ISSN: 0973-7510
E-ISSN: 2581-690X
The present study investigates the cellulolytic potential of three prokaryotic strains belonging to genus Bacillus isolated from the benthic soil of two aquaculture farms in East Kolkata Wetlands. For the purpose of identification and phylogenetic analyses, biochemical characterization of these three strains were performed followed by molecular characterization of 16S rRNA sequences. Maximum cellulolytic potential was found in Bacillus cereus Strain SWA6a for both Exo – b,1-4- glucanase and Endo – b,1-4- glucanase with an activity of 0.08214 ± 0.00412 and 0.11263 ± 0.00478 IU ml-1min-1 respectively. The enzyme activity ranged from 0.06112 to 0.08214 IU ml-1min-1 for Exo – b,1-4- glucanase and 0.10018 to 0.11263 IU ml-1min-1 for Endo – b,1-4- glucanase. Degradation of free insoluble cellulose ranged from 55 to 63% within a time span of 96 hours in pH 6.8 – 7.2 when incubated at 37°C. Overall, the cellulolytic activity of the isolated strains may have strong implications in the field of biological conversion of cellulose-containing municipal wastes, agricultural and forestry wastes. These isolates maintain the equilibrium of the aquaculture pond by maintaining the biogeochemical cycle of carbon and hence has the prospect to be used for bioaugmentation purposes as a step towards a microbial-based aquaculture method.
Aquaculture benthos, Bacillus, Cellulolysis, 16S rDNA Analysis, Cellulase
The process of photosynthesis converts light energy into chemical energy and thus results in the production of plant biomass with cellulose as a major component 1. Thus as one of the most abundant materials on Earth, cellulose plays a key role in the material flow of this planet. Cellulose is one of the major renewable resources and is considerably available in agricultural and municipal wastes, however, most practiced disposal method of this waste is by burning which not only eliminates the possibility of recycling but also pollutes the environment. 1,2. Hence, research has been directed towards the study of the natural process of cellulose decomposition. The biogeochemical cycle of carbon completes with the degradation of the cellulose by microorganisms of the soil and guts of animals. Cellulase, an inducible enzyme is synthesized and secreted extracellularly by a diversity of microorganisms, including bacteria and fungi during their growth in cellulosic materials 3. Cellulose is decomposed by the microorganisms with a multi-enzyme system which converts it into its monomeric form of glucose. The enzyme system comprises Endo – b – 1,4 glucanase which randomly cleaves b 1,4 linkage and extracts elementary fibrils from cellulose crystals, followed by Exo– b – 1,4 glucanase which cleaves the non-reducing end of the cellulose fibrils and converts it into Cellobiose or Cellodextrose which is finally hydrolysed to glucose by b – Glucosidases 4. In the present study, the focus is drawn towards the prokaryotic genera only. There are numerous references of cellulose degrading bacteria from soil sources such as manure compost 5, tea garden soils 6, the soil of paddy field 7, etc. These studies report several genera of bacteria like Bacillus, Acinetobacter, Clostridium, Paenibacillus, Trichonympha, Actinomycetes, etc. 7,8 with various degradation ability and enzymatic activity. Cellulose degrading bacteria have also been isolated from animal guts such as insect caterpillar and snail 1, termites 9 and grass carp 10.
This phenomenon of cellulose decomposition is important not only ecologically but also economically from the human perspective. Deeper insight into the cellulolytic microcosm can provide plausible remedies in the management of pollution caused by Solid Municipal Wastes 11. On the other hand, depletion of non-renewable sources of energy is a matter of utmost concern to humanity. To solve this problem, plant biomass is the only foreseeable source that has a sustainable future. Cellulosic materials in this context are enticing because of its low cost and plentiful supply. Apart from these, the cellulolytic microorganisms are also found convenient in their industrial application where cellulose is used as a raw material. The most important of the industrial application may be the prospective use of lignocellulosic biofuels, where cellulosic materials would be used as a raw material for the production of sugars that can be converted into ethanol and other liquid fuels. The whole process would incorporate the use of microorganisms especially the prokaryotes to facilitate this conversion 12. Other notable industrial application of the cellulose degrading bacteria are found in the composting, paper industry, breweries and food industries 13
However, data of the prokaryotes with cellulolytic potential from the benthic soil of aquatic ecosystems are scanty. The only related study has been done on mangrove soils of Mahanadi river delta, India which reports genera of bacteria like Micrococcus spp., Bacillus spp., Pseudomonas spp., Xanthomonas spp. and Brucella spp. as potential cellulolytic strains 14. The study of the prokaryotic microcosm of the benthic soil of aquatic ecosystems is of utmost importance as they play a wide array of roles in the maintenance of the ecosystem. Recent advancements in the development of the aquaculture systems promote the idea of microbial-based culture systems over traditional culture methods. The Microbial-based culture system involves the study of the benthic soil which can lead to a better understanding of the roles of the members of the prokaryotic community in the microcosm of the aquaculture system 15. Banking on this idea, this study investigates the benthic soil of two aquaculture farms in East Kolkata Wetlands in search of prokaryotic strains with cellulolytic potentials which can be used in the designing of a sustainable aquaculture system.
(Data in this paper are from first author’s thesis to be submitted shortly for the partial fulfillment of the requirements for the Degree of Doctor of Philosophy in the Department of Zoology, The University of Burdwan, West Bengal, India.)
Culture Media
All reagents used in the study were of analytical grades procured from HIMEDIA (India) and Sigma Aldrich Chemical (USA). Minimal salt media (MSM) was used for culture of bacterial strains as a source of essential inorganic nutrients. Composition of the MSM (per litre) was (NH4)2SO4, 2g; Na2HPO4, 3.61 g; KH2PO4, 1.75g; MgSO4.7H2O, 0.2g; CaCl2.H2O, 50mg; FeSO4.7H2O, 1mg; CuSO4.5H2O, 50µg; H3BO3, 10µg; MgSO4.5H2O,10µg; ZnSO4. 7H2O, 70 µg; MoO3, 10µg; pH adjusted to 7.0.
Regular Culture and subculture were done with nutrient agar medium having Peptone 5 g, NaCl 5 g, Beef extract 1.5 g, Yeast extract 1.5 g, Agar 2%, pH 7.4±0.2 (per litre) and Luria agar having Casein enzymic hydrolysate 10g, Yeast extract 5g, Sodium chloride 5g, Agar 2%, pH 7.0±0.2 (Per litre) used after autoclaving (120°C temperature and 20 psi pressure, 15 minutes).
Collection
Soil (~5g) was collected from two Aquaculture farms in East Kolkata Wetlands (22.5528° N, 88.4501° E). Soil samples were collected from the top layer of the benthic soil under 60 cm of water . Soils were aseptically transferred to sterile containers. The containers were labelled and taken to laboratory for further processing.
Isolation
Bacterial isolation was done following the protocol of Gupta et al., 2012, 1 with minor modifications. 2g of soil was inoculated in 50 ml Minimal salt media (MSM) with 2 % w/v Carboxymethylcellulose (CMC) as the sole carbon source. The culture was maintained for 7 days at 37°C with constant shaking at 120 rpm.
The culture media was then subjected to serial dilution up to 106 times and was spread on Congo Red agar media KH2PO4 0.5 g, MgSO4 0.25 g, CMC 20 g, agar 15 g, Congo-Red 0.2 g, and gelatin 2 g; distilled water 1 L and at pH 6.8– 7.2. This media is a fast process to test the cellulolytic activity via discoloration of Congo red around the colonies as halos. After incubation for 12 hours at 37°C, single colonies of the isolates were streaked in Congo red agar media. The colony size and diameter of the halo are noted. The Hydrolysing Capacity (HC) of the isolates were calculated using the ratio of the diameter of the halo and diameter of the colony. This method was used to determine the best isolates with considerable cellulolytic potential and were selected for further studies. In this study three strains outperformed other isolates and were selected for further enzymatic analyses.
The three selected isolates were cultured in an enzyme production media composed of MSM supplemented with sterilised Filter paper (@ 0.05g / 20 ml, cut into 1cm2 pieces) for Exo– b – 1,4 glucanase assay and the same media containing CMC (0.5 g / 20 ml), at pH 6.8–7.2 for Endo– b – 1,4 glucanase assay, and the cultures were incubated for 96 hours at 37°C in a shaker incubator at 120 rpm.
After the incubation period, the cultures were subjected to centrifugation at 8000 rpm for 10 mins at 4°C. The supernatant was collected, filter sterilised and was stored at 4°C for future enzymatic assays. The pellet of the media supplemented with Filter paper was collected to analyze the degradation percentage by Gravimetric Analysis 1.
Enzyme Assay
The end product of the cellulose digestion is reducing sugar 4, thus, total cellulase activity was estimated by measuring the amount of reducing sugar formed from CMC and filter paper. The enzyme activity was determined according to the methods recommended by the International Union of Pure and Applied Chemistry (IUPAC) Commission on biotechnology following the protocol given by Rajoka and Malik, 1994 16 with minor modifications. The Exo– b – 1,4 glucanase activity of the bacterial isolates were determined by incubating 0.5ml of the sterilised supernatant collected as mentioned earlier, with 1ml of 0.1M citrate buffer pH-4.8 and sterilised Whatman no1 filter paper (50mg) in the form of 1cm2 pieces for 1hour at 50°C. The reaction was stopped by adding 3 ml of the 3,5-dinitrosalicylic acid (DNS) reagent to the reaction mixture. DNS added reaction mixture was boiled for 15 minutes and 40% sodium potassium tartrate was added to it. After cooling to room temperature, absorbance was measured at 550 nm (Systronics R2202 Double Beam Spectrophotometer) against blank with same constituents as above except the supernatant which was replaced with sterile triple distilled water of same volume.
Endo– b– 1,4 glucanase activity was assayed by measuring the amount of reducing sugar from Carboxymethylcellulose and was determined by a similar process as that of Exo–b–1,4 glucanase, except the absorbance, was measured at 540 nm against blank 16. The OD value thus obtained was then put into the equation as given below to obtain the activity of the enzyme in IU ml-1min-1.
Enzyme Activity (IU ml-1min-1) = Absorbance x Standard factor
A standard estimation was done using known concentration gradient of glucose (0.1 – 1.0 mg ml-1, 10 samples at 0.1 mg ml-1 interval) and following the same principle as stated above. The OD value thus obtained was used to calculate the standard factor. The same procedure was followed once for Endo– b – 1,4 glucanase at 540nm and for Exo–b– 1,4 glucanase at 550 nm. The Standard factor was calculated according to the following formula and finally, the mean value of the Standard factor was taken for enzyme activity estimation.
Standard Factor = Concentration of Glucose standard (mg ml-1) / Absorbance (at 540 or 550nm)
pH and Temperature stability
Cell-free supernatants containing the crude enzymes were tested for their optimal pH and thermostability. The same process of enzymatic analysis was done by varying the pH of the enzyme production media and temperature of the enzyme prior to incubation. The pH was tested within a range of 4 – 10 (7 samples with an interval of 1 pH) and temperature were tested from 20°C to 70°C (6 samples with 10°C interval).
Biochemical Characterisation
Biochemical characterisation of the bacterial isolates were done by an array of tests including MR, VP, Motility, Gram’s Staining, Citrate, Amylase, Oxidase, Catalase etc. were done to characterize the strains according to Bergey’s Manual of Systematic Bacteriology 17.
Molecular Characterisation
Genomic DNA isolation and molecular characterization of the strains
For the purpose of molecular characterization, genomic DNA was isolated from the strains by Bacterial Genomic DNA isolation kit (GCC Biotech, India). PCR amplification of the 16S rRNA gene DNA was carried out in a thermal cycler (Biorad Gradient Thermal Cycler) using 200 ml PCR tubes with a reaction mixture volume of 50 µl. Each of the reaction mixtures contained 2 µl of template DNA, 1 µl of forward primer 27f (5′-AGAGTTTGATCMTGGCTCAG-3′), 1 µl of reverse primer 1492r (5′-GGTTACCTTGTTACGACTT-3′), 25 µl of 2X PCR MasterMix (Thermo Fisher Scientific), and nuclease-free sterile water to 50 µl. The reaction mixture was subjected to an amplification of 30 cycles, each of which consisted of three steps in the following order, denaturation of template DNA at 94°C for 1 min, annealing of the template DNA at 50°C for 1 min and extension of the primers at 72°C for 2 min. The PCR products were then subjected to Agarose Gel Electrophoresis at 1.5% gel using Ethidium Bromide stain and were subsequently visualized under UV light. Amplicons were purified by Agarose Gel purification kit (New England Biolabs) and sequenced by using the paid sequencing facility from Xceleris (Ahmedabad, India).
Identification and Phylogenetic analyses
The Sequences obtained were analyzed using BLASTn in the NCBI database (http://blast.ncbi.nlm.nih.gov/Blast.cgi). The results thus obtained were screened and ten nearest neighbors were chosen based on the Highest Max Score and Total Score. The aligned sequence files were downloaded and further alignment and phylogenetic analyses were done in MEGA7 18. The alignment of the sequences with their nearest neighbors was done using CLUSTALW. To eliminate the possibility of incorporation of potential chimeras, the sequences were checked using online webtool DECIPHER 19. The phylogenetic tree was constructed based on the evolutionary distances calculated from the number of base substitutions per site of all the three strains by the Neighbor-joining method 20 using Maximum Likelihood Composite model21. The difference in composition bias was considered in evolutionary comparisons during the construction of the Phylogenetic tree 22.
Statistical analysis
All experiments were performed at least in triplicate and values are expressed as mean values ± standard deviation (S.D.)]. Statistical analyses were done and means were compared by Student’s t-test using SPSS 17.0 (SPSS Inc., Chicago, USA). Differences between means at (P ≤ 0.05) level were considered significant.
Characterisation of Cellulose degrading Strains
The three isolates were names as strains SWA4, SWA13a, and SWO6. Results of morphological characterization of these strains like colony shape, cellular morphology, gram character and detailed biochemical characteristics are provided in Table 1. The Results showed that SWO4 and SWO13a as a Gram-negative, non – motile strain whereas SWA6a is a gram-positive, motile strain. All the three strains were capable of producing enzymes like gelatinase, amylase, urease, citratase, catalase and oxidase other than cellulase. The strains were capable of fermenting sugars with varying potentiality.
Table (1):
Biochemical Tests Result of the Isolates. The Positive result is represented with + symbol and negative with – symbol. Y and R represents Yellow and Red colour of the indicator in TSI media. Yellow indicates that the strains are able to ferment sugars in aerobic (Slant) or anaerobic (Butt) condition whereas Red indicates negative result. W stands for white colony in MacKonkey media which indicates these strains were able to utilise lactose.
| Biochemical Tests | Strains | |||
|---|---|---|---|---|
| SWO4 | SWO13a | SWA6a | ||
| TSI | Slant | Y | Y | R |
| Butt | R | Y | Y | |
| H2S | – | – | – | |
| Gas | – | – | – | |
| Oxidase | + | + | + | |
| Catalase | + | – | – | |
| Gram Staining | – | – | + | |
| KOH String Test | – | – | + | |
| MR | + | – | – | |
| VP | – | – | + | |
| Gelatinase | – | – | + | |
| Indole | – | – | – | |
| Motility | – | – | + | |
| DNase | – | – | – | |
| MacKonkey | W | W | – | |
| Urease | – | + | + | |
| Citrate | + | – | – | |
| Amylase | + | + | – | |
Molecular Characterisation
BLASTn analysis results are depicted in Table 2. Isolates SWO13a and SWA6a were identified to be as Bacillus cereus with 97% and 99% homology respectively. Isolate SWO4 was Bacillus flexus with 98% sequence homology. The Phylogenetic tree constructed is depicted in Figure 3. The analysis of the tree shows that the isolates are in separate lineages and are different from one another with SWO4 in a separate lineage altogether.
Table (2):
The Closest Relative, Max Score and Identity % obtained from the BLASTn analysis of the three isolates. The GenBank accession numbers of the strains are also mentioned.
Isolate |
Closest Relative |
Max Score |
Identity (%) |
Accession Number (GenBank) |
|---|---|---|---|---|
SWO4 |
Bacillus flexus |
1753 |
98% |
MG461558 |
SWO13a |
Bacillus cereus |
1430 |
97% |
MF554692 |
SWA6a |
Bacillus cereus |
1555 |
99% |
MF554643 |
Screening test results
The Screening test result performed in Congo Red agar media revealed a glimpse of the comparative account of cellulolytic potential of the three strains involved and is given in Table 3. The highest HC value was found in Isolate SWA6a and the lowest was given by SWO4. However, it is mention worthy that these three strains excelled other strains that were subjected to the same test and hence was selected for the enzymatic assay.
Table (3):
The results of screening tests in Congo Red Agar. The Diameter of the Clearing Zone and that of the Colony are shown. The HC gives an estimation of the cellulolytic potential of the three strains.
Strain |
Diameter of the Clearing Zone (Halo)_(in mm) |
Diameter of the Colony (in mm) |
Hydrolysing Capacity (HC) |
|---|---|---|---|
SWO4 |
12.1 |
20.8 |
1.71 |
SWO13a |
8.7 |
17.9 |
2.05 |
SWA6a |
9.3 |
24.2 |
2.6 |
Enzyme Assay
The enzyme assay confirms the result obtained in the screening test and is given in Table 4. Isolate SWA6a scores the best enzymatic activity for both Exo – b,1-4- glucanase and Endo – b,1-4- glucanase with an activity of 0.08214 ± 0.00412 and 0.11263 ± 0.00478 IU ml-1min-1 respectively. The Overall enzyme activity ranged from 0.06112 to 0.08214 IU ml-1min-1 for Exo – b,1-4- glucanase and 0.10018 to 0.11263 IU ml-1min-1 for Endo -b,1-4- glucanase. Gravimetric analysis revealed that isolates SWO4, SWO13a, and SWA6a were able to degrade 55.1%, 61.9% and 63.4% of raw cellulose given in the form of filter paper respectively, in a span of 96 hours under the given conditions.
Table (4):
The Enzymatic activities and Cellulose Degrading ability of the isolates. The Mean values for each strain is given with SD mentioned. The Degradation of the Filter paper indicates the Cellulolytic potential of the strains.
| Strains | Activity (IU ml-1min-1) | Degradation of Filter Paper (%) | |
|---|---|---|---|
| Exo – β,1-4- glucanase | Endo-β,1-4- glucanase | ||
| SWO4 | 0.06112 ± 0.00326 | 0.10018 ± 0.00269 | 55.1 ± 2.34 |
| SWO13a | 0.07924 ± 0.00510 | 0.10301 ± 0.00156 | 61.9 ± 1.25 |
| SWA6a | 0.08214 ± 0.00412 | 0.11263 ± 0.00478 | 63.4 ± 0.98 |
Results of pH and Temperature stability estimation of both the enzymes of the three isolates are depicted graphically in Figure 1 and 2 respectively. Both the analyses showed a common pattern for both the enzymes. For pH, the maximum activity of the enzymes obtained from the strains showed the highest activity within a range of 6 – 8 and diminishing at either extremities. In case of temperature the highest activity was recorded within 30 – 40°C with the same trend of diminishing activity with either increase or decrease of the temperature. The isolate SWA6a not only showed the highest activity under optimum pH and temperature conditions but also was more temperature and pH tolerant when compared to the other two isolates.
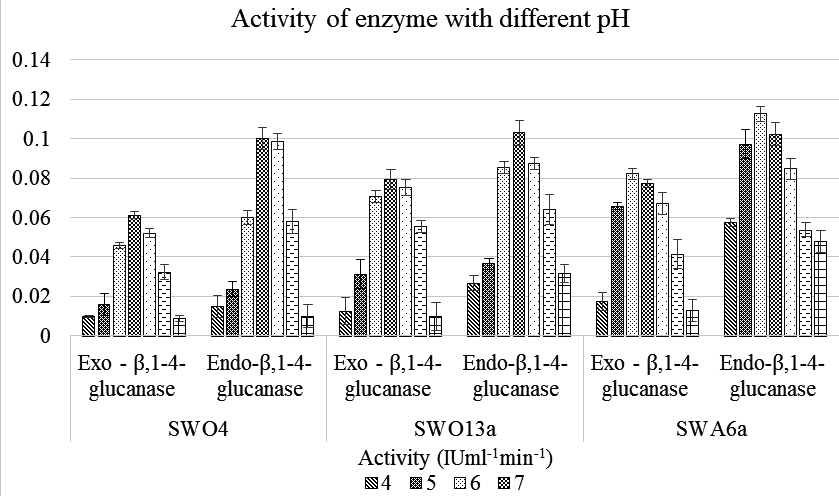
Fig. 1. Graphical representation of the variation of activity of Endo– β – 1,4 glucanase and Exo– β – 1,4 glucanase of the three Strains with varying pH ranging from pH 4 -10
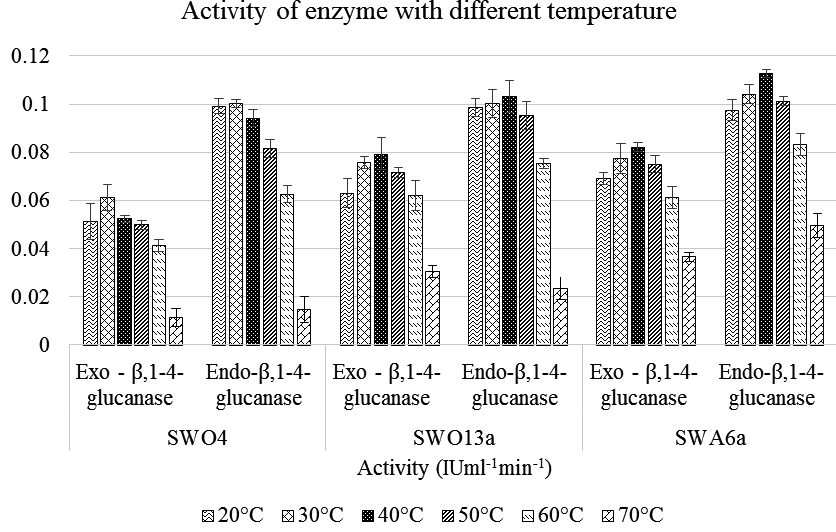
Fig. 2. Graphical representation of the variation of activity of Endo– β – 1,4 glucanase and Exo– β – 1,4 glucanase of the three Strains with varying Temperature ranging from 20°C – 70°C
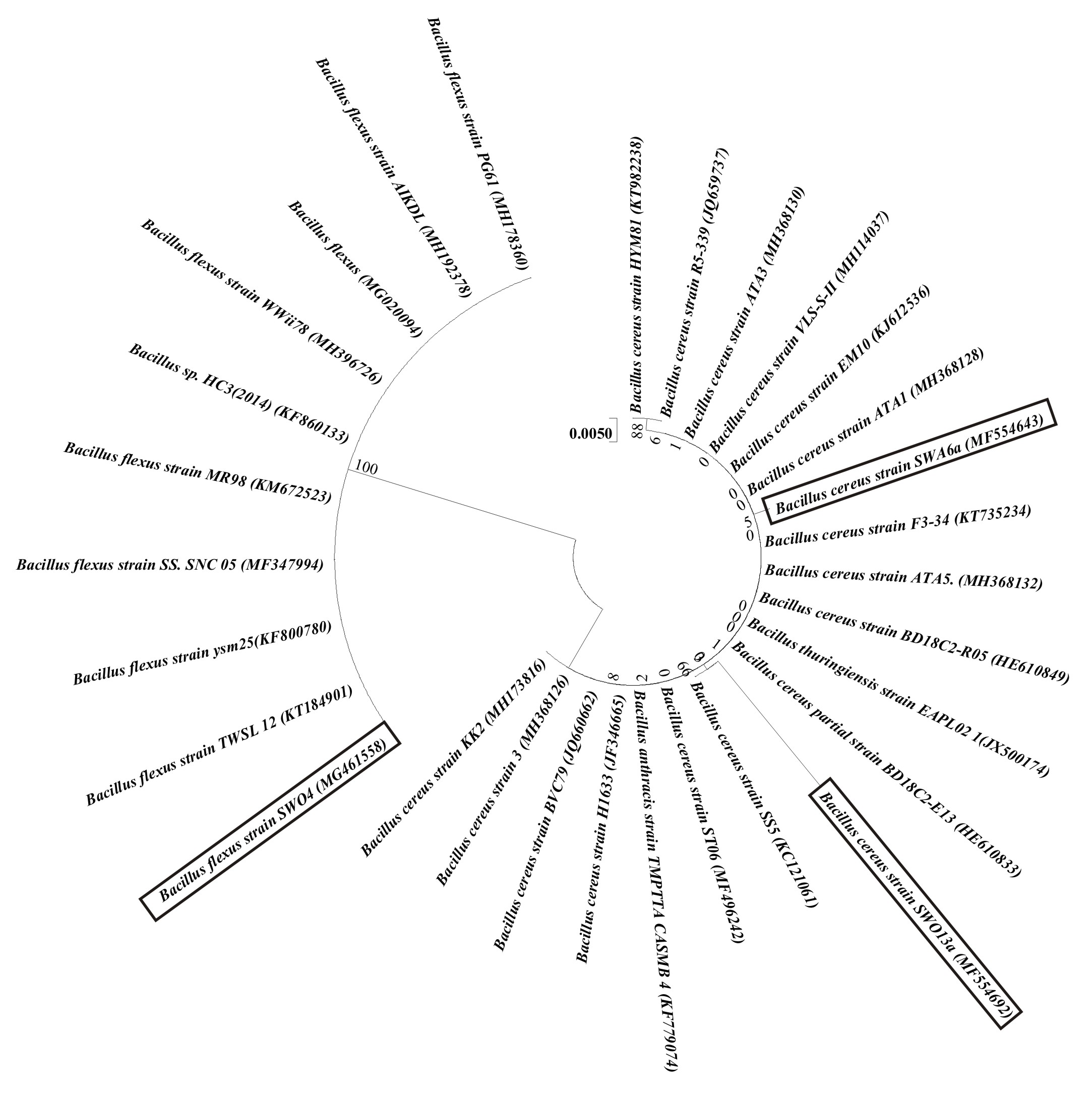
Fig. 3. Evolutionary relationships of the three strains. The Genbank Accession Numbers for each strains used in this analysis is mentioned in parenthesis. The evolutionary history was inferred using the Neighbor-Joining method. The optimal tree with the sum of branch length = 0.10455038 is shown. The tree is drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phy-logenetic tree. The evolutionary distances were computed using the Maximum Composite Likelihood method and are in the units of the number of base substitutions per site. The differences in the composition bias among se-quences were considered in evolutionary comparisons. The analysis involved 30 nucleotide sequences. All posi-tions with less than 95% site coverage were eliminated. That is, fewer than 5% alignment gaps, missing data, and ambiguous bases were allowed at any position. There were a total of 829 positions in the final dataset. Evolution-ary analyses were conducted in MEGA7
Cellulose is indeed considered as a viable option for the supply of raw materials for several industries. Several research indicates its potentiality in this regard. However, the most environment friendly way to decompose and utilise this complex biomolecule is to degrade it using microorganisms like bacteria as a tool 23. The past references showed varied degree of cellulolytic potential from unidentified strains with ranges of 0.012 to 0.196 IU ml-1min-1 for Exo – b,1-4- glucanase and 0.1622 to 0.400 IU ml-1min-1 for Endo – b,1-4- glucanase was estimated from bacteria inhabiting the gut of cellulose – feeding organisms 1. Another study demonstrated the cellulolytic potential of Bacillus licheniformis isolated from Indian Hot springs to be 0.542 IU ml-1min-1 for Exo – b,1-4- glucanase and 0.120 IU ml-1min-1 for Endo – b,1-4- glucanase 24.These two results correspond to the data obtained in the study. On the other hand, present findings do not correspond to the observations of Behera et al. who described a very high cellulolytic potential (2.471 to 98.253 IU ml-1min-1 for Endo – b,1-4- glucanase) produced by several genera of bacteria like Micrococcus spp., Bacillus spp., Pseudomonas spp., Xanthomonas spp. and Brucella spp. isolated from the benthic soil of Mahanadi river 14.
The results from various experimental protocols explored revealed that all the three isolates were capable of utilizing cellulose as a sole carbon source and their efficacy in the breakdown of cellulose is also mention worthy. Similar results were also obtained in the estimation of cellulolytic potential by the earlier workers 1,24 . The data seems to be the first report of prokaryotes with cellulolytic potential from the benthic soil of Aquaculture farms of East Kolkata Wetlands. It can be said that these bacteria are efficient in the decomposition of cellulosic materials in the aquatic ecosystem and hence maintains the ecological balance of the aquaculture ponds by maintaining a proper biogeochemical cycle of carbon providing an ecologically healthy environment for fish culture. This study also illuminates the prospective use of these strains in the cellulose dependent industry and that these strains have the potential efficacy. They can also be used in Solid Waste Management via composting method 11. From the perspective of aquaculture, identification of such strains with cellulolytic potential paves the way for future implementation for microbial – based culture methods. This strains can also be bioaugmented in the natural aquaculture ponds with high level of plant debris and will ultimately lead to the sustenance of the pond.
ACKNOWLEDGMENTS
The authors gratefully acknowledge Department of Zoology, The University of Burdwan and Department of Life Sciences, Presidency University for providing infrastructural facilities to carry out the work.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
- Gupta P, Samant K, Sahu A. Isolation of cellulose-degrading bacteria and determination of their cellulolytic potential. International Journal of Microbiology. 2012.
- Rastogi G, Muppidi GL, Gurram RN, Adhikari A, Bischoff KM, Hughes SR, Apel WA, Bang SS, Dixon DJ, Sani RK. Isolation and characterization of cellulose-degrading bacteria from the deep subsurface of the Homestake gold mine, Lead, South Dakota, USA. Journal of Industrial Microbiology and Biotechnology. 2009; 36(4), 585–598.
- Lynd LR, Weimer PJ, van Zyl WH, Pretorius IS. Microbial Cellulose Utilization: Fundamentals and Biotechnology. Microbiology and Molecular Biology Reviews. 2002; 66(4), 739–739.
- Béguin P, Aubert JP. The biological degradation of cellulose. FEMS Microbiology Reviews. 1994; 13(1),25–58.
- Lu WJ, Wang HT, Yang SJ, Wang ZC, Nie YF. Isolation and characterization of mesophilic cellulose-degrading bacteria from flower stalks-vegetable waste co-composting system. The Journal of General and Applied Microbiology. 2005; 51(6), 353–360.
- Balamurugan A, Jayanthi R, Nepolean P, Pallavi RV, Premkumar R, . Studies on cellulose degrading bacteria in tea garden soils. African Journal of Plant Science. 2011; 5, 22–27.
- Chin K jeong, Hahn D, Hengstmann U, Janssen PH, Hengstmann ULF, Liesack W. Characterization and Identification of Numerically Abundant Culturable Bacteria from the Anoxic Bulk Soil of Rice Paddy Microcosms Characterization and Identification of Numerically Abundant Culturable Bacteria from the Anoxic Bulk Soil of Rice Paddy Micr. Am Soc Microbiol. 1999
- Wang CM, Shyu CL, Ho SP, Chiou SH. Characterization of a novel thermophilic, cellulose-degrading bacterium Paenibacillus sp. strain B39. Letters in Applied Microbiology. 2008; 47(1), 46–53.
- Wenzel M, Schönig I, Berchtold M, Kämpfer P. Aerobic and facultatively anaerobic cellulolytic bacteria from the gut of the termite Zootermopsis angusticollis. Journal of Applied Microbiology. 2002; 92(1), 32–40.
- Wu S, Wang G, Angert ER, Wang W, Li W, Zou H. Composition, diversity, and origin of the bacterial community in grass carp intestine. PLoS ONE. 2012; 7(2), e30440.
- Bayer EA, Lamed R, Himmel ME. The potential of cellulases and cellulosomes for cellulosic waste management. Current Opinion in Biotechnology. 2007; 18, 237
- Rubin EM. Genomics of cellulosic biofuels. Nature. 2008; 454(7206), 841–845.
- Bhat MK, Bhat S. Cellulose degrading enzymes and their potential industrial applications. Biotechnology Advances. 15, 583 (1997)
- Behera BC, Parida S, Dutta SK, Thatoi HN. Isolation and Identification of Cellulose Degrading Bacteria from Mangrove Soil of Mahanadi River Delta and Their Cellulase Production Ability. American Journal of Microbiological Research. 2014; 2(1), 41–46.
- Martínez-Córdova LR, Emerenciano M, Miranda-Baeza A, Martínez-Porchas M. Microbial-based systems for aquaculture of fish and shrimp: An updated review. Reviews in Aquaculture. 2015; 7(2), 131–148.
- Rajoka MI, Malik KA. Cellulase production by Cellulomonas biazotea cultured in media containing different cellulosic substrates. Bioresource Technology. 1997.
- Garrity GM, Holt JG. Bergey’s Manual® Systemic Bacteriology. (Springer New York, 2001, New York, NY), pp. 119–166
- Kumar S, Stecher G, Tamura K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Molecular biology and evolution. 2016; 33(7),1870–1874.
- Bragin E, Chatzimichali EA, Wright CF, Hurles ME, Firth H V., Bevan AP, Swaminathan GJ. DECIPHER: Database for the interpretation of phenotype-linked plausibly pathogenic sequence and copy-number variation. Nucleic Acids Research. 2014; 42(D1),
- Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Molecular Biology and Evolution. 1987; 4(4), 406–425.
- Tamura K, Nei M, Kumar S. Prospects for inferring very large phylogenies by using the neighbor-joining method. Proceedings of the National Academy of Sciences. 2004; 101(30), 11030–11035.
- Tamura K, Kumar S. Evolutionary distance estimation under heterogeneous substitution pattern among lineages. Molecular Biology and Evolution. 2002.
- Sukumaran RK, Singhania RR, Pandey A. Microbial cellulases – Production, applications and challenges. Journal of Scientific and Industrial Research. 2005; 64(11), 832–844.
- Acharya S, Chaudhary A. Optimization of fermentation conditions for cellulases production by Bacillus licheniformis MVS1 and Bacillus sp. MVS3 isolated from Indian hot spring. Brazilian Archives of Biology and Technology. 2012; 55(4), 497–503.
© The Author(s) 2018. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.


