Central nervous system (CNS) infection is a serious illness that can lead to death. CNS infections include meningitis, encephalitis, brain abscesses and myelitis. These diseases are caused by causative agents like bacteria, fungi, parasites, and protozoa, but most commonly by viral infections. To combat this issue, accurate diagnosis of etiological agents at an early stage is crucial for appropriate treatment, control of the disease and prevent from becoming life-threatening to the patients. This review paper summarises the main laboratory diagnostic methods for CNS infections caused by viruses ranging from conventional to molecular methods. Conventional isolation methods are considered the ‘gold standard’ as they provide accurate evidence, but require highly skilled personnel, are time-consuming, critical in cell type selection and are useless for non-cultivable viruses. Electron microscopy allows recognition of viral morphology and ultrastructural details as the principle of virus identification through negative staining or thin section technique (suitable for tissue or cell specimens). However, it offers low sensitivity and requires at least 106 virions per millilitre or milligram in the specimen to be detectable by microscopy. Immunological-based methods have been extensively applied for viral diagnosis by detecting the antiviral antibodies or viral antigens in clinical samples. While these methods provided high sensitivity and specificity, the incubation and window period of an infection may give false-negative results. Lastly, molecular detections have many advantages such as high sensitivity, specificity, rapid, require a small amount of sample, simultaneous detection of multiple different viruses, and produce both qualitative and quantitative results.
Viral infections, CNS, Virus isolation, Electron microscopic, Serological methods, PCR
The central nervous system (CNS) comprises the brain, spinal cord and retina, and is considered the control panel for our bodies responsible for cognition, movement, senses, and emotions.1 The CNS has complex and unique anatomical and immunological characteristics that help identify the pathogenesis and detection of infections.2 It is protected by bony structure (skull and vertebrae) and reserved with the blood-brain barrier (BBB), which serves as a selective barrier for pathogen entry.3 However, the breach of BBB can allow the microbial invasion due to the damaged surrounding tissue, obstruction of microvessels by parasitised leukocytes, platelets, or red blood cells, over secretion of cytokines that affect the tight junction proteins, or interactions of microbe with BBB that facilitate the passage of microorganism.4
Acute infection of CNS depends on the anatomic localisation in the meninges, brain or spinal cord, which results in meningitis, encephalitis, brain abscesses and myelitis, respectively.5 Additionally, the infections can also occur in multiple sites, resulting in a combination of diseases such as meningoencephalitis. Meningitis is an inflammation (swelling) located at the membranes protecting the brain and spinal cord from bacterial or viral infection.6 Encephalitis is an inflammation of brain parenchyma caused commonly by viruses.7 While, brain abscess is a focal area of necrosis with surrounding membrane within brain parenchyma and is usually caused by bacteria, fungi infection, or traumatic injury.8 Myelitis is inflammation across the spinal cord with one or more levels and in the absence of compressive lesion.9 Few etiological agents responsible for causing CNS infections are bacteria, fungi, parasites, protozoa or viruses. The clinical manifestations include fever, headache, vomiting, stiff neck, seizure, altered mental status and behaviour changes.10
Common viruses that lead to CNS infections are the human enterovirus (HEV), herpes simplex virus type 1 (HSV-1), herpes simplex virus type 2 (HSV-2), varicella-zoster virus (VZV), cytomegalovirus (CMV) and Epstein-Barr virus (EBV).11 Among HEV, coxsackie A16 (CA16), enterovirus 71 (EV71) and echovirus (ECHO) are the most common.12 CNS infections by HSV were estimated with 2.3 cases/million in population yearly, and HSV-1 contributed to 95% of all cases.13 Mumps virus (MuV) and measles virus (MeV) infections are reported to be resurgent worldwide.14,15 John Cunningham (JC) virus, which causes progressive multifocal leukoencephalopathy (PML), has often been excluded from routine detection.16 Amongst CNS infections by the arthropod-borne virus (arbovirus), the Japanese Encephalitis virus (JEV) is the leading cause of viral encephalitis in Asia and the Western Pacific; others include Dengue virus (DENV), Zika virus (ZIKV), West Nile virus (WNV) and Chikungunya virus (CHIKV).11,17
CNS infection is ten times more frequent in developing countries than in developed countries, in which many cases were diagnosed clinically and empirically treated without confirming the etiological agents.18 These scenarios can be life-threatening when confronted with cases that present non-specific symptoms at an early stage of the disease.19 Thus, a reliable laboratory diagnosis in the early stage of a CNS infection is vital for properly managing the infected patient.10,20 Currently, radiological investigations are used to diagnose CNS diseases using neuroimaging tests such as electroencephalogram (EEG), computerised tomography scans (CT) and magnetic resonance imaging (MRI).21 However, these tests only detect the pathology of the infection but not the causative agents.
The main laboratory investigations to identify the causative agents are (1) virus isolation in cell culture technique, (2) visualisation of viral structures using electron microscopy, (3) detection of viral antibody or antigen using immunological methods, and (4) detection of viral nucleic acids using molecular methods.22 Isolation of viruses is the ‘gold standard’ method. However, it is laborious, time-consuming, and requires high cost and skill.23 Electron microscopy provides images at a near-atomic level, but it is less sensitive as it requires a higher sample limit for detection than other methods.24 Immuno-based techniques, on the other hand, provide better sensitivity but is highly dependent on the disease progression in the patient, i.e. antibodies may be undetectable during the window period.25 Molecular methods such as PCR-based assay are rapid, highly sensitive and specific, and suitable for early and accurate diagnosis. Still, they are more costly in requiring specialised laboratory equipment, reagents and trained staff.26 This review will explore the current laboratory methods used to diagnose CNS viral infections.
Viral culture
Virus isolation is maintained as the ‘gold standard’ for laboratory diagnosis of viral infections as it provides accurate evidence for the presence of infectious viruses in clinical samples such as cerebrospinal fluid (CSF) and blood, especially when CNS disease is suspected.27 Some cultivable viruses include HSV-1, HSV-2, CMV, VZV, respiratory syncytial virus (RSV), influenza A and B viruses, parainfluenza viruses, HEV, MeV and respiratory adenoviruses.22 Detection of viruses can be done in various cell lines by means of cytopathic effects (CPE), hemadsorption (HAD), or immunologic testing.28 There are three basic cell lines that can be used in cell culture, including primary, diploid and continuous or heteroploidy.29
Conventional viral cultures usually utilise different cell lines to identify which cell will be infected by the virus from the sample.23 The appearance of a typical CPE will be observed up to 30 days of incubation.30 CPE is the morphological alterations of the cells after being invaded by specific viruses, and the characteristics are different between viruses.22 In addition, CPE result also depends on the virus and cell line involved, time of incubation and type of clinical specimen, which gives prediction to determine the type of virus.23,30 Enteroviruses are claimed to be cultured easily with 75% sensitivity and take a 3-8 days turnaround.31 On the other hand, HSV has been cultured from CSF for less than 5% of encephalitis cases, probably due to culture technique that did not provide a diagnosis within the time frame of clinical decision making.32 After all, a confirmatory examination using light microscopy after fixation and staining is needed to observe additional diagnostic features such as inclusion bodies and syncytium formation.33
In the case of the absence of CPE, hemadsorption is another alternative to be considered when involving the influenza virus and parainfluenza virus.34 Hemadsorption is the ability of some virus-infected cells to bind with red blood cells on their surface due to the expression of the hemagglutinating protein on the infected cell’s plasma membrane that is attracted to the ligand of the erythrocyte.35 By that, if haemadsorbing virus is present, erythrocytes will adhere in a clump and show agglutination under a microscope.23 Similar to CPE, a confirmation test such as the immunologic method or PCR test is needed to confirm the identity of the virus.
Rapid culture has been preferably used in laboratory diagnosis over conventional culture as it can increase the speed and specificity of various viruses such as HSV, CMV, VZV, influenza virus, RSV, adenovirus, and parainfluenza virus.2,34 Shell vial or spin-amplified culture is a modified technique where the clinical sample will be inoculated onto the grown monolayer cell on the coverslip at the bottom of the shell vial culture tube, proceed with slow centrifugation and incubation, which can cause minor trauma to the cell surface and enhance the viral entry to infect the cells.36 After 2-3 days, coverslips are fixed and stained with the fluorescent-labelled antibody of a specific virus and examined under a fluorescence microscope.33 This technique allows the detection of a pathogen before CPE occurs.
Another advanced technique of rapid cell culture is called mixed vial, where different cell lines will be grown on a vial with different monoclonal antibodies so that they can detect different types of viruses within the same vial at one time.29 Previously, the combination of MRC-5 and A549 cell lines were used as monolayer cells on vial for diagnosis of CMV, HSV, and adenoviruses, and proven to produce 88.9%, 100%, and 93.8% of sensitivity, respectively.37 But recently, other mixed-cells are available to be used, such as R-Mix for detection of adenoviruses, parainfluenza virus types 1, 2, and 3, influenza virus types A and B; respiratory syncytial virus H&V-Mix for diagnosis of HSV, CMV, VZV, and E-Mix for enteroviruses diagnosis.22,30 Interestingly, genetically modified cell lines have been developed which utilised enzyme linked-inducible system (ELVIS) for diagnosis, whereby a specific viral receptor will be transfected into the cell lines to enhance virus infection towards susceptible cell, thus stimulating the production of enzyme which then allows visible staining of infected cells.38
Despite the advantages of rapid culture, this technique has the weakness of only detecting targeted viruses (less sensitive).29 On the other hand, conventional culture provides detection of unknown viruses, and the viral isolates produced allow for additional investigations such as antiviral testing, epidemiological study and serotyping. However, conventional culture techniques require highly skilled-personnel, are time-consuming, critical in terms of cell type selection, and unusable for the non-cultivable viruses.23,24 Undoubtedly, cell culture technique has its own strengths, however, to diagnose CNS diseases, the CSF sample is often small in volume and difficult to obtain, thus making the cell culture method unfavourable over other methods.2 A study in 2011 found that two out of three samples were positive for echovirus in cell culture, while the other was inconclusive due to insufficient sample amount.39 Hence, larger CSF volumes are usually requested to improve the sensitivity of cell culture methods.31
Direct detection using electron microscopy
Back in the 1970s, electron microscopy was used in the discovery of several unknown viruses which cannot be cultivated, such as human rotaviruses, caliciviruses, astroviruses, hepatitis A virus, some unknown types of adenoviruses and coronaviruses.24,40 Electron microscopic allows recognition of viral morphology and ultrastructural details of cellular infection as a principle in viral identification.41 The two methods employed in electron microscopy are negative staining and thin section microscopy. Negative staining is the most widely used in which the fluid containing the virus will be ultracentrifuged to concentrate the virus in the sample before being placed on a carbon or formvar-coated grid, followed by the addition of electron-dense fluid, which leads to the virions adhering to the grid.24 The discovery of the Nipah virus in Malaysia was made with the application of negative staining on CSF samples that revealed the enveloped structures with surface projections, nucleocapsid with unique helical and herringbone patterns, which were characteristically similar to other paramyxoviruses.42
Thin section microscopy is used when tissues or cells are thick and difficult for the electron beam to penetrate whole thickness, such as after cell culture isolation of viruses from samples.43,44 In the United States, patients bitten by mosquitoes or ticks have been diagnosed with bunyavirus infection using transmission electron microscope (TEM) and showing an encephalitis syndrome.45 In another case, a virus from Arenaviridae was detected using ultrathin section of grown Vero cells containing CSF sample, and was identified as lymphocytic choriomeningitis virus (LCMV), which is an arenavirus transmitted by rodents.46 TEM is also possible to directly detect brain tissues infected with polyomavirus, which can cause progressive multifocal leukoencephalopathy (PML) in the absence of DNA virus in the CSF sample.44,47 However, the limitation of thin sectioning method is that if the infection is focal, the sampling process might not recover tissue samples that contain viruses.43
The most recent method used is the cryo-electron microscopy (cryo-EM) where the sample is observed at cryogenic temperature (less than -160°C) by which the water molecules will crystallise and transforms the biomolecules such as virus particle into the rigid state to obtain a high-resolution image.48 Major advantage of cryo-EM is its ability to examine the virus at near-atomic resolution, which allows examine the structure of macromolecules and their complexes, proteins interactions and emergence of unusual features of viruses that are often linked to increased virulence and stability of the virus.49,50 Recent studies have utilised the cryo-EM technique to reveal the mechanism related to Eastern Equine Encephalitis Virus (EEEV) entry, disassembly, antibody neutralisation and information on capsid genome binding segment structure.51 Besides, cryo-EM can detect the structure of JEV capsid protein that consists of conserved dimeric protein fold made by the anti-parallel pairing of the α1-a1’, a2-a2’, and a4-a4’ helices to distinguish from other flavivirus such as DENV, WNV and ZIKA capsid proteins.52 However, the cryo-EM method is too technically challenging for use solely for diagnostic purposes.
Overall, the biggest limitations of electron microscopy are low sensitivity and requiring at least 106 virions per millilitre or milligram in the specimen to be detected by microscope.24 But, it has the advantage of high magnification and resolution, and wide amplitude focuses, thus able to view the morphology, and chemical-physical composition of the sample, examine the contaminants in tissue, and assess the protein expression at the subcellular scope.53 In some cases, the combination of diagnosis methods can be applied to detect the specific virus causing CNS disease. For instance, a reovirus strain in urine and throat swab samples from two children showing unexplained neurologic symptoms of encephalitis has been cultured in MRC5 cells, followed by molecular technique methods and TEM to confirm the results and contributed to findings of new reovirus strain as an etiologic agent for encephalitis.54 Immuno-electron microscopy (IEM) is another method which utilises a serological principle such as ELISA and followed by virus identification using TEM diagnosis.55 Immune gold labelling is one of the IEM methods where an electron-dense marker is made from a specific antibody labelled with particles of colloidal gold used to detect whole virus antigens or antigens in thin section tissues or cells.56 Hence, the application of mixed methods can increase the accuracy and specificity of detection.
Serology
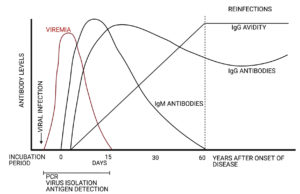
Demonstration of specific antibody response is also another efficient diagnostic method. Serological diagnosis of CNS infections is obtained by detecting the IgM antibodies or investigating neutralising antibody titres to at least a fourfold increase between two consecutive samples taken during the acute and convalescent phase.57 Figure 1 shows the standard graph indicating the progress of antibody levels following viral infection.58 After the incubation period, the virus starts to multiply, and symptoms appear for about a few days to weeks. Antibodies gradually began to produce until it reaches the level of detection and the amount of infectious viruses started to decrease. During primary infection, the immunoglobulin M (IgM) level is high, whereas the avidity of immunoglobulin G (IgG) is low but increases gradually, and the IgG level is positive but much lesser than IgM. Measurement of avidity is important to distinguish the status of infections, whether acute or chronic/past infections. Secondary infection or reinfection by the same antigen will make changes which IgG avidity levels increase rapidly, reaching the highest level, IgG antibodies are still positive, while IgM may not be able to detect.58 At this moment, detection of IgG is useful to monitor the amount of antibodies against antigen in the human body. Few methods can be used to detect the virus antibody; neutralising antibody assay, hemagglutination inhibition test, immunoblotting, enzyme immunoassays and immunofluorescent.25,59
Figure 1. Typical antibody response after viral infections, and recommended detection methods (Created with BioRender.com)
Neutralising antibody assay
A neutralising antibody is an antibody that will reduce the level of infectious capacity of virus by neutralising cell-free virus, thereby reducing the spread and eliminating the possibility of cell infection.60 The virus will be detected by observing the cytopathic effect (killing cells) or the production of antigen after cells infection and detected by immunofluorescent or immunohistochemistry.59 Neutralising antibody assay is highly sensitive and specific for diagnosis. Still, it is time-consuming, requires infectious virus in the test, and is laborious and high cost for cell culture work, thus rarely applied.58,61 This assay is more beneficial in assessment studies such as examining the vaccine’s effectiveness, assessing therapeutic strategy, controlling viral persistence and predicting immune protection.60,62,63
Hemagglutination inhibition (HI) test
Certain viruses contain hemagglutinin molecules on their viral envelop that will aggregate erythrocytes when in contact and form an agglutination. This effect will be inhibited when specific antiviral antibodies are present in the serum. Diagnosis of viral infections is achieved by detecting such antibodies that will inhibit the clumping of erythrocytes when challenged with said viral particles.48 The benefits of this test are sensitive, specific, less technical, have fast results, and is cost-effective. However, it is limited to hemagglutinate viruses such as arthropod-borne viruses, influenza viruses, and parainfluenza viruses.35 Serum sample will be serially diluted with twofold steps into each well of a 96-well microtitre plate, followed by the addition of a constant amount of virus about four hemagglutinating units, incubation, and addition of an equal volume of red blood cell into each well.59,64 This method is usually applied to influenza A virus to detect the hemagglutinin subtype from an unknown isolate by subtype-specific antibody, identify the specificity of the antibody and quantify the antibodies that allow the study of antigenic reactivity between influenza isolates.65,66 However, a study in Japan reported that diagnosis of dengue using the HI test may produce false-positive results due to cross-reactive antibodies among dengue serotypes leading to non-specific inhibitors in both acute and convalescent samples. This can be minimised by using different animals’ red blood cells.64,67
Immunoblotting
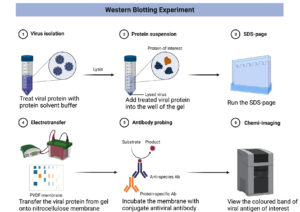
Immunoblotting also known as western blotting, is used to diagnose viral infections by detecting viral protein or antiviral antibodies. The concentrated virus will be solubilised, and the proteins are separated by sodium dodecyl sulphate-polyacrylamide gel electrophoresis (SDS-PAGE), then the separated proteins are transferred onto nitrocellulose membrane or PVDF membrane electrophoretically. Lastly, viral protein will be incubated with enzyme-label antibody, and if the specific binding is present, formation of a coloured band appears after the enzyme substrate is added.68 To detect antiviral antibodies in a patient, the serum (considered a primary antibody) will first be allowed to bind with separated viral proteins on nitrocellulose and then incubate with a secondary enzyme-labelled anti-human antibody. This complex is detected by observing visual bands after adding the enzyme-substrate.25 Figure 2 summarises the overall process of western blot.
The immunoblotting assay was applied for detection of anti-chikungunya virus antibody in human serum. It resulted in high sensitivity (83.3%) and specificity (96.7%) when tested on 30 confirmed infected patients and 30 normal sera.69 Historically, this method achieved high sensitivity, which detected 53% of acute hepatitis C virus among enzyme immunoassay’s negative samples.70 Overall, Western blotting allows the demonstration of antibodies to specific or all viral proteins, and can be used to identify the presence of different antibodies to different viral antigens at a different stage of infections. However, the results produced are more qualitative than quantitative in nature and difficult to be standardised across from different laboratory setups.59
Enzyme immunoassays
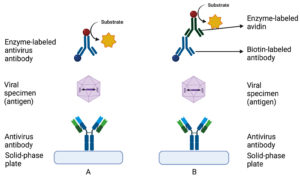
Enzyme immunoassays (EIA) are reaction between enzyme-conjugated monoclonal antibodies with viral antigen protein and visualised by enzymatic colour reaction.71 Enzyme-linked immunosorbent assay (ELISA) is the most common approach to detect antiviral antibodies. Several types of enzymes used in ELISA are alkaline phosphatase, horseradish peroxidase (HRP), and β-galactosidase.72 Antibody-capture ELISA is called indirect ELISA, while antigen-capture ELISA is called direct ELISA (Figure 3).24
Figure 3. Detection of antiviral antibody or antigen by enzyme immunoassays. A: Direct method. Antivirus antibody is ‘locked’ at the solid-phase plate, specimen containing viral antigen will bind specifically to the antibody. The antigen-antibody complex will be detected by an enzyme-labelled antivirus antibody. B: Indirect method. The viral antigen is bound specifically to the biotinylated antiviral antibody and the antigen-antibody complex is detected by adding enzyme-label avidin (enhance sensitivity), followed by enzyme substrate to produce a colour reaction that can assess by eyes or spectrophotometer (Created with BioRender.com)
In a study conducted in Southern Vietnam, ELISA was used to detect Japanese encephalitis virus (12%), Epstein-Barr virus (8.2%), dengue virus (6.5%), enterovirus (2.7%), rubella virus (2%), varicella-zoster virus (1.7%) and cytomegalovirus (1%) among 291 viral meningitis/encephalitis patients.73 ELISA was also able to detect the presence of JEV IgM in patient CSF sample months after the acute phase.74 IgM detection also proved to be crucial for diagnosis of neuroinvasive WNV compared to CSF PCR which contributes a lesser sensitivity (57%).75 However, serological investigations for antibodies might not be accurate in scenarios such as in immunocompromised patients where IgM may be false-negative because antibody response had not mounted yet.2 Repeat IgM assay after seven to ten days is recommended as antibody production is delayed.76 In some reports, serological investigations provided more sensitivity than PCR.77,78 However cross-reactivity with other species of viruses and related vaccines can be an issue.79 The use of monoclonal antibodies and production of recombinant antigens can help alleviate this issue by enhancing the sensitivity and specificity.59
Immunofluorescent assay
Unlike ELISA, immunofluorescent has proved to be able to identify the viral antigens from patients earlier in the disease process.24 This method can be used on infected tissue, nasopharyngeal aspirates, vesicle fluid, faecal specimens and serum samples.58 The immunofluorescent assay mainly consists of two methods; direct immunofluorescent assay (DFA) and indirect immunofluorescent assay (IFA).68 For DFA, the infected cells specimen collected from the patient is centrifuged to obtain the pellet, fixed on glass slides, and stained with fluorescent-labelled virus specific antibodies. The unbound antibody is removed by washing, and the viral antigen will be observed under fluorescent microscope to view the virus-specific staining.29 While for IFA, the antiviral antibody is untagged and it is recognised by the secondary fluorescent-conjugated antiviral antibody.68 The most favourable fluorescent dye is fluorescein isothiocyanate (FITC) which gets excited by blue light (490 nm) and produces green light (519 nm).25
Antigen detection methods are usually applied for cases of virus reactivation such as HSV and VZV where the antibody response may be weaker.58 IF is often used for rapidly diagnosing infections by associated respiratory viruses such as paramyxoviruses, adenoviruses and orthomyxoviruses.24 A study stated that DFA showed much higher specificity (99-100%) for detection of respiratory syncytial virus (RSV) in children, compared to RT-qPCR method.80 Interestingly, one study utilised IFA method for subtyping of influenza A and turned out to have the same accuracy as RT-PCR.81 Immunofluorescent assay test kit has been commercialised for detection of ZIKV (Anti-ZIKV IIFT). The researchers tested the performance of the kit on 126 positive and 102 negative samples and showed 96.8% sensitivity and 72.5% specificity. The lower specificity might be due to cross-reactions with other flaviviruses.82 Thus, the viral antigen and monoclonal antiviral antibody should be carefully developed to prevent cross-reactivities.
The advantages of immunofluorescent assay are having excellent sensitivity, reduced cost, and at the same time, multiple viruses can be detected simultaneously and rapidly within 2 hours, compared to other antigen detection methods.83 Factors that impact the interpretation include the colour and intensity of stain used, distribution of proteins in the cells, the infections of cell types and number of positive cells appeared.84 These limitations can be overcome by using specialised equipment (fluorescent microscope), skilled slides preparation, and interpretation of results in order to differentiate between specific and non-specific staining.85
Molecular diagnostic technique
Detection of IgM and IgG antibodies undoubtedly give useful information on individual immune protection level, but the test requires five to seven days after infection makes unsuitable for certain conditions especially for rapid diagnosis in most cases.25 In addition, turnaround time for culture technique requires 1-28 days and the results may appear negative as patient received antiviral drug.86 While, PCR can still maintain its sensitivity if tested on the same condition sample.32 Molecular diagnostic method involves amplification of viral genomic material which known to be rapid, enable to detect different viruses at the same time with high sensitivity and specificity.68,87 The main purpose of conducting nucleic acid test is to produce timely results which is important for the improvement of patient care at a reasonable cost.88 Nucleic acid detection is crucial when involving the following conditions: (1) uncultivated or difficult to culture viruses; (2) sample contains inactivated virus due to extend storage, fixation of tissue or transport; (3) latent phase which viral genome is dormant and infectious virus is not available; and (4) virus with antibody found in acute infections or during continuous viral infection.59
Conventional Polymerase Chain Reaction (PCR)
The principle of PCR starts with extraction and purification of DNA molecules, amplification of targeted region with presence of DNA polymerase, primers and nucleotides, followed by detection of amplified product using few techniques such as gel electrophoresis, colourimetric method such as intercalating dyes or fluorescently labelled probes, and sequencing.26,88,89 PCR technique has been widely used for identifying the viral infection with great sensitivity ranges from 77.8% to 100% and specificity ranges from 89% to 100%.26 The main processes involve in PCR include denaturation, annealing and extension. Firstly, denaturation phase of DNA template by thermocycler that heats DNA usually at 95°C that enable to break the hydrogen bonds holding the double-stranded DNA to break into a single strand. Second, as the temperature reduced (55°C), primers will allow to bind complementary at each separated strand and flank region to be amplified. Lastly, thermocycler will raise until optimum temperature (commonly at 72°C) for Taq DNA polymerase extends the primers by adding nucleotides complementary to the template, thus creating new copies of DNA product.71,90 These cycles will be repeated about 35-40 times to increase number of copies.58
Usually, PCR products will be visualised by gel electrophoresis to check the presence of DNA band which facilitated by known DNA standard besides the unknown sample.91 Then, the sizes between amplified product and control amplicons are compared in the same gel. PCR product also can be sent for sequencing and compare the unknown-product sequences with known virus sequences to allow the identification of species, strain or subtype.58 Diagnosis of herpes simplex encephalitis using PCR has led to 98% sensitivity and 94% specificity.31 PCR sensitivity depends on the type of DNA stain used and type of system to visualise the gel.92 While, specificity can be increased by thermostable polymerase because cycling can be done under firm annealing conditions to reduce mismatch base pairing that can cause false-positive results.59 Conventional PCR technique is undoubtedly able to detect many viruses causing CNS infections such as VZV, enterovirus, human herpesvirus-6, EBV, CMV, JC virus, and WNV. However, issue arises involving this technique is when the samples contain low concentrations of DNA due to delayed collection sample or rapid clearance caused by neutralising antibody response.26,32 For example, HSV encephalitis results may appear false-negative CSF PCR when specimen is collected too early or too late.93 If the patient is highly suspect for HSV, thus repeat CSF PCR is required despite the early negative results.94 Another thing is conventional PCR has the possibility to cause cross-contamination during post-amplification process such as gel electrophoresis technique.95 This brings to another favourable technique to be used which is real-time PCR.
Reverse Transcription Polymerase Chain Reaction (RT-PCR)
RT-PCR is specially used for diagnosis of CNS infection by RNA viruses. RNA is first transcribed into cDNA using reverse transcriptase, then followed by amplification of cDNA using conventional PCR.68,96 Conventional RT-PCR is claimed as an excellent technique as it able to achieve high sensitivity range from 73-100% and specificity up to 99-100% for diagnosis of viral infection.97,98 Previously, RT-PCR is done with two-steps reaction, where after synthesized cDNA is produced in the first reaction tube, it will then be transferred into second tube for PCR.99 Nowadays, RT-PCR is done within a single tube where all components are included for both reactions (reverse transcription and amplification) and reactions will run at one time, hence produce reliable results since no possibility of contamination occurred.59 However, this technique is rarely used for diagnosis of clinical specimen due to high cost and time-consuming.100 Another alternative is the combination of reverse transcription with real-time PCR (RT-qPCR). This could be more beneficial than conventional PCR as it can eliminate the contamination, able to quantify the amplicons and quicker since absence of post-PCR activities.88 Recently, many researchers applied RT-qPCR for detection of RNA viruses such as ZIKV, HCV, RSV, dengue virus, and influenza A virus.101,102
Real-time Polymerase Chain Reaction
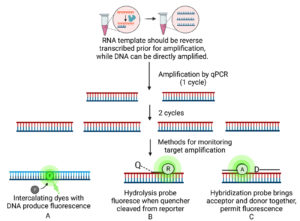
Real-time PCR or known as qPCR is a quantitative method that able to monitor the amplification of targeted amplicon in the reaction tube which facilitated by dyes intercalate with DNA molecules and fluorescent-labelled probes.103 SYBR green dye is a sensitive DNA binding dye as it will bind to any double stranded PCR products, thus the same master mix can be used for different PCR application.32 But, to use this dye, the primer must be highly optimised to prevent amplification of non-specific amplicons such as primer dimer as this will produce signal as well.103,104 Next, hydrolysis TaqMan® probes will tag to specific region of target DNA. As the primer extends by DNA polymerase, the probe is broken down and the linkage between reporter and quencher is cleaved thus resulted in the production of fluorescence.25 The benefits of using TaqMan® probes are able to eliminate nonspecific signal by specific hybridisation-based detection and different dyes are produce to monitor different sequences of amplicon in one tube.95 While, molecular beacon is a small DNA molecule with reporter (fluorescent dye) at 5’ end and quencher at 3’ end.68,105 It will hybridise to the DNA template at annealing step and as polymerase extends the sequence it will directly displace the molecular beacons, returning to stem-loop shape.106 The advantages of molecular beacons are simple, fast, sensitive, accurate, produce high-throughput format and can be used for multiplexing detection using different labelling probes in one tube.106 The amplification of PCR products can be monitored at each cycle by real-time monitoring intensity of fluorescence at a specific time.96 Figure 4 below shows a process of amplified DNA is detected using dyes and different types of probes.
Figure 4. Real-time PCR amplification. A: Dyes produce fluorescence as it binds to any double-stranded PCR product. B: Hydrolysis probe produces fluorescence as the quencher cleaved from reporter. C: Binding of probes to target region in close proximity to produce fluorescence (Created with BioRender.com)
Overall, real-time PCR provides rapid and high-throughput, quantification and detection of target DNA molecules within a shorter time.95 Besides, it can reduce the cross contamination because once reactants are added into tubes, it will never be opened for any further post-PCR handlings.88 This can reduce laborious work, cost, less technical and timely efficient to obtain the results.106 In addition, real-time PCR also has high sensitivity compared to gel system, and high specificity is achieved by the use of specific detection probes that requires it to bind to the target sequence only to produce the signal.59 Another advantage of qPCR is a wide dynamic range of quantification (7–8 log10) and multiplex amplification for several target molecules in a single reaction.107 However, to compare with serological and culture method, PCR requires high cost for primers and reagents, complicated operation and require reverse transcriptase for RNA virus detection.71
Singleplex qPCR and Multiplex qPCR
Singleplex PCR refers to amplification of single target sequences in a single reaction tube. While, multiplex PCR refers to simultaneous detection of multiple target sequences in a single reaction tube.108-110 Singleplex reaction is widely used. Recently many researchers apply multiplex CSF PCR assay to amplify nucleic acid from panels of viruses in their study to reduce the turnaround time, but still maintaining the sensitivity and specificity.111 Similar principle as singleplex qPCR which using intercalating dye and probes to indicate the amplification process. But as for multiplex qPCR, the probes-based multiplex must have multiple fluorescent channels to detect different viruses individually.112 Same goes to intercalating dye-based multiplex that require primers for each virus and having different melting temperatures.113
Many laboratories apply singleplex real-time PCR for detection of RNA and DNA viruses. Diagnosis of cytomegalovirus (CMV) using qPCR in saliva sample had resulted with 100% sensitivity and 99.9% specificity as compared to rapid culture technique.114 Besides, detection of Epstein Barr virus (EBV) as one of potential viral encephalitis also gave high sensitivity (95.7%) and specificity (100%), compared to serological assays.115 Employing this method also can provide information on the viral load in herpes simplex encephalitis patients which important to estimate the period for acyclovir therapy and clinical outcome.116 Next, Taq-Man real-time PCR has proved to successfully diagnose and differentiate five Japanese encephalitis virus genotypes.117 Interestingly, detection of respiratory viruses using singleplex qPCR provides better analytical sensitivity by observing the limit of detection, as compared to multiplex qPCR which resulted in loss of sensitivity.118 Additionally, the sensitivity of human enterovirus 71 in multiplex PCR assay resulted with 10-fold lower (103 copies) than singleplex assay (102 copies), however, detection of pan-enterovirus and coxsackie A16 assay using both approaches lead to identical sensitivity (102 copies). This may be due to the higher viral load than limit of detection in multiplex PCR assay.119 These data show that detection of viruses causing CNS infections using singleplex real-time PCR is highly accurate and reliable.
Multiplex PCR assay is widely used by many clinical diagnoses especially during pandemic or endemic outbreak of viral infections.26 However, it is said to have lower specificity as compared to singleplex assay.96,120 Thus, it is critical during designing the probes and primers which must be specifically able to detect specific virus only, otherwise interactions between them will affect the efficiency of multiplex PCR assay.113 A research made to assess performance of duplex RT-qPCR for detection of DENV and ZIKV showed a great analytical sensitivity as the limit of detection was similar when tested using singleplex assay. Plus, it also produced high specificity as no signals were measured from Flaviviridae family such as WNV, JEV, HCV, YFV and CHIKV.110 Interestingly, singleplex assay is usually produced great sensitivity on EV detection than multiplex assay, but one study proved to have higher sensitivity for EV-71 detection using FilmArray Meningitis/Encephalitis multiplex panel, compared to standard singleplex assay.121 This might be due to the high genetic variability of EV and variation of viral loads in CSF samples.122,123 Comparison findings on multiplex real-time PCR and multiplex conventional PCR was made to test the efficiency on diagnosis of CMV, EBV, HSV-1, HSV-2, JEV, and VZV in CSF samples. It suggested that multiplex qPCR able to detect neurologic viruses from 88 samples out of 147 CSF, while multiplex conventional PCR only detect six samples.124
Overall, detection of several viruses by multiplex PCR has brought to cost-effective, labour-saving and low turnaround time which require up to two hours compared to seven to ten days for virus isolation and immunofluorescence antibody detection.110,125 Rapid results will allow for easier clinical management in terms of decrease health cost and reduction of antimicrobial therapy duration as the disease can be treated earlier before the condition become life-threatening.111 Despite the advance technique of multiplex real-time PCR assay, this assay may require higher limit of detection as compared to conventional singleplex which may lead to reduce the performance in detecting viruses causing CNS infections.122 Moreover, when the number of viruses involved is higher, it will increase the number of detectable colours and making the system complex and high cost.113 Furthermore, issue related to multiplex is associated with competition of primers to bind with target sequence, also competition between multiple targets towards shared reagents.120 Thus, development of multiplex assay requires optimisation of reactions such as by adjusting the concentration of reagents, primers and annealing temperature set up so that each target sequence can be amplified with a similar manner without amplifying the nonspecific product.90 After all, multiplex panels are designed for qualitative results only.111 Most of pathogens will have high variability of nucleic acid load in CSF sample, without any relation with severity of disease.126 In this case, identification using qualitative method is sufficient to give guidance in clinical management. But, nucleic acid quantification using qPCR is beneficial to assess the antiviral therapy efficiency in immunocompromised patients.127 In cases if diagnosis using multiplex assay appears negative in certain target, confirmation test is needed to verify the results.111
Next Generation Sequencing
Molecular techniques such as PCR is still a relatively new approach as compared to the conventional methods, however, technologies have been evolving and changing at a rapid pace to replace this current method.128 Next generation sequencing (NGS) or also called as deep sequencing is the latest technique commonly used in detection of viruses causing CNS diseases by analysing the samples and generate a single sequence from each DNA and cDNA contain in the specimen. By analysing the sequences, researchers able to differentiate between the origin of sequence fragments, whether from human, specific bacterial species or a particular virus.129 Researchers claimed that it is difficult to identify the etiological agents by using traditional microbiological methods such as culture, morphological identification, serological and even PCR.130 The diagnosis is largely depending on clinical experience and prior knowledge of treating physician in predicting the pathogens, followed by selection of detection methods.131 This proved that traditional methods are target-dependent and limited in detection of rare and novel viruses.132 NGS also allows analysis of several genomic loci simultaneously, at the same time revealing the sequence changes hence beneficial in terms of detection of the mutated sequence or location.133 There are several types of NGS applications which are (1) target amplicon NGS that involves sequencing of specifically enriched targets using several methods of enrichment such as amplification or probe hybridization, (2) whole genome sequencing which assembles all DNA fragments across entire pathogen genome, and (3) metagenomic NGS (mNGS) which enables sequencing of all nucleic acids (DNA/RNA) for both pathogens and host within a specimen in a massive parallel way.134,135
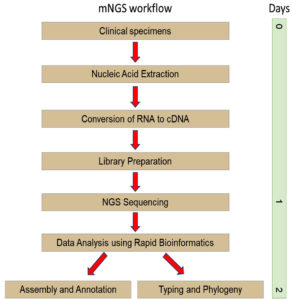
In recent years, many researchers preferred to use mNGS in clinical diagnosis of viruses causing CNS diseases such as encephalitis/encephalopathy, meningitis and meningoencephalitis, using mainly CSF and serum samples.136,137 mNGS is a broad and unbiased technique that able to identify universal pathogens causing CNS diseases within a single sample, simultaneously.138 The principle is quite similar with multiplex PCR, but mNGS provides wider detection as it is target-independent. Furthermore, this technique has the capability to combat limitations of current diagnostic tests which allowing for hypothesis-free, culture independent, and detection of universal pathogens directly from specimen within 2 days.139 According to the researchers, mNGS is frequently applied in growing numbers of laboratories as secondary test for negative result samples obtained using conventional methods.132,140 This is to improve the diagnosis results by detecting other viruses that might not be able to be detected using target-specific traditional methods which aimed only the most common pathogens.141-144 Figure 5 shows the main steps involves in the workflow of mNGS in detection of pathogens directly from clinical sample.
Figure 5. The overall steps involve in metagenomic next generation sequencing in detection of universal pathogens within 2 days’ time
Many findings had resulted with increase yield of viral detection causing CNS diseases by using mNGS technique. For example, VZV IgM antibodies has been resulted with 16.1% of CSF samples, while VZV PCR obtained 16.1% and 9.7% of positive rates using two different real-time PCR protocols. Interestingly, VZV mNGS had resulted with a maximal identification which is 51.6% (16/31) of the same CSF samples. Not only that, an unexpected detection has appeared which is human betaherpesvirus 6A.131 This reflects that agnostic mNGS is an important technique to be applied when the treating physician is not familiar with rare clinical situation. Researchers claimed that with the additional diagnostic yield percentage using mNGS had collected data over 500,000 meningoencephalitis cases worldwide.145 Hence, widespread implementation of mNGS is expected to lead to an increase in the number of identified etiologies and correctly diagnosed cases.132 Another study also applied mNGS in diagnosis of a four-year-old boy having an uncertain symptoms, after the conventional methods portray negative results towards both CSF and blood samples. mNGS showed the presence of herpes simplex virus 1 from brain tissue specimen, confirming the diagnosis of herpes complex virus encephalitis.146 This proved that mNGS is able to give an accurate result. The patient received acyclovir for three weeks, his condition was improved and discharged from the hospital. Furthermore, the efficiency of mNGS is not affected if the patient received empirical antimicrobial treatment compared to the culture method that can reduced approximately 20% of recovery.147
On the other hands, some published reports stated that mNGS detection is limited to individual patients or small retrospective case studies.139 Recent study showed that mNGS was only produced 22% positive rates among 58 infections. And among 26 patients, mNGS had resulted with negative results although the conventional microbiological testing exposed the infections.142 The false negative results might due to the low titers of pathogens in CSF or low sequencing depth of specimen, or could be diagnoses from samples other than CSF probably does not meet the standardized protocol in order to be detected using mNGS. To support the statement, published reports mentioned that currently there is no optimal and highly sensitive steps for library preparation established yet. Lack of standardization protocol and automation in mNGS has an impact on the selection of procedure that can be introduced into routine diagnostic testing to be able to achieve timely-detection within clinically relevant timeframe.132,134,144 Type and efficiency of extraction methods for different types of clinical samples are critical steps to achieve purely unbiased sequences.148 Another large prospective study also found that mNGS did not significant in predicting viral encephalitis and/or meningitis, and suggesting that DNA and RNA extraction methods should be improved in order to improve both nucleic acids positive detection rates.130 Sufficient sequences generated are important to ensure adequate sensitivity for detecting non-human sequences against the human background.149 Sensitivity of mNGS has been compromised in few studies, compared to amplification-based assay due to the low viral loads in the specimen which is a common problem.143,150 A recent study was conducted to counter this problem by enhanced the laboratory technique using hybrid capture (HC) of viral nucleic acids and methylated DNA depletion (MDD) to improve the sequencing analytical sensitivity. As the results, HC able to increase the viral read recovery in almost universally. However, MDD has mixed results whereby it only detected VZV in two samples that are found to be negative by standard mNGS, but also decrease the yield of detection on some viruses suggesting that its role in mNGS was uncertain.141 In contrary, the accuracy, sensitivity and specificity of mNGS in sufficient viral load clinical samples were 100%, 95% and 96%, respectively.144 Hence, more researches should be done using mNGS in order to obtain the best solution for the sensitivity issue in case of low viral load specimen. Although the sensitivity of mNGS is lower than conventional ones, both methods might complement each other to improve etiology diagnosis.
After all, implementation of mNGS in detection of viruses causing CNS infections is really useful especially involving detection of novel or rarely detected pathogens. Other advantages of mNGS include provide opportunities to reduce dependence on target-specific diagnostics, produce more pathogen genomic data and offer shorter turnaround times than serology.141 Apart from diagnostic purpose, this technique also capable in identifying the strain type directly from specimens, study of virulence, study of antimicrobial resistance genes, and assess the host immune response.134 Some disadvantages related to mNGS are expensive in terms of technologies and machines used, time-consuming compared to PCR, require an expertise in performing, testing and analyzing data.134 Even though mNGS has proven its ability in detecting CNS diseases, it cannot eliminate the needs of conventional methods. Hence, more studies are needed to search for standardized guidelines of mNGS in order for the technique to be stand-alone.
This review paper compiled and discussed a few laboratory diagnosis methods together with its findings for researchers to make a proper decision in the detection of viruses causing CNS disease. Isolation of virus via cell culture technique can be applied to cultivatable viruses such as HSV-1, HSV-2, CMV, VZV, RSV, influenza A and B viruses, parainfluenza viruses, enteroviruses, measles virus and respiratory adenoviruses. The detection can be determined by the appearance of typical CPE, which is time-consuming depending on the cell line involved, time of incubation and type of clinical specimen collected. However, rapid culture is able to speed up the results within 2-3 days before CPE occurs. Next, electron microscopy technique can be applied on uncultivable virus, and it gives a clear viral morphology for an easy and rapid detection. However, observer must have special skills for interpretation of the image outcome. Cryo-EM promotes advance technique examining the virus at near atomic resolution which provides much information for various studies. Serological diagnosis is depending on the stage of infection, preferable at acute (IgM) and convalescent phase (IgG). It promotes high sensitivity and specificity, but detection could be delay due to late emergence of antibody/antigen and tendency to cross react with viruses from the same family. Thus, the viral antigen and monoclonal antiviral antibody should be carefully chosen to prevent cross-reactivities. Real-time PCR is more sensitive and specific compared to the others, it also provides timely detection, but the assay is high cost in terms of reagents and specialised equipment, the designation of primers and probes should be carefully developed to avoid inaccurate detection. The application of mNGS in diagnosis of CNS diseases has been proven by many researchers to increase the yields in recent years. However, it is advised to implement this technique in conjunction with the other conventional methods in order to confirm and validate the accuracy of results. Lastly, the selection of laboratory test needs close interaction between clinician and laboratory personnel. An inexpensive yet great quality test combines with rapid results will enable healthcare workers to give suitable treatment to combat this CNS infections.
ACKNOWLEDGMENTS
The authors would like to thank Institute for Medical Molecular Biotechnology and Faculty of Medicine, Universiti Teknologi MARA for providing all the essential facilities and support to complete this article.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial contribution, direct, and intellectual contribution to the work, and approved it for publication.
FUNDING
This work was supported by the Malaysia Ministry of Education Research Grant Scheme 600-RMC/PRGS 5/3 (018/2019).
DATA AVAILABILITY
Not applicable.
ETHICS STATEMENT
Not applicable.
- Tuladhar A, Mitrousis N, Fuhrmann T, Shoichet MS. Central Nervous System. Translational Regenerative Medicine. 2015:415-435.
Crossref - He T, Kaplan S, Kamboj M, Tang YW. Laboratory Diagnosis of Central Nervous System Infection. Curr Infect Dis Rep. 2016;18(11):35.
Crossref - Ballios BG, Baumann MD, Cooke MJ, Shoichet MS. Central Nervous System. In Atala A, Lanza R, Thomson JA, Nerem R, 2nd Ed. (eds.), Principles of Regenerative Medicine. 2011:1023-1046.
Crossref - Archibald LK, Quisling RG. Central Nervous System Infections. Textbook of Neurointensive Care. 2013;427-517.
Crossref - Parikh V, Tucci V, Galwankar S. Infections of the nervous system. Int J Crit Illn Inj Sci. 2012;2(2):82-97.
Crossref - Meningitis. Centres for Disease Control and Prevention. https://www.cdc.gov/meningitis/index.html. Accessed 14 February, 2022.
- Ellul M, Solomon T. Acute encephalitis – diagnosis and management. Clin Med (Lond). 2018;18(2):155-159.
Crossref - Bokhari MR, Mesfin FB. Brain Abscess. In StatPearls. Treasure Island (FL): StatPearls Publishing; 2022 Jan.
- Lim PAC. Transverse Myelitis. Essentials of Physical Medicine and Rehabilitation. 2020;952-959.
Crossref - Bookstaver PB, Mohorn PL, Shah A, et al. Management of Viral Central Nervous System Infections: A Primer for Clinicians. J Cent Nerv Syst Dis. 2017;9:179573517703342.
Crossref - Shi X, Wu R, Shi M, et al. Simultaneous detection of 13 viruses involved in meningoencephalitis using a newly developed multiplex PCR Mag-array system. Int J Infect Dis. 2016;49:80-86.
Crossref - Giulieri SG, Chapuis-Taillard C, Manuel O, et al. Rapid detection of enterovirus in cerebrospinal fluid by a fully-automated PCR assay is associated with improved management of aseptic meningitis in adult patients. J Clin Virol. 2015;62:58-62.
Crossref - Moon SM, Kim T, Lee EM, Kang JK, Lee SA, Choi SH. Comparison of clinical manifestations, outcomes and cerebrospinal fluid findings between herpes simplex type 1 and type 2 central nervous system infections in adults. J Med Virol. 2014;86(10):1766-1771.
Crossref - Rubin S, Eckhaus M, Rennick LJ, Bamford CG, Duprex WP. Molecular biology, pathogenesis and pathology of mumps virus. J Pathol. 2015;235(2):242-252.
Crossref - Reuter D, Schneider-Schaulies J. Measles virus infection of the CNS: human disease, animal models, and approaches to therapy. Med Microbiol Immunol. 2010;199(3):261-271.
Crossref - Babi MA, Pendlebury W, Braff S, Waheed W. JC Virus PCR Detection Is Not Infallible: A Fulminant Case of Progressive Multifocal Leukoencephalopathy with False-Negative Cerebrospinal Fluid Studies despite Progressive Clinical Course and Radiological Findings. Case Rep Neurol Med. 2015;2015:643216.
Crossref - Bhatt S, Gething PW, Brady OJ, et al. The global distribution and burden of dengue. Nature. 2013;496(7446):504-507.
Crossref - Ngo CC, Katoh S, Hasebe F, et al. Characteristics and biomarkers of patients with central nervous system infection admitted to a referral hospital in Northern Vietnam. Trop Med Health. 2021;49(1):42.
Crossref - Dorsett M, Liang SY. Diagnosis and Treatment of Central Nervous System Infections in the Emergency Department. Emerg Med Clin North Am. 2016;34(4):917-942.
Crossref - Shin SY, Kwon KC, Park JW, Kim JM, Shin SY, Koo SH. Evaluation of the Seeplex® Meningitis ACE Detection kit for the detection of 12 common bacterial and viral pathogens of acute meningitis. Ann Lab Med. 2012;32(1):44-49.
Crossref - Li S, Nguyen IP, Urbanczyk K. Common infectious diseases of the central nervous system-clinical features and imaging characteristics. Quant Imaging Med Surg. 2020;10(12):2227-2259.
Crossref - Mazzulli T. Laboratory diagnosis of infection due to viruses, chlamydia, chlamydophila, and mycoplasma. Principles and Practice of Pediatric Infectious Diseases. 2018;1434-1447.e5.
Crossref - Leland DS, Ginocchio CC. Role of cell culture for virus detection in the age of technology. Clin Microbiol Rev. 2007;20(1):49-78.
Crossref - Burrell CJ, Howard CR, Murphy FA. Laboratory Diagnosis of Virus Diseases. Fenner and White’s Medical Virology. 2017:135-154.
Crossref - Louten J. Detection and Diagnosis of Viral Infections. Essential Human Virology. 2016:111-132.
Crossref - Cassedy A, Parle-McDermott A, O’Kennedy R. Virus Detection: A Review of the Current and Emerging Molecular and Immunological Methods. Front Mol Biosci. 2021;8:637559.
Crossref - Slater J. Equine Herpesviruses. In Sellon DC, Long MTBT-EID, 2nd Ed. (eds.), Equine Infectious Diseases. Saunders, Philadelphia. 2014:151-168.e8.
Crossref - Bennett JE, Dolin R, Blaser MJ. Basic Principles in the Diagnosis and Management of Infectious Diseases. In Bennett J, Dolin R, Blaser MJ (eds.), Principles and Practice of Infectious Diseases, 9th Ed. Elsevier, Amsterdam. 2020.
- Peaper DR, Landry ML. Laboratory diagnosis of viral infection. Handb Clin Neurol. 2014;123:123-147.
Crossref - Hematian A, Sadeghifard N, Mohebi R, et al. Traditional and Modern Cell Culture in Virus Diagnosis. Osong Public Health Res Perspect. 2016;7(2):77-82.
Crossref - Ramachandran PS, Wilson MR. Diagnostic Testing of Neurologic Infections. Neurol Clin. 2018;36(4):687-703.
Crossref - Debiasi RL, Tyler KL. Molecular methods for diagnosis of viral encephalitis. Clin Microbiol Rev. 2004;17(4):903-925.
Crossref - Sykes JE, Rankin SC. Isolation in Cell Culture. Canine and Feline Infectious Diseases. 2014;2-9.
Crossref - Mike JL. Rapid Diagnosis of Viral Infections. Laboratory Medicine. 2002;33(4):283-286.
Crossref - MacLachlan NJ, Dubovi EJ. Laboratory Diagnosis of Viral Infections. In MacLachlan NJ, Dubovi EJ (eds.), Fenner’s Veterinary Virology, 5th Ed. Academic Press, United States. 2016:105-129.
Crossref - Jayakeerthi RS, Potula RV, Srinivasan S, Badrinath S. Shell Vial Culture assay for the rapid diagnosis of Japanese encephalitis, West Nile and Dengue-2 viral encephalitis. Virol J. 2006;3:2.
Crossref - Brumback BG, Wade CD. Simultaneous culture for adenovirus, cytomegalovirus, and herpes simplex virus in same shell vial by using three-color fluorescence. J Clin Microbiol. 1994;32(9):2289-2290.
Crossref - Crist GA, Langer JM, Woods GL, Procter M, Hillyard DR. Evaluation of the ELVIS plate method for the detection and typing of herpes simplex virus in clinical specimens. Diagn Microbiol Infect Dis. 2004;49(3):173-177.
Crossref - Kilic I, Altuglu I, Cicek C, et al. Identification of enteroviruses from central nervous system infections by RT-PCR and cell culture methods. Mikrobiyol Bul. 2011;45(3):468-477. https://pubmed.ncbi.nlm.nih.gov/21935780/.
- Curry A, Appleton H, Dowsett B. Application of transmission electron microscopy to the clinical study of viral and bacterial infections: present and future. Micron. 2006;37(2):91-106.
Crossref - Zhang P, Mendonca L. Analysis of Viruses in the Cellular Context by Electron Tomography. In Bamford DH, Zuckerman M (eds.), Encyclopedia of Virology, 4th Ed. 2021:242-247.
Crossref - Chow VT, Tambyah PA, Yeo WM, Phoon MC, Howe J. Diagnosis of nipah virus encephalitis by electron microscopy of cerebrospinal fluid. J Clin Virol. 2000;19(3):143-147.
Crossref - Goldsmith CS, Miller SE. Modern uses of electron microscopy for detection of viruses. Clin Microbiol Rev. 2009;22(4):552-563.
Crossref - Roingeard P, Raynal PI, Eymieux S, Blanchard E. Virus detection by transmission electron microscopy: Still useful for diagnosis and a plus for biosafety. Rev Med Virol. 2019;29(1):e2019.
Crossref - McMullan LK, Folk SM, Kelly AJ, et al. A new phlebovirus associated with severe febrile illness in Missouri. N Engl J Med. 2012;367(9):834-841.
Crossref - Fischer SA, Graham MB, Kuehnert MJ, et al. Transmission of lymphocytic choriomeningitis virus by organ transplantation. N Engl J Med. 2006;354(21):2235-2249.
Crossref - Broutin, L, Rousselot-Denis C, Dartigeas C, et al. Diagnosis of progressive multifocal leukoencephalopathy in case of JC virus DNA negativity in cerebrospinal fluid. Virologie (Montrouge). 2016;20(5):287-292.
Crossref - Ryu W-S. Molecular Virology of Human Pathogenic Viruses. Department of Biochemistry, Yonsei University, Seoul, Korea. 2017:141-148.
Crossref - Benjin X, Ling L. Developments, applications, and prospects of cryo-electron microscopy. Protein Sci. 2020;29(4):872-882.
Crossref - Wang X, Li SH, Zhu L, et al. Near-atomic structure of Japanese encephalitis virus reveals critical determinants of virulence and stability. Nat Commun. 2017;8(1):14.
Crossref - Hasan SS, Sun C, Kim AS, et al. Cryo-EM Structures of Eastern Equine Encephalitis Virus Reveal Mechanisms of Virus Disassembly and Antibody Neutralization. Cell Rep. 2018;25(11):3136-3147.e5.
Crossref - Poonsiri T, Wright GSA, Solomon T, Antonyuk SV. Crystal Structure of the Japanese Encephalitis Virus Capsid Protein. Viruses. 2019;11(7):623.
Crossref - Scimeca M, Montanaro M, Bonfiglio R, Anemona L, Bonanno E. Electron microscopy in human diseases: diagnostic and research perspectives. Nanomedicine (Lond). 2019;14(4):371-373.
Crossref - Ouattara LA, Barin F, Barthez MA, et al. Novel human reovirus isolated from children with acute necrotizing encephalopathy. Emerg Infect Dis. 2011;17(8):1436-1444.
Crossref - Richert-Poggeler KR, Franzke K, Hipp K, Kleespies RG. Electron Microscopy Methods for Virus Diagnosis and High Resolution Analysis of Viruses. Front Microbiol. 2019;9:3255.
Crossref - Schramlova J, Arientova S, Hulinska D. The role of electron microscopy in the rapid diagnosis of viral infections–review. Folia Microbiol (Praha). 2010;55(1):88-101.
Crossref - Kuroda H. Update on Herpes Simplex Encephaliti. Brain Nerve. 2015;67(7):931-939.
Crossref - Vainionpaa R, Leinikki P. Diagnostic Techniques: Serological and Molecular Approaches. Encyclopedia of Virology. 2008;29-37.
Crossref - MacLachlan NJ, Dubovi EJ. Laboratory Diagnosis of Viral Infections. In MacLachlan NJ, Dubovi EJ (eds.), Fenner’s Veterinary Virology, 4th Ed. Academic Press, United States. 2011:101-123.
Crossref - Ramakrishna C, Bergmann CC, Atkinson R, Stohlman SA. Control of central nervous system viral persistence by neutralizing antibody. J Virol. 2003;77(8):4670-4678.
Crossref - Balingit JC, Phu Ly MH, Matsuda M, et al. A Simple and High-Throughput ELISA-Based Neutralization Assay for the Determination of Anti-Flavivirus Neutralizing Antibodies. Vaccines (Basel). 2020;8(2):297.
Crossref - Liao PH, Yang HH, Chou PT, et al. Sufficient virus-neutralizing antibody in the central nerve system improves the survival of rabid rats. J Biomed Sci. 2012;19(1):61.
Crossref - Kurane I. Japanese Encephalitis. In Barrett ADT, Stanberry LR (eds.). Vaccines. 2009:527-535.
Crossref - Kaufmann L, Syedbasha M, Vogt D, et al. An Optimized Hemagglutination Inhibition (HI) Assay to Quantify Influenza-specific Antibody Titers. J Vis Exp. 2017;(130):55833.
Crossref - Spackman E, Sitaras I. Hemagglutination Inhibition Assay. Methods Mol Biol. 2020;2123:11-28.
Crossref - Pedersen JC. Hemagglutination-inhibition assay for influenza virus subtype identification and the detection and quantitation of serum antibodies to influenza virus. Methods Mol Biol. 2014;1161:11-25.
Crossref - Anantapreecha S, A-Nuegoonpipat A, Prakrong S, et al. Dengue virus cross-reactive hemagglutination inhibition antibody responses in patients with primary dengue virus infection. Jpn J Infect Dis. 2007;60(5):267-270.
- Reta DH, Tessema TS, Ashenef AS, et al. Molecular and Immunological Diagnostic Techniques of Medical Viruses. Int J Microbiol. 2020;2020:8832728.
Crossref - Yang Z, Lee J, Ahn HJ, Chong CK, Dias RF, Nam HW. Western Blot Detection of Human Anti-Chikungunya Virus Antibody with Recombinant Envelope 2 Protein. Korean J Parasitol. 2016;54(2):239-241.
Crossref - Yeh CT, Han CM, Lo SY, et al. Early detection of anti-HCc antibody in acute hepatitis C virus (HCV) by western blot (immunoblot) using a recombinant HCV core protein fragment. J Clin Microbiol. 1994;32(9):2235-2241.
Crossref - Dronina J, Samukaite-Bubniene U, Ramanavicius A. Advances and insights in the diagnosis of viral infections. J Nanobiotechnology. 2021;19(1):348.
Crossref - Sakamoto S, Putalun W, Vimolmangkang S, et al. Enzyme-linked immunosorbent assay for the quantitative/qualitative analysis of plant secondary metabolites. J Nat Med. 2018;72(1):32-42.
Crossref - Tan le V, Thai le H, Phu NH, et al. Viral aetiology of central nervous system infections in adults admitted to a tertiary referral hospital in southern Vietnam over 12 years. PLoS Negl Trop Dis. 2014;8(8):e3127.
Crossref - Mawuntu AHP, Bernadus JBB, Dhenni R, et al. Detection of central nervous system viral infections in adults in Manado, North Sulawesi, Indonesia. PLoS One. 2018;13(11):e0207440.
Crossref - Lanciotti RS, Kerst AJ. Nucleic acid sequence-based amplification assays for rapid detection of West Nile and St. Louis encephalitis viruses. J Clin Microbiol. 2001;39(12):4506-4513.
Crossref - Wilson MR, Zimmermann LL, Crawford ED, et al. Acute West Nile Virus Meningoencephalitis Diagnosed Via Metagenomic Deep Sequencing of Cerebrospinal Fluid in a Renal Transplant Patient. Am J Transplant. 2017;17(3):803-808.
Crossref - Nagel MA, Forghani B, Mahalingam R, et al. The value of detecting anti-VZV IgG antibody in CSF to diagnose VZV vasculopathy. Neurology. 2007;68(13):1069-1073.
Crossref - Tiwari RI, Bhullar SS, Chandak NH, et al. Diagnosis of viral central nervous system infections using antipeptide antibody against viral antigen by ELISA. Acta Virol. 2018;62(4):386-393.
Crossref - Chabierski S, Barzon L, Papa A, et al. Distinguishing West Nile virus infection using a recombinant envelope protein with mutations in the conserved fusion-loop. BMC Infect Dis. 2014;14:246.
Crossref - Shafik CF, Mohareb EW, Youssef FG. Comparison of direct fluorescence assay and real-time rt-PCR as diagnostics for respiratory syncytial virus in young children. J Trop Med. 2011;2011:781919.
Crossref - Johnson J, Higgins A, Navarro A, et al. Subtyping influenza A virus with monoclonal antibodies and an indirect immunofluorescence assay. J Clin Microbiol. 2012;50(2):396-400.
Crossref - De Ory F, Sanchez-Seco MP, Vazquez A, et al. Comparative Evaluation of Indirect Immunofluorescence and NS-1-Based ELISA to Determine Zika Virus-Specific IgM. Viruses. 2018;10(7):379.
Crossref - Chan EL, Brandt K, Horsman GB. Comparison of Chemicon SimulFluor direct fluorescent antibody staining with cell culture and shell vial direct immunoperoxidase staining for detection of herpes simplex virus and with cytospin direct immunofluorescence staining for detection of varicella-zoster virus. Clin Diagn Lab Immunol. 2001;8(5):909-912.
Crossref - Landry ML. Developments in immunologic assays for respiratory viruses. Clin Lab Med. 2009;29(4): 635-647.
Crossref - Landry ML, Ferguson D. Cytospin-enhanced immunofluorescence and impact of sample quality on detection of novel swine origin (H1N1) influenza virus. J Clin Microbiol. 2010;48(3):957-959.
Crossref - Busson L, Bartiaux M, Brahim S, et al. Prospective evaluation of diagnostic tools for respiratory viruses in children and adults. J Virol Methods. 2019;266:1-6.
Crossref - Dwivedi S, Purohit P, Misra R, et al. Diseases and Molecular Diagnostics: A Step Closer to Precision Medicine. Indian J Clin Biochem. 2017;32(4):1-25.
Crossref - Cobo F. Application of molecular diagnostic techniques for viral testing. Open Virol J. 2012;6:104-114.
Crossref - Gupta N. DNA Extraction and Polymerase Chain Reaction. J Cytol. 2019;36(2):116-117.
Crossref - Lorenz TC. Polymerase chain reaction: basic protocol plus troubleshooting and optimization strategies. J Vis Exp. 2012;(63):e3998.
Crossref - Dhull D, Sharma V, Sharma Y, Kaushik S. Applicability of molecular assays for detection and typing of herpes simplex viruses in encephalitis cases. Virusdisease. 2019;30(4):504-510.
Crossref - Motohashi K. Development of highly sensitive and low-cost DNA agarose gel electrophoresis detection systems, and evaluation of non-mutagenic and loading dye-type DNA-staining reagents. PLoS One. 2019;14(9):e0222209.
Crossref - Davis LE, Tyler KL. Molecular diagnosis of CNS viral infections. J Neurol Neurosurg Psychiatry. 2005;76(1):10.
Crossref - Puchhammer-Stockl E, Presterl E, Croy C, et al. Screening for possible failure of herpes simplex virus PCR in cerebrospinal fluid for the diagnosis of herpes simplex encephalitis. J Med Virol. 2001;64(4):531-536.
Crossref - Kralik P, Ricchi M. A Basic Guide to Real Time PCR in Microbial Diagnostics: Definitions, Parameters, and Everything. Front Microbiol. 2017;8:108.
Crossref - MA Mustafa, MQ Al-Samarraie, MT Ahmed. Molecular techniques of viral diagnosis. Science Archives. 2020;1 (3), 98-101.
Crossref - Maignan M, Viglino D, Hablot M, et al. Diagnostic accuracy of a rapid RT-PCR assay for point-of-care detection of influenza A/B virus at emergency department admission: A prospective evaluation during the 2017/2018 influenza season. PLoS One. 2019;14(5):e0216308.
Crossref - Falsey AR, Formica MA, Walsh EE. Diagnosis of respiratory syncytial virus infection: comparison of reverse transcription-PCR to viral culture and serology in adults with respiratory illness. J Clin Microbiol. 2002;40(3):817-820.
Crossref - Jalali M, Zaborowska J, Jalali M. The Polymerase Chain Reaction: PCR, qPCR, and RT-PCR. Basic Science Methods for Clinical Researchers. 2017:1-18.
Crossref - Shen M, Zhou Y, Ye J, et al. Recent advances and perspectives of nucleic acid detection for coronavirus. J Pharm Anal. 2020;10(2):97-101.
Crossref - Del Pilar Martinez Viedma M, Puri V, Oldfield LM, Shabman RS, Tan GS, Pickett BE. Optimization of qRT-PCR assay for zika virus detection in human serum and urine. Virus Res. 2019;263:173-178.
Crossref - Behzadi MA, Ziyaeyan M, Alborzi A. A diagnostic one-step real-time reverse transcription polymerase chain reaction method for accurate detection of influenza virus type A. Arch Med Sci. 2016;12(6):1286-1292.
Crossref - Navarro E, Serrano-Heras G, Castano MJ, Solera J. Real-time PCR detection chemistry. Clin Chim Acta. 2015;439:231-250.
Crossref - Zhang Y, Li Y, Guan Z, et al. Rapid Differential Detection of Japanese Encephalitis Virus and Getah Virus in Pigs or Mosquitos by a Duplex TaqMan Real-Time RT-PCR Assay. Front Vet Sci. 2022;9:839443.
Crossref - Farrell RE. Quantitative PCR Techniques. In: Farrell RE, RNA Methodologies, 5th Ed. Academic Press, United States. 2017:283-328.
Crossref - Wang C, Yang CJ. Application of Molecular Beacons in Real-Time PCR. Molecular Beacons. 2013;45-59.
Crossref - Klein D. Quantification using real-time PCR technology: applications and limitations. Trends Mol Med. 2002;8(6):257-260.
Crossref - Eigner U, Reucher S, Hefner N, et al. Clinical evaluation of multiplex RT-PCR assays for the detection of influenza A/B and respiratory syncytial virus using a high throughput system. J Virol Methods. 2019;269:49-54.
Crossref - Hwang KA, Ahn JH, Nam JH. Development and validation of multiplex real-time PCR assays for rapid detection of cytomegalovirus, Epstein-Barr virus, and polyomavirus BK in whole blood from transplant candidates. J Microbiol. 2018;56(8):593-599.
Crossref - Ou TP, Yun C, Auerswald H, et al. Improved detection of dengue and Zika viruses using multiplex RT-qPCR assays. J Virol Methods. 2020;287:113862.
Crossref - Vetter P, Schibler M, Herrmann JL, Boutolleau D. Diagnostic challenges of central nervous system infection: extensive multiplex panels versus stepwise guided approach. Clin Microbiol Infect. 2020;26(6):706-712.
Crossref - Hwang K-A, Ahn JH, Nam J-H. Diagnosis of Viral Infection Using Real-time Polymerase Chain Reaction. jbv. 2018;48(1):1-13.
Crossref - Zhang H, Yan Z, Wang X, et al. Determination of Advantages and Limitations of qPCR Duplexing in a Single Fluorescent Channel. ACS Omega. 2021;6(34):22292-22300.
Crossref - Boppana SB, Ross SA, Shimamura M, et al. Saliva polymerase-chain-reaction assay for cytomegalovirus screening in newborns. N Engl J Med. 2011;364(22):2111-2118.
Crossref - Gartzonika C, Vrioni G, Priavali E, et al. Utility of Real-Time PCR in the Diagnosis of Primary Epstein-Barr Virus Infection. J Med Microb Diagn. 2012;2(1):118.
Crossref - Bhullar SS, Chandak NH, Purohit HJ, Taori GM, Daginawala HF, Kashyap RS. Determination of viral load by quantitative real-time PCR in herpes simplex encephalitis patients. Intervirology. 2014;57(1):1-7.
Crossref - Shao N, Li F, Nie K, et al. TaqMan Real-time RT-PCR Assay for Detecting and Differentiating Japanese Encephalitis Virus. Biomed Environ Sci. 2018;31(3):208-214.
Crossref - Parker J, Fowler N, Walmsley ML, et al. Analytical Sensitivity Comparison between Singleplex Real-Time PCR and a Multiplex PCR Platform for Detecting Respiratory Viruses [published correction appears in PLoS One. 2018;13(10):e0205483]. PLoS One. 2015;10(11):e0143164.
Crossref - Thao NT, Ngoc NT, Tu PV, et al. Development of a multiplex polymerase chain reaction assay for simultaneous identification of human enterovirus 71 and coxsackievirus A16. J Virol Methods. 2010;170(1-2):134-139.
Crossref - da Cunha M de LR de S. Molecular Biology in Microbiological Analysis. Reference Module in Food Science. 2019.
Crossref - Launes C, Casas-Alba D, Fortuny C, Valero-Rello A, Cabrerizo M, Munoz-Almagro C. Utility of FilmArray Meningitis/Encephalitis Panel during Outbreak of Brainstem Encephalitis Caused by Enterovirus in Catalonia in 2016. J Clin Microbiol. 2016;55(1):336-338.
Crossref - Piccirilli G, Chiereghin A, Gabrielli L, et al. Infectious meningitis/encephalitis: evaluation of a rapid and fully automated multiplex PCR in the microbiological diagnostic workup. New Microbiol. 2018;41(2):118-125.
- Volle R, Bailly JL, Mirand A, et al. Variations in cerebrospinal fluid viral loads among enterovirus genotypes in patients hospitalized with laboratory-confirmed meningitis due to enterovirus. J Infect Dis. 2014;210(4):576-584.
Crossref - Ramamurthy M, Alexander M, Aaron S, et al. Comparison of a conventional polymerase chain reaction with real-time polymerase chain reaction for the detection of neurotropic viruses in cerebrospinal fluid samples. Indian J Med Microbiol. 2011;29(2):102-109.
Crossref - Shen C-H. Chapter 9 – Amplification of Nucleic Acids. Diagnostic Molecular Biology. 2019:215-247.
Crossref - Ruzek D, Piskunova N, Zampachova E. High variability in viral load in cerebrospinal fluid from patients with herpes simplex and varicella-zoster infections of the central nervous system. Clin Microbiol Infect. 2007;13(12):1217-1219.
Crossref - Frange P, Boutolleau D, Leruez-Ville M, et al. Temporal and spatial compartmentalization of drug-resistant cytomegalovirus (CMV) in a child with CMV meningoencephalitis: implications for sampling in molecular diagnosis. J Clin Microbiol. 2013;51(12):4266-4269.
Crossref - Lindeman NI, Fletcher JA & Longtine JA. Chapter 31: Application of Modern Techniques. Diagnostic Histopathology of Tumors. 5th Edition. Elsevier; 2021:2288-2310.
- Brown JR, Bharucha T, Breuer J. Encephalitis diagnosis using metagenomics: application of next generation sequencing for undiagnosed cases. J Infect. 2018;76(3):225-240.
Crossref - Xing X-W, Zhang J, Ma Y-B, et al. Metagenomic Next-Generation Sequencing for Diagnosis of Infectious Encephalitis and Meningitis: A Large, Prospective Case Series of 213 Patients. Front Cell Infect Microbiol. 2020;10:88.
Crossref - Zhu Y, Xu M, Ding C, et al. Metagenomic Next-Generation Sequencing vs. Traditional Microbiological Tests for Diagnosing Varicella-Zoster Virus Central Nervous System Infection. Front Public Heal. 2022;9:738412.
Crossref - Carbo E, Blankenspoor I, Goeman J, Kroes A, Claas E, Vries J. Viral metagenomic sequencing in the diagnosis of meningoencephalitis: a review of technical advances and diagnostic yield. Expert Rev Mol Diagn. 2021;21.
Crossref - Cheng Y-W, Stefaniuk C, Jakubowski MA. Real-time PCR and targeted next-generation sequencing in the detection of low level EGFR mutations: Instructive case analyses. Respir Med Case Reports. 2019;28:100901.
Crossref - Li C, Wang Y. Progress in the application of metagenomic next-generation sequencing in pediatric infectious diseases. Pediatr Neonatol. 2022;63(5):445-451.
Crossref - Ramachandran PS, Ramesh A, Creswell F V, et al. Integrating central nervous system metagenomics and host response for diagnosis of tuberculosis meningitis and its mimics. Nat Commun. 2022;13(1):1675.
Crossref - Nguyen A, Le Nguyen TN, Hong N, et al. Viral Metagenomic Analysis of Cerebrospinal Fluid from Patients with Acute Central Nervous System Infections of Unknown Origin, Vietnam. Emerg Infect Dis. 2021;27:205-213.
Crossref - Zanella M-C, Lenggenhager L, Schrenzel J, Cordey S, Kaiser L. High-throughput sequencing for the aetiologic identification of viral encephalitis, meningoencephalitis and meningitis. A narrative review and clinical appraisal. Clin Microbiol Infect. 2019;25(4):422-430.
Crossref - Forbes JD, Knox NC, Ronholm J, Pagotto F, Reimer A. Metagenomics: The Next Culture-Independent Game Changer. Front Microbiol. 2017;8.
Crossref - Simner P, Miller S, Carroll K. Understanding the Promises and Hurdles of Metagenomic Next-Generation Sequencing as a Diagnostic Tool for Infectious Diseases. Clin Infect Dis. 2017;66.
Crossref - Ramchandar N, Coufal N, Warden A, et al. Metagenomic Next-Generation Sequencing for Pathogen Detection and Transcriptomic Analysis in Pediatric Central Nervous System Infections. Open Forum Infect Dis. 2021;8.
Crossref - Piantadosi A, Mukerji S, Ye S, et al. Enhanced Virus Detection and Metagenomic Sequencing in Patients with Meningitis and Encephalitis. MBio. 2021;12.
Crossref - Wilson M, Sample H, Zorn K, et al. Clinical Metagenomic Sequencing for Diagnosis of Meningitis and Encephalitis. N Engl J Med. 2019;380:2327-2340.
Crossref - Perlejewski K, Bukowska I, Rydzanicz M, et al. Next-generation sequencing in the diagnosis of viral encephalitis: sensitivity and clinical limitations. Sci Rep. 2020;10:16173.
Crossref - Hasan M, Sundararaju S, Tang P, et al. A metagenomics-based diagnostic approach for central nervous system infections in hospital acute care setting. Sci Rep. 2020;10:11194.
Crossref - Granerod J, Ambrose H, Davies N, et al. Causes of encephalitis and differences in their clinical presentations in England: A multicentre, population-based prospective study. Lancet Infect Dis. 2010;10(12):835-844.
Crossref - Liu L, Wang JW, Wang CJ, et al. Meta genomic next-generation sequencing of a case of intracranial hemorrhage with positive herpes simplex virus and literature review. Chin J Appl Clini Pediatr 2020;35:1263-1266.
- Zhang Y, Cui P, Zhang H-C, et al. Clinical application and evaluation of metagenomic next-generation sequencing in suspected adult central nervous system infection. J Transl Med. 2020;18(1):199.
Crossref - Schlaberg R, Chiu C, Miller S, Procop G, Weinstock G. Validation of Metagenomic Next-Generation Sequencing Tests for Universal Pathogen Detection. Arch Pathol Lab Med. 2017;141(6):776-786.
Crossref - Ramachandran P, Wilson M. Metagenomics for neurological infections – expanding our imagination. Nat Rev Neurol. 2020;16:1-10.
Crossref - Chen J, Zhang R, et al. Clinical usefulness of metagenomic next-generation sequencing for the diagnosis of central nervous system infection in people living with HIV. Int J Infect Dis. 2021;107:139-144.
Crossref
© The Author(s) 2022. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.