ISSN: 0973-7510
E-ISSN: 2581-690X
This study was conducted to isolate actinomycetes from forest areas of Tamilnadu, to check the antibacterial efficacy of the biomolecules produced by the isolates and their application as antibacterial textile finish. The cross streak plate method and disc diffusion methods were used for screening the actinomycetes isolates for determining antibacterial activity. Chromatographic techniques were used for the partial purification of biomolecules. Partially purified biomolecules were analysed by spectroscopy. Standard procedure was followed for the impregnation of biomolecule on cotton cloth and antibacterial activity studies. Twenty two actinomycetes were isolated from forest soil and screened for antibacterial activity.. On secondary screening the strains KUAJ1 and KUAJ5 exhibited antibacterial activity against the test organisms. KUAJ1 extract produced a maximum zone of inhibition of 21mm against Klebsiella sp and KUAJ5 extract produced a maximum zone of inhibition of 22mm against Bacillus subtilis. The products were partially purified and further analyzed by spectroscopy. On UV – visible spectroscopy maximum absorption spectra of 218 and 240 nm were shown respectively by KUAJ1 and KUAJ5.FTIR results exhibited O-H, C-H and CH2-S groups in KUAJ1 and N-H, CH3-S groups in KUAJ5 extract. Extracted compounds were coated on cotton fabric and its antibacterial activities were tested. This study proves that the actinomycetes isolated from Tamilnadu forest soil shows antibacterial activites and the active biomolecules produced by isolates could be used for antibacterial textile finish on 100% cotton material
Actinomycetes, screening, TLC, Spectrometric analysis, textile, biomolecule coating.
Actinomycetes, the filamentous, gram positive bacteria, are the most economically and biotechnologically valuable prokaryotic microorganisms. They are virtually unlimited sources of novel compounds with many therapeutic applications. Due to diversity and proven ability to produce new structures they hold a prominent position in the field of pharmaceutics1. They were found to produce nearly half of the discovered bioactive secondary metabolites include antibiotics, antitumor agents, immunosuppressive agents and enzymes2. Actinomycetes provided about two third part (more than 4,000) naturally occurring antibiotics discovered including many of those important in medicine3. Almost 80% of the world’s antibiotics are known to come from the genera Streptomycetes and Micromonospora4. According to world health organization, regular use of antibiotics has led to the generation of antibiotic resistance in many bacterial and fungal pathogens. Because of the fast emergence of newer infections and organisms developing resistance which render already existing antibiotics a less effective one. Therefore constant search for new antibiotics to overcome these problems is a matter of study5. Thus drug resistant strains of pathogens emerge quickly than the rate of discovery of new antibiotics6. The present study is an attempt to isolate actinomycetes, producing antibiotics from forest soil, one of the under explored area with a hope that actinomycetes from unexplored area may produce some novel bioactive molecules against emerging resistant pathogens.
Increase concern over the damage caused by exposure to microbes, chemical pesticides, UV light and pollutants in last few years heightened the demand for protective garments. Clothing today is expected to be water proof, flame resistant, self – cleaning, insect repellant and antimicrobial to protect human beings from infection, UV light, chemical and biological agents7. With the growing public health awareness of the pathogenic effects, malodors and stain formations caused by microorganisms, there is an increasing need for antibacterial materials in many application areas like medical devices, health care, hygienic application, water purification systems, hospital, dental surgery equipment, textiles, food packaging, and storage8. In order to impart permanent functionality to textile surfaces, biocompatible polymers like chitosan, carrageenan and their derivatives can be used by some chemical strategy using anchoring chemical such as cyanuric chloride and hydroxydichlorotriazine9. The study also focuses on the application of bioactive molecule extracted from actinomycetes isolated from forest soil in textile finish. An attempt was made to coat 100% cotton fabric with chitosan and sodium alginate and the bioactive biomolecule produced by the potent actinomycetes isolate.
Collection of soil sample
Three soil samples were collected at the depth of 5-m10 cm in sterile plastic bags from three different forest areas, Tamil Nadu, India. All samples were transported to the laboratory, labeled and refrigerated for further study10.
Sample pretreatment
Two different pretreatments were done on soil samples. The soil samples were air dried for 3-4 days at room temperature. And wet heating was performed at a temperature of 70°C for 15 minutes in a water bath11, 12.
Isolation of actinomycetes
1 g of pretreated soil samples were suspended in 10 ml distilled water and vortexed for 10 minutes and serially diluted to 10-6. 0. 1 ml of aliquots was taken from each dilution and spread plating was done on Actinomycetes isolation agar and incubated at room temperature for 7 days13.
Morphological characterization
Morphological characters such as colony characteristics, type of aerial hyphae, growth of vegetative hyphae, soluble pigment production were observed14.
Microscopic characterizations
Gram staining was performed and phase contrast microscopy was done after performing agar slide culture
Bacterial strains used
Four bacterial strains were used for this study. They were Escherichia coli(MTCC 443), Bacillus subtilis (MTCC 441), Staphylococcus aureus (MTCC 737)and Klebsiella sp (MTCC 418).
Primary screening
Primary screening of isolate was done to ascertain the potential of strains with respect to the production of antimicrobial secondary metabolites. On the basis of primary screening results, isolates showing substantial antimicrobial activities are selected for subsequent secondary screening programme15. Primary screening was done by right angle streak method16. During the primary screening, isolates were screened against selected bacterial strains by using perpendicular streak. In this method nutrient agar medium was used and each plate was streaked with individual isolate at the center / diameter of the plate and incubated at 30°C for 7 days. Later 24 h old fresh subculture of test organisms were prepared and streaked perpendicular to the isolate and incubated at 37°C for 24 h. zone of inhibition was observed after incubation.
Preparation of crude extract
The active isolate resulted from primary screening were used for their extracellular antibiotic production capabilities under submerged fermentation conditions. The production medium containing starch 1%, potassium nitrate 0.2%, sodium chloride 0.2%, Di- potassium hydrogen phosphate 0.2%, MgSO4 0.005%, CaCO30.002%, ferrous sulphate 0.001%, casein 0.03% and sea water 50% was used for antibiotic production. 7 days old slant culture of selected isolates was used for inoculum preparation. Sterile distilled water was added to each slant and 5 ml of this spore suspension was added to 45 ml of the production media and incubated at 28°C on a rotary shaker (120 rpm) for 4 days. Then samples were collected in to sterile centrifuge tube and centrifuged at 3000 rpm for 15 minutes. The supernatant was filtered through Whatman No.1 filter paper. The filtrate was aseptically transferred in to a sterile conical flask and stored at 4°C for further assay.
Extraction of the antimicrobial compound from culture supernatant using different solvents and purification by column chromatography
The extraction of active compounds from crude culture broth was done using different solventslike n-butanol, n-hexane, ethyl acetate, petroleum ether, methanol and chloroform. The solvents were mixed in the proportion of 1:1. Aqueous layer and organic layers were separated and tested for the presence of antibiotic using disc diffusion method. The partially purified form of active compounds was subjected to further purification by column17. The dried extract was eluted with chloroform – methanol solvent system. Stepwise elution was done by decreasing the concentration of chloroform and increasing the concentration of methanol. In each successive elution concentration of chloroform was reduced 10% and that of methanol was increased by 10% until the concentration of methanol reached 100%. All the fractions each of 20ml were collected and further analyzed.
Thin layer chromatography
The active compound obtained after solvent extraction was spotted on pre-coated thin layer chromatography plates. The plates were eluted with chloroform: methanol solvent system (4:1)18. The spotted TLC plates were kept in iodine chamber for the development of spots which corresponds to the antimicrobial compounds.
Secondary screening
Secondary screening was done by disc diffusion method19. The media used was Muller Hinton Agar. The test organisms were prepared by inoculating few colonies in to sterile nutrient broth and incubated at 37°C until the visible turbidity matches that of 0.5 McFarland solution which is equivalent to 106 – 108 CFU/ml. On the medium, test organisms were swabbed by sterile swab and left at room temperature for 3 – 5 minutes for the surface moisture to be absorbed before applying the discs with crude extract. Sterile discs were loaded with 100µl of the crude extract re for the assay of antagonistic activity and filter paper soaked in sterile distilled water was used as control. The plates were incubated at 37°C for 24 h and the zone of inhibition (mm) was measured.
Spectroscopic analysis
The fractions which showed the antimicrobial activity were subjected to spectroscopic analysis. UV-visible spectra were obtained on Hitachi U2900 spectrometer and infrared spectra were recorded20.
Coating of cotton material with bioactive biomolecule
100% cotton fabric was used for biomolecule coating purpose21.
Pre- treatment of cotton material – scouring and bleaching
The cotton fabric was scoured using 3.0% sodium hydroxide and 2% sodium carbonate and all other auxiliaries prescribed by standard procedure. It was then bleached with 4% hydrogen peroxide and all other auxiliaries as per the standard procedure22.
Chitosan and sodium alginate polymer coating
Chitosan solution was prepared by stirring a dispersion of chitosan (8g) in 2.0% (v/v) aqueous acetic acid solution for 1hr at 60ºC and then 2g of sodium alginate polymer was added to the chitosan solution and stirred for 10 min. The fabric was padded twice with solution of same concentration and was dried at 80ºC for 5 min and cured at 140ºC for 3 min.
Immobilization of bioactive biomolecule
Active fractions from column chromatography were used as antibacterial agents for this purpose. The column eluents were dried and dissolved in sterile distilled water. 2.0 ± 0.1 cm fabric samples were taken from the polymer coated cotton fabric and immersed separately in the solutions and allowed to remain stand till 24h. Then they were dried at room temperature23.
Antibacterial activity assessment
The antibacterial activity was evaluated against Staphylococcus aureus (MTCC 737) and Escherichia coli (MTCC 443). Bioactive material coated fabric samples were placed at the centre of bacteria seeded agar plate. After 24h the plates were observed for the zone of inhibition24.
A total of 22 actinomycetes were isolated from three different forest soil samples. Microbes were isolated in mixed culture by spread plate method. Based on colony morphology individual actinomycetes were isolated from the plates. [Fig 1]
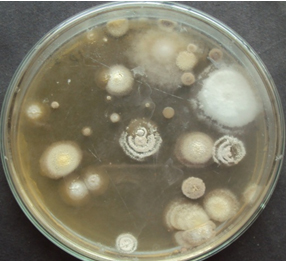
Fig. 1. Actinomycete colonies on actinomycetes isolation agar
Morphological and cultural characteristics
Morphological characters such as colony morphology, colour of aerial mycelium, substrate mycelium and production of soluble pigment were studied and tabulated [Table 1]. Gram staining revealed that all the isolates are gram positive in nature.
Table (1):
Morphological characteristics and gram reaction of actinomycetes isolates.
S.No. |
Actinomycete Isolate |
Aerial mycelium |
Reverseside pigment |
Soluble pigment |
Gram staining |
|---|---|---|---|---|---|
1 |
KUAJ1 |
White |
Creamy white |
– |
+ |
2 |
KUAJ2 |
Cottony White |
White |
– |
+ |
3 |
KUAJ3 |
Green |
Whitish green |
– |
+ |
4 |
KUAJ5 |
Creamy white |
Yellow |
Yellow |
+ |
5 |
KUAJ6 |
Brownish white |
Grey |
– |
+ |
6 |
KUAJ22 |
Golden yellow |
Deep brown |
– |
+ |
‘+’ positive reaction
‘ -’ negative reaction
Agar slide culture and Phase contrast microscopy
Agar slide culture was performed and after incubation time cover slips were removed and observed under phase contrast microscope after staining with lacto phenol cotton blue. Results are shown in [Fig. 2].
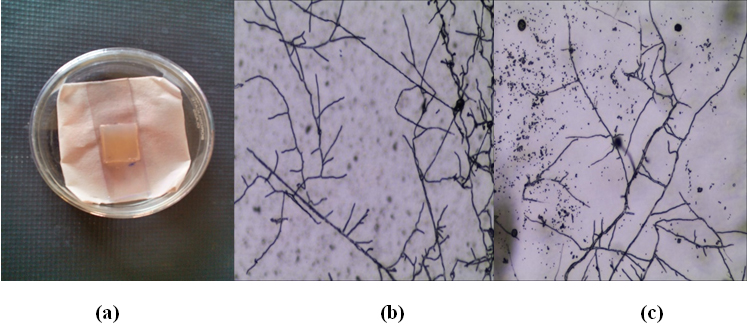 (a) Actinomycetes growth on agar slide culture,
(a) Actinomycetes growth on agar slide culture,(b) Phase contrast image of KUAJ1 and (c) Phase contrast image of KUAJ5
Fig. 2. Agar slide culture and phase contrast microscopy
Primary screening
Among the 22 actinomycetes isolated from soil sample, six isolate showed antibacterial activity against at least one of the test bacteria. In perpendicular streak method, the result revealed that the isolates KUAJ1 and KUAJ5 possess broad spectrum activities against tested bacteria as shown in [Table2].
Table (2):
Antibacterial activity of various isolate against test organisms.
S. No |
Actinomycetes Isolate |
E coli |
Klebsiells sp |
B. subtilis |
S. aureus |
|---|---|---|---|---|---|
1 |
KUAJ1 |
+ |
+ |
+ |
+ |
2 |
KUAJ2 |
+ |
– |
– |
– |
3 |
KUAJ3 |
+ |
– |
– |
– |
4 |
KUAJ5 |
+ |
– |
+ |
+ |
5 |
KUAJ6 |
+ |
– |
– |
+ |
6 |
KUAJ22 |
+ |
– |
– |
+ |
‘+’ shows antibacterial activity
‘-’ No antibacterial activity
Primary screening by right angle streak method. Pathogens used are Escherichia coli
(MTCC 443), Bacillus subtilis (MTCC 441), Staphylococcus aureus(MTCC 737) and Klebsiella sp(MTCC 418).
Extraction of active compound
Different solvents were tested for the extraction of active compound. In case of KUAJ1 and KUAJ5 active compound yield observed in the residue extracted with methanol.
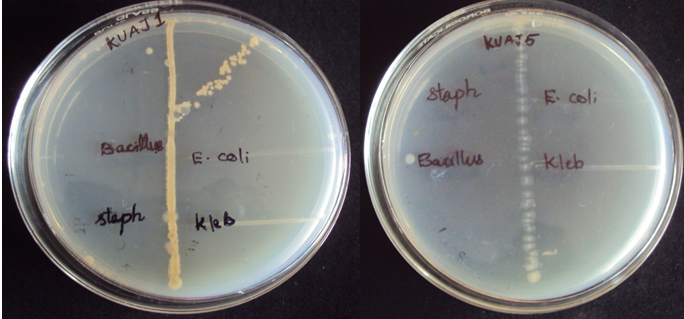
Fig. 3. Primary screening results KUAJ1 and KUAJ5
Column chromatography
The bioactive compounds from methanolic extract were further purified by column chromatography. Twenty eluents were collected and analyzed for antimicrobial activities. Eleventh fraction showed activity against both gram negative and gram positive bacteria in KUAJ1 extract and fifteenth fraction showed activity against test organisms in KUAJ5.
Secondary screening
The partially purified bioactive compounds prepared from two isolates were subjected to secondary screening by disc diffusion method in triplicates. The isolate KUAJ1 showed maximum activity against Klebsiella sp and KUAJ 5 exhibited maximum activity against Bacillus sp. Mean zone of inhibition against pathogens are tabulated.
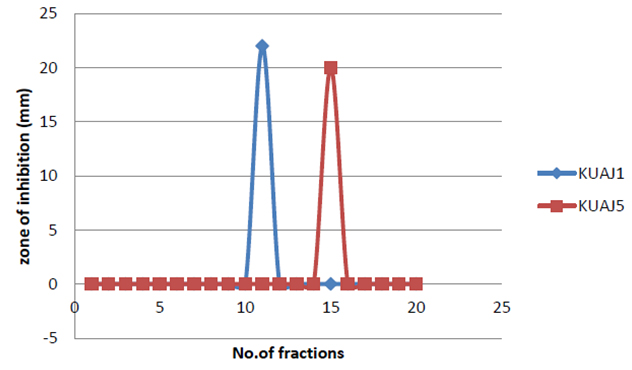
Fig. 4. Inhibitory effect of column eluents
The partially purified antimicrobial compound obtained after solvent extraction was analyzed using pre- coated silica gel plates. Chromatograms were developed in iodine chamber. Rf values were found to be 0.75 and 0.48 for KUAJ1 and KUAJ5 respectively.
Spectroscopic characterization of crude extract
The UV – visible spectra for methanolic extract showed maximum absorption spectra at 218 and 240 nm respectively for KUAJ1 and KUAJ5. The IR spectrum exhibited 3414/cm (broad) and1020.34 (strong) for KUAJ1 and 3433.29, 3404.29, 339.64 (broad) and 1022.27 (strong) for KUAJ 5[25].
Coating of bioactive biomolecule on cotton
100% cotton cloth was coated with chitosan and sodium alginate polymer. 2.0±0.1 cm diameter cloth pieces were cut from the polymer coated cotton fabric and subjected for bioactive molecule coating. Coated material was then subjected for antibacterial assessment. The results are tabulated.
Table (3):
Antibiogram of KUAJ isolates against various pathogens.
| S.No. | Test organism | Mean zone of inhibition(mm) | |
|---|---|---|---|
| KUAJ1 | KUAJ5 | ||
| 1 | E.coli | 20 | 18 |
| 2 | Klebsiella sp | 21 | 20 |
| 3 | Bacillus subtilis | 18 | 22 |
| 4 | Staphylococcus sp | 18 | 16 |
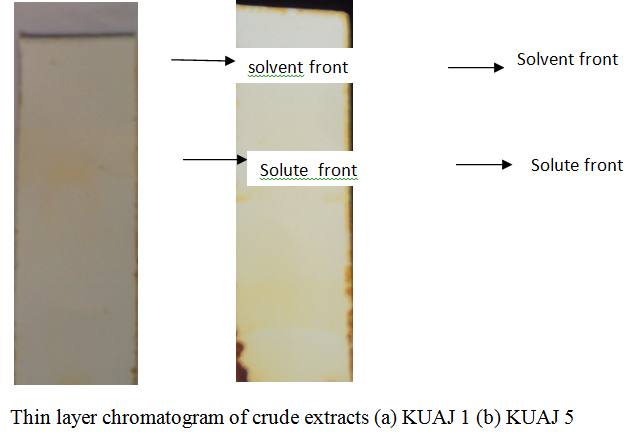 Fig. 5. Infrared spectrum of the partially purified antimicrobial compound produced by KUAJ1
Fig. 5. Infrared spectrum of the partially purified antimicrobial compound produced by KUAJ1Table (4):
Evaluation of antibacterial activity of polymer and immobilized bioactive biomolecule loaded cotton sample.
| S.No. | Sample | Zone of inhibition(cm) | |
|---|---|---|---|
| Staphylococcus sp | E. coli | ||
| 1. | CP | 3.6 | 3.8 |
| 2. | CP1 | 4.6 | 4.8 |
| 3. | CP5 | 4.2 | 4.4 |
Polymer coated fabric sample produced a zone of inhibition of 3.6cm and 3.8cm against the test pathogen Staphylococcus sp and E.coli respectively. Both CP1 and CP5 shown more activity against E.coli, the gram negative test pathogen.
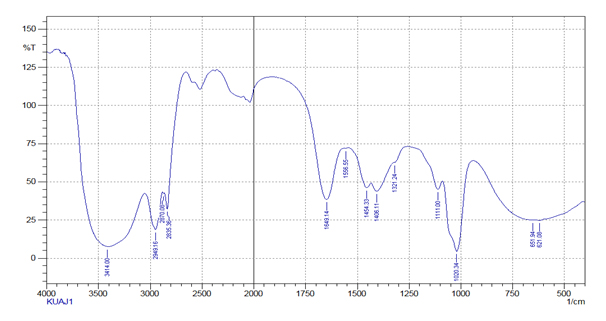
Fig. 6.Thin layer chromatography
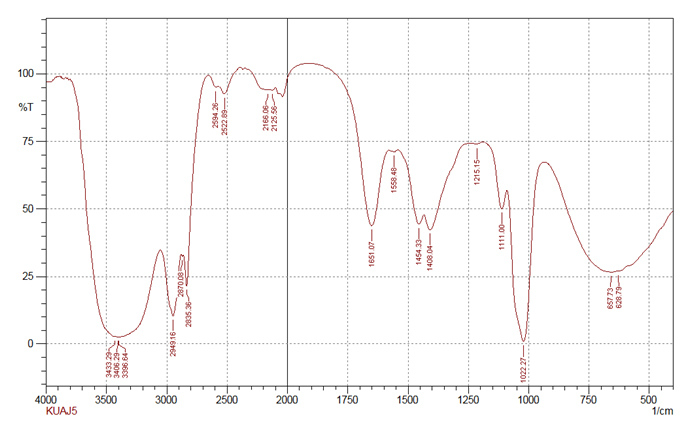 Fig. 7. Infrared spectrum of the partially purified antimicrobial compound produced by KUAJ5
Fig. 7. Infrared spectrum of the partially purified antimicrobial compound produced by KUAJ5Forest soil samples from Ooty, Tamil Nadu were subjected for investigation and 22 actinomycetes were isolated. On primary screening six isolates showed activity against one or the other of the test organisms in right angle streak method. Potential isolates from primary screening were subjected for secondary screening by disc diffusion method. Among the screened isolates KUAJ1and KUAJ5 showed intense activity. The active isolates of primary screening when subjected to secondary screening showed different activity from that of primary screening; some of the active isolates did not show the activity in the secondary screening while some showed little activity. According to Bushell (1997)26, during the screening of novel secondary metabolite, actinomycetes isolates often show antibiotic activity on agar but not in liquid culture. TLC results of KUAJ 1 extract shows an Rf value of 0.75 which is similar to that of kanamycin and KUAJ5 extract with Rf value 0.48 shows similarity with thienamycin whose Rf values is0. 49 according to Monisha khanna et al. Maximum UV absorbance peak for KUAJ1 is 218 nm and KUAJ5 is 240 nm which are in the range of 215- 217 nm which indicates a highly polyene nature. From UV- visible spectroscopic analysis it was found that both the strains are able to produce broad spectrum antibacterial agents which absorbs at UV range. FTIR results exhibited O-H , C-H and CH2-S groups in KUAJ1 and N-H, CH3-S groups in KUAJ5 extract. Antibacterial activities of polymer coated cotton impregnated with bioactive molecules were also studied. Chitosan and sodium alginate coated cotton fabric produced zone of inhibition as the polymer itself has got some antimicrobial property according to O.L Shanmugasundaram. Antibacterial activity results resemble that of secondary screening results. Both the samples were able to inhibit E.Coli, one of the test pathogen to the maximum. So the results indicate that the bioactive molecules produced by the actinomycetes isolated from forest soil can be applied on textile for antibacterial finish.
Actinomycetes were isolated from forest soil and their antibacterial efficacies were tested. Selected strains KUAJ1 and KUAJ5 exhibited a broad spectrum activity. Antibacterial biomolecules produced by these strains coated on cotton fabric showed antibiogram. Based on these results it has been shown that the forest soils possess various antibiotic producing actinomycetes and may be considered as one of the potential source of bioactive compounds producing actinomycetes which can be used in textile industry for antibacterial finish.
ACKNOWLEDGMENTS
The authors acknowledge the management of Karpagam University for providing all necessary facilities to carry out the study.
- Venkata Ratna., Ravikumar Dasari., Murali Yugandhar Nikku., Sri Rami Reddy Donthireddy. Screening of marine actinomycetes: Optimization of process parameters for the production of novel antibiotics by Amycolatopsis Alba Var.nov DVR D4. J. Microb. Biochem .Technol. 2011., 3(5): 092-098.
- Mani Jayaprakashvel.Therapeutically active biomolecules from marine actinomycetes. Journal of modern biotechnology., 2012; 1(1):1-7..
- Jahir Alam., Khan,Anand., Singh Patel.Extraction and purification of antibacterial metabolite from actinomycetes spp. Isolated from soil samples. International journal of pharmaceutical research and development., 2011; 3(10): 63-71.
- AmitPandey., Imran Ali., Kailash Singh Butola., Tanushri Chatterji., Vidyottama Singh. Isolation and characterization of Actinomycetes from Soil and evaluation of antibacterial activities of Actinomycetes against pathogens. Int. J. App. Boil.Pharm.l Technol., 2011; 2: 384 – 392.
- Sunil LaxmanAttimarad., Gaviraj N., Ediga., AsifAbdulrahimanKarigar., RavindraKaradi., NageshChandrashekar., Chandrashekara Shivanna.Screening. isolation and purification of antibacterial agents from marine actinomycetes. International journal of current Pharmaceutical Journal., 2012; 1(12):394-402.
- GarimaArya,P., Koushik,Padma Singh.Screening of actinomycetes from biodegraded buiding material anad their antibacterial potential. Int. J. Curr. Microbiol.Sci 2013; 2(6): 113-116.
- Gulrajani, M., L,Deepti, Gupta.Emerging technique for functional finishing of textiles. Indian. J. Fibre Text . Res., 2011; 36: 388-397.
- Shahidi, S., Ghoranneviss, M., Moazzenchi, B., Rashidi A., Mirjalili M.Investigation of Antibacterial Activity on Cotton Fabrics with Cold Plasma in the Presence of a Magnetic Field.Plasma Process and Polym. 2007; 4(S1) S1098–S1103.
- Dierk, Knittel., Eckhard Schollmeyer. Chitosans for permanent antimicrobial finish on textiles. Lenzinger Ber. 2006; 85: 124-130.
- JemimahNaine, S.N., Nasimunislam, B., Vaishnavi,V., Mohanasrinivasn,C., Subathra Devi, C. Isolation of soil actinmycetes inhabiting Amirthi forest for the potential source of bioactive compounds. Asian J. Pharm . Clin. Res., 2012; 5(3);189-192.
- Baskaran, R.,Vijayakumar, R., Mohan, P.M. Enrichment method for the isolation of bioactive actinomycetes from mangrove sediments of Andaman Island India. Malays. J. Microbiol., 2011; 7(1) :22-28s.
- Jennifer Emelda, E.N., Vijayalakshmi,T., Santhanakrishnan..Antibacterial activity of antibiotic producing Streptomyces macrospores. Int.J. Pharm.Biol. Sci., 2012; 2(3):20-23.
- Nanjwade Basavaraj, K.,Chandrasekhara, S., Ali. M., Shamarez, Prakash S., Goundanavar., Fakirappa, Manvi, V.Isolation and morphological characterization of antibiotic producing actinomycetes. Tropi. J.Pharm .Res., 2009; 9(3): 231-236.
- Raja, R., Seetha, S., Sahana, V.Preliminary studies on antifungal activity of actinomycete isolated from ice point. International journal of advanced research., 2013; 1: 17- 22.
- Monisha, Khanna., Renu, Solanki., Rup, Lal. Selective isolation of rare Actinomycetes producing novel antimicrobial compounds. Int.J.Adv.Biotechnol. Res., 2011; 2(3):357-375.
- Gebreselema, Gebreyohannes., FelekeMoges., Samuel, Sahile., Nagappa, Raja,.Isolation and characterization of potential antibiotic producing actinomycetes from water and sediments of Lake Tana, Ethiopia. Asian Pac. J.Trop. Biomed., 2013; 3(6):426-435.
- Houssam, Atta, M., Maged, S., Ahamad. Antimycin –A Antibiotic Biosynthesis produced by Sreptomyces sp. AZ-AR -262: Taxonomy, Fermentation, Purification and Biological Activities. Aust. J.Basic Appl. Sci., 2009; 3(1):126-135.
- Bhagabati, Pandey., Prakash, Ghimire., Vishwanath, Prasad., Agarwal.Studies on the antibacterial activity of the actinomycetes isolated from the khimbu region of Nepal.
- Kariminik A,Baniasadi F.Pageantagonistic activity of actinomycetes on some Gram Negative and Gram positive bacteria.World appl. Sci. J; 2010; 8(7): 828-832.
- Rajendren, R., Abirami, M., Jagadeeswari, S., Prabhavathi, P.Production, optimization and partial purification of antimicrobial compound from Streptomyces exfoliates. A journal of science and technology, 2(1):58-62.
- Usha, R., Prabu, E., Palaniswami, M.,Venil, C.K., Rajendran, R.Synthesis of metal oxide Nano particles by Streptomyces sp for development of antibacterial textiles. Global J. Biotechnol. Biochem., 2010; 5(3):153-160.
- Mirajilili, M., Nasirian, S.S., Karimiu, L. Effect of corona discharge treatement on some properties of wool fabrics. Afr. J. Biotec., 2011; 10: 19436-19443.
- Shanmugasundaram, O.L. Development and characterization of cotton and organic cotton gauze fabric coated with biopolymer and antibiotic drug for wound healing. Indian. J. Fibre Text . Res., 2012; 37:146-150.
- Badji, B., Zitouni, A., Mathieu, F., Lebrihi, A., Sabaou, N . Antimicrobial compounds produced by Actinomadura sp. AC 262 isolated from an Algerian Saharan soil. Can.J.Microbiol., 2006; 52(4):373-382.
- John Coates. Interpretation of Infrared Spectra, A Practical Approach. Encyclopedia of Analytical Chemistry . R.A. Mayers (Ed) pp 10815-10837. John Wiley & Sons Ltd, Chichester, 2000.
- Bushell M. E,Smitht V,Lynchs H.C.1997. A physiological model for the control of erythromycin production in batch and cyclic fed batch culture. Microbiology 1997; 143:475-480.
© The Author(s) 2018. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.


