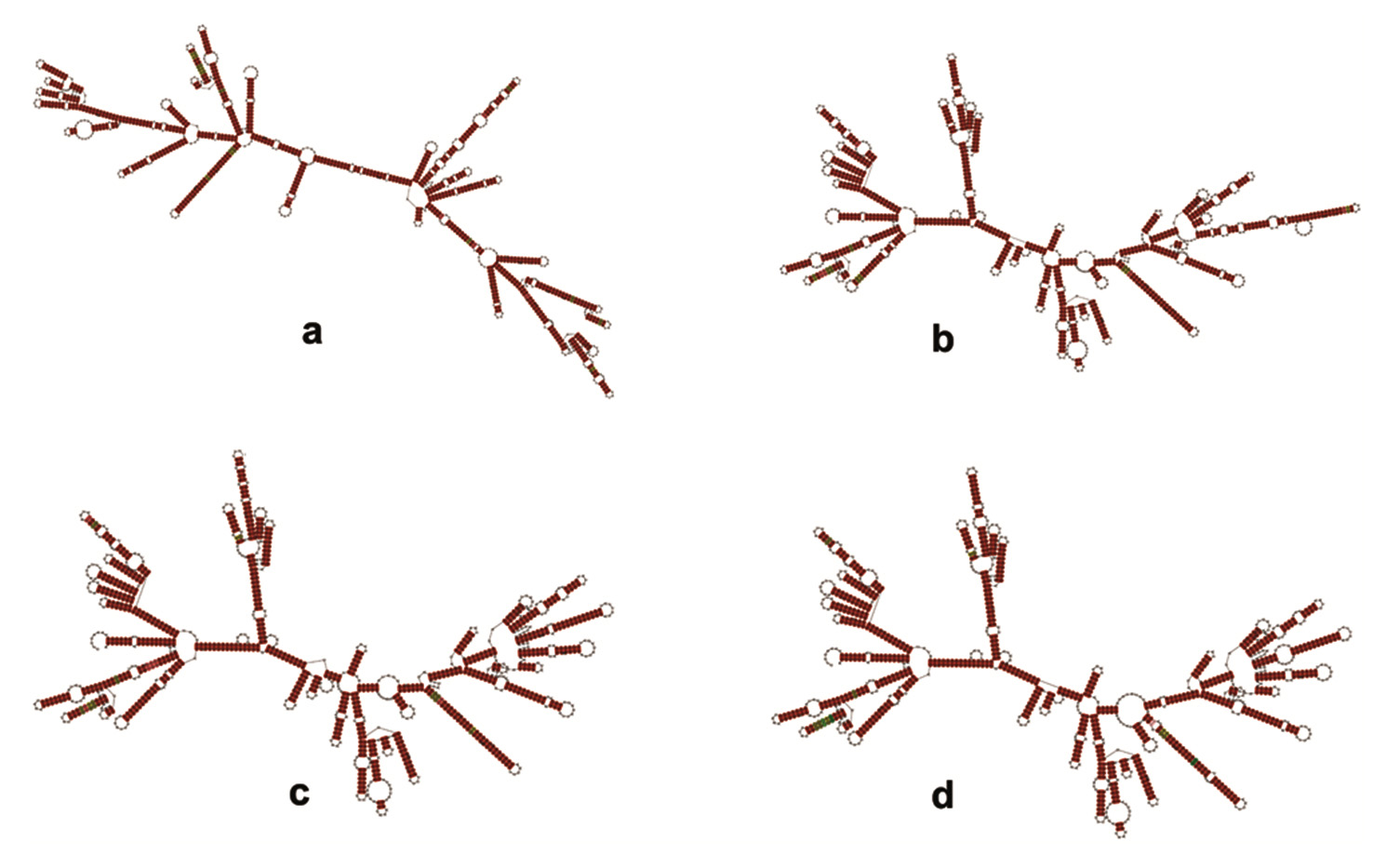 Fig. 5. RNAalifold predicted consensus 16S rRNA secondary structures of a Streptomyces LD-21, b Streptomyces LD-29, c Streptomyces LD-34 and d Streptomyces LD-39 with closely-related type strains based on minimum free energy. Regions shaded in brown denote highest degree of conservation with a single number of base pair type.
Fig. 5. RNAalifold predicted consensus 16S rRNA secondary structures of a Streptomyces LD-21, b Streptomyces LD-29, c Streptomyces LD-34 and d Streptomyces LD-39 with closely-related type strains based on minimum free energy. Regions shaded in brown denote highest degree of conservation with a single number of base pair type.In present work, actinobacteria were isolated from a limestone mining site. The soil environment at the sample collection site was having minimal nutritional conditions (oligo-trophic). Based on morphological, biochemical and chemotaxonomic characteristics, Strepto-myces was found to be the dominant cultivable actinobacterium isolated from this environment. This, however, also suggests the use of more specific methods of isolation to recover the rarer genera of actinobacteria. Four bioactive isolates all identified as Streptomyces showed considerable antibacterial activity against Gram-positive bacteria and they were also found to harbor at least one of the three biosynthetic gene clusters viz. PKS-I, PKS-II and NRPS which are related to synthesis of bioactive metabolites. More work is needed to assess their activity over a range of the Gram-positive bacterium including multi-drug resistant strains. Optimization of bioactive metabolites production by the bioactive isolates is under way. This under-explored niche from Meghalaya is important for mining of actinobacteria for obtaining novel bioactive metabolites that might have applications in various fields.
Acknowledgments
The authors sincerely thank the Department of Botany, Gauhati University, Guwahati, Assam for providing facilities under SAP (DRS-I) and FIST to complete this work.
Conflicts Of Interest
The authors declare that there is no conflict of interest.
Authors’ Contribution
All authors have made substantial, direct and intellectual contribution to the work and approved it for publication.
Funding
None
Data Availability
All datasets generated or analyzed during this study are included in the research work.
Ethics Statement
This article does not contain any studies with human participants or animals performed by any of the authors.
- Goodfellow M. Phylum XXVI. Actinobacteria phyl. nov., p.33. In Goodfellow M, Kampfer P, Busse H-J, Trujillo ME, Suzuki K-i, Ludwig W, Whitman WB (eds.), Bergey’s Manual of Systematic Bacteriology, Volume Five: The Actinobacteria, 2nd Ed. Springer, 2012.
- Genilloud O. Actinomycetes: Still a source of novel antibiotics. Nat. Prod. Rep., 2017; 34(10): 1203-1232.
- Berdy J. Bioavtive microbial metabolites – A personal view. J. Antibiot. (Tokyo), 2005; 58(1): 1-26.
- Wright G.D. Something old, something new: revisiting natural products in antibiotic drug discovery. Can. J. Microbiol., 2014; 60(3): 147-154.
- Fischbach M.A., Walsh C.T. Antibiotics for emerging pathogens. Science, 2009; 325(5944): 1089-1093.
- Lawrence J. Drug discovery returns to the wild. Pharm. J., 2015; 294(7849): online. doi:10.1211/PJ.2015.20067820
- Mohite O.S., Weber T., Kim H.U., Lee S.Y. Genome-scale metabolic reconstruction of actinomycetes for antibiotics production. Biotechnol. J., 2019; 14(1): 1800377 (1-9).
- Nimaichand S., Devi A.M., Tamreihao K., Ningthoujam D.S., Li W.J. Actinobacterial diversity in limestone deposit sites in Hundung, Manipur (India) and their antimicrobial activities. Front Microbiol., 2015; 6: 413.
- Maciejewska M., Adam D., Martinet L., Naome A., Calusinska M., Delfosse P., Carnol M., Barton H.A., Hayette M.P., Smargiasso N., De Pauw E., Hanikenne M., Baurain D., Rigali S. A phenotypic and genotypic analysis of the antimicrobial potential of cultivable Streptomyces isolated from cave moonmilk deposits. Front Microbiol., 2016; 7: 1455.
- Adam D., Maciejewska M., Naome A., Martinet L., Coppieters W., Karim L., Baurain D., Rigali S. Isolation, characterization and antibacterial activity of hard-to-culture actinobacteria from cave moonmilk deposits. Antibiotics, 2018; 7(2): 28.
- Belyagoubi L., Belyagoubi-Benhammou N., Jurado V., Dupont J., Lacoste S., Djebbah F., Ounadjela F.Z., Benaissa S., Habi S., Abdelouahid D.E., Saiz-Jimenez C. Antimicrobial activities of culturable microorganisms (Actinomycetes and fungi) isolated from Chaabe Cave, Algeria. Int. J. Speleol., 2018; 47(2): 189-199.
- Covington B.C., Spraggins J.M., Ynigez-Gutierrez A.E., Hylton Z.B., Bachmann B.O. Response of secondary metabolism of hypogean actinobacterial genera to chemical and biological stimuli. Appl. Environ. Microbiol., 2018; 84(19): e01125-18.
- Hamedi J., Kafshnouchi M., Ranjbaran M. A Study on actinobacterial diversity of Hampoeil cave and screening of their biological activities. Saudi J Biol Sci, 2018; In Press. doi:10.1016/j.sjbs.2018.10.010
- Yasir M. Analysis of bacterial communities and characterization of antimicrobial strains from cave microbiota. Brazilian J. Microbiol., 2018; 49(2): 248-257.
- Myers N., Mittermeier R.A., Mittermeier C.G., da Fonseca G.A.B., Kent J. Biodiversity hotspots for conservation priorities. Nature, 2000; 403(6772): 853-858.
- Allen S.E., Grimshaw H.M., Parkinson J.A., Quarmby C. Chemical Analysis of Ecological Materials. Blackwell Scientific, Oxford, 1974.
- Anderson J.M., Ingram J.S.I. Tropical Soil Biology and Fertility: A Handbook of Methods. 2nd Ed. C.A.B International, Wallingford, 1993.
- Jackson M.L. Soil Chemical Analysis. Prentice Hall of India Pvt. Ltd., New Delhi, 1973.
- Hayakawa M., Nonomura H. Humic acid-vitamin agar, a new medium for the selective isolation of soil actinomycetes. J. Ferment. Technol., 1987; 65(5): 501-509.
- Cross T. Growth and examination of actinomycetes: Some guidelines, pp. 605-609. In Holt JG, Sneath PH, Krieg NR (eds.), Bergey’s Manual of Determinative Bacteriology, 9th Ed. Williams & Wilkins, Baltimore, 1994.
- Berd D. Laboratory identification of clinically important aerobic actinomycetes. Appl Microbiol., 1973; 25(4): 665-681.
- Hasegawa T., Takizawa M., Tanida S. A rapid analysis for chemical grouping of aerobic actinomycetes. J. Gen. Appl. Microbiol., 1983; 29(4): 319-322.
- Staneck J.L., Roberts G.D. Simplified approach to identification of aerobic actinomycetes by thin-layer chromatography. Appl. Environ. Microbiol., 1974; 28(2): 226-231.
- Chen X., Jiang Y., Li Q., Han L., Jiang C. Molecular Phylogenetic Identification of Actinobacteria, 2016, pp. 141-174. In Dhanasekaran D., Jiang Y. (eds.), Actinobacteria: Basics and Biotechnological Applications. ExLi4EvA. doi:10. 5772/62029
- Monciardini P., Sosio M., Cavaletti L., Chiocchini C., Stefano D. New PCR primers for the selective amplication of 16S rDNA from different groups of actinomycetes. FEMS Microbiol. Ecol., 2002; 42: 419-429.
- Yoon S.H., Ha S.M., Kwon S., Lim J., Kim Y., Seo H., Chun J. Introducing EzBioCloud: A taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int. J. Syst. Evol. Microbiol., 2017; 67(5): 1613-1617.
- Kumar S., Stecher G., Tamura K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol., 2016; 33(7): 1870-1874.
- Tamura K. Estimation of the number of nucleotide substitutions when there are strong transition-transversion and G+C-content biases. Mol. Biol. Evol., 1992; 9(4): 678-687.
- Felsenstein J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution (N.Y.), 1985; 39: 783-791.
- Ghosh A., Sutradhar S., Baishya D. Delineating thermophilic xylanase from Bacillus licheniformis DM5 towards its potential application in xylo-oligosaccharides production. World J. Microbiol. Biotechnol., 2019; 35(2): 34.
- Gruber A.R., Lorenz R., Bernhart S.H., Neubock R., Hofacker I.L. The Vienna RNA websuite. Nucleic Acids Res., 2008; 36: W70-W74.
- Bernhart S.H., Hofacker I.L., Will S., Gruber A.R., Stadler P.F. RNAalifold: Improved consensus structure prediction for RNA alignments. BMC Bioinformatics, 2008; 9: 474.
- Otoguro M., Hayakawa M., Yamazaki T., Iimura Y. An integrated method for the enrichment and selective isolation of Actinokineospora spp. in soil and plant litter. J. Appl. Microbiol., 2001; 91(1): 118-130.
- Holder I.A., Boyce S.T. Agar well diffusion assay testing of bacterial susceptibility to various antimicrobials in concentrations non-toxic for human cells in culture. Burns, 1994; 20(5): 426-429.
- Ayuso-Sacido A., Genilloud O. New PCR primers for the screening of NRPS and PKS-I systems in actinomycetes: Detection and distribution of these biosynthetic gene sequences in major taxonomic groups. Microb. Ecol., 2005; 49(1): 10-24.
- Shankar U., Tripathi R.S., Pandey H.N. Structure and seasonal dynamics of humid tropical grasslands in Meghalaya, India. J. Veg. Sci., 1991; 2(5): 711-714.
- Lamare R.E., Singh O.P. Changes in soil quality in limestone mining area of Meghalaya, India. Nat Environ. Pollut. Technol., 2017; 16(2): 545-550.
- Misra A., Tyler G. Influence of soil moisture on soil solution chemistry and concentrations of minerals in the Calcicoles Phleum phleoides and Veronica spicata grown on a limestone soil. Ann. Bot., 1999; 84(3): 401-410.
- Cohen-Fernandez A.C., Naeth M.A. Erosion control blankets, organic amendments and site variability influenced the initial plant community at a limestone quarry in the Canadian Rocky Mountains. Biogeosciences, 2013; 10(7): 5243-5253.
- Sarwanto D., Prayitno C.H. The diversity and productivity of indigenous forage in former limestone mining quarry in karst mountain of Southern Gombong, Central Java Indonesia. Anim. Prod., 2015; 17(2): 69-75.
- Hanief S.M., Thakur S.D., Gupta B. Vegetal profile of natural plant succession and artificially re-vegetated limestone mines of Himachal Pradesh, India. J. Trop. For., 2007; 23: 128-135.
- Seong C.N., Choi J.H., Baik K. An improved selective isolation of rare actinomycetes from forest soil. J. Microbiol., 2001; 39(1): 17-23.
- Hayakawa M., Nonomura H. A new method for the intensive isolation of actinomycetes from soil. Actinomycetologica, 1989; 3(2): 95-104.
- Cao C., Jiang J., Sun H., Huang Y., Tao F., Lian B. Carbonate mineral formation under the influence of limestone-colonizing actinobacteria: Morphology and polymorphism. Front Microbiol, 2016; 7: 366.
- Fang B-Z, Salam N., Han M-X, Jiao J-Y, Cheng J., Wei D-Q, Xiao M., Li W-J. Insights on the effects of heat pretreatment, pH, and calcium salts on isolation of rare actinobacteria from karstic caves. Front Microbiol., 2017; 8: 1535.
- Nimaichand S., Zhu W.Y., Yang L.L., Ming H., Nie G.X., Tang S.K., Ningthoujam D.S., Li W.J. Streptomyces manipurensis sp. nov., a novel actinomycete isolated from a limestone deposit site in Manipur, India. Antonie Van Leeuwenhoek, 2012; 102(1): 133-139.
- Li W-J, Nimaichand S., Jiang Z., Liu M-J, Khieu T-N, Kim C-J, Hozzein W.N., Park D-J, Wadaan M.A.M., Ningthoujam D.S. Streptomyces canchipurensis sp. nov., isolated from a limestone habitat. Antonie Van Leeuwenhoek, 2014; 106: 1119–1126.
- Cao C.L., Zhou X.Q., Qin S., Tao F.X., Jiang J.H., Lian B. Lentzea guizhouensis sp. nov., a novel lithophilous actinobacterium isolated from limestone from the Karst area, Guizhou, China. Antonie Van Leeuwenhoek, 2015; 108(6): 1365-1372.
- Hezbri K., Ghodhbane-Gtari F., Montero-Calasanz M.C., Sghaier H., Rohde M., Schumann P., Klenk H.P., Gtari M. Geodermatophilus sabuli sp. nov., a g-radiation-resistant actinobacterium isolated from desert limestone. Int. J. Syst. Evol. Microbiol., 2015; 65(10): 3365-3372.
- Take A., Inahashi Y., Omura S., Takahashi Y., Matsumoto A. Streptomyces boninensis sp. nov., isolated from soil from a limestone cave in the Ogasawara Islands. Int. J. Syst. Evol. Microbiol., 2018; 68: 1795-1799.
- Deka D., Jha D.K. Optimization of culture parameters for improved production of bioactive metabolite by endophytic Geosmithia pallida (KU693285) isolated from Brucea mollis Wall ex. Kurz, an endangered medicinal plant. J. Pure Appl. Microbiol., 2018; 12(3): 1205-1213.
© The Author(s) 2019. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.


